Abstract
Brain dynamics can be simulated using virtual brain models, in which a standard mathematical representation of oscillatory activity is usually adopted for all cortical and subcortical regions. However, some brain regions have specific microcircuit properties that are not recapitulated by standard oscillators. Moreover, magnetic resonance imaging (MRI)-based connectomes may not be able to capture local circuit connectivity. Region-specific models incorporating computational properties of local neurons and microcircuits have recently been generated using the mean field (MF) approach and proposed to impact large-scale brain dynamics. Here, we have used a MF of the cerebellar cortex to generate a mesoscopic model of the whole cerebellum featuring a prewired connectivity of multiple cerebellar cortical areas with deep cerebellar nuclei. This multi-node cerebellar MF was then used to substitute the corresponding standard oscillators and build up a cerebellar mean field virtual brain (cMF-TVB) for a group of healthy human subjects. Simulations revealed that electrophysiological and fMRI signals generated by the cMF-TVB significantly improved the fitness of local and global dynamics with respect to a homogeneous model made solely of standard oscillators. The cMF-TVB reproduced the rhythmic oscillations and coherence typical of the cerebellar circuit and allowed to correlate electrophysiological and functional MRI signals to specific neuronal populations. In aggregate, region-specific models based on MF and pre-wired circuit connectivity can significantly improve virtual brain simulations, fostering the generation of effective brain digital twins that could be used for physiological studies and clinical applications.
Similar content being viewed by others
Introduction
Structure, function and dynamics, the organising principles dominating brain activity1, have recently been recapitulated by virtual-brain technology2,3,4. Virtual brains leverage structural magnetic resonance imaging (MRI) data to reconstruct brain connectivity, place mathematical representations of neural activity in grey matter nodes, and generate emerging dynamics through simulations5,6,7. In this way, microscale neuronal dynamics can be integrated into mesoscale population models, which act then as generative models for macroscale signals such as Blood Oxygen Dependent (BOLD) signals, acquired with functional MRI (fMRI), or neural activity generated by electrical currents recorded with electroencephalography (EEG) or magnetoencephalography (MEG)3,8,9. Since virtual-brain models can be tailored to single subjects by integrating subject-specific MRI recordings with the generative models of neuronal activity, they are opening the perspective of brain digital twins10,11 applicable to personalised medicine for understanding brain function and enabling the personalised treatment of diseases. In brain modelling landscape, ‘The Virtual Brain’ (TVB) is a commonly used neuroinformatic platform designed to simulate whole brain dynamics in physiological and pathological states2,3,4,11,12,13,14,15,16,17. TVB operates under the assumption that a neural mass model, collapsing neuronal heterogeneity into two populations (one excitatory and the other inhibitory), can capture local microcircuit dynamics all over the brain18,19. Typical neural mass models include the Wong-Wang (WW), the Wilson-Cowan (WC), and the Generic Oscillator (GO) model19,20,21,22. Recent advancements of TVB technology include more complex models tailored to specific pathologies such as epilepsy and/or designed to generate specific macroscale recordings such as EEG signals12,23,24. However, by using the same models for all nodes, TVB misses the opportunity to capture local microcircuit functional diversity, limiting its applicability to investigate the pathophysiological mechanisms underlying disrupted brain oscillations. The resolution of local microcircuit structure and function is critical to accurately simulate brain dynamics and extract physiological information at higher scales. Spiking neural networks (SNN) are a computational representation of the region-specific microcircuitry, embedding thousands of neurons interconnected according to the anatomo-physiology of a specific region. While ideally, the best resolution should be achieved for each node, in practice, the mathematical representation of nodal activity must be compatible with the computational load of a whole-brain simulation, and simulating large-scale dynamics with SNN is computationally intractable. Mean-field (MF) approximation simplifies the dynamics of complex systems by approximating the interactions of individual elements with an averaged interaction field. When applied to brain modelling, MF approximation replaces the neuron-neuron interaction with neuron-mean field ones, providing a statistical description of the local microcircuit25,26. The MF model representing a specific brain region can be formulated using the SNN as a functional ref. 27. This approach preserves the physiological structural organisation and functional properties of local neuronal microcircuits at a neuronal population level of granularity, which is computationally efficient for being used as a generative model of large-scale dynamics26,27,28,29,30,31.
The cerebellum is a large brain structure accounting for more than half of all brain neurons18, is tightly interconnected with the rest of the brain32,33, and has a feed-forward organisation that is not represented by standard neural masses34. The cerebellar cortex features four major neuronal populations, i.e. granule cells (GrCs), Golgi cells (GoCs), molecular layer interneurons (MLIs), and Purkinje cells (PCs), whose physiological function and connectivity are oversimplified (or overtly wrong in physiological terms) in neural mass models. Moreover, the cerebellar output from PCs is inhibitory on deep cerebellar nuclei (DCN)18,35, while communication between TVB nodes normally uses excitatory synapses only3. Finally, communication between cerebellar nodes cannot be fully resolved by MRI tractography in humans in vivo36,37,38. All these issues can severely compromise the anatomo-physiological consistency of virtual-brain simulations. The cerebellar mean field model (CRBL MF) is tailored on salient features of the cerebellar neuronal populations based on precise cellular-level reconstructions of the microcircuit29,34,39. Structural and functional parameters of the CRBL MF, such as the synaptic convergence, quantal synaptic conductance and synaptic time decay, were extracted from a computational representation of the cerebellar microcircuit, i.e. a cerebellar SNN, previously validated using experimental recordings34,39,40,41. For methodological consistency, the MF theory used to develop the CRBL MF follows that of the MF of the cerebral cortex, relying on a transfer function formalism that allows to transfer the microscale biophysical parameters to a mesoscopic domain26,27. Specificity of the cerebellar cortex neuronal architecture was preserved, including recurrent excitation and inhibition amongst GrC, GoC, MLI and PC18. The population-specific transfer functions return postsynaptic activity as firing rate (Hz) from presynaptic inputs (i.e. firing rates), accounting for neuronal discharge frequencies and membrane potential fluctuations.
Here, we developed a multi-node CRBL MF and integrated it into TVB, generating a cerebellar mean field TVB (cMF-TVB) model. This required first to develop a multi-node CRBL MF by assembling interconnected MF (one for each cerebellar node) with nodes of DCN. Then, the multi-node CRBL MF was connected to the rest of the brain, in which nodes are typically represented as standard neural masses. To maintain the physiological inter-regional communication, the connectivity between cerebellar cortical nodes and DCN was imposed a priori, as it was not fully resolved on the microscopic and mesoscopic level by MRI tractography: inhibitory connections were set between PCs and DCN, 10% feedback was introduced from DCN to PCs, and parallel fibres were added to connect adjacent cerebellar regions42,43. Finally, simulated electrical activity and the BOLD signal were referred to the specific MF neuronal populations most likely to generate them and were validated against empirical data recorded in humans in vivo. The cMF-TVB was validated against standard TVB simulations, constructed by associating the same two-population model (i.e. Wong-Wang mode (WW) l19,20) to all the nodes.
In aggregate, the cMF-TVB incorporating a region-specific model, i.e. the multi-node CRBL MF, proved capable to improve local circuit and whole brain simulations, and to capture high-level physiological features of cerebellar activity like large-scale oscillations on multiple frequency bands44,45. Consequently, cMF-TVB would enable a neuronal population-specific characterisation of disrupted cerebellar oscillations in neurodegenerative diseases such as Parkinson’s disease. Moreover, the cMF-TVB framework is deployed as a ‘plug and play’ tool, ensuring a seamless integration of other region-specific MF models. This work stands as the first brick to construct a brain avatar based on a physiological-grounded representation of the inter-region heterogeneity. When applied to pathology, the patient brain avatar will allow us to explore the neural basis underpinning pathology-driven dynamics.
Results
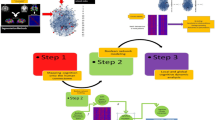
In this work, we addressed how to integrate different types of models with MRI data into the TVB platform using the CRBL MF as a prototype. Our framework treats brain modelling with a multiscale approach that employs a mesoscale MF model to link the microscale neuronal circuitry information to the macroscale observed dynamics. The diagram of the multiscale brain modelling pipeline is presented in Fig. 1, showing our integrative approach applied to the cerebellum and spanning from the microscopic to macroscopic domain. The technical implementation of the multi-node CRBL MF and its integration into TVB, ending up with the cMF-TVB, are detailed in Methods and schematized in Supplementary Fig. 1. We provide a list of acronyms in the section ‘Abbreviations’ to facilitate readability by summarising the main acronyms used throughout the paper.
An integrative approach to model cerebellar activity, bridging the gap between microscale and macroscale signals. The round panels illustrate the activity generated by GrC at different scales as an example of a multiscale modelling framework. Microscale experimental recordings (e.g. action potentials recorded in the GrC) are used to develop point-neuron models. These are wired into highly specific microcircuits, e.g. a cerebellar spiking neural network reproducing the spiking activity of cerebellar neurons, such as GrCs. The spiking neural network serves as the backbone of the CRBL MF, scaling up to the mesoscale by reproducing population activity in terms of firing rate (Hz). The cerebellar mean field model is then integrated into a virtual-brain simulator, ending up with the cMF-TVB that provides effective multiscale simulations.
The nature of the CRBL MF, which is made of 4 neuronal populations, required a specific curation of the structural connectivity (SC) to account for cerebellar-specific connectivity in the construction of the cMF-TVB circuitry. Figure 2 illustrates the axonal fibres that establish cerebellar connectivity, including parallel fibres, which link adjacent regions (i.e. nodes) of the cerebellar cortex through excitatory long-range synapses, and PC axons, which project from the cerebellar cortex to the DCN, forming inhibitory long-range synapses (Fig. 2A). The curated cerebellar SC incorporates both elements, along with the feedback projections from DCN, creating a structural backbone that includes both excitatory and inhibitory long-range connections (Fig. 2B). The SC curation pipeline is detailed in detail in Supplementary Fig. 2. The subject-specific whole brain SC includes integrates curated cerebellar SC, with the cerebro-cerebellar structural weights derived using tractography (details in the section ‘Structural connectivity (SC)’), and it is used as structural backbone to build up the cMF-TVB architecture. The cMF-TVB circuitry is constructed by connecting the multi-node CRBL MF, representing the cerebellar cortex, with the WW, representing the DCN and the cerebrum. Inter-model connections were set according to SC weights, resulting in the circuitry schematized in Fig. 3, which represents the functional backbone of cMF-TVB. Simulations of cerebellar dynamics were used to assess the constructive and predictive validity, specifically (i) to assess the robustness of the multi-node CRBL MF and cMF-TVB in simulating neuronal activity (section ‘Constructive and predictive validity’); (ii) to investigate the similarity between empirical BOLD (empBOLD) and simulated BOLD (simBOLD) computed by convolving the neuronal activity with the TVB-built-in hemodynamic response function, i.e. the Balloon-Wind Kessel model46 (sections ‘Multi-node CRBL MF simulations (open loop)’ and ‘cMF-TVB simulations (closed loop)’); and (iii) to determine the ability of the cMF-TVB in capturing the multiple oscillatory bands of the cerebellum44 (section ‘Emergent cerebellar rhythms and coherence in the cMF-TVB’).
