Abstract
Probiotics have been widely used as an adjuvant therapy to treat or prevent various diseases. However, therapeutic effect is usually contingent on the ability of the probiotic cells to adhere to and colonize the gastrointestinal tract. Pediococcus pentosaceus Li05 has previously been shown anti-inflammatory effect, however, its mucoadhesive ability limited further application. In this study, MUC2 antibody was used as an adhesin and conjugated to the surface of Li05 (M-Li05) and our results demonstrated that the adhesion and colonization resistance were significantly improved in vitro. Moreover, competitive adhesion experiments in mice demonstrated that Li05 outcompeted Citrobacter rodentium; an effect even more pronounced with M-Li05. The study also showed that M-Li05 significantly reduced colonic histological injury, alleviated inflammation and decreased pathogens colonization in the C. rodentium-infected and DSS-induced colitis mouse model. In summary, the results demonstrated that M-Li05 showed promise for future application as a therapeutic strategy for colitis.
Similar content being viewed by others
Introduction
Inflammatory bowel disease (IBD) is a group of diseases including ulcerative colitis (UC) and Crohn’s disease (CD), characterized by chronic inflammation of the gastrointestinal (GI) tract1. Many factors including diet, genetics, host immunity, and intestinal microbiome, are involved in the development of IBD2,3,4,5. The composition of the gut microbiota of patients with IBD differs markedly from that of healthy individuals6,7. Many studies have shown that IBD patients demonstrated a reduced diversity of their gut microbiota, with the lack of diversity among Firmicutes being especially notable1.
Probiotics are defined as “live microorganisms that, when given in sufficient amounts, provide health benefits to the host”8. Many reports have demonstrated the anti-inflammation efficacy of various probiotic bacteria for IBD in clinical trials and animal experiments9,10,11,12. The probiotic strain Pediococcus pentosaceus Li05 can strengthen the intestinal epithelial barrier function, modulate host immunity and gut microbiota in Clostridiodies difficile infected colitis, dextrate sulfate sodium (DSS)-induced colitis and CCl4-induced cirrhosis13,14,15. However, the application of Li05 is limited due to the strain’s susceptibility to harsh conditions exacerbated by chemical, physical, and biological factors during its transmission through the gastrointestinal tract, and poor adhesion to the intestinal mucosa15,16.
We have previously successfully enhanced the viability of Li05 in harsh conditions through encapsulating Li05 in microcapsules15, but its adhesion in the colon remains to be improved. Successful colonization in the intestine is a necessary criterion for evaluating whether probiotics can have a long-term effect17. Mucosal adhesion is considered a critical step in probiotic colonization which is achieved by stable binding of the probiotic cells to the mucus through specific interactions between adhesins and complementary receptors18, like mucin 2 (MUC2), a predominant component of the intestinal mucosa19. One way to enhance the adhesive ability of the probiotic to the mucosa is by directly increasing the amount of adhesins on the bacterial surface. Synthetic adhesins in the form of antibodies (anti-MUC2) (aMUC2) can be conjugated to the probiotic cell surface to promote the binding to MUC2 and facilitate adhesion17. Besides, this surface modification process is rapid and mild to probiotics, avoiding long-term exposure in aerobic environment.
Our work intended to elucidate the effect of aMUC2 surface modification on the mucoadhesive properties. We also used Citrobacter rodentium infected and DSS-induced acute colitis mouse models to evaluate the enhanced function of Li05 after aMUC2 modification in treating IBD. As a mouse-specific pathogen, C. rodentium shares 67% genes with both human pathogenic enteropathogenic Escherichia coli (EPEC) and enterohaemorrhagic E. coli (EHEC)20, and can simulate infectious diarrhoea and IBD21. C. rodentium infection in mice is accompanied with transmissible colonic crypt hyperplasia (CCH), intestinal barrier destruction, intestinal inflammation and gut dysbiosis22. The gut microbiota has a major part in host interactions with C. rodentium infection23,24. The administration of probiotics can facilitate removal of C. rodentium and relief of inflammation by modulating the composition of the gut microbiota25,26. DSS is wildly used to induce acute colitis in mice by causing damage to the intestinal mucosa and activating the inflammatory response. Many researches have demonstrated that the gut microbiota in DSS-induced colitis mice is severely imbalanced, accompanied by the proliferation of pathogens, including Escherichia coli13. We hypothesized that treatment with M-Li05 would enhance the ability of the probiotic to adhere in the gut and outcompete pathogens, thereby ameliorate colitis.
Results
Surface modification of Li05
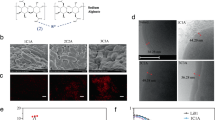
Biotin is a vitamin that can bind with high affinity to avidin and streptavidin. Because of its light molecular weight (244 Da) and stable chemical properties, biotin can be conjugated to many proteins without changing their biological activities27. In this study, biotin was conjugated to the primary amines on the surface of Li05 by N-hydroxysuccinimide ester (NHS). The biotinylation of Li05 was then evaluated through binding with fluorescent streptavidin. Confocal fluorescence images (Fig. 1A) showed that modified Li05 cells were surrounded by a ring of red fluorescence. In contrast, non-biotinylated bacteria showed little signal under red fluorescence. To test the stability of the modification in GI tract, the modified Li05 was incubated with simulated gastric fluids and simulated intestinal fluids, and fluorescence showed that there was no obvious shedding of the fluorescent streptavidin from the bacterial surface when compared to untreated Li05 (Fig. 1B).
A Confocal fluorescence images of unmodified and biotinylated Li05 after incubation with Alexa Fluor Streptavidin Conjugate. B Scanning electron microscopy images of Li05 and M-Li05. C Confocal fluorescence images of streptavidin-conjugated Li05 after 30 or 60 min of exposure to stimulated gastric fluid or stimulated intestinal fluid. D Colonies of Li05 and M-Li05 after adhesion on Caco-2/HT29-MTX cells, and each adhesion rate. E Imaging of bioluminescent C. rodentium in 12- well plate. F Imaging of bioluminescent C. rodentium in MAC agar medium. G C. rodentium adhesion rate and bioluminescence (p/s/cm2/sr) in each group. H Schematic diagram of the colonization study in germ-free rats, and CFUs of P. pentosaceus in feces at indicated timepoints for unmodified Li05 and M-Li05 groups. Data are presented as mean ± SEM, n = 3, *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
Streptavidin can be conjugated with aMUC2 by targeting primary amine groups on the IgG antibodies. By the strong affinity of streptavidin and biotin, aMUC2 can be successfully conjugated on the bacterial surface and M-Li05 was prepared. The morphology of modified bacteria was characterized by Scanning electronic microscopy (SEM). The size of bacteria did not change after modification and no obvious morphological difference between unmodified Li05 and M-Li05 was observed (Fig. 1C). Results also showed that modification did not have any impact on the growth rate and viability of bacteria (Fig. S2D, E).
Bacterial attachment and colonization resistance assay
To determine the mucoadhesion effect of Li05 before and after surface modification, an in vitro Caco-2/HT29-MTX cells monolayer model was used. Surface modification of aMUC2 significantly increased the adhesion rate of Li05 (Fig. 1D) and the adhesion ratio of M-Li05 was more than twice the ratio of the unmodified Li05. Probiotics may resist the external invading pathogenic by preventing their colonization in GI tract. To assess the colonization resistance of Li05 and M-Li05 toward pathogens, we chose C. rodentium as a model pathogen and conducted competitive adhesion experiment in vitro. The results showed that colonization resistance of Li05 after aMUC2 modification against C. rodentium was significantly enhanced, as treatment with M-Li05 remarkably decreased the adhesion rate and bioluminescence radiance (Fig. 1E-G) of C. rodentium compared to the CR and Li05 groups.
