Abstract
COVID-19 vaccine adaptation is critical to respond to continuously emerging SARS-CoV-2 variants with enhanced immune evasion. The ARVAC protein subunit vaccine, based on the receptor binding domain of the spike protein of SARS-CoV-2, has been adapted to XBB.1.5 and JN.1 variants, as monovalent and bivalent formulations. Preclinical studies in mice showed that ARVAC XBB.1.5 and JN.1 monovalent vaccines induced strong neutralizing antibodies against XBB and JN.1 lineages, though with limited efficacy against phylogenetically distant variants. By contrast, bivalent formulations combining Gamma antigen with either XBB.1.5 or JN.1 antigens demonstrated superior cross-neutralizing activity, covering variants from Ancestral to JN.1. Additionally, Gamma-containing bivalent vaccines elicited neutralizing antibodies against SARS-CoV-1, highlighting their potential for broad-spectrum immunity. Cellular immune studies confirmed robust CD4+ T cell activation across all formulations. These findings support the continued adaptation of ARVAC to current circulant variants and propose ARVAC bivalent vaccines containing the Gamma antigen as a strategy for induction of pan-sarbecovirus immunity.
Similar content being viewed by others
Introduction
COVID-19 remains a threat to public health as SARS-CoV-2 virus continues to evolve resulting in multiple variants that increasingly evade immunity acquired through vaccination and natural infection. Since the emergence of Omicron lineage in late 2021 several subvariants have developed. The emergence of Omicron XBB-sublineages and then the new JN.1 lineage has dramatically reduced the efficacy of both SARS-CoV-2 wild-type monovalent and bivalent (wild-type + Omicron BA.5) mRNA and protein subunit vaccines1,2.
The reduced immunity due to ongoing virus evolution has highlighted the need for update vaccine antigen composition. In the fall of 2023, the US Food and Drug Administration (FDA) authorized the use of updated COVID-19 vaccines from Moderna, Pfizer-BioNTech, and Novavax which incorporate a monovalent component targeting the XBB.1.5 variant3. Data from these manufacturers indicate that monovalent XBB-containing COVID-19 vaccines enhance the immune response against XBB-sublineage variants4,5. In 2024, JN.1 lineages become predominant around the world. However, several reports demonstrate that XBB.1.5-adapted COVID-19 vaccines are less effective against JN.1 in comparison to XBB lineages2,6,7. In May 2024 new recommendations from FDA and EMA supported the change to antigen composition to JN.1 lineage8. The JN.1 lineage circulation was subsequently being replaced by JN.1 sublineages including KP.2, KP.3 and XEC. Given the rapid emergence of subsequent variants with the parallel chance to immune escape, development of a broad vaccine formulation that can targets several SARS-CoV-2 variants as well as SARS-CoV-1 is urgent. We have proof of concept results showing that the addition of the Gamma antigen to a bivalent formulation increases the breadth of the induced neutralizing antibody (nAb) response9.
Protein-based vaccines, particularly those targeting the receptor-binding domain (RBD) of SARS-CoV-2, have gained significant attention due to their favorable safety profile and ability to elicit a targeted and effective immune response. Notably, several subunit COVID-19 vaccines utilizing the RBD of the SARS-CoV-2 spike protein have been approved for use in different countries (Soberana plus, ZF2001, Covaccine and Bimervax)10,11,12 Consequently, numerous studies have explored strategies to further enhance the immunogenicity and efficacy of RBD-based vaccines13.
ARVAC is a recombinant protein subunit COVID-19 vaccine based on a dimeric receptor binding domain (RBD) from SARS-CoV-2. The bivalent ARVAC vaccine, based on the Gamma and Omicron BA.4/5 SARS-CoV-2 variants, received authorization for use in Argentina in October 2023 following a phase 3 study that confirmed a broad immune response against Omicron lineages. Preclinical and phase 1 studies supported the selection of Gamma RBD as vaccine antigen, as it broadened the nAb responses across different variants of concern (VOC)9,14. ARVAC has been developed by an Argentinian pharmaceutical company being the first COVID-19 vaccine fully developed in South America. According to recommendations from the WHO, FDA, and EMA, the ARVAC vaccine has been updated to include monovalent and bivalent formulations containing XBB.1.5 and JN.1 derived antigens. To our knowledge, among approved subunit RBD based vaccines, ARVAC is the only that has been adapted to the JN.1 variant, and this version is currently approved for use in Argentina. This work presents the evaluation of updated ARVAC immunogenicity in preclinical studies.