A Cerebellar SC is curated by integrating microscale data, specifically the cerebellar spiking neural networks, with macroscale recordings, namely the subject-specific anatomical MRI acquisition. To quantify the connections between pairs of cerebellar cortical nodes, the convergence of parallel fibres from GrC to GoC, MLI, and PC is extracted from a cerebellar spiking neural network. Volumes of adjacent cerebellar regions are computed from subject-specific T13D images and summed to weight the population-specific synaptic convergences (i.e. GrC-GoC, GrC-MLI and GrC-PC). Volume-weighted synaptic convergences are normalised for the Intracranial Volume. The connectivity between cerebellar cortex and DCN is derived from anatomically constrained tractography computed on the pre-processed subject-specific diffusion-weighted images. The connection weights from cerebellar cortical nodes to the DCN nodes are made inhibitory and the feedback from DCN to cerebellar cortical nodes was reduced at 10% of the feedforward, accordingly to the physiology of the cerebellar circuit. B Curated cerebellar SC: Green edges represent inhibitory connections (from PC to DCN), while red edges represent excitatory connections made by parallel fibres between adjacent cerebellar cortex nodes. The thickness of the edges is proportional to the SC weights.
The cMF-TVB is built by interconnecting different models according to the spatial map defined by the SC. The cMF-TVB includes the multi-node CRBL MF, made of 27 MF models in the cerebellar cortex, hardwired with 6 WW models in DCN (yellow box), plus 93 WW models in the other brain nodes (light blue boxes). After segmentation and remapping in an atlas virtual space, each cerebellar region is attributed to a node and is represented with an MF. Connectivity between adjacent CRBL MF laying on the transverse plane corresponds to parallel fibres (parallel fibres) generated by GrCs in a source node (m) projecting its activity to GoC, MLI and PC in a target node (k), so that the coupling strength [Hz] from m to k through parallel fibres is Cppfs(k,m). Node k forwards the inhibitory activity of its PCs to the excitatory neurons of a DCN (j), which then sends back connections to node k, with coupling strength CPC(j,k) and Cmfs(k,j), respectively. Node k also receives the excitatory activity of a cerebral node (i), so that the coupling strength [Hz] from j to k through mossy fibres is Cmfs(k,i). DCN and the rest of the brain are coupled through the activity of their excitatory neurons (coupling C between standard oscillators is omitted for simplicity).
Constructive and predictive validity
Simulations of cerebellar neuronal activity were performed both in open loop, i.e. isolated cerebellar cortex, represented by the multi-node CRBL MF (Fig. 4) and in closed loop, considering the cerebellar cortex wired in a network with DCNs and the cerebrum, represented by the cMF-TVB (Fig. 5). The CRBL MF showed population activity within physiological ranges for GrC, GoC, MLI and PC, demonstrating that the biologically grounded features of a single CRBL MF29 was preserved after the multi-node integration in TVB both in open and closed loop (Supplementary Figs. 3A and 4A). Simulations of the BOLD signal in open and closed loops were performed using the simulated neuronal dynamics as generative activity. The performance of the cMF-TVB was assessed either by representing the CRBL with standard neural mass models or with the CRBL MF model and comparing the simBOLD with the empBOLD. Importantly, there is evidence that the BOLD signal of the cerebellum mostly reflects the on-off activity of neurons at the input stage (PC activity is almost stationary over the integration time length of fMRI), thus, we specifically computed the simBOLD signal from GrC activity, capturing the biological origin of BOLD signals47,48.
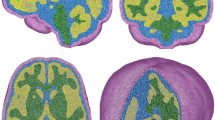
A Cerebellar cortex neuronal activity, reported for left crus II as an example, is simulated using the CRBL MF integrated in the TVB platform for a randomly chosen subject taken from the Human Connectome Project dataset. The curated cerebellar structural connectivity was used to set the connections between different nodes (i.e. regions). To simulate the cerebellar cortex neuronal activity, namely the population-specific firing rate (\({\nu }_{{\rm{p}}}\) with p = [GrC, GoC, MLI, PC]), one CRBL MF was associated with each node belonging to the cerebellar cortex. For each node, the simulated activity lies within the physiological range for each neuronal population. The inset panels show the first 50 ms of the simulation, revealing the different temporal fluctuations in firing activity of each population. Notably, the trend of MLI and PC is opposite, showing the inhibitory effects of MLI on PC. B BOLD signals are extracted from fMRI data (empBOLD) and simulated using, as generative model, either the CRBL MF or one of the generic unspecific models already available in the TVB platform (WW Wong-Wang, WC Wilson-Cowan, GO Generic Oscillator). The flat maps show the MAE between empirical and simulated BOLD, averaged across subjects for each cerebellar cortex region (cold colours correspond to a low MAE). The CRBL MF, as a generative model for BOLD signals, remarkably reduces the MAE in each cerebellar cortex region. C The single-subject MAE is computed for each subject by averaging the region-specific MAE over regions. The shaded areas represent the SEM, indicating inter-regional variability. When using the CRBL MF as a generative model for simulating BOLD signals, the MAE between simulated and empirical signals (~0.20) is significantly reduced compared to unspecific models (Mann–Whitney p value < 0.001).
Cerebellar cortex dynamics are evaluated from whole-brain dynamics simulated with cMF-TVB and compared to a simulation performed using the standard TVB. A Cerebellar cortex neuronal activity, namely the population-specific firing rate (\({\nu }_{{\rm{p}}}\) with p = [GrC, GoC, MLI, PC]) reported for left crus II as an example, is simulated using the CRBL MF integrated in the TVB platform for a subject randomly chosen from the Human Connectome Project dataset. Curated whole-brain structural connectivity is used to map the connections between whole-brain nodes. To simulate the cerebellar cortex neuronal activity, a CRBL MF is associated with each node belonging to the cerebellar cortex, while WW is used for cerebral cortical and subcortical nodes. For each node, the simulated activity is within the physiological ranges for each neuronal population. Furthermore, the interconnection with cerebral nodes results in a heterogeneous activity of MLIs and PCs, driven by the heterogeneous weight input from the cerebral regions to the GrCs. The inset panels display the first 50 ms of the simulation, illustrating that temporal fluctuations in firing activity increase variability across all populations due to input from the cerebral region. B The flat maps show the MAE between empirical BOLD signals (extracted from fMRI data) and the simulated BOLD averaged over the subjects for each cerebellar cortex region (cold colour corresponds to low MAE) extracted from the whole brain simulation. cMF-TVB shows a better performance compared to the standard TVB (WW models in all nodes), reducing the MAE in each cerebellar cortex region. C The single-subject MAE is computed for each subject by averaging the cerebellar cortex region-specific MAE over cerebellar cortex regions. The shaded areas represent the SEM, indicating inter-regional variability. When using the cMF-TVB for simulating whole brain activity, the MAE between simulated and empirical signals (~0.25) is significantly reduced compared to the standard TVB (Mann–Whitney p value < 0.001).
Multi-node CRBL MF simulations (open loop)
In open-loop simulations, the multi-node CRBL MF was activated by 4 Hz random activity on mossy fibres of all the nodes, mimicking the background noise frequency typically observed at rest. For each subject, the simulated firing rate of GrCs was convolved with the Balloon-Wind Kessel model to generate an in-silico BOLD signal of the cerebellar cortical regions (i.e. cerebellar simBOLD). The mean absolute error (MAE) between simBOLD and empBOLD was reduced when the multi-node CRBL MF was used instead of WW, WC or GO (Fig. 4A). The same occurred with the average MAE across subjects (Fig. 4B). The MAE was not normally distributed for WW, WC, and GO, with a non-homogeneous variance in each model (Table 1). The MAE of the CRBL MF against other models was statistically different (Mann–Whitney test, p < 0.0001) for each pair of comparisons (Fig. 4C), namely: CRBL MF vs WW; CRBL MF vs WC; CRBL MF vs GO (Table 2).
cMF-TVB simulations (closed loop)
In closed-loop simulations, the cerebellum was fed by activity transmitted by the other brain nodes of the cMF-TVB. For each subject, simBOLD was computed by following the procedure detailed in the section ‘Multi-node CRBL MF simulations (open loop)’ for the cerebellar cortex, while for the rest of the brain, it was obtained by convolving the simulated firing rate of the excitatory populations of the WW models with the Balloon-Wind Kessel model. The MAE between simBOLD and empBOLD, was calculated both for the TVB and cMF-TVB considering the cerebellar cortical subnetwork (Fig. 5), the DCNs subnetwork (Fig. 6A) and the whole brain (Fig. 6B). For each subject, the MAE between simBOLD and empBOLD in the cerebellar cortex, was reduced when the cMF-TVB was used instead of TVB (Fig. 5B). The same occurred with the average MAE across subjects (Fig. 5C). Interestingly, the MAE improvement was observed also for the DCN nodes (Mann–Whitney test, p < 0.001) (Fig. 6A) and for the whole brain (Mann–Whitney test, p < 0.001) (Fig. 6B), indicating that the improvement brought the CRBL MF propagated across the brain networks.
Whole-brain activity was simulated with cMF-TVB (one CRBL MF associated with each cerebellar cortical node and one WW associated with each deep cerebellar nucleus and cerebrum node). MAE across subjects for cMF-TVB (yellow) and TVB (blue) models, shown as line plots with dots representing individual data points. The shaded areas represent the SEM, indicating inter-subject variability. A simBOLD signals of DCN were extracted from whole-brain simulations performed with either cMF-TVB or TVB. For each subject, simBOLD was averaged over regions and compared to empBOLD by computing the subject-specific MAE for cMF-TVB and TVB. cMF-TVB improves DCN simulations for all the subjects remarkably (except for subject 4, where cMF-TVB and TVB are comparable). Overall, the MAE is significantly reduced when using cMF-TVB (Mann–Whitney p value < 0.001), showing that the inclusion of a region-specific model (i.e. CRBL MF) improves local dynamics compared to non-specific models (e.g. WW). B The subject-specific MAEs between empBOLD and simBOLD using cMF-TVB and TVB are also computed for the whole brain simulation. For all the subjects, cMF-TVB improves the simulation performance significantly (Mann–Whitney p value < 0.001), highlighting that region-specific models can also improve global dynamics.
Emergent cerebellar rhythms and coherence in the cMF-TVB
An emergent property of the cMF-TVB concerns its ability to reveal specific EEG/MEG bands in the cerebellar activity. The cerebellum showed prominent activity in the theta band with relevant components also in other EEG bands (Fig. 7A). During resting-state cMF-TVB simulations, activity was recorded from all neuronal populations in the multi-node CRBL MF. Indeed, for each neuronal population, the Power Spectrum Density (PSD) showed activation in all bands (see Supplementary Fig. 5 for the pipeline and the result of a subject-specific PSD). At the group level, theta band activity predominated in GrC, MLI, and PC, while in GoC the delta, theta, and alpha bands had almost the same power (Fig. 7A). The predominant frequencies averaged across populations were 3 Hz for delta, 6 Hz for theta, 11 Hz for alpha, 20 Hz for beta, and 64 Hz for gamma (Fig. 7B). All bands were represented in the nodes of the multi-node CRBL MF, although some regional variations could be observed most probably reflecting the specific connectivity of each cerebellar node in the cMF-TVB and the different dynamics engaged in the local cerebellar circuits (Fig. 7C). For each region, population-specific carrier-frequency is reported in Supplementary Table 1.
