Abstract
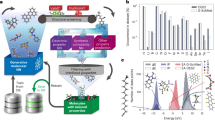
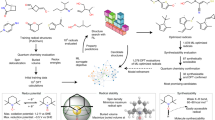
An interactive web tool, PredPotS, has been developed for predicting one-electron standard reduction potentials of organic molecules in aqueous solutions. The predictions are generated using deep learning models trained and validated on a chemically diverse database comprising reduction potentials of approximately 8000 organic compounds. The reduction potentials of this database were computed using a composite computational protocol that combines the semiempirical quantum chemical method (GFN2-xTB) and a well-established DFT approach (M06-2X functional along with the SMD solvent model). While this computational approach is cost-effective, it is subject to certain limitations, which are nonetheless duly accounted for in the development of the database. The applied graph-based deep learning methods perform remarkably well in terms of the standard performance metrics. By entering or uploading the SMILES codes of the molecules, PredPotS provides fast and sensible predictions for one-electron standard reduction potentials for a diverse set of organic molecules also in the range compatible with the electrochemical stability of aqueous electrolytes. The PredPotS web tool is particularly well-suited for screening redox-active candidates for aqueous organic redox flow batteries, but it may also prove useful in a variety of other electrochemical applications.
Similar content being viewed by others
Introduction
Materials that undergo reversible oxidation-reduction reactions (redox-active materials) play a crucial role in numerous research fields and applications, such as energy storage and conversion1,2,3,4,5, photoredox catalysis6,7,8,9, electrochemical sensing10,11,12, and molecular electronics13,14,15, for instance. Redox potentials represent one of the most basic properties of redox-active materials because they display the relative stability of distinct oxidation states, and therefore provide valuable information regarding the direction and feasibility of redox reactions. The knowledge of redox potentials is thus essential in the design of new materials and electrochemical systems. Redox potentials measured by commonly used experimental methods (i.e. potentiometry or cyclic voltametry) are available for a large number of inorganic, organometallic, and organic compounds, mostly in aqueous solutions, and they are usually reported in the form of standard reduction potentials (Eo), or as midpoint potentials at given pH (Em), if prototropic equilibria are involved16,17,18,19,20. These compilations are, however, from multiple sources and they contain data with varying levels of accuracy. Obtaining accurate redox potential data from electrochemical measurements pose several challenges that include the involvement of protonation processes, irreversibility, stability issues, slow reaction kinetics, just to name a few limitations.
The identification of new redox-active compounds and related developments in various applications of redox chemistry can be greatly accelerated via computational screening tools21,22. This approach has been recently well exploited in search for new organic redox couples as potential candidates for next-generation redox flow batteries (RFBs), which are considered as a viable solution to large-scale storage of renewable energy23,24,25,26,27,28,29,30,31. Density functional theory (DFT) calculations provide reasonably accurate redox potentials32,33,34,35,36,37,38,39,40,41,42,43, and the obtained data are often used to analyze structure-property correlations44,45,46,47,48,49, or to build extended databases for virtual screening50,51,52,53,54,55,56,57,58,59,60. Computational screening becomes particularly efficient when quantum chemical (QC) methods are combined with machine learning (ML) techniques61,62,63,64,65,66. Generating molecular databases using DFT or other QC calculations followed by regression or neural network ML analysis has been successfully used to predict redox potentials of various reduction processes relevant to RFB developments67,68,69,70,71,72,73,74,75,76,77,78,79. ML models trained on extended databases and employing molecular-input line-entry system (SMILES) strings80,81,82 as molecular representations provide sound predictions orders of magnitude faster than pure DFT calculations. ML techniques also allow for simultaneous optimization of other targeted properties (i.e. solubility, stability, synthesizability) along with redox potentials. The utilization of multiobjective ML tools in the optimization of desired properties redox-active materials is a promising concept in discovering feasible candidate redox pairs for new RFBs. 67,69,72
We have recently proposed a cost-effective computational protocol that combines semiempirical (GFN2-xTB) and DFT (M06-2X) quantum chemical methods to predict 1e− standard reduction potentials for an enlarged set of organic molecules83. This composite computational approach has been validated via benchmark studies showing satisfactory correlations with experimental data and also with those obtained from more demanding full DFT calculations. The proposed protocol has been applied to generate a molecular database of N-functionalized pyridinium derivatives, and the analysis of substituent effects on the reduction potentials served as a starting point in the exploration of vitamin B6-based redox-active benzoyl pyridinium salts as possible electrolytes in aqueous organic redox flow batteries (AO-RFBs)84. In our present work, we used a very similar computational protocol to construct a diverse database of organic molecules and we applied various deep learning models85 with the aim of developing an easy-to-use predictive tool for 1e− standard reduction potentials in aqueous phase. This tool is made available as an open-access web application, which is introduced and described herein.
Results
Molecular database RP-ChEMBL
Molecules from the ChEMBL molecular library86 were selected to build a database (referred to as RP-ChEMBL) that contains the 3D structures of \({\rm{A}}/{{\rm{A}}\cdot }^{-}\) redox pairs and the computed 1e− standard reduction potentials of the organic species A. ChEMBL is a publicly available large-scale database that provides information on bioactive molecules and proteins with emphasis on drug discovery. This database is not specifically related to redox chemistry, but it encompasses a wide array of already synthesized organic compounds that span a broad chemical space, therefore we think it is appropriate to build a diverse molecular database for screening purposes87,88,89,90. As an initial proof of concept, we focused on relatively small organic molecules. Namely, we have downloaded all molecules with molecular weight (M) less than 200 g/mol and number of heavy atoms greater than 6 from the available ChEMBL compound entries. From these ~38,000 molecules, we selected approximately 8000 molecules with the goal of maintaining high chemical diversity within the selected dataset. The pairwise Tanimoto similarity index91 was used for that purpose. We used the maxmin and sphere-exclusion algorithms92 for dissimilarity-based selection as implemented in Canvas93. Most of the selected compounds are neutral, but some of them are charged species because they are derived from salts. In these latter cases, the counterions were omitted in the subsequent calculations. For species, where the protonation state is not evident (e.g. for tautomeric pairs or anions), the Epik module of the Schrödinger package94 was used to estimate the protonation sites corresponding to pH = 7 in aqueous phase. The Open Babel toolbox [https://openbabel.org/] was used to convert the downloaded SDF structures of the molecules into SMILES codes, which were standardized by using the RDKit package95.