Results
Adaptation of RBD antigen to XBB.1.5 and JN.1 in monovalent and bivalent vaccine formulations
Production of the recombinant Ancestral or Gamma variant RBD antigens has been previously shown9. In this work new RBD antigens derived from Omicron BA.4/5, Omicron XBB.1.5 and JN.1 SARS-CoV-2 variants were expressed in CHO cells using the same previously reported methodology. Each RBD antigen consists of a single-chain dimeric repeats of amino acids 319R to 537K of the Spike protein of the corresponding SARS-CoV-2 variant. Antigen purity was confirmed by SDS-PAGE (Supplementary Fig. 1). The binding of the Omicron BA.4/5, XBB.1.5 and JN.1 RBD versions to the hACE2 receptor was assessed using a ligand-receptor binding ELISA. All antigens demonstrated comparable binding levels to hACE2 (Supplementary Fig. 1). The purified antigens were then adsorbed to aluminum hydroxide (Alum) to make the final formulations.
Adapted ARVAC XBB.1.5 monovalent and bivalent vaccines induce neutralizing antibodies against XBB and JN.1 lineages in primary vaccination schemes
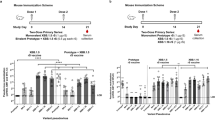
First, XBB.1.5-adapted vaccines were compared to earlier Omicron BA.4/5 vaccine versions. Naïve mice received a two-dose primary immunization scheme with either monovalent vaccines (XBB.1.5 or Omicron BA.4/5) or bivalent vaccines (Gamma + XBB.1.5 or Gamma + Omicron BA.4/5). Serum samples were collected 28 days after the second dose and analyzed for antigen specific binding IgG and SARS-CoV-2 nAbs.
All vaccine groups elicited robust specific IgG responses against XBB.1.5, Omicron BA.4/5, Gamma and Ancestral derived RBD antigens (Supplementary Fig. 2). nAb titers against JN.1, XBB.1.18, Omicron BA.5, Gamma and Ancestral were assessed using a live virus assay. The geometric mean of nAb titers (GMTs) indicated that neutralizing activity against XBB lineage were higher in animals that received either a monovalent or a bivalent vaccine containing the XBB.1.5 antigen (GMT 127.5 and 66.1, respectively) than in those animals that received a monovalent or a bivalent vaccine containing the Omicron BA.4/5 antigen (GMTs 7.3 and 11.4 respectively, Fig. 1a). Mice vaccinated with bivalent formulations containing the Gamma antigen induced strong neutralizing responses against both the Gamma and Ancestral variants. In contrast, mice vaccinated with monovalent vaccines targeting XBB.1.5 or Omicron BA.4/5 exhibited low neutralizing antibody titers against the Ancestral and Gamma viruses. Notably, all vaccine formulations elicited nAbs against the genetically divergent JN.1 variant, being the monovalent and bivalent formulations containing XBB.1.5 the versions that induced the highest levels (Fig. 1a).
BALB/c mice were i.m. immunized on days 0 and 14 with ARVAC XBB.1.5 (n = 6), ARVAC Omicron BA.4/5 (n = 6), ARVAC Gamma/Omicron BA.4/5 (n = 6) or ARVAC Gamma/XBB.1.5 (n = 6). a Neutralizing antibody titers against JN.1, XBB.1.18, Omicron BA.5, Gamma and ancestral SARS-CoV-2 were evaluated at 42 days post prime (dpp). Neutralization titer was defined as the serum dilution that reduces 50% the cytopathic effect (NT50). Bars are GMT ± SD. *p < 0.05, ***p < 0.001. Kruskal–Wallis test. b Radar charts were drawn based on the geometric mean titers (GMTs) of nAbs against live viruses at 42 dpp. c Antigenic map was generated from the nAb titers in serum after the second dose of vaccination. Squares represented nAb titers from mice vaccinated with XBB.1.5 (red squares), Omicron BA.4/5 (green squares), Gamma/BA.4/5 (blue squares) and Gamma/XBB.1.5 (purple squares) formulations. Black, blue, green, red and purple circles correspond to the Ancestral, Gamma, Omicron BA.5, XBB.1.18 and JN.1 SARS-CoV-2 variants, respectively. Each grid square corresponds to a two-fold dilution in the neutralization assay. The antigenic distance is interpretable in any direction. d Splenocytes from immunized mice were obtained 1 month after last dose and were stimulated with medium or Omicron BA.4/5 RBD-peptides pool plus recombinant Omicron BA.4/5 RBD for 18 h and then brefeldin A was added for 5 h. Afterward, cells were harvested and stained with anti-CD4 specific Abs, fixed, permeabilized, and stained intracellularly with anti-IFN-γ and anti-TNF-α. Results are presented as percentage of IFN-γ or TNF-α-producing CD4+ T lymphocytes. Bars are means ± SEM. Each dot is an individual mouse. *p < 0.05, **p < 0.01 vs. placebo. Kruskal–Wallis. e Splenocytes from immunized mice were stimulated with complete medium or recombinant Omicron BA.4/5 RBD for 5 d and then IL-5 was measured in the supernatant by ELISA. Results are presented as concentration of IL-5 in pg/ml. Bars are means ± SEM. Each dot is an individual mouse. *p < 0.05, **p < 0.01 vs. medium. Kruskal–Wallis. f Lung cells from immunized mice were stimulated with medium or recombinant Omicron BA.4/5 RBD for 3 d and then IL-5, IFN-γ and TNF-α was measured in the supernatant by ELISA. Results are presented as concentration of the cytokine in pg/ml. Bars are means ± SEM. Each dot is an individual mouse. *p < 0.05, **p < 0.01 vs. medium. Kruskal–Wallis.