A For each population, the PSD is averaged across subjects to define the bands of circuit oscillations (delta = [0.4 4) Hz, theta = [4 8) Hz, alpha = [8 13) Hz, beta = [12 30) Hz, gamma = [30 100) Hz) for each neuronal population. The theta band is predominant in GrCs, MLIs, and PCs. B Predominant frequencies for each band (only frequencies with a PSD ≥ 0.4 are reported) computed on the cerebellar activity (HZ) averaged across populations. 64 Hz is the predominant frequency with a PSD = 0.87, revealing that a remarkable synchronisation of cerebellar populations’ activity occurs in the middle of the gamma band. C Activity bands in different cerebellar regions. The weight of each band is computed by summing the population-specific power (i.e. the normalised PSD area under the curve, (AUC)). The gamma band is around 20% in all regions, the beta band is usually less than 20% (except in vermis VI and lobules X), the delta, theta, and alpha bands show higher variability across regions.
Another emergent property of the cMF-TVB concerns its ability to reveal coherence between cerebellar nodes during resting-state fMRI recordings. For all subjects, the coherence between cerebellar nodes was more similar to the empirical data when using the cMF-TVB instead of the TVB. The difference between the two models was statistically significant (see Frobenius norm, cosine similarity, Kullback–Leibler (KL) divergence and Pearson Correlation Coefficient (PCC) in Table 3).
Discussion
In the present work, for the first time, a multi-node CRBL MF model was developed and integrated into the cMF-TVB, allowing us to face a set of physiological issues that remained unaddressed so far. First, we have surpassed the generic representation of node activity in the TVB, accounting for the cerebellar microcircuit organisation that does not conform to, e.g. WW, WC, or GC neural masses. Moreover, we have faced the inability of diffusion-weighted imaging (DWI) to resolve fibres interconnecting cerebellar regions and DCN by pre-assembling the CRBL MF and DCN and by weighting their connectivity afterwards. Finally, according to physiology, we have modified the cerebellar output and made it inhibitory on DCN. It should be noted that the extra-cerebellar connectivity was also curated to obviate the inability of tractography in selecting out ipsilateral cerebello-thalamo-cortical spurious tracts32,33,49. We will consider below how the integration of anatomo-physiological properties of a specific brain circuit into large-scale simulation advances the biomimetic capacity of virtual brains toward the generation of effective brain digital twins.
Architecture and function of the multi-node CRBL MF model
The CRBL MF has been designed to recapitulate the main anatomical and physiological features of the cerebellar cortex–DCN circuit and to simulate cerebellar dynamics. The cerebellar cortex features a unique anisotropic geometry supporting a forward architecture with recurrent excitation and inhibition based on 4 main neuronal populations (GrC, GoC, MLI, and PC). These properties, along with the specific transfer function of neurons, have been incorporated into an MF model providing a biomimetic representation of a cerebellar region, such as a node in a virtual brain29. Multiple such MF models have been interconnected to generate the multi-node CRBL MF model, which has been hardwired with DCN. This mesoscale assembly accounts for parallel fibres that cross the cerebellum on the transverse axis50,51. Moreover, it accounts for inhibitory projections from PC to DCN52 and for feedback projections from DCN to the cerebellar granule layer18,53,54,55. These connections set up intra-cerebellar communication, allowing information to be transmitted in feed-forward and feedback loops. This organisation is instrumental to generate cerebellar cortex activity of GrCs, GoC, MLIs, and PCs in the physiological frequency range, along with the emergence of rhythmic oscillations and coherence in the circuit55. The identity of different cerebellar regions was ensured by the independence of extra-cerebellar inputs. In summary, we observed the following:
-
1.
Average discharge frequency. The oscillation frequency of cerebellar neurons in the multi-node CRBL MF, both in open and in closed loop, was compatible with that observed experimentally39. This ensured that the MF model was working within its natural operating regime.
-
2.
Oscillations. Neuronal discharge showed rhythmic components on the main frequency bands revealed by MEG and EEG recordings in humans with a prevalence of the theta band44. This is an important aspect of validation since the CRBL MF, although tuned on mice data, proved able to capture ensemble properties typical of the human brain29. As an ancestral structure of the hindbrain, the cerebellum features a largely preserved microcircuitry across species, allowing the CRBL MF model to maintain the same architecture for both mice and humans and enabling functional results that can be readily translated across species50,56.
-
3.
Coherence. Oscillations were synchronised across cerebellar regions, revealing a coherence similar to empirical recordings. The cerebellar connectivity and specific neuronal dynamics of the multi-node CRBL MF model support the resting-state cerebellar network revealed by fMRI recordings.
The CRBL MF allowed the analysis and interpretation of physiological signals and their correlation with specific neuronal populations. Indeed, by using the CRBL MF as the generative model, which differentiates neuronal dynamics with a state variable for each population, the BOLD signal of the cerebellar cortex is directly computed from the simulated GrC activity. GrC clusters undergo sharp on-off transitions that are capable of controlling neurovascular coupling through nitric oxide production and diffusion both in the granular and molecular layer48. Conversely, PC activity is almost stationary over the integration time of fMRI. This aspect can impact resting-state measurements (like here), but it would become even more relevant in task-dependent fMRI simulations. Secondly, the PSD can be directly related to PCs and MLIs. PCs and MLIs, given their elongated dendritic arborisation and ordered spatial orientation, generate the signals revealed by MEG and by (the rare) hd-EEG recordings available for the cerebellum57. Conversely, GrCs are organised in closed fields and are unlikely to contribute to these electrical signals remarkably. Therefore, signals recorded from the cerebellum can be connected to the activity of specific cell populations that are more likely to generate them.
The reconstruction and personalisation of intra-cerebellar connectivity
A crucial step to generate a microcircuit-specific MF model of the cerebellum was the curation of the intra-cerebellar SC. Even though tractography represents the gold standard for computing the in vivo human SC and parallel fibres have been observed with histological MRI36,50, detection of parallel fibres pathways in vivo is impractical due to their sub-millimetre size and the specific location in the outermost layer of the cerebellar cortex. Moreover, although MRI tractography may reveal U-fibres connecting adjacent lobules, these are spurious reconstructions of under-resolved bifurcating mossy fibres. Therefore, MRI tractography alone would result in false-negative and false-positive connections between adjacent regions of the cerebellar cortex. Here, we first anticipated the general network connectivity and then inferred the connection weights by implementing an across-scale procedure. At a microscale level, region-specific microcircuit representations of the cerebellar network give the unique opportunity to extract synaptic convergences of parallel fibres for each connection projecting from GrC of one node to GoC, MLI, and PC of transversally connected nodes (e.g. from lobule VI right/left to vermis V as shown in Supplementary Fig. 2). This microscale reconstruction reflects a general circuit organisation that needs to be scaled for the single subjects. To do so, the macroscale connectivity strength between interconnected nodes was calculated by weighting synaptic convergence values by the corresponding cerebellar volumes. In this way, the microscopic organisation of the cerebellar cortical circuit was maintained and made subject-specific. A similar reasoning was applied to connections between the cerebellar cortex and DCN. The corresponding axons travelling from PC to DCN and from DCN to the cerebellar granule layer cannot be detected by MRI tractography. Again, the synaptic connectivity between cerebellar cortex and DCN was taken from microscale representations, using convergence weights that were then scaled for the corresponding regional cerebellar volumes in single subjects. To implement the inhibitory action that PCs exert on DCN, though, the sign of cerebellar cortex–DCN communication was inverted. It should be noted that CRBL MF in cortical nodes and WW models in DCN naturally communicate through spike frequencies. Thus, the cMF-TVB incorporating the multi-node CRBL MF effectively solves the major anatomical and physiological issues of signal communication in the cerebro-cerebellar loop.
Emergent brain rhythms in CMF-TVB simulations
Rhythmicity and multi-node coherence in cerebellar signals emerged on multiple bands during resting-state cMF-TVB simulations, in line with experimental recordings. All MF elements (i.e. GrC, GoC, PC, MLI) showed oscillations on multiple bands, although with some differences. It should be noted that oscillations in an MF element imply synchronisation of the entire neuronal population in the corresponding spiking neuronal network. This population-level detail enables us to separate the contribution of each neuronal type (i.e. MF element) to specific frequency bands, providing a mechanistic understanding of how distinct cerebellar populations shape the cerebellar oscillatory activity.
The theta band was observed both in the granular and molecular layer with a dominant frequency at 6 Hz, which corresponds to the resonance frequency measured in GrCs and GoCs as well as in the whole granular layer45,58,59. In the MF, theta band oscillations are an emergent property of the cerebellar model in response to inputs from the rest of the brain. Interestingly, theta band oscillations have been revealed during control of continuous movements operated by the cerebellar-thalamic-cortical loop44. Moreover, theta band oscillations are associated with memory, opening a new perspective to investigate the cerebellar role in memory encoding and retrieval55,60,61,62.
The alpha band was more visible in GoC and MLI with a dominant frequency at 11 Hz. This is around the spontaneous oscillation frequency of these neurons and is characteristic of the µ-wave associated with sensorimotor activity. The cMF-TVB can thus detect a fundamental correlate of the role of the cerebellum-DCN in motor timing, which indeed occurs around 10 Hz44,63.
The beta band emerged in all cell populations, with a higher contribution in PCs, with a primary frequency of 20 Hz, demonstrated the capability of cMF-TVB to detect higher rhythms emerging in both motor and cognitive functions64,65. Although cMF-TVB simulations are at resting state, the capability of detection of beta band can be used to understand the neuronal mechanisms of each cerebellar population underlining complex cognitive tasks such as semantic prediction61. The gamma band shows the lowest power in MF elements but the highest PSD value, which is centred at 64 Hz, suggesting that the maximum degree of cerebellar activity synchronisation occurs around the middle of the gamma band. The combination of theta and gamma waves is critical for cognitive processing, in which the cerebellum is now known to be involved66,67,68.