Colonization of M-Li05 in GF rats
After a single gavage of Li05 and M-Li05 separately, the feces of the rats in different groups were collected at different time points to evaluate dynamics of Li05 in GIT (Fig. 1H). No P. pentosaceus or other bacteria were detected in the feces of rats in either group prior to gavage. Subsequently, around 1*103 CFUs / g P. pentosaceus were detected at 1 h and 4 h after oral gavage in the Li05 and M-Li05 groups. After 8 h, the fecal bacterial counts in the M-Li05 group greatly increased and reached the plateau stage (8*108 CFUs / g). While in the Li05 group, the fecal concentration of Li05 is only around 106 CFUs / g. Besides, rats treated with M-Li05 had significantly higher concentrations of P. pentosaceus in their feces at 24 h and 48 h after gavage when compared with the Li05 group. However, there was no difference in the concentration of P. pentosaceus in feces on the 3rd, 4th, 5th, 6th day after gavage (Fig. 1H).
Effect of M-Li05 on attenuating C. rodentium-induced colitis
Figure 2A showed the schematic diagram of evaluating the effect of M-Li05 treatment on C. rodentium-induced colitis. The weight of mice in C. rodentium infected groups had reduced significantly compared to the NC group on day 12 (Fig. 2B). Through macroscopic examination, C. rodentium-infected mice in the CR group had obviously shorter colon length than the NC and ML groups (Fig. 2C). Histologic examination of colon sections revealed the development of colonic inflammation and crypt hyperplasia in C. rodentium infected mice (Fig. 2D). ML group showed a significantly lower colonic damage score. Immunohistochemical results revealed the intensities of Ki67 in the ML group decreased obviously (Fig. 2E). Meanwhile, shorter crypt length was found in the L and ML groups when compared to the CR group (Fig. 2G). However, treatment of unmodified Li05 or biotin-streptavidin-aMUC2 did not prevent effectively colon shortening or reduce the pathological score compared to the CR group. In addition, we detected the expression of inflammatory cytokines and significantly lower levels of TNF-α, IFN-γ transcripts were found in the ML group compared to the CR and BM groups (Fig. 2H).
A Schematic diagram of the mouse experiment with a C. rodentium-induced colonic inflammation model system. B Body weight of mice recorded during experimental period. C Representative images of the colon tissues from mice and colon length from the five groups. D Representative images of H&E staining of the colon tissue. E Representative images of Ki67 immunohistochemistry-stained preparations of the colon tissue. F Representative transmission electron microscopy images of colon tissues. G Histopathology scores and quantification of crypt hyperplasia by measuring crypt length (μm) of mice. H, I Colonic TNF-α, IFN-γ, IL-17, ZO1, occludin and MUC2 expression were determined by qRT-PCR. Data are presented as mean ± SEM, n = 6, *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
The transmission electron microscopy (TEM) images showed fewer number of C. rodentium adhesion on the intestinal microvilli in the ML group compared to the CR and BM groups so that the microvilli were denser and more intact (Fig. 2F), while there were still lesions of colon tissues in the L group. The relative expression of tight junction protein 1 (ZO1) was decreased in the CR and BM groups compared to the NC group. Treatment with M-Li05 dramatically restored the mRNA levels of ZO1, occludin and MUC2 when compared to the CR group (Fig. 2I).
Colonization competition between Li05 and C. rodentium in gut
To evaluate whether M-Li05 had an advantage compared to unmodified Li05 on colonization resistance to C. rodentium in the murine gut, levels of C. rodentium in the colon and feces were monitored and quantified (Fig. 3). It was found that the quantities of C. rodentium were similar amongst mice in the CR, BM, L and ML groups during the first 10 days post infection (p.i.). However, at day 12 and 14, fecal C. rodentium level was obviously reduced in the ML group compared to the CR and BM groups. Furthermore, the colonic number of C. rodentium was lower in the ML group compared to the CR and BM groups (Fig. 3C). Although lower level of C. rodentium was also observed in mice treated with unmodified Li05 compared to the CR group, no significant difference was found. 2D bioluminescent imaging result was consistent with those shown in Fig. 3B, showing that decreased level of C. rodentium bioluminescence was observed in the ML group in comparison with the CR, BM and L groups at day 12 p.i.
A In vivo optical imaging of bioluminescent C. rodentium infection in mice in the CR, BM, L, and ML groups at day 0, 6, 12, and 14 p.i. B Dynamics of C. rodentium bioluminescence (p/s/cm2/sr) in each experimental group and fecal C. rodentium concentration from fecal samples collected at each time point. C Quantification of C. rodentium concentration in feces and colon at day 14. Data are presented as mean ± SEM, n = 5–6, * Compared to CR, # compared to BM, ^ compared to L group. *P < 0.05, **P < 0.01 and ***P < 0.001.
Effects of M-Li05 on gut microbiota composition and metabolism in C. rodentium-induced colitis
The number of P. pentosaceus excretion in the feces was quantified, and the result showed that the abundance of P. pentosaceus in the ML group was remarkably higher than the L group at day 7, 14 and 21 (Fig. 4A).
A Quantification of P. pentosaceus in feces at day 0, 7, and 14. B PCOA of the gut microbiota composition of mice in the five groups. C, D Relative abundances of the most abundant taxa at phylum level. E Relative abundances of the most abundant taxa at genus level. F Relative abundances of Citrobacter, Lactobacillus, Turicibacter and Akkermansia. G SCFAs concentrations in the fecal samples in the five groups. Data are presented as mean ± SEM, n = 6, *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
16S rRNA gene sequencing was used to explore the changes of the gut microbiota composition among these five groups. Although Chao1 and Shannon indices of the ML group had a downward trend, both indices showed no significant difference in mice among these five groups (Fig. S3A). The β-diversity was assessed by analysis of similarities (Adonis) and PCoA. PC1 and PC2 axes of PCoA plot exhibited 39.17% and 29.16% of the variance respectively. (Fig. 4B). Adonis result demonstrated significant differences between mice in the NC and CR, ML and CR, ML and L, and ML and NC groups. At the phylum level, the relative abundance of Firmicutes was elevated whereas Bacteroidota was depressed in the ML group compared to the NC group (Fig. 4C, D). At the family level, the relative abundance of Bacteroidaceae was elevated in the L group compared to the CR, L and ML groups. The relative abundance of Prevotellaceae was decreased among mice in the ML group compared to the NC and L groups (Fig. S3B). At the genus level (Fig. 4E, F), the relative abundance of Lactobacillus was dramatically higher in the ML group compared to the NC, CR and L groups. The relative abundance of Turicibacter and Citrobacter (P = 0.053) was, however, lower among mice in the ML group compared to mice in the CR group.
The concentrations of SCFAs in the feces, including acetic acid, propionic acid, butyric acid, isobutyric acid and valeric acid were measured (Fig. 4G). Mice treated with unmodified Li05 or M-Li05 showed elevated levels of all these SCFAs compared to the mice in the NC group. Amongst five groups, the ML group exhibited highest levels of SCFAs with doubled concentrations of acetic acid, propionic acid and butyric acid and tripled concentrations of isobutyric acid and valeric acid compared to the NC and CR groups.