Spider graphs in Fig. 1b further illustrate that vaccine formulations containing XBB.1.5 are more effective at inducing nAbs against XBB and JN.1 lineages, while those bivalent versions containing the Gamma antigen showed a good performance neutralizing Ancestral and Gamma SARS-CoV-2 viruses. Of note, bivalent Gamma/XBB.1.5 vaccine demonstrated the greatest cross-neutralizing activity covering distant variants from Ancestral to JN.1 lineages (Fig. 1b). Antigenic maps were created to visualize the relationships between the sera and the nAbs against SARS-CoV-2 variants. The positioning of each serum sample on the maps is indicative of its efficacy in neutralizing a particular variant. One grid line on the antigenic map corresponds to a twofold dilution in the virus neutralization assay. Results obtained after antigenic distance analysis show that XBB.1.5 monovalent vaccine is closely to JN.1 and Omicron BA.5 subvariants and more distant to variants like Ancestral or Gamma (Fig. 1c). Interestingly, the bivalent Gamma/XBB.1.5 vaccine demonstrated the broadest activity, effectively reducing the antigenic distance between the Ancestral or Gamma variants and the XBB or JN.1 lineages (Supplementary Table 1).
T cell responses against Omicron BA.4/5 RBD antigen were evaluated following ARVAC vaccination in the spleen and lung of immunized mice. In spleen, RBD specific-IFN-γ and -TNF-α producing CD4+ T cells were induced 1 month after administration of monovalent (XBB.1.5 or Omicron BA.4/5) and bivalent (Gamma/XBB.1.5 or Gamma/Omicron BA.4/5) ARVAC vaccines (Fig. 1d). Antigen-specific CD8+ T cells responses were not observed after splenocytes stimulation (Supplementary Fig. 3). Additionally, significant levels of IL-5 were observed after antigenic stimulation of splenocytes from mice immunized with any vaccine version (Fig. 1e).
In addition, mucosal/lung T cell responses were evaluated. Consistent with systemic findings, IFN-γ and TNF-α levels were significantly increased in antigen-stimulated lung cells compared to unstimulated controls in all ARVAC vaccine formulations (Fig. 1f). Although IL-5 production was detected upon antigen stimulation in cells from all vaccinated groups, a statistically significant increase was only observed in the bivalent ARVAC Gamma/Omicron BA.5 group. These results suggest that ARVAC formulations induce predominantly a Th1-biased immune response at lungs following immunization.
ARVAC adapted XBB.1.5 monovalent and bivalent vaccines as boosters induce nAbs against XBB and JN.1 variants
The adapted ARVAC vaccine versions were evaluated as booster doses. To this end, mice were first primed intramuscularly with two doses of a bivalent mRNA Wuhan/Omicron BA.5 based vaccine (Spikevax) and 2 months later (day 67) boosted with either a dose of a monovalent or a bivalent ARVAC vaccine or an additional dose of Spikevax. Serum samples were collected on days 67 (before booster—2 months after primary series—) and 97 (4 weeks after booster dose) to evaluate nAb responses.