The role of the cerebellum as an adaptive filter, refining movements through plasticity-driven learning, had been largely investigated, with recent advancements showing that cerebellar microcircuits contribute to predictive coding and the temporal coordination of neural signals across brain networks69,70,71. Our model, by explicitly preserving the physiological structure of the cerebellar microcircuit, stands as a tool to investigate how these oscillatory dynamics emerge and how specific neuronal populations contribute to functional processes such as movement smoothing and error correction. By resolving the contributions of each MF element to the oscillatory behaviour of the cerebellum separately, the cMF-TVB permits the differentiation of the population-specific mechanisms through which cerebellar rhythms interact with cortical and subcortical circuits. Extending the application of cMF-TVB to pathological states, this model provides a mechanistic framework for investigating pathology-driven disrupted cerebellar rhythms driven by specific neural populations and their propagation across brain networks. Abnormal cerebellar rhythms are implicated in several neurodegenerative and psychiatric conditions. In Parkinson’s disease, reduced alpha band correlates with motor impairment, while altered gamma band synchronisations correlate with the severity of tremor72,73,74. In Parkinson’s disease, a critical role of the cerebellum in cortical beta band oscillations has been suggested75. Furthermore, gamma and theta band alterations are also observed in schizophrenia patients76,77. In dementia and encephalopathy, there are also distinctive EEG signatures (https://emedicine.medscape.com/article/1138235-overview?form=fpf). Therefore, the cMF-TVB embedding multi-node CRBL MF could be used to investigate the physiology and pathophysiology of cortico-cerebellar interactions in resting-state simulations as well as in sensorimotor and cognitive tasks.
Considerations on the performance of simulations using the multi-node CRBL MF
The multi-node CRBL MF surpassed the corresponding models using neural masses in BOLD fMRI simulations, determining a ~50% MAE reduction compared to WW, WC, and GO (both on average and in single subjects). It should be noted that, with the multi-node CRBL MF either in open or in closed loop, neuronal populations operated within their physiological activity range with a dynamical heterogeneity driven by the specific node connectivity. Thus, the maintenance of cerebellar biological properties of neurons and microcircuit connectivity impacted favourably on the resolution of local circuit activity. Interestingly, the CRBL MF also impacted on connected regions, reducing the MAE in DCN and in the whole brain, although nodes outside the cerebellum were modelled with unspecific WW neural masses. It is noteworthy that the performance improvement of whole-brain simulated signals is not driven by any parameter’s optimisation, but solely due to the inclusion of region-specificity in the generative model. This demonstrates the remarkable impact of integrating even a single region-specific model into virtual-brain models on local and whole-brain dynamics. As a next step, a multiparametric optimisation will be performed against macroscale subject-specific recordings, such as BOLD signals, to extract subject-specific physiological information from the fine-tuned parameters of the mesoscale region-specific models78. When applied to pathological states, this approach allows the extraction of a subset of biophysical parameters (i.e. synaptic convergence, quantal synaptic conductance, and synaptic time decay) for each neuronal population within the CRBL MFs associated with the cerebellar regions affected by the pathological condition. As an example, in the case of Parkinson’s Disease, it would be possible to extract the quantal synaptic conductance of PCs, which is modulated by pathological states like dopamine depletion, and influences the propagation of disrupted signals from the cerebellar cortex to DCN. Therefore, it will be possible to explore the response of different neuronal populations to various pathological conditions. Moreover, the importance of synaptic kinetics and neuromodulator effects in shaping network oscillations has been demonstrated79,80,81. Advancing region-specific MFs to incorporate neuromodulator mechanisms provides the computational substrate for evaluating the effects of treatments that modulate neurotransmitter release, with implications for synaptic plasticity at both regional and whole-brain levels.
Since TVB is data-driven, the quality of SC is crucial both for the accuracy and the interpretation of simulations. Thus, any improvements in MRI data or the combination with other datasets (e.g. EEG or MEG) are likely to improve the simulations. We have introduced two elements of novelty: curation of the intra-cerebellar SC using a general architecture derived from anatomy a priori and subject-specific connection weighting using morphological parameters extracted from standard 3DT1 images. This new procedure, which resolves the inability of MRI in detecting details of local network connectivity but still allows for personalisation, could be easily applied to any local networks, improving the digital twin technology. Further improvements are expected from the application of ultra-high field DWI methods, which could enhance the characterisation of intra-cerebellar connectivity.
This work shows that integrating anatomo-physiological properties of a specific brain circuit into large-scale simulations can advance current technology toward the generation of effective brain digital twins7. The use of MF models to substitute specific regional nodes is promising as it improves model performance toward a closer matching with subject data. Moreover, the cMF-TVB allows to identify physiological properties of the circuit, like rhythmic oscillations and coherence, and to improve the identification of region-specific activities. The mechanisms of circuit activity could be directly related to specific neuronal populations, further improving the interpretation of electrophysiological and MRI signals. Although our work is deployed as a component of a larger simulation framework, it constitutes a critical first brick for building a comprehensive brain avatar based on region-specific modelling. Indeed, among possible developments, the reconstruction process reported here for the multi-mode CRBL MF and cMF-TVB could be applied to pathological conditions, in which patient’s circuits could be modelled according to both pathology-specific and subject-specific abnormalities. And by integrating other region-specific MFs28,31,82,83 one can envisage the generation of an all-MF virtual brain that could be used both for resting-state and task-dependent fMRI analysis. Eventually, cMF-TVB models embedding advanced MF of specific brain regions could be used for the investigation of sensorimotor and cognitive processes and to generate virtual twins of patient’s brain in pathological conditions, opening new opportunities for diagnosis and treatment.
Methods
TVB simulates subject-specific brain dynamics at different scales, starting from SC matrices, usually derived by applying tractography to DWI data2. TVB allows simulating the neuronal activity of interconnected regions by resolving computational models of local circuit function in each node. By convolving the resulting neuronal activity with a built-in hemodynamic response function, TVB provides a simulation of the BOLD signal for each node3. Applications of TVB in clinical trials and research studies are growing, and include the improvement of pre-operative planning in epilepsy and the investigation of subject-specific metabolic changes in neurodegenerative diseases4,11,12,13,14,15,16,17.
MRI data were downloaded from the Human Connectome Project (http://db.humanconnectome.org). Cerebral and cerebellar nodes were identified based on an ad-hoc atlas and were used to compute the empBOLD timeseries, from resting state fMRI data (section ‘Dataset and data preparation’), and the whole brain SC, from DWI data.
The workflow is reported in Supplementary Fig. 1, and all the steps are explained in detail in the sections ‘Structural connectivity (SC)’ to ‘Validation’. Briefly:
-
A.
Curation of the cerebellar SC to account for cerebellar-specific connectivity (section ‘Structural connectivity (SC)’). We differentiated parallel fibres axonal pathways, connecting adjacent cerebellar cortex regions, from the mossy fibres pathways, connecting the cerebellar cortex with DCN and/or with the cerebrum.
-
B.
Integration of CRBL MF equations in TVB (sections ‘Multi-node CRBL MF model’ and ‘Construction of the cMF-TVB’). We associated one CRBL MF to each node of the cerebellar SC, and a generic model to DCNs and cerebral nodes. We updated the model equations to simulate the propagation of the neuronal activity from one region to another, and we updated the input/output relations to enable generic models (e.g. WW) receiving inhibitory input coming from the CRBL MF.
-
C.
TVB simulations (section ‘Simulations’). We tested the framework in two conditions: first, the cerebellum in open loop (i.e. isolated cerebellar cortex activity) to assess the performance of the multi-node CRBL MF; then, the cerebellum in closed-loop (i.e. whole brain activity) to assess the performance of the cMF-TVB, including different models for different regions.
-
D.
Validation (section ‘Validation’). Multi-node CRBL MF and cMF-TVB simulations were evaluated in terms of constructive and predictive validity.
Dataset and data preparation
High-quality pre-processed MRI data (Siemens 3T Connectome Skyra MRI scanner with a 32-channel receive head coil) was downloaded from the Human Connectome Project database (https://db.humanconnectome.org/), WU-Minn Consortium (Principal Investigators: David Van Essen and Kamil Ugurbil; 1U54MH091657), funded by the 16 NIH Institutes and Centers that support the NIH Blueprint for Neuroscience Research. Data was used in compliance with the HCP Open Access Data Use Terms84. The dataset included the following imaging data: DWI (1.25 mm isotropic resolution, b = 1000, 2000, 3000 s/mm2, 90 isotropically distributed directions/b-value and 18 b0 images), rs-fMRI (2 mm isotropic resolution, TR/TE = 720/33.1 ms, 1200 volumes) and 3DT1-weighted images (0.7 mm isotropic resolution). We selected the same 8 healthy subjects (2 males and 6 females; 30.6 ± 4.1 years) used in our previous work on the impact of cerebellar connectivity on whole brain dynamic simulations10. Preprocessing of fMRI data follows the steps of the Human Connectome Project Functional ICA-FIX Pipeline. The main steps include motion correction using 24 parameters, distortion corrections and non-aggressive denoising of timeseries. Full details can be found in Glasser et al.85.
Brain parcellation
Brain regions were identified using an already defined ad-hoc atlas including cerebellar parcellations from a spatially unbiased atlas template of the cerebellum and brainstem (SUIT), cerebral parcellations from the automated anatomical labelling atlas, and deep grey matter structures using FSL-first10. Whole-brain parcellations resulted in 126 regions with 27 cerebellar cortical regions, 6 DCN, and 93 cerebral regions. Each grey matter parcellation was considered as a node for the SC extraction and for the assignment of a computational model to run TVB.
Structural connectivity (SC)
For each subject, the SC matrix was computed using the anatomically constrained tractography with 30 million of streamlines implemented in MRtrix386,87. SC weights were defined using probabilistic streamline tractography (second-order integration over fibre orientation distributions) and by assigning each streamline to the nearest node in a 2-mm spherical neighbourhood88. Elimination of spurious ipsilateral cerebro-cerebellar tracts and the selection of contralateral efferent and afferent cerebellar connections were performed from whole-brain tractograms as described in refs. 32,33,49. Directionality of the cerebro-cerebellar loop connections was included in the SC matrix by imposing the assumption that cerebellar efferent streamlines are via the superior cerebellar peduncle, while the cerebellar afferent streamlines are via the middle cerebellar peduncle10. For each subject, the whole-brain SC results in in a 126 × 126 matrix. Moreover, the intra-cerebellar SC was curated to account for the anatomical a priori knowledge on the intra-cerebellar connectivity pathways missed by tractography. The methodological details on the curation of the intra-cerebellar SC are presented in the following section.
Curation of the intra-cerebellar SC
The cerebellar cortex SC was curated to introduce the missing connectivity between adjacent cerebellar cortical regions, which cannot be detected from DWI tractography (Fig. 2). Therefore, we introduced weights driven by known properties of parallel fibres (Fig. 2A). Moreover, to provide a realistic connectivity weight for the cerebellar cortex–DCN loop, we used known properties of PC axonal bundles (Fig. 2B)89,90. These steps are provided in detail below and schematized in Supplementary Fig. 2:
Cerebellar cortex connectivity
Connectivity weights between pairs of adjacent regions, driven by parallel fibres, were quantified with a multiscale approach by extracting the parallel fibres synaptic convergence from a previously validated cerebellar cortex SNN and weighting it with subject-specific morphological constraints40,41,50. Specifically, the synaptic convergence values from GrCs on GoCs, on MLIs, and on PCs were weighted by the volume of each pair of interconnected cerebellar regions to account for inter-subjects’ variability, and then normalised for the intracranial volume to prevent any bias. Volumes of cerebellar regions were computed by overlapping the SUIT atlas with the anatomical 3DT1. The SUIT atlas provides separate parcellations for the Vermis and the right and left lobules, thus the topology of the cerebellar cortex SC was defined for adjacent regions k (with k from I to X): lobule(k) left – vermis(k) and lobule(k) right – vermis(k) (e.g. lobule IX right – vermis IX and lobule IX left – vermis IX; see Supplementary Fig. 2).