Effect of M-Li05 on attenuating DSS-induced acute colitis
The schematic diagram illustrating the treatment of M-Li05 on DSS-induced colitis is shown in Fig. 5A. To assess the therapeutic potential of M-Li05, we established a DSS-induced acute colitis model. Treatment with M-Li05 significantly alleviated DSS-induced weight loss, DAI score, and colon length shortening (Fig. 5B, C, D).
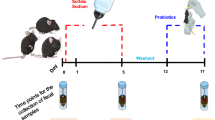
A Schematic diagram of the mouse experiment with a DSS-induced colonic inflammation model system. B Weight change (%) during the experimental period. C DAI score recorded every day after DSS drinking. D Representative images of colon tissues and colon length. E Representative images of ZO1, occludin and claudin1 immunofluorescence-stained preparations of the colon tissue. F Expression level of tight conjunction protein genes (ZO1, occludin and claudin1) and MUC2. G LBP concentration in serum in the five groups. Data are presented as mean ± SEM, n = 8, *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
Colonic immunofluorescence staining revealed enhanced distribution of tight junction proteins ZO-1, occludin, and claudin1 in the ML group compared to controls (Fig. 5E). Additionally, analysis of tight junction protein expression showed that M-Li05 treatment significantly restored the transcription levels of ZO-1, occludin, claudin1, and MUC2, while Li05 treatment alone only elevated ZO-1 and occludin expression (Fig. 5F). Inflammation-induced intestinal barrier disruption leads to bacterial translocation and elevates serum lipopolysaccharide-binding protein (LBP) level. Notably, M-Li05 administration significantly reduced serum LBP level, while neither Li05 nor biotin-streptavidin-aMUC2 treatment did not inhibit LBP elevation (Fig. 5G). These findings suggested that M-Li05 effectively ameliorated DSS-induced acute colitis.
Mucosal injury is a hallmark of acute colitis, characterized by crypt destruction, goblet cell loss, and inflammatory cell infiltration. In the P and BM groups, extensive mucosal damage was observed, while the ML group exhibited less severe histopathological changes and lower histopathology score (Fig. 6A). The mucosal damage triggered a robust inflammatory response, as evidenced by reduced infiltration of macrophages (F4/80-positive cells) and neutrophils (MPO-positive cells) in the ML group compared to the P and BM groups (Fig. 6B, C). Flow cytometry analysis showed that M-Li05 inhibited the increase in macrophages and M1 macrophages in the colonic lamina propria, whereas Li05 treatment only exhibited a slight reduction in macrophage number without reaching statistical significance (Fig. 6D, E). Serum levels of inflammatory cytokines IL-1α, IL-6, IL-12, P40, and MCP1 were significantly reduced in the ML group, while IL-10 level increased following M-Li05 treatment (Fig.6F). qPCR analysis of colon tissues further confirmed the downregulation of IL-1β, IL-6, and TNF-α after M-Li05 treatment (Fig. 6G).
A Representative images of H&E staining of the colon tissue and Histopathology scores. B, C Representative images of F4/80 and MPO immunohistochemistry-stained preparations of the colon tissue and respective positive area %. D, E Flow analysis of macrophages and M1 macrophages in colonic lamina propria. F Cytokines (IL-1α, IL-6, IL-10, IL-12, MCP1, MCP1β) level in the serum. G IL-1β, IL-6 and TNF-α expression level in the colon tissue. Data are presented as mean ± SEM, n = 8 for pathology scoring, cytokines and transcription level, n = 6 for immunohistochemistry scoring, *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
M-Li05 ameliorated gut microbiota composition disorder in DSS-induced acute colitis
To study the effect of Li05/M-Li05 on the gut microbiota structure in DSS-induced colitis, we conducted 16s rRNA sequencing analysis. Chao1 was used to assess α diversity and no significant differences were found between the five groups (Fig. 7A). PCoA analysis revealed a distinct alteration in gut microbiota composition in the DSS-induced colitis groups (Fig. 7B). At the phylum level, there was a significant decrease in relative abundance of Proteobacteria, Deferribacterota, and Actinobacteriota in the ML group when compared to the P group (Fig. 7D). At the genus level, an increase in relative abundance of Pediococcus, Bacteroides, Prevotellaceae, and Eubacterium was observed while pathogenic bacteria such as Escherichia_Shigella, Mucispirillum and Anaerostipes exhibited decreased abundance in the ML group compared to the P group (Fig. 7E). However, the treatment of Li05 or biotin-streptavidin-aMUC2 did not showed obvious decrease in the relative abundance of pathogenic bacteria.
A α-diversity of the faecal microbiota in the five groups. B PCoA plot among the five groups. C Relative abundance of P. pentosaceus in the fecal samples of L and ML groups. D, E Relative abundance in the most abundant taxa at phylum and genus level and the difference in P and ML groups. Data are presented as mean ± SEM, n = 8, *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
LEfSe analysis was used to further reveal prognostic microbial markers. As depicted in Fig. S4, based on a LDA score (log10) > 4.0, we observed a significant enrichment of Proteobacteria phylum, Enterobacteriaceae family and Escherichia_Shigella genus in the P group compared to the NC and ML groups. Conversely, family Lactobacillaceae and genus Pediococcus were found to be enriched in the ML group compared to the P and L groups.
Colonic transcriptome analysis of M-Li05 effect on DSS-induced colitis
To assess the impact of M-Li05 on DSS-induced colitis, we performed transcriptome analysis on colon tissues. A total of 6364 differentially expressed genes (DEGs) were identified between the P and NC groups (3371 up-regulated and 2993 down-regulated), and 4101 DEGs were identified between the ML and P groups (1972 up-regulated and 2129 down-regulated), with P < 0.05 and a fold change > 1.0 (Fig. S5A, B). Principal component analysis (PCA) showed distinct alterations in gene expression across the five groups (Fig. S5C).
KEGG enrichment analysis revealed that, compared to the NC group, the P group exhibited up-regulation of several key pathways, including cytokine–cytokine receptor interactions, NF-κB signaling, and TNF signaling. Notably, M-Li05 treatment significantly suppressed these pathways, with down-regulation of genes such as Ccl2, Ccl4, Cxcl2, IL-1α, IL-1β, Icam1, TNF, TLR2, and TLR5 (Fig. S5D-E). These findings suggested that M-Li05 exerted an anti-inflammatory effect by modulating cytokine and immune receptor signaling pathways.
Discussion
Adhesion and colonization in the intestine are prerequisites for probiotics to have long-term probiotic effects. Although Li05 has been shown to have promise as an adjuvant therapy in treating colitis and cirrhosis, the limited mucoadhesive effect of the probiotic limits its function. To enhance the mucoadhesive properties of Li05, we developed Li05 with aMUC2 modification. By conjugation with fluorescent streptavidin, we confirmed that biotin was successfully linked to the surface of Li05. Meantime, treatment of simulated gastric acid or simulated intestinal fluids did not affect the stability of the conjugation of biotin to Li05. Growth curves of biotinylated Li05, M-Li05 and unmodified Li05 further demonstrated that biotinylation and surface modification with aMUC2 did not affect the growth of Li05. SEM imaging of M-Li05 further showed no bacterial cell damage.