A booster dose with the monovalent ARVAC XBB.1.5 vaccine significantly increased nAb titers against JN.1 (9.8-fold), XBB.1.18 (14.7-fold), and Omicron BA.5 (14.6-fold). However, it did not enhance nAb titers against the Gamma and Ancestral SARS-CoV-2 variants compared to pre-booster levels (Fig. 2a). The monovalent ARVAC Omicron BA.4/5 vaccine exhibited a similar profile but with lower efficacy, it induced increases in nAb titers against JN.1 (6.8-fold), XBB.1.18 (4.6-fold), and Omicron BA.5 (6-fold) but it did not enhance nAb titers against the Gamma and Ancestral SARS-CoV-2 variants compared to pre-booster titers.
BALB/c mice were i.m. immunized on days 0 and 14 with bivalent mRNA vaccine Spikevax (Prototype/Omicron BA.5) and boosted on day 67 with: ARVAC XBB.1.5 (n = 6), ARVAC Omicron BA.4/5 (n = 6), ARVAC Gamma/Omicron BA.4/5 (n = 6), ARVAC Gamma/XBB.1.5 (n = 6) or Spikevax (Prototype/Omicron BA.5) (n = 6). a Neutralizing-antibody titers against XBB.1.18, JN.1, Omicron BA.5, Gamma and Ancestral SARS-CoV-2 were evaluated at 63 (preboost) and 93 dpp (postboost). Neutralization titer was defined as the serum dilution that reduces 50% the cytopathic effect (NT50). Bars are GMT ± SD. *p < 0.05, ***p < 0.001. Kruskal–Wallis test. b Radar charts were drawn based on the GMTs of nAb in serum pre and post boost against live viruses. c Antigenic maps were generated from nAb titers in serum pre and after the booster dose. Squares represented nAb titers from mice vaccinated with XBB.1.5 (red squares), Omicron BA.4/5 (green squares), Gamma/BA.4/5 (blue squares) and Gamma/XBB.1.5 (purple squares) formulations. Black, blue, green, red and purple circles correspond to the Ancestral, Gamma, Omicron BA.5, XBB.1.18 and JN.1 variants, respectively. Each grid square corresponds to a two-fold dilution in the neutralization assay. The antigenic distance is interpretable in any direction.
Interestingly, only bivalent vaccines containing the Gamma antigen elicited a broader cross-reactive Ab response, exhibiting high nAb titers not only against Ancestral and Gamma variants but also against phylogenetical distant variants such as Omicron BA.5 and JN.1 (Fig. 3a). Radar charts of nAb titers highlighted superior cross-neutralization capacity of the bivalent Gamma/XBB.1.5 vaccine formulation compared to monovalent XBB.1.5 and bivalent Gamma/Omicron BA.5 vaccines (Fig. 2b).
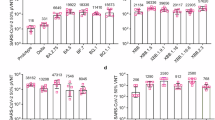
Titers of nAbs to SARS-CoV-1 (a) and SARS-CoV-2 (b) virus by PSVNA at day 93 (28 days post boost) after administration of ARVAC vaccines. Positive control in a is a human IgG1 anti SARS-CoV-2 Spike CR3022 antibody and the positive control in b is the International Standard (NIBSC, code: 21/340). Bars are GMT ± SD. The GMT value is indicated inside bars. *p < 0.05, ***p < 0.001. Kruskal–Wallis test.
Neutralizing antibody titers obtained before and after booster doses were used to construct antigenic maps. Analysis of antigenic distances revealed a reduction in the distances between XBB.1.18 and JN.1 across nearly all vaccinated groups, with the greatest reduction observed in the monovalent XBB.1.5 group (from 4.3 AU pre boost to 1.8 AU post boost, Fig. 2c and Supplementary Table 2). Furthermore, antigenic maps indicate that the newly emergent variants XBB.1.18 and JN.1 are clustered together but remain distant from Ancestral, Gamma and Omicron BA.5 variants following booster doses.
ARVAC bivalent vaccine formulations induce cross-reactive antibodies against SARS-CoV-1
Finally, to assess the broadening of the nAb response, serum samples were analyzed for nAbs against SARS-CoV-1 using a pseudovirus neutralization assay (PSVNA). Notably, boosting the animals with monovalent Omicron BA.4/5 and bivalent ARVAC vaccine formulations containing Gamma resulted in significative nAb titers against SARS-CoV-1 in comparison to pre boosting titers (two doses of Spikevax—bivalent Wuhan/Omicron BA.5—, Fig. 3a). SARS-CoV-2 neutralization titers were determined in the same assay, showing that monovalent Omicron BA.4/5 and bivalent ARVAC vaccine formulations also induced significant nAbs against Wuhan SARS-CoV-2 (Fig. 3b).