Cerebellar cortex–deep cerebellar nuclei connectivity
Connectivity weights from cerebellar cortex regions to DCN, driven by PC axons were quantified as explained in the section ‘Structural connectivity (SC)’ and assigned to the efferent (i.e. from cerebellar cortex to DCN) bundles. Specifically, a streamline was assigned to all the nodes encountered along its whole length88. The weight of afferent bundles (i.e. from DCN to cerebellar cortex) was computed as 10% of the correspondent forward projections90. In order to correctly link the cerebellar cortex with other brain regions, it is important to make a functional consideration: while models of cortical regions are connected to each other through excitatory fibres, similarly to what happens between two adjacent CRBL MF transmitting excitatory signals, each CRBL MF has an inhibitory connection to the DCN52,91. As in TVB, the connectivity between functional models is weighted by the corresponding SC, to capture the inhibitory/excitatory nature of the signals, we multiplied all connections from the cerebellar cortex to the DCN by the value −1.
Functional data
The time-course of the BOLD signal was extracted from resting-state fMRI for each region, resulting in a 14-min signal, namely the empBOLD signal. The empBOLD was used as ground-truth to evaluate the simBOLD signals (section ‘Validation’).
Multi-node CRBL MF model
The multi-node CRBL MF model was implemented by associating a CRBL MF with each node of the cerebellar cortex, to test a network of CRBL MF, i.e. the multi-node CRBL MF model, integrated into the TVB neuroinformatic platform in an open-loop configuration. The architecture of the multi-node CRBL MF consisted in 27 models (i.e. CRBL MF), one per each node of the cerebellar cortical regions parcelled with the SUIT atlas (section ‘Brain parcellation’). Note that DCNs are not included in the multi-node CRBL MF since their neuronal microcircuitry is deeply different from that of the cerebellar cortical regions, and therefore they are represented with a WW instead of a CRBL MF. The structural connections within the multi-model framework were set according to the cerebellar cortical SC (section ‘Curation of the intra-cerebellar SC’).
The fundamental unit of the multi-node framework is the CRBL MF, derived through a multiscale approach that begins at the microscale with the definition of a single-cell model. This model is used to construct an SNN embedding thousands of these cells, representing a micrometric portion of the brain. The process ends with the formulation of an MF model that describes the mesoscopic neuronal population activity, making it suitable for integration into a whole-brain dynamics simulator.
Here, we outline the key methodological steps for developing an MF model that bridges micro- and macroscale dynamics, ultimately enabling the construction of a network of MF models. A detailed validation of the CRBL MF is provided in our previous work29.
Single-cell model
The dynamics of cerebellar neurons were modelled using the extended Generalized Leaky Integrate-and-Fire (E-GLIF) equation, which describes each neuron as a single-compartment system, leading to a point-neuron representation. E-GLIF models the time evolution of membrane potential (Vm) depending on two intrinsic currents to generate slow adaptation (Iadap) and fast depolarisation (Idep)40,41:
where Isyn = synaptic current, Cm = membrane capacitance; τm = membrane time constant; Erev = reversal potential; Ie = endogenous current; kadap and k2 = adaptation constants; k1 = decay rate of Idep.The cell parameters reported in the equations are specific for each cerebellar cortical neuron, i.e. GrC, GoC, PC and MLI (values reported in Supplementary Table 3), resulting in one set of E-GLIF system for each cerebellar neuron type.
Synaptic model
Connecting different single-cell models required the definition of a synaptic model. Specifically, the connections between E-GLIF neurons were modelled as conductance-based synapses. The synaptic current Isyn is defined as follows:
when a spike occurs at tspk, the conductance gsyn changes according to an alpha function:
where \({G}_{{syn}}\) is the conductance weight (nS) and \({\tau }_{{syn}}\) is the synaptic time decay (ms), specific for each synapse type. E-GLIF neuron models (1) are integrated with conductance-based synaptic models (2) to construct the microcircuit, which represents the functional reference of cerebellar spiking activity for CRBL MF development34,92.
Spiking neural network model
Connecting different E-GLIF neurons using the conductance-based synaptic model yields a definition of a cerebellar SNN. Population-specific synaptic parameters (i.e. synaptic convergence, quantal synaptic conductance [nS] and synaptic time decay [ms]), reported in Supplementary Table 2 were defined accounting for the specific synaptic properties for each neuron type. The cerebellar SNN describes a micrometric portion of the cerebellar cortex by including 28615 GrC, 70 GoC, 446 MLI and 99 PC, resulting in ~3 × 104 E-GLIF equations plus and ~1.5 × 106 alpha-shaped conductance-based synapses34.
Derivation of the mean field model
The CRBL MF describes the population activity in terms of firing rate (Hz) resulting in the following 20 differential equations system29:
where \({\nu }_{{\rm{p}}}\) is the activity of population p; \({{\rm{c}}}_{\lambda \eta }\) is the (co)variance between population λ and \(\eta\); T is the time constant, \({N}_{\lambda }\) is the number of cells included in population λ defined in the section ‘Spiking neural network model’, and Fp is the population-specific transfer function with p = {GrC, GoC, MLI, PC}. The mean field formalism used here is based on the derivation of the transfer function (F in the equations) that is a mathematical construct that relates a neuron’s output, expressed as firing rate, to its presynaptic input, also expressed in terms of firing rates.
The population-specific transfer function is derived as a function of the presynaptic activity (\({\nu }_{s}\)) by using a semi-analytic approach, which provides a formulation in terms of average membrane potential (\({\mu }_{V})\), its standard deviation (\({\sigma }_{V})\), its autocorrelation time decay (\({\tau }_{V})\), and a phenomenological threshold (\({V}_{{thre}}^{{eff}})\), modelling the single neuron non-linearities26,27:
The multiplicative factor α also adapts the equations to high input frequency regimes29,93. As an example of population-specific dependency, for p = GrC, then \({F}_{{GrC}}({\nu }_{mossyfibres},{\nu }_{GoC})\) with mossy fibres and GoC carrying the presynaptic excitatory and inhibitory inputs, respectively.
The average membrane potential is calculated as a function of the average synaptic input generated by firing rates. This input determines the average population conductance (\({\mu }_{{Gp}}\)) expressed as:
Where p is the target population, νs is the presynaptic population activity in Hz, gL is the leak conductance for each population and \({\mu }_{{Gs}}\) are presynaptic population-specific conductances defined as:
Ks−p, τs−p, Qs−p are the population-specific synaptic parameters, i.e. synaptic convergence, and quantal conductances of the connection and synaptic decay times for a population s (values are reported in Supplementary Table 2). Therefore, considering p = GrC \({\mu }_{{Gp}}\) = \({K}_{{mossy\; fibers}-{grc}}{Q}_{{mossy\; fibers}-{grc}}{{\tau }}_{{mossy\; fibers}-{grc}}+{K}_{{goc}-{grc}}{Q}_{{goc}-{grc}}{{\tau }}_{{goc}-{grc}}\) + gLp.
The mean input conductance (μG) and the effective and membrane time constant (\({\tau }_{m}^{{eff}}\)) are given by:
and
The standard deviation (σV) and time constant (τV) of voltage fluctuations read as follow:
where \({U}_{s}=\frac{{Q}_{s-p}}{{\mu }_{G}}\left({E}_{s}-{\mu }_{V}\right)\)
The phenomenological threshold is expressed as a linear combination of the membrane potential fluctuations properties:
where \({\tau }_{V}^{N}=\frac{{\tau }_{V}{g}_{{Lp}}}{{C}_{m}}\) (i.e. normalised autocorrelation time), P are the polynomial coefficients fitted on a numerical template extracted from SNN activity (values reported in Supplementary Table 4), and the normalisation values are set as accordingly to our previous work29: \({\mu }_{V}^{0}=-60{\rm{mV}}\), \(\partial {\mu }_{V}^{0}={\rm{mV}}\), \({\sigma }_{V}^{0}=4{\rm{mV}}\), \(\partial {\sigma }_{V}^{0}=6{\rm{mV}}\), \({\tau }_{V}^{N0}=0.5\), \(\partial {\tau }_{V}^{N0}=1\).
Transfer function modifications to build a network of CRBL MFs
In multi-node CRBL MF, the first-order CRBL MF is included with ad-hoc modifications to the population-specific transfer function to develop a network of CRBL MFs interconnected according to the spatial map defined by the cerebellar SC. Each CRBL MF embedded in the multi-node framework was customised as follows, considering that for intra-cerebellar cortical connections, the input of a CRBL MF needs to be driven by the parallel fibres, while the background noise mimicking the input from the cortex and DCN is driven by mossy fibres.
Population-specific transfer functions (F in equations) of GoC, MLI and PC also integrate the input carried by the parallel fibres (pfs in the equations) of the GrC to connected CRBL MF. As an example, from the single-node CRBL MF29 to the multi-node CRBL MF, the PC transfer function was updated from:
to
where \(\nu\) is the presynaptic firing rate. Specifically, \(\nu\)GrC is the activity of the GrC of the same module,\(\nu\)MLI the activity of the MLIs of the same node, and \(\nu\)pfsext is the external activity with respect to the module, i.e. the activity of the GrC population of the adjacent modules driven by parallel fibres.
Following the same rationale, MLI Transfer Function and GoC Transfer Function resulted in:
where \(\nu\)mfsext is the external input carried by mossy fibres (mfs in the equations) projecting from cortical regions and/or DCNs towards GoC. It is worth noting that, although the presynaptic activity input of MLI and PC is coming from the same neuronal population, resulting in a similar form of Eqs. (14) and (15), the parameters regulating the postsynaptic behaviour of MLI and PC are different (Supplementary Table 2), yielding two different transfer functions29,41 (Supplementary Table 4).
The GrC transfer function was invariant since the GrC population receives an external driven only from the mossy fibres:
Input configuration
In TVB the total input c (Hz) for a target node k is computed as the summation of the activity of all the afferent nodes weighted by the SC and resulting in:
where N is the total number of nodes, wkm is the SC weight from node m to the target node k, Γ is the coupling function between node m and target node k and \({\nu }_{k}{,\nu }_{m}\), are the activity of node k and node m, respectively.
It is worth noting that among the coupling functions available in the TVB framework2,3, here, Γ was chosen as a simple scaling factor = 1. Given this, any improvement in TVB performance is related to the specific computational properties of nodes (e.g. a region-specific model) rather than to the coupling between nodes. Activity of the target node m at the time instant t is computed by summing the activity of the target node recorded at the previous time instant \(\tau\) and the input c from afferent nodes:
To differentiate the contribution of mossy fibres and parallel fibres, c was weighted for the mossy fibres convergence and the parallel fibres convergence estimated from a previously validated SNN used to construct the CRBL MF29,34,40,41, resulting in cmf and cpf (Table 4). To account for the convergence heterogeneity across cerebellar neuronal populations (e.g. GrC receives only cMF, while GoC receives both cMF and cpf), cMF and cpf were further split into contributions specific for each population as reported in Table 5.