As noted in our previous work83, the semiempirical GFN2-xTB method does not always provide reliable structures during geometry optimizations, and these uncertainties were taken into account when constructing the RP-ChEMBL database as well. Our earlier benchmark calculations revealed that structural transformations—such as bond dissociation, full or partial intramolecular ring closure, and tautomerization—observed during geometry optimizations were typically artifacts of the GFN2-xTB method. It should be noted, however, that structural transformations occurring during the reduction process can also be chemically plausible. Nonetheless, assigning 1e− standard reduction potentials to such molecules is not straightforward, either experimentally or computationally. The analysis of electrochemically induced reactivity, which is certainly an important aspect of redox chemistry, was beyond the scope of the present work. In this study, structural changes occurring upon 1e− reduction were identified through connectivity analysis of the original and reduced states, and the affected molecules were excluded from the database. The final RP-ChEMBL database includes 8033 molecular entries involving heavy atoms C, N, O, F, P, S, Cl, and Br, and the molecular weight of these compounds varies in the range of 78–200 g/mol. The computed 1e− standard reduction potentials span a potential range between –4.3 and +0.8 V versus SHE. The distribution of molecules with respect to the computed potentials is plotted in Fig. 1. It is important to note that the potential window compatible with the practical electrochemical stability of aqueous electrolytes (from –1 V to more positive values) is less populated as a result of diversity requirement, but this potential region is considerably covered as well. All information relevant to the RP-ChEMBL molecular database (computed potentials and structural data) are provided in the Supporting Information (section S1).
Distribution of one-electron (1e−) standard reduction potentials computed for molecules in the RP-ChEMBL database, showing the overall range of redox values across the dataset.
DeepChem models and their performance
Deep learning models, as implemented in the open-source Python-based DeepChem framework [https://deepchem.io/] 96 (version 2.8.0), were used in our present work to train predictive models on the RP-ChEMBL molecular database. We applied five different deep learning architectures, namely the graph convolutional network models GCN and Graph Conv, graph attention network models GAT and Attentive FP, and the directed acyclic graph model DAG. These models were specifically developed for graph property predictions, therefore, they are expected to be suitable for handling larger datasets of organic molecules.
The primary inputs for all these models were the standardized SMILES codes of the molecules included in RP-ChEMBL, but the SMILES representations were converted to graph structures using the built-in featurizers (for details, see the Supporting Information, section S2). For model training and monitoring the performance, we followed the conventional randomized splitting with the ratio of 0.8:0.1:0.1 to divide the dataset into training, validation and test sets. The basic metrics used to validate the performance of the trainings are the mean absolute error (MAE), the root mean squared error (RMSE), and the determination coefficient of the least squares linear fitting (R2), as referenced to the potentials computed with the GFN2-xTB/M06-2X protocol. The default hyperparameters specific for the neural network of models used in DeepChem proved to perform well for the present purposes; however, some additional parameters regarding the model training and validation were optimized as described in the Supporting Information (sections S2 and S3).
The performance of the trained models is compared in Table 1. All the applied deep learning methods perform remarkably well in terms of the basic metrics. For instance, the MAE values of predictions on the test dataset are smaller than 0.2 V, the RMSE values are between 0.20–0.25 V, and there is a reasonably good correlation between the predicted and the computed potentials as indicated by the R2 data (≥0.9). The level of uncertainty of predictions is actually comparable to that of the GFN2-xTB/M06-2X computational protocol83.
Of the five deep learning models, we find the Attentive FP model to give the best performance with impressive statistical metrics on all three datasets. The parity plot displaying the correlation between the predicted and computed data for this particular model is shown in Fig. 2; the analogous plots obtained for the other models are reported in the Supporting Information (section S4).
Comparison of predicted one-electron (1e−) standard reduction potentials obtained using the Attentive FP model with computed reference values from GFN2-xTB/M06-2X calculations, illustrating the correlation between predicted and calculated data. Red, orange and green circles represent data from training, validation and test sets.
Despite the small MAE value obtained for the Test dataset with the Attentive FP model (MAE = 0.14 V), it is apparent from Fig. 2 that the deviation of the predicted potentials from the computed values covers a much broader energy range. The distribution of signed errors (Fig. 3) shows that the uncertainty of predictions is within ±0.25 V for the majority of the molecules. There are only a few outliers with notable discrepancies (>0.6 V), and the analysis of these cases reveals that some of these deviations originate from the uncertainties of the GFN2-xTB method. Although the RP-ChEMBL database was built by being aware of the limitations of this semiempirical QC method, for a few particular structural units of molecules involved in RP-ChEMBL the geometry optimization gave inaccurate structures resulting in somewhat flawed reduction potentials with the GFN2-xTB/M06-2X protocol. Some of the exceptional deviations could also be related to limited capacity of deep learning method to generalize across the diverse set of molecular structures. A detailed analysis of the origin of the outlier data is presented in the Supporting Information (section S5). Nevertheless, the overall performance of the Attentive FP model is quite acceptable, definitely sufficient for computational screening purposes, and the predictability of the other deep learning models is satisfying as well (see details in the Supporting Information, section S6).
Distribution of signed errors for predictions using the AttentiveFPModel method, with bars colored by data split (train, valid and test). Overlaid kernel density estimate curves provide a smoothed view of the error distribution for each set. Percentage value indicates the fraction of test molecules with errors within the –0.25 to 0.25 V range.