ARVAC monovalent and bivalent adapted JN.1 vaccines induce nAbs against JN.1 lineages
Following international regulatory agencies recommendations, ARVAC was adapted to JN.1 variant and its immunogenicity was tested. Naïve BALB/c mice received a two-dose primary immunization series with either monovalent (JN.1 or XBB 1.5), bivalent (Gamma + JN.1 or Gamma + Omicron BA.4/5) or mRNA XBB.1.5 (Spikevax) vaccines.
Neutralizing activity was assessed in sera against a broad panel of SARS-CoV-2 variants, ranging from the Ancestral strain to the JN.1 lineage, including the KP.2 and KP.3 sublineages. The monovalent JN.1-adapted vaccine elicited significantly higher levels of nAbs against KP.3.1.1, KP.2.3, and JN.1 compared to the monovalent XBB.1.5 and bivalent Gamma/Omicron BA.4/5 vaccines, and comparable levels to the bivalent Gamma/JN.1 vaccine (Fig. 4a). Notably, the bivalent Gamma/JN.1 vaccine elicited higher nAb levels against Omicron BA.5, Gamma and the Ancestral variants than the monovalent JN.1-adapted formulation, demonstrating robust coverage across all tested viral variants (Fig. 4a).
BALB/c mice were immunized at day 0 and day 14 via i.m. with: ARVAC JN.1 (n = 6), ARVAC XBB.1.5 (n = 6), ARVAC Gamma/BA.4/5 (n = 6), ARVAC Gamma/JN.1 (n = 6) and Spikevax XBB.1.5 mRNA vaccine. a Neutralizing antibody titers against KP.3.1.1, KP.2.3, JN.1, XBB.1.5, Omicron BA.5, Gamma and Ancestral SARS-CoV-2 were evaluated at 42 dpp. Neutralization titer was defined as the serum dilution that reduces 50% the cytopathic effect (NT50). Bars are GMT ± SD. *p < 0.05, ***p < 0.001. Kruskal–Wallis test. b Radar charts were drawn based on the GMTs of nAb in serum 42 dpp against live viruses. c Splenocytes from immunized mice were obtained a month after last immunization and stimulated with medium or JN.1 RBD-peptides pool plus recombinant JN.1 RBD for 18 h and then brefeldin A was added for 5 h. Afterward, cells were harvested and stained with anti-CD4 and anti-CD8 specific Abs, fixed, permeabilized, and stained intracellularly with anti-IFN-γ and anti-TNF-α. Results are presented as percentage of IFN-γ or TNF-α-producing CD4+ or CD8+ T lymphocytes. Bars are means ± SEM. Each dot is an individual mouse. *p < 0.05, **p < 0.01, ***p < 0.001 vs. placebo. Kruskal–Wallis. d Splenocytes from immunized mice were stimulated with medium or recombinant JN.1 RBD for 3 d and then IL-5 was measured in the supernatant by ELISA. Results are presented as concentration of IL-5 in pg/ml. Bars are means ± SEM. Each dot is an individual mouse. *p < 0.05, ***p < 0.001 vs. placebo. Kruskal–Wallis. e Lung cells obtained from immunized mice were stimulated with medium or recombinant JN.1 RBD for 3 days and then IFN-γ, TNF-α and IL-5 were measured in the supernatant by ELISA. Results are presented as cytokine concentration in pg/ml. Bars are means ± SEM. Each dot is an individual mouse. *p < 0.05, **p < 0.01 vs. placebo. Kruskal–Wallis.
The breadth of the immune response is further illustrated in the spider plots of Fig. 4b, which highlight the broad neutralizing capacity of the bivalent Gamma/JN.1 vaccine against phylogenetically diverse SARS-CoV-2 variants.
Furthermore, T cell immune responses were determined in spleen and lung cells from immunized mice after stimulation with JN.1 antigen. At systemic level (spleens), ARVAC JN.1, XBB.1.5 and Gamma/JN.1 vaccines induced significantly higher frequencies of CD4+ T cells producing IFN-γ and TNF-α, as compared to placebo (Fig. 4c). ARVAC Gamma/Omicron BA.4/5 and Spikevax XBB.1.5 vaccines did not induce RBD-JN.1 specific CD4+ T effector cells. Regarding CD8+ T cell responses, immunization with monovalent JN.1 and XBB.1.5 vaccines induced a significant increase in IFN-γ+ CD8+ T cells compared to placebo while the other groups did not (Fig. 4c). Besides, IL-5, a marker of Th2-type responses, was detected in all vaccinated groups, supporting also the presence of a Th2 response (Fig. 4d).