This estimation was necessary in the open loop configuration only, because the external input was superimposed to all the nodes as a background noise mimicking the input from the cerebrum and/or the DCN. In the closed-loop scenario, this superimposition was not required because the external input from the cerebrum and/or the DCN was simulated.
Construction of the cMF-TVB
The cMF-TVB was implemented considering the cerebellum wired in a closed loop with the DCN and the cerebrum. The CMF-TVB was built by interconnecting 126 models (one per region), divided into 27 CRBL Mossy fibres, corresponding to the cerebellar cortex, and 99 WW models, corresponding to the cerebral cortical and subcortical regions, as well as the DCN20,29. A schema of the inter-model connectivity is reported in Fig. 3. The cMF-TVB inherits the population-specific transfer functions, and the input configurations described in the sections ‘Single-cell model’ and ‘Synaptic model’ for the multi-node CRBL MF. In addition, it requires further modifications to the input/output relations of the CRBL MF (Cerebellar mean field model (CRBL MF)) and the WW models (Wong-Wang model (WW)) to allow for inter-model connections. Notably, despite the different mathematical formulations used for the WW and CRBL MF, both representations generate firing rates within the same frequency range. The two representations are coupled through synaptic projections, ensuring that firing rate remains the only parameter transmitted between cerebral and cerebellar regions.
Cerebellar mean field model (CRBL MF)
Cerebellar MF equations were modified in terms of population-specific transfer function to include the coupling contribution following the same rationale detailed in the section ‘Multi-node CRBL MF model’. With this configuration, the contribution carried by parallel fibres (i.e. from adjacent CBL MF) was separated from that carried by other region nodes (i.e. from a connected WW), resulting in a physiological mapping of the input/output relations. Indeed, the input carried by parallel fibres projecting from GrCs of adjacent nodes became part of the input to GoC, MLI and PC of the target module, while the granular layer of the target module was driven by the input from mossy fibres from the connected DCN and/or cerebral models.
In detail, with this configuration, each population of a CRBL MF could receive an additive ‘external input’ coming from adjacent modules. The coupling is specifically set up according to the origin of the external input: in the case it comes from a cerebellar cortex module to another cerebellar cortex module it goes from the GrC of the source module to the GoC, MLI and PC of the target module; on the other hand, in the case it comes from a DCN or cerebral node (e.g. from a WW model) the input goes to the granular layer of the target cerebellar module, namely to the GrC and GoC. Therefore, in a mathematical form, the coupling towards any cerebellar cortex node can be written as:
where k is the cerebellar target node (e.g. vermis X), wkm is the curated SC weight from cerebellar source node m (e.g. right lobule X), vm(GrC) is the activity of the GrC population in node m, wkj is the SC weight that can be either from a cerebral node or a DCN node (e.g. right fastigial), and Sj(E) is the activity coming from the WW excitatory population of such cerebral or DCN node.
It is worth noting that within the cMF-TVB, the differentiation between mossy and parallel fibres is achieved by segregating the input activity to a cerebellar cortex target node, which is driven separately by parallel and mossy fibres. Consequently, the estimation of mossy fibre parameters and coupling strength (section ‘Synaptic model’) is not required.
Wong-Wang model (WW)
The excitatory population of the WW model (E) receives input both from other Es belonging to other modules of connected WWs, and from PCs of connected CRBL MF. Therefore, it was necessary to modify the WW equation to separate the excitatory contribution from Es and the inhibitory contribution of PCs. The WW coupling expression (couplingj) for a target node j (e.g. the right fastigial), was modified from Deco et al.20 as follows, by adding the inhibitory input from PCs:
where G is the global coupling parameter, JN is the synaptic parameter associated with the NMDA channel, cji(E) is the excitatory activity of cerebral source node i (Si(E)), weighted by the structural weight wji between target node j and source node j. cjk(PC) is the PC activity from the cerebellar source node m (\({\nu }_{{{\rm{k}}}^{({\rm{PC}})}}\)) weighted by the structural weights wjk to target node j from source node k (e.g. to right fastigial nucleus from Vermis X, see Fig. 3). Note that the indicial notation subtends a summation over index i and k.
With the addition of cjk(PC), it is possible to introduce the contribution of a cerebellar-specific inhibitory activity into the WW equations20. Both external excitatory and inhibitory input for a target node k (Ij(E) and Ij(I), respectively) are dependent on specific coupling terms (cjk(E) and cji(PC)):
where WE and Wi are scaling parameters of the overall effective external input (I0 = 0.382 nA), w+ is the local recurrent excitation, JN is the parameter associated to NMDA channel, S denotes the average synaptic gating variable of target node k, Ji is local inhibitory current, Iexternal is the external stimulation for simulating task evoked activity (set at 0 for resting-state simulation), and λ indicates the long-range feed-forward inhibition. The superscripts (E), (I) and (PC) indicate the populations as introduced above.
The coupling term for the cerebellar nodes was differentiated into contribution from parallel and contribution from mossy fibres (cpfs(k−m) and cmfs(k−j) respectively), resulting in the following expression for a cerebellar target node k (e.g. Vermis X), receiving input from a cerebellar source node m (e.g. right lobule X), from a DCN source node j (e.g. right fastigial nucleus), and from a cerebral node (e.g. thalamus):
where m = cerebellar cortical node, and j = DCN and/or cerebral node, and cpfs(k−m) and cmfs(k−j) defined as in section ‘Cerebellar mean field model (CRBL MF)’.
Simulations
Simulations lasting 14 min, with an integration step (dt) of 0.1 ms, were performed to investigate cerebellar activity in both open-loop and closed-loop configurations. Neuronal activity and BOLD signals were simulated for each node following the setup reported in Table 6. To simulate BOLD signals, the TVB-built-in function tvb_analysers_fmri_balloon3 was used, which takes neuronal activity as input and generates a BOLD signal through convolution with the Balloon-Wind Kessel model46.
In the closed-loop simulations, the impact of cerebellar activity on whole-brain dynamics was also analysed.
Multi-node CRBL MF simulations (open loop)
The cerebellar cortex SC was used to map the connections strength between CRBL Mossy fibres constructing the multi-node CRBL MF. Each node of the cerebellar cortex SC was associated with a model, resulting in a network of 27 interconnected models. The cerebellar cortex activity was simulated comparing the performance of generic models already available in TVB, namely WW model, WC model and GO, against the performance of the region-specific CRBL MF, purposely integrated in TVB.
cMF-TVB simulations (closed loop)
Whole-brain SC was used to map the connection strength between models in cMF-TVB. Whole-brain activity was simulated either with a standard virtual brain (TVB), using the WW model for all nodes, or with the cMF-TVB, combining 93 WW models covering the cerebrum and the DCN with 27 CRBL MF models for the cerebellar cortex.
Validation
Multi-node CRBL MF and cMF-TVB simulations were evaluated both in terms of constructive and predictive validity. Moreover, cMF-TVB simulations were used to explore the cerebellar emergent brain rhythm, assessing the contribution of each population to specific frequency bands and their propagation over the cerebella regions. Finally, emergent synchrony amongst the cerebellar nodes was computed in terms of coherence.
Constructive validity
Simulated cerebellar neuronal responses were compared with the physiological ranges of each neuronal population, both in open and in closed loop, to assess whether the constructive validity, already demonstrated for the CRBL MF model itself, was maintained in both the multi-node CRBL Mossy fibres framework and cMF-TVB.
Predictive validity
The TVB platform was applied to simulate BOLD signals using neuronal models as generative models. The ability of the cMF-TVB to simulate BOLD signals using region-specific models as generative sources was quantified by computing the MAE between the empBOLD and the simBOLD signal as the sum of their time point-wise absolute differences, divided by the total number of time points (mean_absolute_error.py of scikit-learn package—python 3.10). Specifically, we computed the overall MAE for each cerebellar cortical region (i.e. regional MAE averaged on subjects) and the subject-specific MAE (i.e. averaged on cerebellar cortical regions). Additionally, we provided a measure of the data dispersion by computing the standard error of the mean (SEM), defined as the ratio between the standard deviation and the square root of the number of observations. For the overall MAE, the corresponding SEM characterised the variability across subjects (i.e. the ratio between across-subject standard deviation and root square of the number of subjects), while for the subject-specific MAE, the corresponding SEM quantified the inter-region variability (i.e. the ratio between across-region standard deviation and root square of the number of regions).
Shapiro–Wilk and Levene’s tests were applied to check the normal distribution of the overall MAE for each model (i.e. CRBL MF, WW, WC and GO), and the homogeneity of variance across models. Based on the outcome of the Shapiro–Wilk and the Levene’s test, Mann–Whitney was selected as the most appropriate statistical test to compare the distributions of the overall MAE between pairs of models. The statistical significance threshold was set to 0.001.
Emergent cerebellar rhythm: a frequency band exploration
Power Spectrum Analysis was implemented on the cerebellar neuronal simulated activities (i.e. the state variable of the CRBL MF) to test whether CRBL MF output variable can provide physiological insight into the cerebellar oscillations at different frequency band, i.e. delta = [0.4 4) Hz, theta = [4 8) Hz, alpha = [8 13) Hz, beta = [12 30) Hz, gamma = [30 100) Hz. Specifically, we investigated the predominant frequency band of the activity of each cerebellar neuronal population and, consequently, the overall cerebellar dominant frequency spectrum. For each subject, the cerebellar population-specific Power Spectrum Density (PSD) was computed with the Fourier transform, using the Welch method to reduce the PSD variance. Sampling frequency was set to 250 Hz. PSD was analysed firstly (i) considering the cerebellum as a unique region, and then (ii) considering the cerebellar cortical regions separately:
(i) For each brain rhythm band, PSD was averaged over the cerebellar regions. The area under the curve (AUC) of each population-specific PSD was computed and normalised for the number of frequencies in each band (AUCnorm), then it was averaged across subjects to quantify the total predominant activity band for each population. A band-specific predominant frequency was then extracted from the average activity across all the cellular populations to identify the cerebellar peak frequency within each band.
(ii) For each brain rhythm, for each region, AUCsnorm were computed to explore the frequency bands propagation. The rate of brain rhythm bands within each region was obtained by summing the population-specific AUCnorm, providing the overall brain rhythm band distributions across cerebellar regions.
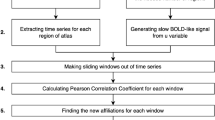
Synchrony in the cerebellar subnetwork
For each subject, the coherence was computed on the CMF-TVB simBOLD, on the TVB simBOLD and on the empBOLD of the cerebellar cortex. For each pair of cerebellar cortical nodes, the coherence was computed as:
where C is the coherence between node i and node j, Pij is the cross-spectral density estimate from the BOLD signal recorded from x and y, Pii and Pjj are the PSD of the BOLD signals of node i and j, respectively. PSDs were computed by applying the Fourier transform to the BOLD signals divided into windows of length = 512, with an overlap of half window length (i.e. 512/2) and a sampling frequency as the TR (0.72 s).