PredPotS web tool
The trained deep learning models provided the foundation for developing a predictive tool for the estimations of 1e− standard reduction potentials of small or medium-sized organic molecules in aqueous phase. This tool was implemented as an interactive web application accessible through any web browser. The web tool PredPotS (Predicting reduction potentials from SMILES codes) is hosted on a webserver at https://predpots.ttk.hu/. A detailed description of the usage of PredPotS is available in the Supporting Information (section S7), and also under the Help tab of the application. Herein, we only summarize the basic features of the web tool.
Potential predictions in PredPotS are made by entering or uploading the SMILES codes of molecules, and the results are displayed in a table that lists the predictions of all five deep learning models. Model predictions that deviate significantly from the average of these data are treated as outliers and are excluded from the calculation of the mean prediction, which is also displayed in the table. A confidence interval is calculated from the predictions of the individual models and a corresponding confidence rating (1–5) is assigned based on the interval width and the number of contributing predictions. This measure, referred to as prediction confidence, is also shown in the table. This filtering is controlled by a built-in z-score based threshold (for details, see the Supporting Information, section S7). This option can be switched off, but we suggest using it to enhance the confidence level of potential predictions.
The Similarity feature of the web application identifies the molecule included in the RP-ChEMBL database that is structurally most similar to the entered species in terms of molecular fingerprints. The similarity is measured by means of Tanimoto coefficients obtained by using RDKit. The similarity scores and the reduction potentials of the most similar molecules are displayed as a result of similarity search.
The performance metrics of the five deep learning models trained on the RP-ChEMBL molecular database and the corresponding parity plots are available under the Performance tab of the application. The parity plots embody detailed information about the molecules involved in the database (computed and predicted potentials, Lewis structure, etc.), which can be accessed interactively as described in the Help menu. All datasets discussed in the paper are available under the Datasets tab.
Illustrative examples
To illustrate the applicability of PredPotS, we first present the predictions for a few redox-active organic compounds considered recently as potential anolytes for AO-RFBs (Scheme 1). Benzoyl-pyridinium 1+ was shown to undergo reversible one-electron redox process in aqueous solutions, albeit the second redox event was found to be irreversible97. Several substituted variants of this framework have been synthesized and electrochemically characterized recently84, from which we selected 2+ as a representative. Methyl-viologen 32+ is the simplest member of the bipyridinium-based compounds, which are most commonly used active species for anolytes98,99,100,101,102,103,104,105. Extending the π-conjugation of the bipyridinium framework by incorporating a spacer between two pyridinium rings, such as in 44+, is a successful strategy to achieve a two-electron storage anolyte100,106.
Illustrative examples for the applicability of PredPotS.
The experimentally determined 1e− reduction potentials of these pyridinium derivatives (as referenced to SHE) are compiled in Table 2 along with the predictions provided by PredPotS. None of these species are involved in the RP-ChEMBL database, but similar pyridinium ions are present as demonstrated by similarity analysis (see the Supporting Information, Fig. S12). The mean predictions are fairly accurate for species 1+ and 2+, but they are reasonable for species 32+ and 44+ as well, and this seems to correlate with the level of confidence of predictions. On the other hand, the order of experimental potentials (i.e. the trend along this series of structurally related species) is not reproduced by most of the models (except Attentive FP). However, such a high level of accuracy cannot be expected for models that were trained over a much broader potential range (between –4.3 and +0.8 V versus SHE) than that covered by the potentials of the four pyridinium compounds (between –0.76 and –0.45 V). Reduction potentials computed with the GFN2-xTB/M06-2X protocol are also reported in Table 2 (last column, and they are indeed more consistent with the experimental data.
One-electron couples from Wardman compilation
For further illustration of the applicability of the PredPotS tool, we consider a larger set of compounds, for which experimental data are available in aqueous solutions. Namely, we have used the Wardman compilation of experimental reduction potentials of one-electron couples17 as a reference and analyzed the quality of PredPotS predictions. The reduction potentials reported in ref. 17 are midpoint potentials of (\({\rm{A}}/{{\rm{A}}\cdot }^{-}\)) couples, which in many cases can be considered as good estimates for 1e− standard reduction potentials. This actually depends on whether prototropic equilibria are coupled or not with reduction processes, but no pH dependence is noted for the compiled reduction potentials. The majority of experimental data refer to pH = 7, but for some entries other pH conditions are given, or even not specified. In the Wardman compilation, the organic species include various quinone, nitroaryl, bipyridinium compounds, as well as some other organic molecules, and the potentials are given versus the SHE.
Molecules with M < 300 g/mol molecular weight, altogether 313 molecules, were selected from the Wardman compilation, and potential predictions were obtained via the PredPotS tool. Although the RP-ChEMBL training database includes molecules with M < 200 g/mol, herein we extended the molecular weight range to assess the applicability of PredPotS tool for larger molecules. Radical oxidant species were not considered in our analysis, as the RP-ChEMBL database involves only closed-shell molecules. The SMILES codes of this molecular set were collected in a csv text file format and uploaded in the online application. The potential predictions were ready within 11 s after the submission. The related database is available under the Datasets tab of PredPotS.
The parity plot showing the correlation between the mean predictions and the experimental 1e− reduction potentials is displayed in Fig. 4. The level of correlation as measured by the R2 metrics is clearly reduced compared to the overall performance of the deep learning methods, but the MAE = 0.21 V value is still reasonably small. The reduced performance is partially due to the significantly narrowed range of potentials of these oxidants as compared to that of the entire RP-ChEMBL database (–1.2 to +0.2 V versus –4.3 to +0.8 V, respectively), but it is also related to the fact that larger molecules (with M > 200) were also included in the present analysis. This latter argument is apparent in Fig. 4, which shows a larger scatter of predictions for M > 200 molecules (highlighted in light blue), and also from the improved metrics found for the M < 200 set in dark blue (R2 = 0.74, MAE = 0.13 V). The parity plot for the M < 200 set of compounds is depicted in Fig. 5, where we highlighted a few classes of compounds, such as quinones, nitrobenzenes, and bipyridimium ions. These compounds form distinct groups along the trendline of the plot providing further support for the reliability of predictions.
Data corresponding to M < 200 molecules are higlighted in dark blue.