At lungs, a T helper response with significative production of IFN-γ and IL-5 was observed in RBD-JN.1 stimulated cells from mice immunized with monovalent JN.1, bivalent Gamma/BA.4/5 or Gamma/JN.1 ARVAC vaccines and Spikevax XBB.1.5 in comparison to placebo group (Fig. 4e). These data demonstrate that ARVAC-adapted vaccines, especially those including JN.1 and Gamma antigens, elicit strong antigen specific Th1-Th2 cellular immune responses at both systemic and local/mucosal levels.
ARVAC adapted JN.1 monovalent and bivalent vaccines as boosters induce neutralizing antibodies against JN.1, KP2, KP.3 and XEC variants
Adapted JN.1 ARVAC versions were also evaluated as booster doses. To this end, mice were first primed intramuscularly with two doses of the mRNA XBB.1.5 based vaccine (Spikevax) and 2 months later (day 75) boosted with either a dose of a monovalent or bivalent ARVAC vaccine or an additional dose of Spikevax XBB.1.5. Serum samples were collected on days 75 (before booster—2 months after last dose of the primary series—) and 103 (4 weeks after booster dose) to evaluate nAb responses.
A booster dose with the monovalent ARVAC JN.1 vaccine significantly increased nAb titers against JN.1 (11.3-fold), KP.2 (126.8-fold), KP.3 (12.3-fold) and XEC (71.5-fold) variants. However, it did not enhance nAb titers against distant variants like XBB.1.5, Omicron BA.5, Gamma or Ancestral SARS-CoV-2 compared to pre-booster levels (Fig. 5a). Of note, bivalent vaccine Gamma/JN.1 elicited a broader cross-reactive response, exhibiting high nAb titers against phylogenetical distant variants such as Ancestral, Gamma and Omicron in addition to JN.1, KP and XEC lineages (Fig. 5a). The radar charts in Fig. 5b illustrate the cross-neutralizing capacity of the bivalent vaccine, highlighting the broader neutralization profile of the Gamma/JN.1 formulation compared to both monovalent vaccines and the bivalent Gamma/Omicron BA.4/5 formulation.
BALB/c mice were i.m. immunized on days 0 and 14 with mRNA XBB.1.5 vaccine (Spikevax) and boosted on day 75 with: ARVAC JN.1 (n = 6), ARVAC XBB.1.5 (n = 6), ARVAC Gamma/Omicron BA.4/5 (n = 6) and ARVAC Gamma/JN.1 (n = 6) or Spikevax XBB.1.5 (n = 6). a Neutralizing-antibody titers against XEC, KP.2, KP.3, JN.1, XBB.1.5, Omicron BA.5, Gamma and Ancestral SARS-CoV-2 were evaluated at 75 (preboost) and 103 dpp (postboost). Neutralization titer was defined as the serum dilution that reduces 50% the cytopathic effect (NT50). Bars are GMT ± SD. *p < 0.05, **p < 0.01, ***p < 0.001. Kruskal–Wallis test. b Radar charts were drawn based on the geometric mean titers (GMTs) of neutralizing antibody in serum pre and post boost against live viruses for each vaccinated group.
Discussion
The constant emergence of SARS-CoV-2 variants and sublineages that evade neutralizing antibodies induced upon infection, vaccination or both requires the use of adapted vaccines. A vaccine adapted to the Omicron XBB.1.5 variant became available in Europe in September 2023 and in September 2024 a new adapted vaccine based in the JN.1 or KP.2 lineage has been released to the market following the recommendations for updating authorized/approved vaccines given by the WHO, EMA, and FDA15.
Adaptation of ARVAC vaccine to new variants has been shown during preclinical and clinical development of the vaccine where two monovalent versions (Gamma and Omicron BA.4/5) and a bivalent version were tested9,14,16. Due to its superior performance in eliciting broader protective nAbs, the bivalent formulation (Gamma + Omicron BA.4/5) received approval17. Following regulatory agency recommendations, the vaccine was subsequently updated to monovalent formulations targeting XBB.1.5 and JN.1 variants. In addition, based on previous results showing that addition of Gamma antigen increases the breadth of nAb responses9, in this work the bivalent counterparts containing Gamma antigen plus either the XBB.1.5 or the JN.1 antigen were tested and compared to the monovalent versions. We demonstrated that immunization with either monovalent or bivalent recombinant RBD protein adjuvanted with alum (ARVAC vaccine) adapted to the XBB.1.5 or JN.1 variants induce higher nAbs against XBB and JN.1 lineages compared to the previous approved bivalent Gamma/Omicron BA.4/5 ARVAC version. These results support the change in the vaccine strain composition to target the most current circulating variant following the recommendations of international regulatory agencies.