For each subject, coherence matrices for cMF-TVB, TVB and empirical fMRI data were computed by averaging the coherence resulting for each frequency in the spectrum.
To quantify the discrepancy between simulated and empirical data, cMF-TVB and TVB coherence matrices were compared to that resulting from the empirical BOLD using the following scores: (i) MAE to provide a measure of how close the matrices are in terms of coherence strength, (ii) Frobenius Norm to quantify the overall deviation between the two matrices, (iii) cosine similarity to capture the overall difference in the orientation of the two matrices, (iv) KL divergence to compare the distribution of the two matrices, (v) PCC and Spearman correlation coefficient (SCC) to compare the trend of the coherence values. Significant differences were computed using a paired t-test on cMF-TVB vs empirical data and TVB vs empirical data, separately. The significance threshold was set at 0.001.
Data availability
Our pipeline is entirely available, including the curation of the intra-cerebellar SC and the integration and interconnection of the CRBL MF. Implementation of intra-cerebellar SC curation is available at https://github.com/RobertaMLo/tvb_data_preprocessing. Our modeling framework is built as a python package consistently with the AdEx TVB previously implemented and it is available under requests at https://github.com/RobertaMLo/cMF-TVB which includes the cMF-TVB equations and the configuration files for the model-specific parameters. The datasets generated and analyzed during the current study are available on Zenodo in the cMF-TVB dataset (https://zenodo.org/records/15181392).
Abbreviations
- GrC:
-
Granule cells
- GoC:
-
Golgi cells
- MLI:
-
Molecular layer interneurons
- PC:
-
Purkinje cells
- DCN:
-
Deep cerebellar nuclei
- SNN:
-
Spiking neural network
- CRBL MF:
-
Cerebellar mean field model
- WW:
-
Wong Wang model
- WC:
-
Wilson-Cowan model
- GO:
-
Generic Oscillator
- TVB:
-
The Virtual Brain
- cMF-TVB:
-
Cerebellar mean field virtual brain
- SC:
-
Structural connectivity
- fMRI:
-
Functional magnetic resonance imaging
- BOLD:
-
Blood Oxygen Level Dependent (signal)
- empBOLD:
-
empirical BOLD
- simBOLD:
-
simulated BOLD
- DWI:
-
Diffusion-weighted imaging
- MAE:
-
Mean absolute error
- SEM:
-
Standard error of the mean
- PSD:
-
Power spectrum density
- AUC:
-
Area under the curve
- mfs:
-
mossy fibres
- pfs:
-
parallel fibres
References
Arbib, M. A. Spanning the levels in cerebellar function. Behav. Brain Sci. 19, 434–435 (1996).
Sanz Leon, P. et al. The virtual brain: a simulator of primate brain network dynamics. Front. Neuroinform. 7, 385–430 (2013).
Sanz-Leon, P., Knock, S. A., Spiegler, A. & Jirsa, V. K. Mathematical framework for large-scale brain network modeling in The Virtual Brain. Neuroimage 111, 385–430 (2015).
Wang, H. E. et al. Virtual brain twins: from basic neuroscience to clinical use. Natl Sci. Rev. 11, nwae079 (2024).
Breakspear, M. Dynamic models of large-scale brain activity. Nat. Neurosci. 20, 340–352 (2017).
Deco, G., Jirsa, V. K. & McIntosh, A. R. Emerging concepts for the dynamical organization of resting-state activity in the brain. Nat. Rev. Neurosci. 12, 43–56 (2011).
D’Angelo, E. & Jirsa, V. The quest for multiscale brain modeling. Trends Neurosci. xx, 1–14 (2022).
Ritter, P., Schirner, M., Mcintosh, A. R. & Jirsa, V. K. The virtual brain integrates computational modeling and multimodal neuroimaging. Brain Connect 3, 121–145 (2013).
Meier, J. M. et al. Virtual deep brain stimulation: multiscale co-simulation of a spiking basal ganglia model and a whole-brain mean-field model with The Virtual Brain. Exp. Neurol. 354, 114111 (2022).
Palesi, F. et al. The importance of cerebellar connectivity on simulated brain dynamics. Front. Cell. Neurosci. 14, 1–11 (2020).
Monteverdi, A. et al. Subject-specific features of excitation/inhibition profiles in neurodegenerative diseases. Front. Aging Neurosci. 1–17. https://doi.org/10.3389/fnagi.2022.868342 (2022).
El Houssaini, K., Bernard, C. & Jirsa, V. K. The Epileptor Model: a systematic mathematical analysis linked to the dynamics of seizures, refractory status epilepticus, and depolarization block. eNeuro 7:ENEURO.0485-18.2019 (2020).
Monteverdi, A. et al. Virtual brain simulations reveal network-specific parameters in neurodegenerative dementias. Front. Aging Neurosci. 15, 1–15 (2023).
Zimmermann, J. et al. Differentiation of Alzheimer’s disease based on local and global parameters in personalized Virtual Brain models. NeuroImage Clin. 19, 240–251 (2018).
Triebkorn, P. et al. Brain simulation augments machine-learning-based classification of dementia. Alzheimer’s Dement.8, e12303 (2022).
Stefanovski, L. et al. Linking molecular pathways and large-scale computational modeling to assess candidate disease mechanisms and pharmacodynamics in Alzheimer’s disease. Front. Comput. Neurosci. 13, 1–27 (2019).
Stefanovski, L. et al. Bridging scales in Alzheimer’s disease: biological framework for brain simulation with the virtual brain. Front. Neuroinform. 15, 630172 (2021).
D’Angelo, E. Physiology of the Cerebellum. Handbook of Clinical Neurology Vol. 154 (Elsevier B.V., 2018).
Wong, K. F. & Wang, X. J. A recurrent network mechanism of time integration in perceptual decisions. J. Neurosci. 26, 1314–1328 (2006).
Deco, G. et al. How local excitation-inhibition ratio impacts the whole brain dynamics. J. Neurosci. 34, 7886–7898 (2014).
Wilson, H. R. & Cowan, J. D. A mathematical theory of the functional dynamics of cortical and thalamic nervous tissue. Kybernetik 13, 55–80 (1973).
Stefanescu, R. A. & Jirsa, V. K. A Low dimensional description of globally coupled heterogeneous neural networks of excitatory and inhibitory neurons. PLOS Comput. Biol. 4, e1000219 (2008).
Jirsa, V. K. et al. The virtual epileptic patient: individualized whole-brain models of epilepsy spread. Neuroimage 145, 377–388 (2017).
Jansen, B. H. & Rit, V. G. Electroencephalogram and visual evoked potential generation in a mathematical model of coupled cortical columns. Biol. Cybern. 73, 357–366 (1995).
Zerlaut, Y. & Destexhe, A. Enhanced responsiveness and low-level awareness in stochastic network states. Neuron 94, 1002–1009 (2017).
El Boustani, S. & Destexhe, A. A master equation formalism for macroscopic modeling of asynchronous irregular activity states. Neural Comput. 21, 46–100 (2009).
Zerlaut, Y., Chemla, S., Chavane, F. & Destexhe, A. Modeling mesoscopic cortical dynamics using a mean-field model of conductance-based networks of adaptive exponential integrate-and-fire neurons. J. Comput. Neurosci. 44, 45–61 (2018).
Goldman, J. S. et al. A comprehensive neural simulation of slow-wave sleep and highly responsive wakefulness dynamics. Front. Comput. Neurosci. 16 (2023).
Lorenzi, R. M. et al. A multi-layer mean-field model of the cerebellum embedding microstructure and population-specific dynamics. PLoS Comput. Biol. 19, e1011434 (2023).
Tesler, F. et al. Multiscale modeling of neuronal dynamics in hippocampus CA1. Front Comput Neurosci. 18, 1432593 (2024).
Overwiening, J., Tesler, F., Guarino, D. & Destexhe, A. A multi-scales study of thalamic state-depemndent responsivness. PLoS Comput. Biol. 20, e1012262 (2024).
Palesi, F. et al. Contralateral cerebello-thalamo-cortical pathways with prominent involvement of associative areas in humans in vivo. Brain Struct. Funct. 220, 3369–3384 (2015).
Palesi, F. et al. Contralateral cortico-ponto-cerebellar pathways reconstruction in humans in vivo: implications for reciprocal cerebro-cerebellar structural connectivity in motor and non-motor areas. 1–13 https://doi.org/10.1038/s41598-017-13079-8 (2017).
De Schepper, R. et al. Model simulations unveil the structure-function-dynamics relationship of the cerebellar cortical microcircuit. Commun. Biol. https://doi.org/10.1038/s42003-022-04213-y (2022).
Ito, M. Cerebellar circuitry as a neuronal machine. Prog. Neurobiol. 78, 272–303 (2006).
Dell’Acqua, F., Bodi, I., Slater, D., Catani, M. & Modo, M. MR diffusion histology and micro-tractography reveal mesoscale features of the human cerebellum. Cerebellum 12, 923–931 (2013).
Lundell, H. & Steele, C. J. Cerebellar imaging with diffusion magnetic resonance imaging: approaches, challenges, and potential. Curr. Opin. Behav. Sci. 56, 101353 (2024).
Habas, C. & Manto, M. Chapter 14—Probing the neuroanatomy of the cerebellum using tractography. in The Cerebellum: From Embryology to Diagnostic Investigations Vol. 154 (eds Manto, M. & Huisman, T. A. G. M.) 235–249 (Elsevier, 2018).
Geminiani, A. et al. Mesoscale simulations predict the role of synergistic cerebellar plasticity during classical eyeblink conditioning. PLOS Comput. Biol. 20, 1–29 (2024).
Geminiani, A. et al. Complex dynamics in simplified neuronal models: reproducing Golgi cell electroresponsiveness. Front. Neuroinform. 12, 1–19 (2018).
Geminiani, A., Casellato, C., D’Angelo, E. & Pedrocchi, A. Complex electroresponsive dynamics in olivocerebellar neurons represented with extended-generalized leaky integrate and fire models. Front. Comput. Neurosci. 13, 1–12 (2019).
Eccles, J. C. & Masao Ito, J. S. The Cerebellum as a Neuronal Machine Vol. 12 (Springer Berlin Heidelberg, 1975).
De Zeeuw, C. I., Koppen, J., Bregman, G. G., Runge, M. & Narain, D. Heterogeneous encoding of temporal stimuli in the cerebellar cortex. Nat. Commun. 14 (2023).
Gross, J. et al. The neural basis of intermittent motor control in humans. PNAS 99, 2299–2302 (2002).
Gandolfi, D., Lombardo, P., Mapelli, J., Solinas, S. & D’Angelo, E. Theta-frequency resonance at the cerebellum input stage improves spike-timing on the millisecond time-scale. Front. Neural Circuits 7, 1–16 (2013).
Stephan, K. E., Weiskopf, N., Drysdale, P. M., Robinson, P. A. & Friston, K. J. Comparing hemodynamic models with DCM. Neuroimage 38, 387–401 (2007).