The enhanced adhesion ability of M-Li05 was verified in Caco-2/HT29-MTX cells, GF rats and C57BL mice. On Caco-2/HT29-MTX cells model, we found significantly higher amount of M-Li05 was attached to the monolayer than the unmodified Li05. In vivo colonization in GF rats experiment, M-Li05 could complete colonization faster than unmodified Li05. Meanwhile, rats treated with M-Li05 had significantly higher concentration of P. pentosaceus in their feces at 24 h and 48 h after gavage. However, due to the amount of anti-MUC2 on the bacteria surface decreased with bacteria proliferation, this improvement of the adhesion ability of M-Li05 would become insignificant in the later stage. Accordingly, the number of P. pentosaceus in the feces showed no difference after 3 days. Continuous administration of Li05/M-Li05 in C57BL/6 mice showed that the number of P. pentosaceus in the feces remarkably increased in the ML group compared to the L group in the early, mid-term, and end of the experiment. Taken together, these results demonstrated that modification of aMUC2 could enhance the adhesion of Li05, accelerate intestinal colonization and increase the abundance of Li05 in the intestinal tract. Colonization resistance against pathogen is an important indicator for evaluating the function of probiotics. Our in vitro colonization resistance test revealed co-incubation with M-Li05 could significantly decrease the attachment of C. rodentium to Caco-2/HT29-MTX cells. The modification of aMUC2 not only enhanced adhesion, but also enhanced colonization resistance to pathogenic bacteria.
To evaluate the function of M-Li05 in the treatment for colitis, C. rodentium infected colitis and DSS-induced acute colitis mouse model were established. C. rodentium infection is widely used to model IBD and colonic cancer21. Gut microbiota critically participated in the establishment and progress of C. rodentium infection23,28. Our in vivo study demonstrated the treatment of M-Li05 could ameliorate C. rodentium infection by clearing C. rodentium, regulating the gut microbiota and metabolism, restoring intestinal barrier and alleviating inflammation. However, these effects were not obvious when biotin-streptavidin-aMUC2 or Li05 were administrated singly. The C. rodentium infection cycle includes four phases: 1) the establishment phase (1–3 p.i.), 2) the expansion phase (4–8 p.i.), 3) the steady-state phase (8–12 p.i.), and 4) The clearance phase (from 12 p.i.)22. Fecal shedding of C. rodentium was monitored by fecal bacterial enumeration and 2D BLI. The shedding of C. rodentium showed distinct reduction at the steady-state and clearance phases (day 12 and 14) in the ML group, demonstrating that M-Li05 accelerated clearing C. rodentium by competitive exclusion. C. rodentium infection can trigger inflammation when C. rodentium breaks the epithelial barrier, whose lipopolysaccharide (LPS), the type III secretion system (T3SS), and peptidoglycan can be recognized as pathogen-associated molecular patterns (PAMPS) by host immune system21. Multiple cytokines (IL-1β, IL-6, TNF-α, IFN-γ, IL-17) are involved in the development of C. rodentium infections21. Consistent with previous findings25,29,30, colonic mRNA levels of TNF-α, IFN-γ, IL-17 were obviously increased in the CR group and the treatment of M-Li05 attenuated the expression of these inflammatory factors and neutrophil infiltration, but mice in the Li05 treatment group only showed a lower level with no significant difference. The C. rodentium pathogenic process can be recognized as attaching and effacing (A/E) lesions which is characterized by intimate contact between bacteria and the intestinal epithelium, effacement of the microvilli and actin accumulation31. The locus of enterocyte effacement (LEE) of C. rodentium can encode several effector proteins including T3SS which participate in disrupting intestinal tight junction integrity32. We observed typical A/E lesions through TEM. A large number of C. rodentium adhered to the surface of microvilli, causing extensive destruction of microvilli and damage to tight junction in the CR and BM groups. We evaluated intestinal barrier damage by detecting tight junction protein RNA levels and found that colonic mRNA levels of ZO1, occludin and MUC2 were increased in the ML group compared to the CR group. Therefore, the administration of M-Li05 attenuated colonic inflammation and restored the intestinal barrier damage caused by infection of C. rodentium.
Numerous studies have shown that gut microbial structure contribute to the individual intestinal susceptibility to C. rodentium infection23,33,34. Intestinal microbiota can fulfil an important colonization resistance to C. rodentium by strengthening intestinal barrier, competing for specific nutrients, and producing microbial metabolites to affect the expression of virulence genes22. Our results described a higher relative abundance of Lactobacillus in M-Li05-fed mice compared to those in the CR and NC groups, but the opposite was true for Citrobacter and Turicibacter. Lactobacillus is considered an important commensal which can enhance the formation of the mucus barrier and provide colonization resistance by limiting pathogen access to intestinal epithelial cells35,36,37. The abundance of Turicibacter appeared to be inversely correlated with intestinal butyrate levels which have an anti-inflammatory effect38. The Citrobacter were less abundant further implying an improved effectiveness of M-Li05 on C. rodentium clearance. Some studies have speculated that resistance to C. rodentium infection is linked to a decreased ratio of Firmicutes / Bacteroidota (F/B)33,39. In contrast, our results showed that the relative abundance of Firmicutes was increased whereas Bacteroidota was decreased in the ML group compared to the NC and L groups, indicating a higher F/B in the ML group. Correspondingly, abundances of Bacteroidaceae and Prevotellaceae, which are common families of the phylum Bacteroidetes, were also decreased among mice in the ML group compared to mice in the L group. Our results indicated that clearing of C. rodentium did not seem to rely on the higher abundance of Bacteroidetes. It can be speculated that M-Li05 may promote the proliferation of probiotics like Lactobacillus37 and Akkermansia25 to accelerate the clearence of C. rodentium.
The intestinal microbiota can activate intestinal immune cells by their production of the SCFAs. Although previous experiments have indicated that Li05 achieved anti-inflammatory effects through SCFAs13, our results showed that stimulation of production of SCFAs was quite limited when unmodified Li05 was used and only the levels of propionate and hexanoic acids were observed to increase. The administration of M-Li05 did, however, significantly increase the concentration of six different SCFAs. Acetate and propionate interact with G protein-coupled receptor 41 (GPR41) and GPR43 on the surface of enterocytes and stimulate the proliferation of TH1 cells and TH17 cells, which in turn respond to C. rodentium and accelerate C. rodentium clearance40. Butyric acid is another important anti-inflammatory metabolite produced by gut microbes41. Studies have demonstrated that butyrate could alter the metabolic activity of macrophages to improve oxidative phosphorylation and promote alternative macrophage activation42,43. Macrophage activity is related with TH17 and TH2 cells which response for pathogen clearance including C. rodentium42. Meanwhile, administration of appropriate concentration of propionate and butyrate can promote MUC2 expression by regulating the promoter44 which may explain why MUC2 expression was elevated in the ML group. However, some studies have found that butyrate could induce expression of the type III secretion system which created contiguous channels between the bacteria and host cell membranes, allowing the injection of virulence factors directly to host cells22,45. Taken together, the increase of intestinal SCFAs appeared to have a beneficial effect on C. rodentium clearance and relief of intestinal lesions.