Monovalent XBB-adapted ARVAC vaccine formulations as primary or booster doses significantly increase nAb titers against the XBB lineage compared to the current bivalent Gamma/BA.4/5 vaccine (Fig. 1a). Indeed, the XBB.1.5 adapted vaccine generates neutralizing antibodies against the JN.1 variant. Similarly, the monovalent JN.1-adapted ARVAC vaccine in primary immunization schedules enhances GMT against JN.1, KP.2.3, and KP.3.1.1 variants compared to the monovalent XBB.1.5 and bivalent Gamma/BA.4/5 vaccines (Fig. 4a). Of note, monovalent and bivalent JN.1-adapted ARVAC vaccine as booster doses of a primary regimen with an XBB.1.5 mRNA vaccine increases the nAbs not only against JN.1 but also against KP.2, KP.3 and XEC subvariants (Fig. 5a). The low cross-neutralization observed in mice for the antigenically distant strain XBB.1.5 with the monovalent and bivalent JN.1 adapted vaccines was also reported for adapted JN.1 and KP.2 mRNA vaccines18.
Notably, the bivalent Gamma/XBB.1.5 and Gamma/JN.1 vaccines generate a robust and broad cross-reactive response, neutralizing not only newly circulant variants as JN.1 and XBB.1.5 but also distant variants such as Omicron BA.5, Gamma, and Ancestral variants, highlighting the potential of incorporating the Gamma antigen to enhance antibody breadth against SARS-CoV-2 variants (Figs. 1b, 4b and 5b).
Moreover, ARVAC variant-adapted vaccines evaluated in this study, elicited robust Th1-Th2 cell responses. Indeed, one month after the final immunization with either monovalent or bivalent XBB.1.5- or JN.1-adapted vaccines, a mixed Th1/Th2 cell response was induced at both systemic and local levels (Figs. 1d–f and 4c–e). Particularly, the local immune responses observed at lungs may contribute to enhanced protection by limiting viral replication in the respiratory tract and reducing disease severity19,20. Of note, T cell responses were also elicited upon stimulation with RBD-based antigens even if they are distinct from those included in the vaccine formulations, indicating a conserved immune response. These results are consistent with previous reports with other variants, where antigenic changes have slight impact on pre-existing T cell-mediated immunity21,22.
Development of a universal vaccine against SARS-CoV-2 and SARS-related coronaviruses is needed to reduce the threat from current and future SARS-related coronaviruses with pandemic potential23. Our previous study has demonstrated that the Gamma antigen can broaden the immune response by inducing nAbs against different variants, likely due to the presence of more immunogenic epitopes in the RBD9. Similar results have been shown with a Beta variant subunit vaccine, which shares the same aminoacid substitutions in its sequence (N501Y, E484K, and K417N)24. Our results with bivalent vaccines (Gamma/XBB.1.5 and Gamma/JN.1) confirm that the Gamma antigen in a bivalent vaccine enhances the breadth of the immune response in immunized animals.
Notably, our results with the bivalent Gamma/XBB.1.5 vaccine demonstrate that immunization with the ARVAC vaccine can induce neutralizing antibodies with cross-reactive capacity against other related viruses, such as SARS-CoV-1 (Fig. 3). Instead, low cross-reactivity against SARS-CoV-1 has been reported following immunization with updated COVID-19 vaccines such as BNT162b2 XBB.1.525. A different strategy, including antigens from MERS-related and SARS-related zoonotic, epidemic, and pandemic coronaviruses showed to induce a broad immune response protecting against Sarbecoviruses and Merbecoviruses in animals26. The ARVAC bivalent vaccine, containing Gamma antigen, has demonstrated broad-spectrum activity against distant SARS-CoV-2 variants and SARS-CoV-1 proposing it as a very strong candidate for developing a pan-sarbecovirus vaccine. The development of a pan-sarbecovirus vaccine is critical for global health, as it can target a broad spectrum of coronaviruses within the Sarbecovirus subgenus, including SARS-CoV and several SARS-CoV-2 variants. A limitation of this study is the absence of long-term immunogenicity data, that can evaluate the durability of immune responses induced by monovalent and bivalent formulations.