Selected classes of compounds are highligted by color codes.
The performance of the five deep learning models regarding their predictions for the M < 200 molecules of the Wardman set is illustrated in Table 3 in terms of the basic metrics. The Attentive FP model shows again the best overall performance with statistical metrics very similar to those of the averaged predictions (for details, see the Supporting Information, section S8).
Predictions for larger molecules
The results obtained for the Wardman compilation of one-electron couples imply that the applicability of the PredPotS tool might be extended to larger molecules than those involved in the RP-ChEMBL training set. To evaluate this hypothesis, we selected additional molecules from the ChEMBL database with molecular weight from the 200 < M < 300 range, and computed the 1e− standard reduction potentials using the composite GFN2-xTB/M06-2X protocol (1719 molecules; for details, see the Supporting Information, section S9). The parity plot of mean predicted versus computed data is presented in Fig. 6 and demonstrates a satisfactory correlation for this new set of molecules. The performance metrics found for the mean predictions are indeed encouraging (R2 = 0.89, MAE = 0.18 V), but the uncertainty of these predictions is clearly higher as illustrated by a relatively large number of outliers, as well as by the error distribution diagram shown in Fig. 7. This latter diagram reveals that the discrepancies between the mean predictions and computed data are within ±0.25 V for 74.9% of molecules, whereas this ratio is notably higher (84.1%) for the Test dataset of the original RP-ChEMBL database (see the Supporting Information, section S9).
Outlier data with discrepancies larger than 0.6 V are highlighted in orange.
An overlaid kernel density estimate (KDE) curve provides a smoothed view of the error distribution, and the percentage value indicates the fraction of molecules with errors within the –0.25 to 0.25 V range.
Based on our analysis, we think that the PredPotS tool could provide informative predictions for larger molecules as well, but the increased level of uncertainty should be taken into account when predictions are made for molecules with M > 200.
Discussion
Our present work builds upon a concept initially formulated within the CompBat project107, which focused on the development of machine learning–assisted high-throughput screening tools for the identification of promising electrolyte materials for aqueous organic redox flow batteries (AO-RFBs). In this context, we have proposed a composite computational approach, specifically the GFN2-xTB/M06-2X protocol, to construct redox potential databases suitable for training various machine learning models. Herein, we developed the RP-ChEMBL database, which comprises one-electron standard reduction potentials of 8033 organic molecules selected from the publicly available ChEMBL molecular library. In constructing our database, we incorporated a structurally diverse set of compounds with molecular weights (M) up to 200 g/mol. While we recognize that this range is below the typical target for balanced AO-RFB design, it still encompasses a diversity of core molecular frameworks that could be suitable for reversible one-electron uptake.
Five graph-based deep learning models were trained and validated on the RP-ChEMBL database, all of which demonstrated remarkable predictive performance. The Attentive FP model was found to be particularly efficient among these architectures. The analysis of error distributions and cases with significant discrepancies between computed and predicted reduction potentials highlighted uncertainties associated with the GFN2-xTB method, though these are limited only to a few and very specific structural motifs. Overall, the trained models demonstrated sufficient reliability for predictive applications, which led to the development of an interactive web-based tool. The resulting platform, PredPotS, requires only the SMILES representation of a molecule as input and provides rapid, reliable predictions of 1e− standard reduction potentials for organic compounds in aqueous solution assuming no chemical transformation in the reduced states of molecules. The predictions are obtained within seconds, which is only a fraction of time needed to compute the potentials with the GFN2-xTB/M06-2X protocol.
The practical utility of the PredPotS tool was evaluated using a set of organic redox couples with experimentally reported reduction potentials available in the literature. The predicted values consistently fell within the potential range compatible with the electrochemical stability window of aqueous electrolytes, and showed a reasonable correlation with experimental data, even for compounds with molecular weights exceeding those represented in the training set. This finding was further supported by an analysis on a subset of molecules from the ChEMBL database with molecular weights in the range of 200–300 g/mol, for which the model yielded predictions with only slightly increased uncertainty.
In conclusion, this study presented the methodology and implementation of a predictive tool intended for the preliminary screening of organic compounds in aqueous solution, based on their one-electron reduction potentials. We recommend the use of this tool for obtaining rapid yet reasonable estimates, particularly for molecules of relatively small size, a limitation that can be addressed through further expansion of the underlying database. Prediction accuracy may also be enhanced by incorporating full DFT-based computational methods in future database developments. Efforts in this direction are currently underway within our research group.
Methods
Computational protocol
The composite GFN2-xTB/M06-2X computational protocol has previously been described in details83, but for the sake of clarity, we outline the main features here as well.
The semiempirical extended tight-binding GFN2-xTB method108,109 as implemented in the xtb program package (version 6.4)110 was used to preoptimize the initial geometries of molecules in aqueous phase. We applied the same method to perform conformational search for both oxidation states of the molecules, which was carried out via the crest program111. The most stable conformer based on the aqueous-phase Gibbs free energies was selected for each species to calculate the reduction potential. The solvent effects were incorporated implicitly via the generalized Born model with surface area contributions (GBSA) as implemented in xtb. The rigid-rotor harmonic-oscillator (RRHO) approximation was used to estimate the thermal and entropic contributions to Gibbs free energies at T = 298 K. The structures optimized at GFN2-xTB level were used to carry out single-point DFT electronic energy calculations using the M06-2X functional112 along with the 6-311 + G(d,p) basis set, wherein the solvation effects were estimated via the SMD implicit solvation model using water as a solvent113. The DFT calculations were performed using Gaussian 16114.