In the DSS-induced colitis mouse model, treatment with M-Li05 resulted in significant improvement in clinical indicators (such as weight loss, DAI, and colon length), intestinal barrier integrity, colonic inflammation, and intestinal microbiota dysbiosis. DSS exerts a toxic effect on the colonic epithelium leading to extensive ulceration of the colon, which causes direct contact between intestinal epithelial cells and antigens, and triggering strong immune response46,47. Treatment with M-Li05 elevated the expression of ZO1, occludin, claudin1 and MUC2 and restored intestinal barrier, decreasing bacterial translocation and the level of serum LBP. Intestinal microbiota plays a crucial role in initiating and maintaining intestinal inflammatory response in patients with IBD by providing antigens or other stimulating factors that trigger immune cell activation48. Studies revealed that harmful bacteria presented overgrowth, including Fusobacterium49 and Enterobacteriaceae50, while probiotics like Faecalibacterium prausnitzii51 were inhibited. Escherichia coli, known for its strong mucosal erosivity52, can trigger a potent immune response by secreting effector proteins like EspG, Map, and Tir53, which increase intestinal permeability54. Escherichia coli, Escherichia_Shigella, and C. rodentium, all belonging to the Enterobacteriaceae family, share significant genetic similarities21. Surface modification of aMUC2 was shown to enhance the adhesion of Li05, improving its resistance to C. rodentium. Correspondingly, in the ML group, we observed a reduced relative abundance of Proteobacteria, Enterobacteriaceae, Escherichia_Shigella, Mucispirillum, and Anaerostipes.
Exposure to LPS activates various immune cells, including neutrophils55 and macrophages56, leading to the production of cytokines such as TNF-α, IL-1α, IL-1β, IL-6, and IL-1257,58. By inhibiting pathogen colonization, M-Li05 reduced antigen exposure to intestinal immune cells, thereby suppressing colonic inflammation. Transcriptome analysis of colon tissues revealed that DSS-induced colitis upregulated inflammation-related genes, including those involved in cytokine–cytokine receptor interactions, NF-κB, and TNF signaling pathways. Treatment with M-Li05 significantly reduced the expression of these inflammatory genes, likely due to its colonization resistance against pathogens and the preservation of the intestinal barrier. Specifically, neutrophil and macrophage activation, particularly M1 macrophages, was reduced, and pro-inflammatory cytokines, such as IL-1α, IL-1β, IL-6, IL-12, and TNF-α, were significantly repressed in the ML group.
In this study, the aMUC2 adhesin was successfully conjugated to the surface of P. pentosaceus Li05, enhancing its mucoadhesion in the gut. The modification process was rapid, non-toxic, and did not affect the growth characteristics of Li05. Treatment with M-Li05 significantly alleviated C. rodentium infection and DSS-induced colitis by clearing pathogens, improving gut barrier integrity, inhibiting excessive inflammation, and modulating metabolic profiles. While M-Li05 shows promise as a therapeutic for colitis, the current modification process is not suitable for large-scale production, limiting its broader application. Future research will focus on optimizing the production process for scalability.
Methods
Bacterial strains and cell lines
Pediococcus pentosaceus Li05 (CGMCC 7049) was cultured in Man Rogosa Sharpe (MRS) medium (Oxoid, Basingstoke, UK) in an anaerobic workstation (37 °C, 24 h) (bioMérieux, Marcy l′Ètoile, France). Bioluminescent Citrobacter rodentium strain ICC180 was inoculated into Luria-Bertani (LB) broth (BD, Franklin Lakes, NJ, USA) in an aerobic incubator (37 °C, 24 h, 200 rpm). Caco-2 cell (TCHu146) and HT29-MTX-E12 cell were purchased from Cell Bank/Stem Cell Bank, Chinese Academy of Sciences. Caco-2 and HT29-MTX cells were cultured using Dulbecco′s modified Eagle′s medium (DMEM) (Gibco, Grand Island, USA) supplemented with 1% penicillin-streptomycin (Gibco, Grand Island, USA) and 10% fetal bovine serum (Gibco, Grand Island, USA) at 37 °C with 5% CO2.
Surface modification of P. pentosaceus Li05
The process of surface modification of Li05 is shown in Fig. S1. At first, Li05 was cultured anaerobically in MRS broth (37 °C, 24 h). Bacteria were harvested for further use via centrifugation (5000 × g, 5 min) and washed twice with phosphate-buffered saline (PBS) solution. Then, 1 ml suspension containing 109 CFUs Li05 cells was incubated with 1 mg sulfo-NHS-functionalized biotin (Thermo Fisher Scientific, MA, USA) on a shaker for 30 min at room temperature (RT). After biotinylation, Li05 were harvested via centrifugation and washed twice with 100 mM glycine PBS solution.
Anti-MUC2 polyclonal antibody (Abcam, Cambridge, UK) was conjugated to streptavidin using a streptavidin conjugation kit (Abcam, Cambridge, UK). Anti-MUC2-streptavidin was then incubated with biotinylated bacteria for 30 min on a shaker. Finally, the aMUC2-targeted Li05 (M-Li05) was harvested via centrifugation (5000 × g, 5 min) and washed twice with PBS.
To evaluate the conjugation of streptavidin to biotinylated Li05, 1 ml of non-biotinylated or biotinylated Li05 (109 CFUs / mL) were incubated with 10 μL fluorescent streptavidin (1 mg/ml) (Alexa Fluor 568 Streptavidin) (Thermo Fisher Scientific, MA, USA) for 30 min, followed by incubation with DAPI staining solution (Servicebio, Wuhan, China) for 10 min in a dark environment and washed twice with PBS. Images of the stained bacteria were taken using a confocal microscope (DMi8, Leica, Wetzlar, Germany).
To evaluate the stability of the modification, 1 ml biotinylated Li05 (109 CFUs / mL) was conjugated with fluorescent streptavidin in the way described above. Simulated gastric fluids and simulated intestinal fluids were prepared according to a protocol59. After DAPI staining, the solution was treated with simulated gastric fluids or simulated intestinal fluids for 0 min, 30 min or 1 h in a dark environment. After treatment, images of the stained bacteria were taken immediately using a confocal microscope (DMi8, Leica, Wetzlar, Germany).
Scanning electronic microscopy of bacterial morphology
1 ml suspension (109 CFUs / mL) of Li05 and M-Li05 were seeded in coverslips and incubated for 1 h at 37 °C. Then, the coverslips were fixed with 2.5% glutaraldehyde solution (24 h, 4 °C) protecting from light. After electron microscopy sample preparation, the samples were subsequently coated with a thin layer of gold by direct current sputtering before visualization at different magnifications (5000× and 30000×) with a Nova Nano 450 SEM (Thermo Fisher Scientific, Czech).
Bacterial attachment to Caco-2 / HT29-MTX cells model
Caco-2 and HT29-MTX cells were seeded in 6-well plates (Corning, NY, USA) in a 4:1 ratio and grown to 90–100% confluence. Before treatment with bacteria, Caco-2 / HT29-MTX cells were triple-washed with PBS to completely remove the remaining FBS and pen-strep. Li05 and M-Li05 were prepared as described above. After the final wash, bacteria were resuspended in DMEM and diluted to 108 CFUs/mL, of which 1 mL was added to the plates and incubated with Caco-25/HT29-MTX cells (37 °C, 1 h). Then, Caco-2 / HT29-MTX cells were washed three times with PBS to remove the unattached bacteria. The remaining cells and bacteria were harvested by cell scrapers (BIOLOGIX, Shandong, China). The numbers of the adhered bacteria were counted by plating gradient dilutions of the mixture of cells and bacterial suspension on MRS agar medium (Oxoid, Basingstoke, UK).