Gagliano, G. et al. Non-linear frequency dependence of neurovascular coupling in the cerebellar cortex implies vasodilation–vasoconstriction competition. Cells 11 (2022).
Mapelli, L. et al. Granular layer neurons control cerebellar neurovascular coupling through an NMDA receptor/NO-dependent system. J. Neurosci. 37, 1340–1351 (2017).
Palesi, F. et al. Reconstructing contralateral fiber tracts: methodological aspects of cerebello-thalamocortical pathway reconstruction. Funct. Neurol. 31, 229–238 (2016).
Wyatt, K. D., Tanapat, P. & Wang, S. S.-H. Speed limits in the cerebellum: constraints from myelinated and unmyelinated parallel fibers. Eur. J. Neurosci. 21, 2285–2290 (2005).
Heck, D. & Sultan, F. Cerebellar structure and function: making sense of parallel fibers. Hum. Mov. Sci. 21, 99–109 (2002).
Ito, M., Yoshida, M., Obata, K., Kawai, N. & Udo, M. Inhibitory control of intracerebellar nuclei by the Purkinje cell axons. Exp. Brain Res. 10, 64–80 (1970).
Mapelli, J. & D’Angelo, E. The spatial organization of long-term synaptic plasticity at the input stage of cerebellum. J. Neurosci. 27, 1285–1296 (2007).
D’Angelo, E. & De Zeeuw, C. I. Timing and plasticity in the cerebellum: focus on the granular layer. Trends Neurosci. 32, 30–40 (2009).
De Zeeuw, C. I., Lisberger, S. G. & Raymond, J. L. Diversity and dynamism in the cerebellum. Nat. Neurosci. 24, 160–167 (2021).
Sepp, M. et al. Cellular development and evolution of the mammalian cerebellum. Nature 625, 788–796 (2024).
Samuelsson, J. G., Sundaram, P., Khan, S., Sereno, M. I. & Hämäläinen, M. S. Detectability of cerebellar activity with magnetoencephalography and electroencephalography. Hum. Brain Mapp. 41, 2357–2372 (2020).
Hoffmann, L. C. & Berry, S. D. Cerebellar theta oscillations are synchronized during hippocampal theta-contingent trace conditioning. Proc. Natl Acad. Sci. USA106, 21371–21376 (2009).
D’Angelo, E. et al. Theta-frequency bursting and resonance in cerebellar granule cells: experimental evidence and modeling of a slow K+-dependent mechanism. J. Neurosci. 21, 759–770 (2001).
Herweg, N. A., Solomon, E. A. & Kahana, M. J. Theta oscillations in human memory. Trends Cogn. Sci. 24, 208–227 (2020).
Dave, S., VanHaerents, S. & Voss, J. L. Cerebellar theta and beta noninvasive stimulation rhythms differentially influence episodic memory versus semantic prediction. J. Neurosci.40, 7300–7310 (2020).
Andreasen, N. C. et al. The cerebellum plays a role in conscious episodic memory retrieval. Hum. Brain Mapp. 8, 226–234 (1999).
Llinás, R., Lang, E. J. & Welsh, J. P. The cerebellum, LTD, and memory: Alternative views. Learn. Mem. 3, 445–455 (1997).
Kaethler, L. B., Brown, K. E., Meehan, S. K. & Staines, W. R. Investigating cerebellar modulation of premovement beta-band activity during motor adaptation. Brain Sci. 13 (2023).
Andersen, L. M. & Dalal, S. S. The role of the cerebellum in timing. Curr. Opin. Behav. Sci. 59, 101427 (2024).
Nguyen, V. T. et al. Distinct cerebellar contributions to cognitive-perceptual dynamics during natural viewing. Cereb. Cortex 27, 5652–5662 (2017).
Popa, D. et al. Functional role of the cerebellum in gamma-band synchronization of the sensory and motor cortices. J. Neurosci. 33, 6552–6556 (2013).
Courtemanche, R., Robinson, J. C. & Aponte, D. I. Linking oscillations in cerebellar circuits. Front. Neural Circuits 7, 1–16 (2013).
Hull, C. Prediction signals in the cerebellum: beyond supervised motor learning. eLife 9, e54073 (2020).
Dean, P., Porrill, J., Ekerot, C.-F. & Jörntell, H. The cerebellar microcircuit as an adaptive filter: experimental and computational evidence. Nat. Rev. Neurosci. 11, 30–43 (2010).
Marr, D. A theory of cerebellar cortex. J. Physiol. 202, 437–470 (1969).
Stoffers, D. et al. Increased cortico-cortical functional connectivity in early-stage Parkinson’s disease: an MEG study. Neuroimage 41, 212–222 (2008).
Brittain, J.-S. & Brown, P. Oscillations and the basal ganglia: motor control and beyond. Neuroimage 85, 637–647 (2014).
Wu, T. & Hallett, M. The cerebellum in Parkinson’s disease. Brain 136, 696–709 (2013).
Mazzoni, P. et al. Differential components of bradykinesia in Parkinson’s disease revealed by deep brain stimulation. J. Neurophysiol. 132, 870–878 (2024).
Faris, P., Pischedda, D., Palesi, F. & D’Angelo, E. New clues for the role of cerebellum in schizophrenia and the associated cognitive impairment. Front. Cell. Neurosci. 18, 1–24 (2024).
Uhlhaas, P. J. & Singer, W. Abnormal neural oscillations and synchrony in schizophrenia. Nat. Rev. Neurosci. 11, 100–113 (2010).
Ziaeemehr, A., Woodman, M., Domide, L. & Petkoski, S. Virtual Brain Inference (VBI): a flexible and integrative toolkit for efficient probabilistic inference on virtual brain models. 1–50 (2025).
Ursino, M., Cona, F. & Zavaglia, M. The generation of rhythms within a cortical region: analysis of a neural mass model. Neuroimage 52, 1080–1094 (2010).
Wendling, F., Bartolomei, F., Bellanger, J. J. & Chauvel, P. Epileptic fast activity can be explained by a model of impaired GABAergic dendritic inhibition. Eur. J. Neurosci. 15, 1499–1508 (2002).
Robinson, P. A., Rennie, C. J., Rowe, D. L., O’Connor, S. C. & Gordon, E. Multiscale brain modelling. Philos. Trans. R. Soc. B Biol. Sci. 360, 1043–1050 (2005).
Tesler, F., Kozlov, A., Grillner, S. & Destexhe, A. A multiscale model of striatum microcircuit dynamics. eLife 2023.12.28.573546. https://doi.org/10.7554/eLife.99402.1 (2024).
Tesler, F. et al. Multiscale modeling of neuronal dynamics in hippocampus CA1. Front. Comput. Neurosci. 18 (2024).
Van Essen, D. C. et al. The WU-Minn Human Connectome Project: an overview. Neuroimage 80, 62–79 (2013).
Glasser, M. F. et al. The minimal preprocessing pipelines for the Human Connectome Project. Neuroimage 80, 105–124 (2013).
Jeurissen, B., Tournier, J.-D., Dhollander, T., Connelly, A. & Sijbers, J. Multi-tissue constrained spherical deconvolution for improved analysis of multi-shell diffusion MRI data. Neuroimage 103, 411–426 (2014).
Tournier, J.-D. et al. MRtrix3: a fast, flexible and open software framework for medical image processing and visualisation. Neuroimage 202, 116137 (2019).
Smith, R. E., Tournier, J.-D., Calamante, F. & Connelly, A. The effects of SIFT on the reproducibility and biological accuracy of the structural connectome. Neuroimage 104, 253–265 (2015).
D’Angelo, E. Cerebellar granule cell. In Handbook of the Cerebellum and Cerebellar Disorders (eds Manto, M., Schmahmann, J. D., Rossi, F., Gruol, D. L. & Koibuchi, N.) 765–791 https://doi.org/10.1007/978-94-007-1333-8_31 (Springer Netherlands, 2013).
Ito M. Cerebellar circuitry as a neuronal machine. Prog Neurobiol. 78, 272–303 (2006).
Ouardouz, M. & Sastry, B. R. Mechanisms underlying LTP of inhibitory synaptic transmission in the deep cerebellar nuclei. J. Neurophysiol. 84, 1414–1421 (2000).
Geminiani, A., Pedrocchi, A., D’Angelo, E. & Casellato, C. Response dynamics in an olivocerebellar spiking neural network with non-linear neuron properties. Front. Comput. Neurosci. 13, 1–15 (2019).
Carlu, M. et al. A mean-field approach to the dynamics of networks of complex neurons, from nonlinear Integrate-and-Fire to Hodgkin-Huxley models. J. Neurophysiol. 123, 1042–1051 (2020).
Acknowledgements
This research has received funding from the European Union’s Horizon 2020 Framework Program for Research and Innovation under Framework Partnership Agreement No. 650003 (HBP FPA) to ED, CGWK. CGWK received funding from BRC (#BRC704/CAP/CGW), MRC (#MR/S026088/1), Ataxia UK, MS Society (#77). This research has also received funding from Centro Fermi project “Local Neuronal Microcircuits” to ED. We acknowledge the use of Fenix Infrastructure resources, which are partially funded from the European Union’s Horizon 2020 research and innovation program through the ICEI project under the grant agreement No. 800858. This research was supported by the EBRAINS research infrastructure, funded from the European Union’s Horizon 2020 Framework Program for Research and Innovation under the Specific Grant Agreement No. 945539 (Human Brain Project SGA3). CC received fundings from CN1 CN00000013, Programma di ricerca CN00000013 “National Centre for HPC, Big Data and Quantum Computing”, finanziato dal Decreto Direttoriale di concessione del finanziamento n.1031 del 17.06.2022 a valere sulle risorse del PNRR MUR – M4C2 – Investimento 1.4 - Avviso “Centri Nazionali” - D.D. n. 3138 del 16 dicembre 2021. Further support was provided by Digital Europe Grant TEF-Health #101100700 and #NEXTGENERATIONEU (NGEU), funded by the Ministry of University and Research (MUR). The Virtual Brain Twin Project received funding from Horizon Europe under grant agreement No. 101137289. This work was also supported by the Italian Ministry of Research through the PNRR projects funded by the European Union – NextGenerationEU: “A multiscale integrated approach to the study of the nervous system in health and disease” (Project PE0000012, CUP F13 C2200124007, “MNESYS”) to CGWK and “European Brain Research Infrastructure—Italy” (Project IR0000011, CUP B51E22000150006, “EBRAINS-Italy”) to ED and EBRAINS2.0 (GA 101147319 — EBRAINS 2.0) to ED. The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
E.D. and C.G.W.K. conceptualised the study. R.L. designed the pipeline and performed the analyses. R.L. and E.D. wrote the paper. F.P. and C.C. provided support and guidance with data analysis. All the authors reviewed the submitted version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lorenzi, R.M., Palesi, F., Casellato, C. et al. Region-specific mean field models enhance simulations of local and global brain dynamics. npj Syst Biol Appl 11, 66 (2025). https://doi.org/10.1038/s41540-025-00543-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41540-025-00543-9