The 1e− standard reduction potentials of the organic species comprising our present molecular database were computed according to the Nernst equation:
where \({G}^{o}\left({\rm{A}}\right)\) and \({G}^{o}\left({{\rm{A}}\cdot }^{-}\right)\) denote the aqueous phase Gibbs free energies of the original and the reduced forms of organic electron acceptor A, F is the Faraday constant, and \({\varDelta E}^{{ref}}\) is the absolute potential of the reference standard hydrogen electrode (SHE) (4.281 V)115. The Gibbs free energies were computed via a composite manner:
where \({E}^{{DFT}}\left({\rm{A}}/{{\rm{A}}\cdot }^{-}\right)\) refers to the electronic energy term with the inclusion of solvation free energy computed at the DFT level, and \({\Delta G}^{{xTB}}\left({\rm{A}}/{{\rm{A}}\cdot }^{-}\right)\) involves all finite temperature contributions estimated with the GFN2-xTB method. Test calculations on a database of experimental half-peak potentials of a variety of organic potentials, as well as benchmark calculations with respect to full DFT computations demonstrate that this composite protocol is suitable to predict redox potentials for a large set of molecules with an accuracy close to that obtained at full DFT level83.
Data availability
The PredPotS web application with tutorial is publicly available at: https://predpots.ttk.hu/, where users can access the training datasets and XYZ coordinate files directly: https://predpots.ttk.hu/datasets. All datasets have also been uploaded alongside the manuscript on the journal’s online platform.
Code availability
All associated resources, including the Python code used for model training, datasets, and XYZ files, are also publicly available on GitHub at https://github.com/AndreaH28/PredPotS.
References
Gür, T. M. Review of electrical energy storage technologies, materials and systems: challenges and prospects for large-scale grid storage. Energy Environ., Sci. 11, 2696–2767 (2018).
Dehghani-Sanij, A. R., Tharumalingam, E., Dusseault, M. B. & Fraser, R. Study of energy storage systems and environmental challenges of batteries. Renew. Sustain. Energy Rev. 104, 192–208 (2019).
Sánchez-Díez, E. et al. Redox flow batteries: Status and perspective towards sustainable stationary energy storage. J. Power Sources 481, 228804 (2021).
Li, Z., Jiang, T., Ali, M., Wu, C. & Chen, W. Recent progress in organic species for redox flow batteries. Energy Storage Mater. 50, 105–138 (2022).
Ragupathy, P., Bhat, S. D. & Kalaiselvi, N. Electrochemical energy storage and conversion: an overview. WIREs Energy Environ. 12, e464 (2022).
Shaw, M. H., Twilton, J. & MacMillan, D. W. C. Photoredox catalysis in organic chemistry. J. Org. Chem. 81, 6898–6926 (2016).
Romero, N. A. & Nicewicz, D. A. Organic photoredox catalysis. Chem. Rev. 116, 10075–10166 (2016).
Bell, J. D. & Murphy, J. A. Recent advances in visible light-activated radical coupling reactions triggered by (i) ruthenium, (ii) iridium and (iii) organic photoredox agents. Chem. Soc. Rev. 50, 9540–9685 (2021).
Cheung, K. P. S., Sarkar, S. & Gevorgyan, V. Visible light-induced transition metal catalysis. Chem. Rev. 122, 1543–1625 (2021).
Baranwal, J., Barse, B., Gatto, G., Broncova, G. & Kumar, A. Electrochemical sensors and their applications: a review. Chemosensors 10, 363 (2022).
Gibi, C., Liu, C.-H., Anandan, S. & Wu, J. J. Recent advances on electrochemical sensors for detection of contaminants of emerging concern (CECs). Molecules 28, 7916 (2023).
Singh, R., Gupta, R., Bansal, D., Bhateria, R. & Sharma, M. A review on recent trends and future developments in electrochemical sensing. ACS Omega 9, 7336–7356 (2024).
Ward, J. S. & Vezzoli, A. Key advances in electrochemically-addressable single-molecule electronics. Curr. Opin. Electrochem. 35, 101083 (2022).
Li, X., Ge, W., Guo, S., Bai, J. & Hong, W. Characterization and application of supramolecular junctions. Angew. Chem. Int. Ed. 62, e2022168 (2023).
Li, T., Bandari, V. K. & Schmid, O. G. Molecular electronics: creating and bridging molecular junctions and promoting its commercialization. Adv. Mater. 35, 2209088 (2023).
Bard, A. J., Parsons, R. & Jordan, J. Standard Potentials in Aqueous Solutions (Marcel Dekker, 1985).
Wardman, P. Reduction potentials of one-electron couples involving free radicals in aqueous solution. J. Phys. Chem. Ref. Data 18, 1637–1755 (1989).
Bratsch, S. G. Standard electrode potentials and temperature coefficients in water at 298.15 K. J. Phys. Chem. Ref. Data 18, 1–21 (1989).
Atkins, P. Inorganic Chemistry 5th edn. (W. H. Freeman, 2010)
Vanýsek, P. in CRC Handbook of Chemistry and Physics 92nd edn (ed. W. M. Haynes) 5-80-5-89 (CRC Press, 2011).
Fornari, R. P. & de Silva, P. Molecular modeling of organic redox-active battery materials. WIREs Comput. Mol. Sci. 11, e1495 (2020).
Yang, X., Zhuang, Y., Zhu, J., Le, J. & Cheng, J. Recent progress on multiscale modeling of electrochemistry. WIREs Comput. Mol. Sci. 12, e1559 (2021).
Wang, W. & Sprenkle, V. Redox flow batteries go organic. Nat. Chem. 8, 204–206 (2016).
Winsberg, J., Hagemann, T., Janoschka, T., Hager, M. D. & Schubert, U. S. Redox-flow batteries: from metals to organic redox-active. Mater. Angew. Chem. Int. Ed. 56, 686–711 (2016).
Leung, P. et al. Recent developments in organic redox flow batteries: a critical review. J. Power Sources 360, 243–283 (2017).
Zhang, C. et al. Progress and prospects of next-generation redox flow batteries. Energy Storage Mater. 15, 324–350 (2018).
Ding, Y., Zhang, C., Zhang, L., Zhou, Y. & Yu, G. Molecular engineering of organic electroactive materials for redox flow batteries. Chem. Soc. Rev. 47, 69–103 (2018).
Chen, R. Toward high-voltage, energy-dense, and durable aqueous organic redox flow batteries: role of the supporting electrolytes. ChemElectroChem 6, 603–612 (2018).