Bacterial colonization in Germ-Free (GF) rats
GF Sprague-Dawley male 10 weeks-old rats were purchased from Shanghai Lab Animal Research Center (Shanghai, China) and nurtured and maintained by the Germ-free Animal Platform of State Key Laboratory for Diagnosis and Treatment of Infectious Diseases. Rats were randomly divided into two groups: treatment of unmodified Li05 (Li05) and treatment of M-Li05 (M-Li05). Li05 and M-Li05 groups were orally administrated once with 1 mL bacterial solution (containing 109 CFUs P. pentosaceus Li05 or M-Li05). Fecal samples were collected at different time points after gavage (1 h, 4 h, 8 h, 24 h, 2 days, 3 days, 4 days, 5 days and 6 days). Numbers of adhered P. pentosaceus were counted by RT-qPCR as described in the following section.
In vitro adhesion competition against C. rodentium
To assess the colonization resistance of Li05 or M-Li05 toward pathogenic bacteria, C. rodentium was selected as a model pathogen. Caco-2 and HT29-MTX cells were seeded in 12-well plates (Corning, NY, USA) in a ratio of 4:1 and grown to 90–100% confluence. Before treatment with bacteria, cells were triple-washed with PBS to completely remove the remaining FBS and pen-strep. For the competitive adhesion experiment, bacteria in the same concentration (about 5*107 CFUs / ml) were suspended in DMEM. Cultures of DMEM / C. rodentium, Li05 / C. rodentium or M-Li05 / C. rodentium were incubated with Caco-2 monolayers (37 °C, 1 h) and washed three times with PBS to remove unattached bacteria. Bioluminescence of C. rodentium was taken immediately by IVIS Spectrum optical imaging system (Caliper Life Sciences, Waltham, MA, USA). Then, the remaining cells and bacteria were harvested with cell scrapers. The numbers of the adhered C. rodentium were counted by plating gradient dilutions of the mixture of cells and bacterial suspension on MacConkey (MAC) agar medium (Solarbio, Beijing, China).
Mice
Specific pathogen-free (SPF) male and female eight-week-old C57/BL6J mice were purchased from Ziyuan Experimental Animal Technology (Hangzhou, China) and housed under 12 h light and dark cycles. Mice were fed with Lab mice diet (Zhejiang Experimental Animal Center, Hangzhou, China).
Animal experiment 1: inhibition of C. rodentium infected colitis in mice
According to a protocol for C. rodentium infection mouse model28, female mice (18-20 g) were chosen and divided randomly into five groups (n = 6 in each group): negative control (NC), C. rodentium infected (CR), treatment of biotin-streptavidin-aMUC2 (BM), treatment of Li05 (L) and treatment of M-Li05 (ML). L and ML groups were orally treated with 200 μl bacterial solution (~109 CFUs) of Li05 or M-Li05 from day -7 to day 14. BM group were orally treated with 200 μl of biotin-streptavidin-aMUC2 from day −7 to day 14. CR and NC groups were intragastrically treated with 200 μl PBS from day −7 to day 14 as control. Before infection, C. rodentium was grown overnight in LB broth and harvested via centrifugation (3000 g, 15 min). At day 1, each mouse in the CR, BM, L and ML groups was orally inoculated with 200 μl of bacterial solution (containing 3*109 CFUs C. rodentium) to induce infection, whereas mice in the NC group received 200 μl of PBS as control. The weight of mice and fecal condition were recorded daily. At day 14, all mice were humanely sacrificed under isoflurane anesthesia.
Animal experiment 2: prevention of DSS-induced colitis in mice
Male mice were divided randomly into five groups (n = 8 in each group): NC, P, BM, L and ML. Mice in the BM group were treated with 200 μl biotin-streptavidin-aMUC2 by oral gavage from day −7 to day 9. Mice in the L and ML groups were treated with 200 μl Li05 or M-Li05 by oral gavage from day −7 to day 9. As control, mice in the NC and P groups received 200 μl PBS daily. Mice in P, BM, L and ML groups were given drinking water containing 2.5% (wt/vol) DSS (molecular weight: 36,000–50,000 Da; MP Biomedicals, California, USA) from day 0 to day 7, followed by distilled water for 3 days. The disease activity index (DAI) score was recorded daily after starting DSS administration to evaluate the colitis severity. The DAI score (0–12) include three parts: weight loss, fecal blood and consistency. Fecal blood was detected by fecal occult blood reagent test (Baso, Zhuhai, China). At day 9, all mice were humanely sacrificed under isoflurane anesthesia.
Quantification of Li05 and C. rodentium in mice
To evaluate the clearance of C. rodentium and colonization of Li05, fecal pellets were collected from mice in each group every two days starting from day 4 to day 14. Previously reported primers were used to quantify C. rodentium60 and P. pentosaceus61. The standard curves of quantification were provided in Fig. S2A, B. DNA was extracted from an accurately weighed fecal sample (50 mg) by the DNeasy PowerSoil Pro kit (Qiagen, Hilden, Germany). Then, RT-qPCR was performed to quantify C. rodentium and P. pentosaceus. The samples were tested in duplicate.
Dynamic change of C. rodentium in GIT using 2D bioluminescent imaging
Mice were anaesthetized with isoflurane and depilated before capture. Bioluminescence imaging (BLI) of C. rodentium was performed by IVIS Spectrum optical imaging system at days 4, 6, 8, 12, and 14. Regions of bioluminescence were identified and quantified (photons/s/cm2/sr) (sr=Steradian) using the Living Image 4.3.1 software (Caliper Life Sciences).
Evaluation of colonic morphology by transmission electron microscopy
Colon tissues were collected and fixed immediately in 2.5% glutaraldehyde solution in a dark environment (24 h, 4 °C). After washed 3 times with PBS, the tissues were fixed with 1% osmic acid and washed with PBS again. After subjecting to ethanol (50%, 70%, 90%, 100%) and absolute acetone for dehydration, the samples were sectioned with ultramicrotome (Leica, Wetzlar, Germany). Lead citrate solution and uranyl acetate solution were used to stain the samples successively. Finally, the prepared tissues were finally observed at different magnifications (11000× and 49000×) under Tecnai G2 Spirit transmission electron microscope (Thermo Fisher Scientific, MA, USA).
Histopathology and immunohistochemistry analysis
Immediately after sacrificing the mice, 1 cm terminal colon tissues were collected and fixed with 4% paraformaldehyde overnight at RT. Then colon tissues were embedded in paraffin and cut into 2μm-thick sections. After deparaffinization and hydration, the colon sections were stained with hematoxylin solution for 3 min. After washing, the colon sections were stained with eosin solution for 2 min. Following dehydration, the colon samples were observed under a light microscope and the degree of colon injury was scored according to a histological scoring standard62. For immunohistochemistry, colon sections were stained with myeloperoxidase (MPO), Ki67 and F4/80 (Abcam, Cambridge, UK). For immunofluorescence, colon sections were stained with ZO-1, occludin and claudin1 (Abcam, Cambridge, UK).
RNA extraction and real-time quantitative PCR analysis
RNeasy mini kit (Qiagen, Hilden, Germany) was used for the extraction of total RNA from terminal colon tissues and the whole process was carried out on ice to minimise RNA degradation. Extracted RNA was immediately reverse transcribed to cDNA using PrimeScript RT master mix (TaKaRa Biomedicals, Kusatsu, Japan) and then stored at -80°C. The relative expression of targeted genes was analysed in duplicate and Gapdh was used as an internal reference. Finally, ΔΔCT was calculated for the final measurement. Primers are presented in Table S1.