Cao, J., Tian, J., Xu, J. & Wang, Y. Organic flow batteries: recent progress and perspectives. Energy Fuels 34, 13384–13411 (2020).
Fang, X. et al. Multielectron organic redoxmers for energy-dense redox flow batteries. ACS Mater. Lett. 4, 277–306 (2022).
Cao, Y. & Aspuru-Guzik, A. Accelerating discovery in organic redox flow batteries. Nat. Comput. Sci. 4, 89–91 (2024).
Isegawa, M., Neese, F. & Pantazis, D. A. Ionization Energies and aqueous redox potentials of organic molecules: comparison of DFT, correlated ab initio theory and pair natural orbital approaches. J. Chem. Theory Comput. 12, 2272–2284 (2016).
Tagade, P. M. et al. Empirical relationship between chemical structure and redox properties: mathematical expressions connecting structural features to energies of frontier orbitals and redox potentials for organic molecules. J. Phys. Chem. 122, 11322–11333 (2018).
Sterling, C. M. & Bjornsson, R. Multistep explicit solvation protocol for calculation of redox potentials. J. Chem. Theory Comput. 15, 52–67 (2018).
Zhang, Q., Khetan, A. & Er, S. Comparison of computational chemistry methods for the discovery of quinone-based electroactive compounds for energy storage. Sci. Rep. 10, 22149 (2020).
Neugebauer, H., Bohle, F., Bursch, M., Hansen, A. & Grimme, S. Benchmark study of electrochemical redox potentials calculated with semiempirical and DFT methods. J. Phys. Chem. A 124, 7166–7176 (2020).
McNeill, A. R., Bodman, S. E., Burney, A. M., Hughes, C. D. & Crittenden, D. L. Experimental validation of a computational screening approach to predict redox potentials for a diverse variety of redox-active organic molecules. J. Phys. Chem. C 124, 24105–24114 (2020).
Hruska, E., Gale, A. & Liu, F. Bridging the experiment-calculation divide: machine learning corrections to redox potential calculations in implicit and explicit solvent models. J. Chem. Theory Comput. 18, 1096–1108 (2022).
Tomaník, L., Rulíšek, L. & Slavíček, P. Redox potentials with COSMO-RS: systematic benchmarking with different Databases. J. Chem. Theory Comput. 19, 1014–1022 (2023).
Achazi, A. J. et al. Development of a multi-step screening procedure for redox active molecules in organic radical polymer anodes and as redox flow anolytes. J. Comput. Chem. 45, 1112–1129 (2024).
Renningholtz, T., Lim, E. R. X., James, M. J. & Trujillo, C. Computational methods for investigating organic radical species. Org. Biomol. Chem. 22, 6166–6173 (2024).
Wang, F., Ma, Z. & Cheng, J. Accelerating computation of acidity constants and redox potentials for aqueous organic redox flow batteries by machine learning potential-based molecular dynamics. J. Am. Chem. Soc. 146, 14566–14575 (2024).
Jinnouchi, R., Karsai, F. & Kresse, G. Correction: Absolute standard hydrogen electrode potential and redox potentials of atoms and molecules: machine learning aided first principles calculations. Chem. Sci. 16, 10061–10062 (2025).
Huynh, M. T., Anson, C. W., Cavell, A. C., Stahl, S. S. & Hammes-Schiffer, S. Quinone 1 e– and 2 e–/2 H+ reduction potentials: identification and analysis of deviations from systematic scaling relationships. J. Am. Chem. Soc. 138, 15903–15910 (2016).
Han, Y.-K. & Jin, C.-S. Computational screening of electroactive indolequinone derivatives as high-performance active materials for aqueous redox flow batteries. Curr. Appl. Phys. 18, 1507–1512 (2018).
Fornari, R. P., Mesta, M., Hjelm, J., Vegge, T. & de Silva, P. Molecular engineering strategies for symmetric aqueous organic redox flow batteries. ACS Mater. Lett. 2, 239–246 (2020).
Schwan, S., Schröder, D., Wegner, H. A., Janek, J. & Mollenhauer, D. Substituent pattern effects on the redox potentials of quinone-based active materials for aqueous redox flow batteries. ChemSusChem 13, 5480–5488 (2020).
de la Cruz, C. et al. New insights into phenazine-based organic redox flow batteries by using high-throughput DFT modelling. Sustain. Energy Fuels 4, 5513–5521 (2020).
Zhang, Q., Khetan, A., Sorkun, E. & Er, S. Discovery of aza-aromatic anolytes for aqueous redox flow batteries via high-throughput screening. J. Mater. Chem. A 10, 22214–22227 (2022).
Hachmann, J. et al. The Harvard Clean Energy Project: large-scale computational screening and design of organic photovoltaics on the world community grid. J. Phys. Chem. Lett. 2, 2241–2251 (2011).
Pyzer-Knapp, E. O., Suh, C., Gómez-Bombarelli, R., Aguilera-Iparraguirre, J. & Aspuru-Guzik, A. What Is high-throughput virtual screening? A perspective from organic materials discovery. Annu. Rev. Mater. Res. 45, 195–216 (2015).
Cheng, L. et al. Accelerating electrolyte discovery for energy storage with high-throughput screening. J. Phys. Chem. Lett. 6, 283–291 (2015).
Er, S., Suh, C., Marshak, M. P. & Aspuru-Guzik, A. Computational design of molecules for an all-quinone redox flow battery. Chem. Sci. 6, 885–893 (2015).
Flores, S. D. P., Martin-Noble, G. C., Phillips, R. L. & Schrier, J. Bio-inspired electroactive organic molecules for aqueous redox flow batteries. 1. Thiophenoquinones. J. Phys. Chem. C. 119, 21800–21809 (2015).
Pelzer, K. M., Cheng, L. & Curtiss, L. A. Effects of functional groups in redox-active organic molecules: a high-throughput screening approach. J. Phys. Chem. C. 121, 237–245 (2017).
Tabor, D. P. et al. Mapping the frontiers of quinone stability in aqueous media: implications for organic aqueous redox flow batteries. J. Mater. Chem. A 7, 12833–12841 (2019).
Kristensen, S. B., van Mourik, T., Pedersen, T. B., Sørensen, J. L. & Muff, J. Simulation of electrochemical properties of naturally occurring quinones. Sci. Rep. 10, 13571 (2020).
Zhang, Q. et al. Data-driven discovery of small electroactive molecules for energy storage in aqueous redox flow batteries. Energy Storage Mater. 47, 167–177 (2022).
Sorkun, E., Zhang, Q., Khetan, A., Sorkun, M. C. & Er, S. RedDB, a computational database of electroactive molecules for aqueous redox flow batteries. Sci. Data 9, 718 (2022).
Khetan, A. High-throughput virtual screening of quinones for aqueous redox flow batteries: status and perspectives. Batteries 9, 24 (2022).
Keith, J. A. et al. Combining machine learning and computational chemistry for predictive insights into chemical systems. Chem. Rev. 121, 9816–9872 (2021).
Meuwly, M. Machine learning for chemical reactions. Chem. Rev. 121, 10218–10239 (2021).
Li, T., Zhang, C. & Li, X. Machine learning for flow batteries: opportunities and challenges. Chem. Sci. 13, 4740–4752 (2022).
Wei, Z., He, Q. & Zhao, Y. Machine learning for battery research. J. Power Sources 549, 232125 (2022).
Fedorov, R. & Gryn’ova, G. Unlocking the potential: predicting redox behavior of organic molecules, from linear fits to neural networks. J. Chem. Theory Comput. 19, 4796–4814 (2023).
Tang, L., Leung, P., Xu, Q. & Flox, C. Machine learning orchestrating the materials discovery and performance optimization of redox flow battery. ChemElectroChem 11, e202400024 (2024).
Janet, J. P., Ramesh, S., Duan, C. & Kulik, H. J. Accurate multiobjective design in a space of millions of transition metal complexes with neural-network-driven efficient global optimization. ACS Cent. Sci. 6, 513–524 (2020).
Barker, J., Berg, L., Hamaekers, J. & Maass, A. Rapid Prescreening of organic compounds for redox flow batteries: a graph convolutional network for predicting reaction enthalpies from SMILES. Batteries Supercaps 4, 1482–1490 (2021).
Agarwal, G., Doan, H. A., Robertson, L. A., Zhang, L. & Assary, R. S. Discovery of energy storage molecular materials using quantum chemistry-guided multiobjective bayesian optimization. Chem. Mater. 33, 8133–8144 (2021).
Ghule, S., Dash, S. R., Bagchi, S., Joshi, K. & Vanka, K. Predicting the redox potentials of phenazine derivatives using DFT-assisted machine learning. ACS Omega 7, 11742–11755 (2022).
Carvalho, R. P., Marchiori, C. F. N., Brandell, D. & Araujo, C. M. Artificial intelligence driven in-silico discovery of novel organic lithium-ion battery cathodes. Energy Storage Mater. 44, 313–325 (2022).
Sowndarya, S. V. S. et al. Multi-objective goal-directed optimization of de novo stable organic radicals for aqueous redox flow batteries. Nat. Mach. Intell. 4, 720–730 (2022).
Duke, R., Bhat, V., Sornberger, P., Odom, S. A. & Risko, C. Towards a comprehensive data infrastructure for redox-active organic molecules targeting non-aqueous redox flow batteries. Digital Discov. 2, 1152–1162 (2023).
Hashemi, A. et al. Density functional theory and machine learning for electrochemical square-scheme prediction: an application to quinone-type molecules relevant to redox flow batteries. Digital Discov. 2, 1565–1576 (2023).
Sorkun, M. C., Ghassemi, E. N., Yatbaz, C., Koelman, J. M. V. A. & Er, S. RedPred, a machine learning model for the prediction of redox reaction energies of the aqueous organic electrolytes. Artif. Intell. Chem. 2, 100064 (2024).
Jia, L. et al. Predicting redox potentials by graph-based machine learning methods. J. Comput. Chem. 45, 2383–2396 (2024).
Du, J. et al. Data-driven discovery of carbonyl organic electrode molecules: machine learning and experiment. J. Mater. Chem. A 12, 12034–12042 (2024).
Gao, P., Kochan, D., Tang, Y.-H., Yang, X. & Saldanha, E. G. Machine learning for the redox potential prediction of molecules in organic redox flow battery. J. Power Sources 629, 236035 (2025).
Si, Z. et al. Data-based prediction of redox potentials via introducing chemical features into the transformer architecture. J. Chem. Inf. Model. 64, 8453–8463 (2024).
Weininger, D. SMILES, a chemical language and information system. 1. Introduction methodology and encoding rules. J. Chem. Inf. Comput. Sci. 28, 31–36 (1988).
Schwaller, P. et al. Molecular transformer: a model for uncertainty-calibrated chemical reaction prediction. ACS Cent. Sci. 5, 1572–1583 (2019).
Ross, J. et al. Large-scale chemical language representations capture molecular structure and properties. Nat. Mach. Intell. 4, 1256–1264 (2022).
Hamza, A. et al. N-alkylated pyridoxal derivatives as negative electrolyte materials for aqueous organic flow batteries: computational screening. Chem. Eur. J. 29, e202300996 (2023).
Nechaev, A. A. et al. Exploration of vitamin B6-based redox-active pyridinium salts towards the application in aqueous organic flow batteries. Chem. Eur. J. 30, e202400828 (2024).
Schmidhuber, J. Deep learning in neural networks: an overview. Neural Netw. 61, 85–117 (2015).
Bento, A. P. et al. The ChEMBL bioactivity database: an update. Nucl. Acids Res. 42, D1083–D1090 (2013).
Das, M., Ghosh, A. & Sunoj, R. B. Advances in machine learning with chemical language models in molecular property and reaction outcome predictions. J. Comp. Chem. 45, 1160–1176 (2024).