16S rRNA DNA analysis
Fecal total bacterial DNA was extracted with DNeasy PowerSoil Pro kit (Qiagen, Hilden, Germany). The V3-V4 region of 16S rRNA gene was amplified using universal bacterial primers (see table S1). After purification, the amplified products were further used to generate the sequencing libraries. Finally, the qualified library was sequenced with NovaSeq-PE250 platform (Illumina). Paired-end reads were assigned to samples based on their unique barcodes. Then Paired-end reads were merged, denoised, and filtered using the QIIME2 software (Version QIIME2-202006) to obtain Amplicon Sequence Variants (ASVs). α diversity was analysed by Chao1 and Shannon indexes. β diversity was visualized by Principal Coordinate Analysis (PCoA) based on weighted UniFrac metrics.
Flow analysis of colonic lamina propria
The intact colons were separated and immerged in PBS solution immediately after the mice were sacrificed. The dissociation and digestion were conducted after cutting and cleaning colon tissue. The immune cells were acquired by centrifugation at 1000 g for 20 min. After staining with L/D, CD45, CD11b, F4/80, CD86 and CD206 (BD Pharmingen, San Diego, CA, USA), the cells were fixed and analysed using LSR Fortessa (BD Biosciences, San Jose, CA, USA). The data were analysed by FlowJo_V10.8.
Transcriptome analysis of colon tissue
Total RNA from the colon was extracted and reverse transcribed to cDNA as previously mentioned. Then cDNA was sequenced and the clean reads aligned to reference genome. The expression level of each transcript was calculated according to the transcripts per million reads. The differential expression genes (DEGs) with FC≧1 and P value < 0.05 were considered to be significantly different expressed genes. Finally, KEGG-enrichment analysis was performed using Python scipy software to identify which DEGs were significantly enriched in specific pathways.
Quantification of short chain fatty acids in colonic content
50 mg fecal sample was suspended in 1 ml 80% methanol-water solution. Mixed the supernatant with 20 μl of 3-NPH and 20 μl of EDC reagent after homogenization and centrifugation (20,000 × g, 15 min, 4 °C). After centrifugation and incubation (40 °C, 30 min), 440 μl initial mobile phase solution was added to the mixture. Then, 200 μl of the mixture was transferred into the injection vial for LC-MS/MS analysis. The analysis was performed on a TSQ Altis mass spectrometer (Thermo Scientific, MA, USA). The external standard method was used for quantification, and the retention time and MRM fragment ions were compared by the methodology standards for characterization.
Statistical analysis
The results are presented as means ± SEM. The Kolmogorov–Smirnov test or Shapiro-Wilk test was used to analyse the normality of the data. Statistically significant differences between groups were evaluated by one-way ANOVA followed by Tukey’s post hoc test or Student’s t-tests. Brown-Forsythe ANOVA followed by Dunnett-T3’s test was applied for the groups that did not show homogeneity of variance. Nonparametric ANOVA (Kruskal–Wallis test) followed by Dunn’s post hoc test was applied when data did not pass normal distribution test. P < 0.05 was considered statistically different. GraphPad Prism (version 8) was used for statistical analysis.
Ethics
The present study was approved by the ethics committee of the First Affiliated Hospital of Zhejiang University (2022-1020).
Data availability
All the data in this study are presented in the paper or the Supplementary Materials. Additional data relevant to the article are available from the corresponding author upon request.
References
Matsuoka, K. & Kanai, T. The gut microbiota and inflammatory bowel disease. Semin Immunopathol. 37, 47–55 (2015).
Khor, B., Gardet, A. & Xavier, R. J. Genetics and pathogenesis of inflammatory bowel disease. Nature 474, 307–317 (2011).
Khan, I. et al. Differential susceptibility of the gut microbiota to DSS treatment interferes in the conserved microbiome association in mouse models of colitis and is related to the initial gut microbiota difference. Adv. Gut Microbiome Res. 2022, 7813278 (2022).
Fritsch, J. et al. Low-fat, high-fiber diet reduces markers of inflammation and dysbiosis and improves quality of life in patients with ulcerative colitis. Clin. Gastroenterol. Hepatol. 19, 1189–1199.e30 (2021).
Rizzello, F. et al., Implications of the westernized diet in the onset and progression of IBD. Nutrients, 11 (2019).
Lavelle, A. & Sokol, H. Gut microbiota-derived metabolites as key actors in inflammatory bowel disease. Nat. Rev. Gastroenterol. Hepatol. 17, 223–237 (2020).
Kudelka, M. R. et al. Intestinal epithelial glycosylation in homeostasis and gut microbiota interactions in IBD. Nat. Rev. Gastroenterol. Hepatol. 17, 597–617 (2020).
Hill, C. et al. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 11, 506–514 (2014).
Sood, A. et al., The probiotic preparation, VSL#3 induces remission in patients with mild-to-moderately active ulcerative colitis. Clin Gastroenterol. Hepatol. 7, 1202-9, 1209.e1 (2009).
Rembacken, B. J. et al. Non-pathogenic Escherichia coli versus mesalazine for the treatment of ulcerative colitis: a randomised trial. Lancet 354, 635–639 (1999).
Zocco, M. A. et al. Efficacy of Lactobacillus GG in maintaining remission of ulcerative colitis. Aliment Pharm. Ther. 23, 1567–1574 (2006).
Qu, S. et al. Akkermansia muciniphila alleviates dextran sulfate sodium (DSS)-induced acute colitis by NLRP3 activation. Microbiol Spectr. 9, e0073021 (2021).
Bian, X. et al. Pediococcus pentosaceus LI05 alleviates DSS-induced colitis by modulating immunological profiles, the gut microbiota and short-chain fatty acid levels in a mouse model. Micro Biotechnol. 13, 1228–1244 (2020).
Shi, D. et al. Administration of Lactobacillus salivarius LI01 or Pediococcus pentosaceus LI05 prevents CCl(4)-induced liver cirrhosis by protecting the intestinal barrier in rats. Sci. Rep. 7, 6927 (2017).
Xie, J. et al. Impact of encapsulating a probiotic (Pediococcus pentosaceus Li05) within gastro-responsive microgels on Clostridium difficile infections. Food Funct. 12, 3180–3190 (2021).
Yao, M. et al. Enhanced viability of probiotics (Pediococcus pentosaceus Li05) by encapsulation in microgels doped with inorganic nanoparticles. Food Hydrocoll. 83, 246–252 (2018).
Han, S. Y. et al. Probiotic gastrointestinal transit and colonization after oral administration: a long journey. Front. Cell. Infect. Microbiol. 11, 12 (2021).
Van Tassell, M. L. & Miller, M. J. Lactobacillus adhesion to mucus. Nutrients 3, 613–636 (2011).
Yao, D. et al. MUC2 and related bacterial factors: Therapeutic targets for ulcerative colitis. EBioMedicine 74, 103751 (2021).
Petty, N. K. et al. The Citrobacter rodentium genome sequence reveals convergent evolution with human pathogenic Escherichia coli. J. Bacteriol. 192, 525–538 (2010).
Collins, J. W. et al. Citrobacter rodentium: infection, inflammation and the microbiota. Nat. Rev. Microbiol 12, 612–623 (2014).
Mullineaux-Sanders, C. et al. Citrobacter rodentium-host-microbiota interactions: immunity, bioenergetics and metabolism. Nat. Rev. Microbiol. 17, 701–715 (2019).
Mullineaux-Sanders, C. et al. Citrobacter rodentium relies on commensals for colonization of the colonic mucosa. Cell Rep. 21, 3381–3389 (2017).
Hopkins, E. G. D. et al., Intestinal epithelial cells and the microbiome undergo swift reprogramming at the inception of colonic Citrobacter rodentium infection. mBio 10 (2019).
Mao, T. et al. Hyaluronan-induced alterations of the gut microbiome protects mice against Citrobacter rodentium infection and intestinal inflammation. Gut Microbes 13, 1972757 (2021).
Collins, J. W. et al. Pre-treatment with Bifidobacterium breve UCC2003 modulates Citrobacter rodentium-induced colonic inflammation and organ specificity. Microbiology 158, 2826–2834 (2012).
Said, H. M. Biotin: biochemical, physiological and clinical aspects. Subcell. Biochem. 56, 1–19 (2012).
Crepin, V. F. et al. Citrobacter rodentium mouse model of bacterial infection. Nat. Protoc. 11, 1851–1876 (2016).
Lebeis, S. L. et al. TLR signaling mediated by MyD88 is required for a protective innate immune response by neutrophils to Citrobacter rodentium. J. Immunol. 179, 566–577 (2007).
Gibson, D. L. et al. MyD88 signalling plays a critical role in host defence by controlling pathogen burden and promoting epithelial cell homeostasis during Citrobacter rodentium-induced colitis. Cell Microbiol. 10, 618–631 (2008).
Frankel, G. & Phillips, A. D. Attaching effacing Escherichia coli and paradigms of Tir-triggered actin polymerization: getting off the pedestal. Cell Microbiol. 10, 549–556 (2008).
Guttman, J. A. et al. Evidence that tight junctions are disrupted due to intimate bacterial contact and not inflammation during attaching and effacing pathogen infection in vivo. Infect. Immun. 74, 6075–6084 (2006).
Kamada, N. et al. Regulated virulence controls the ability of a pathogen to compete with the gut microbiota. Science 336, 1325–1329 (2012).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353.e21 (2016).
Chen, C. M. et al. Lactobacillus plantarum PS128 promotes intestinal motility, mucin production, and serotonin signaling in mice. Probiotics Antimicrob. Proteins 14, 535–545 (2022).
Celebioglu, H. U. & Svensson, B. Dietary nutrients, proteomes, and adhesion of probiotic lactobacilli to mucin and host epithelial cells. Microorganisms 6, 17 (2018).
Wu, H. et al. Lactobacillus reuteri maintains intestinal epithelial regeneration and repairs damaged intestinal mucosa. Gut Microbes 11, 997–1014 (2020).
Kong, Q. et al. Daily intake of Lactobacillus alleviates autistic-like behaviors by ameliorating the 5-hydroxytryptamine metabolic disorder in VPA-treated rats during weaning and sexual maturation. Food Funct. 12, 2591–2604 (2021).
Willing, B. P. et al. Altering host resistance to infections through microbial transplantation. PLoS One 6, e26988 (2011).
Kim, M. H. et al. Short-chain fatty acids activate GPR41 and GPR43 on intestinal epithelial cells to promote inflammatory responses in mice. Gastroenterology 145, 396–406.e1-10 (2013).
Carlsson, A. H. et al. Faecalibacterium prausnitzii supernatant improves intestinal barrier function in mice DSS colitis. Scand. J. Gastroenterol. 48, 1136–1144 (2013).
Scott, N. A. et al., Antibiotics induce sustained dysregulation of intestinal T cell immunity by perturbing macrophage homeostasis. Sci. Transl. Med. 10 (2018).
Trompette, A. et al. Dietary fiber confers protection against flu by shaping Ly6c(-) patrolling monocyte hematopoiesis and cd8(+) t cell metabolism. Immunity 48, 992–1005.e8 (2018).
Burger-van Paassen, N. et al. The regulation of intestinal mucin MUC2 expression by short-chain fatty acids: implications for epithelial protection. Biochem. J. 420, 211–219 (2009).
Burkinshaw, B. J. & Strynadka, N. C. Assembly and structure of the T3SS. Biochim. Biophys. Acta 1843, 1649–1663 (2014).
Wirtz, S. et al. Chemically induced mouse models of acute and chronic intestinal inflammation. Nat. Protoc. 12, 1295–1309 (2017).
Guo, W. et al. Small molecule-driven mitophagy-mediated NLRP3 inflammasome inhibition is responsible for the prevention of colitis-associated cancer. Autophagy 10, 972–985 (2014).
Mirsepasi-Lauridsen, H. C. et al., Escherichia coli pathobionts associated with inflammatory bowel disease. Clin. Microbiol. Rev. 32 (2019).
Ohkusa, T. et al. Fusobacterium varium localized in the colonic mucosa of patients with ulcerative colitis stimulates species-specific antibody. J. Gastroenterol. Hepatol. 17, 849–853 (2002).
Lupp, C. et al. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe 2, 119–129 (2007).
Walker, A. W. et al. High-throughput clone library analysis of the mucosa-associated microbiota reveals dysbiosis and differences between inflamed and non-inflamed regions of the intestine in inflammatory bowel disease. BMC Microbiol 11, 7 (2011).
Reis, R. S. & Horn, F. Enteropathogenic Escherichia coli, Samonella, Shigella and Yersinia: cellular aspects of host-bacteria interactions in enteric diseases. Gut Pathog. 2, 8 (2010).
Hardwidge, P. R. et al. Modulation of host cytoskeleton function by the enteropathogenic Escherichia coli and Citrobacter rodentium effector protein EspG. Infect. Immun. 73, 2586–2594 (2005).
Kaper, J. B., Nataro, J. P. & Mobley, H. L. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2, 123–140 (2004).
Qi, X. et al. Identification and characterization of neutrophil heterogeneity in sepsis. Crit. Care 25, 50 (2021).
Orecchioni, M. et al. Macrophage polarization: different gene signatures in M1(LPS+) vs. classically and M2(LPS-) vs. alternatively activated macrophages. Front Immunol. 10, 1084 (2019).
Pan, X. et al. Macrophage immunometabolism in inflammatory bowel diseases: from pathogenesis to therapy. Pharm. Ther. 238, 108176 (2022).
Danne, C. et al. Neutrophils: from IBD to the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 21, 184–197 (2024).
Brodkorb, A. et al. INFOGEST static in vitro simulation of gastrointestinal food digestion. Nat. Protoc. 14, 991–1014 (2019).
Sagaidak, S. et al. Development of a real-time PCR assay for quantification of Citrobacter rodentium. J. Microbiol. Methods 126, 76–77 (2016).
Dreier, M. et al. SpeciesPrimer: a bioinformatics pipeline dedicated to the design of qPCR primers for the quantification of bacterial species. PeerJ 8, e8544 (2020).
Chen, C. C. et al. Concurrent infection with an intestinal helminth parasite impairs host resistance to enteric Citrobacter rodentium and enhances Citrobacter-induced colitis in mice. Infect. Immun. 73, 5468–5481 (2005).
Acknowledgements
Thanks to Gad Frankel for kindly providing the bioluminescent C. rodentium strain ICC180.
Author information
Authors and Affiliations
Contributions
L.L. and M.Y. conceived the research. S.H., K.W. and K.S. designed, conducted experiments and wrote the manuscript. X.D., Y.H. and Y.L., analyzed the data. S.Z., B.Q. and Y.L. contributed new reagents or analytical tools. H.X., D.M. and B.B. polished this manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, S., Wang, K., Su, K. et al. Improved adhesion and function of Pediococcus pentosaceus Li05 for acute colitis through surface modification. npj Sci Food 10, 5 (2026). https://doi.org/10.1038/s41538-025-00652-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-025-00652-z