Li, H. et al. A knowledge-guided pre-training framework for improving molecular representation learning. Nat. Com. 14, 7568 (2023).
Mayr, A. et al. Large-scale comparison of machine learning methods for drug target prediction on ChEMBL. Chem. Sci. 9, 5441–5451 (2018).
Hoque, A., Surve, M., Kalyanakrishnan, S. & Sunoj, R. B. Reinforcement learning for improving chemical reaction performance. J. Am. Chem. Soc. 146, 28250–28267 (2024).
Bajusz, D., Rácz, A. & Héberger, K. Why is Tanimoto index an appropriate choice for fingerprint-based similarity calculations?. J. Cheminform 7, 20 (2015).
Snarey, M., Terrett, N. K., Willett, P. & Wilton, D. J. Comparison of algorithms for dissimilarity-based compound selection. J. Mol. Graph. Mod. 15, 372–385 (1997).
Schrödinger Release 2017-1: Canvas, Schrödinger, LLC, New York, NY, (2017).
Schrödinger Release 2017-1: Epik, Schrödinger, LLC, New York, NY, (2017).
Schneider, N., Sayle, R. A. & Landrum, G. A. Get your atoms in order—an open-source implementation of a novel and robust molecular canonicalization algorithm. J. Chem. Inf. Model. 55, 2111–2120 (2015).
Ramsundar, B. et al. Deep Learning for the Life Sciences: Applying Deep Learning to Genomics, Microscopy, Drug Discovery, and More (O’Reilly Media, Inc. 2019).
Sevov, C. S., Hendriks, K. H. & Sanford, M. S. Low-potential pyridinium anolyte for aqueous redox flow batteries. J. Phys. Chem. C. 121, 24376–24380 (2017).
Striepe, L. & Baumgartner, T. Viologens and their application as functional materials. Chem. Eur. J. 23, 16924–16940 (2017).
Huang, J. et al. Spatially constrained organic diquat anolyte for stable aqueous flow batteries. ACS Energy Lett. 3, 2533–2538 (2018).
Hu, S. et al. Phenylene-bridged bispyridinium with high capacity and stability for aqueous flow batteries. Adv. Mater. 33, 2005839 (2021).
Burešová, Z. et al. Redox property tuning in bipyridinium salts. Front. Chem. 8, 631477 (2021).
Rak, K. et al. Electrochemical investigation of structurally varied azinium scaffolds. Org. Biomol. Chem. 19, 8830–8839 (2021).
Tang, G. et al. Designing robust two-electron storage extended bipyridinium anolytes for ph-neutral aqueous organic redox flow batteries. J. Am. Chem. Soc. 2, 1214–1222 (2022).
Liu, X. et al. Thienoviologen anolytes for aqueous organic redox flow batteries with simultaneously enhanced capacity utilization and capacity retention. J. Mater. Chem. A 10, 9830–9836 (2022).
Rubio-Presa, R., Lubián, L., Borlaf, M., Ventosa, E. & Sanz, R. Addressing practical use of viologen-derivatives in redox flow batteries through molecular engineering. ACS Mater. Lett. 5, 798–802 (2023).
Luo, J., Hu, B., Debruler, C. & Liu, T. L. A π-conjugation extended viologen as a two-electron storage anolyte for total organic aqueous redox flow batteries. Angew. Chem. Int. Ed. 130, 237–241 (2017).
For more information on the EU funded CompBat project, see: https://compbat.aalto.fi.
Grimme, S., Bannwarth, C. & Shushkov, P. A Robust and accurate tight-binding quantum chemical method for structures, vibrational frequencies, and noncovalent interactions of large molecular systems parametrized for all spd-block elements (Z = 1–86). J. Chem. Theory Comput. 13, 1989–2009 (2017).
Bannwarth, C., Ehlert, S. & Grimme, S. GFN2-xTB—an accurate and broadly parametrized self-consistent tight-binding quantum chemical method with multipole electrostatics and density-dependent dispersion contributions. J. Chem. Theory Comput. 15, 1652–1671 (2019).
User Guide to Semiempirical Tight Binding, https://xtb-docs.readthedocs.io/en/latest/contents.html
Pracht, P., Bohle, F. & Grimme, S. Automated exploration of the low-energy chemical space with fast quantum chemical methods. Phys. Chem. Chem. Phys. 22, 7169–7192 (2020).
Zhao, Y. & Truhlar, D. G. The M06 suite of density functionals for main group thermochemistry, thermochemical kinetics, noncovalent interactions, excited states, and transition elements: two new functionals and systematic testing of four M06-class functionals and 12 other functionals. Theor. Chem. Acc. 120, 215–241 (2007).
Marenich, A. V., Cramer, C. J. & Truhlar, D. G. Universal solvation model based on solute electron density and on a continuum model of the solvent defined by the bulk dielectric constant and atomic surface tensions. J. Phys. Chem. B 113, 6378–6396 (2009).
Frisch, M. J. et al. Gaussian 16, Revision C.01 (Gaussian, Inc., 2016).
Isse, A. A. & Gennaro, A. Absolute potential of the standard hydrogen electrode and the problem of interconversion of potentials in different solvents. J. Phys. Chem. B 114, 7894–7899 (2010).
Acknowledgements
The present project has been funded by the European Union’s Horizon 2020 research and innovation programme under grant agreement No 875565 (CompBat project).
Author information
Authors and Affiliations
Contributions
F.B.N., A.H., Á.M., and I.P. conceptualized the project. F.B.N., A.H., and Á.M. created the database and molecular library. A.H. carried out quantum chemical calculations. F.B.N., B.T., E.A.-A., and A.H. performed machine learning analysis, designed and implemented the web tool. L.S. supervised the machine learning studies. I.P. wrote the original draft manuscript and provided supervision. All authors discussed the results and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Németh, F.B., Hamza, A., Tugyi, B. et al. PredPotS: web tool for predicting one-electron standard reduction potentials for organic molecules in aqueous phase. npj Comput Mater 12, 23 (2026). https://doi.org/10.1038/s41524-025-01890-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41524-025-01890-1