Several studies have shown that updated monovalent vaccines reduce the risk of severe outcomes from infection with SARS-CoV-2 Omicron subvariants like hospitalization and death27,28. Updating COVID-19 vaccines to better match circulating strains is essential for boosting relevant immunity and maintaining effectiveness against a range of serious clinical outcomes. The results from ARVAC adapted vaccines support the use of monovalent vaccines to enhance immune responses to new circulating variants. Additionally, our study also explores the use of multivalent vaccines containing Gamma antigen as a strategy for develop a pan-sarbecovirus vaccine.
Methods
Cloning of antigens
Constructs from Gamma, Omicron BA.4/5, XBB.1.5 and JN.1 SARS-CoV-2 variants of the antigen encoding a single-chain dimer of the receptor binding domain (RBD) comprising amino acids 319R to 537K of the Spike protein were used. The signal peptide sequence from the SARS-CoV-2 Spike protein was used for protein secretion. Coding regions for all used final expression vectors were confirmed by nucleotide sequencing (Macrogen, Korea). The constructs were codon-optimized for CHO cell expression and synthesized by GenScript (Hong Kong Limited, Japan). All constructs were cloned into the pJC3UMCS-4 expression vector (Pablo Cassara Foundation) comprising a CMV promoter and a cis-acting sequence for minimized host gene silencing. Linearized forms of the vectors were transfected into CHO-S cells by lipofection.
After 15–20 days of G418 antibiotic selection, end point dilution procedure was applied for clone isolation. Best clone candidates were selected by specific and volumetric productivity assessed by ELISA using cells culture supernatants. Recombinant protein relative abundance was confirmed by SDS-PAGE and Coomassie Brilliant Blue G-250 staining. Initial antigen was obtained from supernatants from shake flask cultures (from selected polyclones and/or isolated clones). Culture medium used for antigen production was serum free.
Pilot scale manufacture of antigens and vaccines
Pilot batches of antigen and vaccines were manufactured by Laboratorio Pablo Cassara, Argentina. Controlled cell substrates were used to inoculate a 2-L bioreactor with 1.4 L as working volume or a 200-L single use bioreactor with 50–75 L as working volume. An alternate tangential flow system (ATF, Repligen) was used for perfusion. The harvest after 20–25 days was used as starting material for a downstream process based on three chromatographic steps: first an affinity mix-mode capture chromatography step followed by two different ionic chromatography to eliminate residual host cell DNA and host cell proteins and other process-related impurities. TFF (Tangential flow filtration) was used to adjust pH, conductivity, and protein concentrations as needed between successive chromatographic steps and to obtain the concentrated solution of the antigens. RBD samples were run under reducing and non-reducing conditions by SDS-PAGE (10% polyacrylamide). The samples were visualized by staining with Coomassie Brilliant Blue G-250.
ACE2 binding to RBD analyzed by titration ELISA
The hACE-2 receptor was immobilized to ELISA plates (Greiner Bio One F-Bottom, High Binding, Cat: 655061) by incubation with hACE-2 solution (1 μg/ml) overnight at 4 °C. The next day was blocked with 5% free fat milk in PBS/Tween 20 0.05% at 37 °C for 1 h. Serial dilutions of RBD Omicron BA.4/5, XBB.1.5 and JN.1 were added to the plates and incubated for 2 h at 37 °C. Then plates were washed and incubated for 2 h at 37 °C with in-house anti-RBD rabbit polyclonal antibody. After extensively washing plates were incubated for 1 h at 37°C with anti-rabbit goat polyclonal antibody conjugated with peroxidase (Dako cat. P0448). Signal was detected after adding TMB substrate solution (OptEIA, BD, Cat:555214). The reaction was stopped with 10% sulfuric acid solution and the absorbance is measured at 450 nm in Microplate Reader (BIORAD iMark). Three independent assays were performed per sample and the average EC50 was calculated using the GraphPad Prism 9.0 program.
Adjuvant and formulations
Pilot batches of vaccines were manufactured using aluminum hydroxide as adjuvant. Aluminum hydroxide 2% was supplied by CRODA (Croda International plc, United Kingdom). The antigen was adsorbed onto the adjuvant by mixing. Free antigen was controlled to be less than 10% of total antigen in the vaccine. Endotoxin levels of vaccine formulations were determined by (Limulus amebocyte lysate endochrome kit, Endosafe U.S. License No. 1197). All vaccine formulations contained <10 EU/50 µg RBD.
Ethics statement
All experimental protocols with animals were conducted in strict accordance with international ethical standards for animal experimentation (Helsinki Declaration and its amendments, Amsterdam Protocol of welfare and animal protection and National Institutes of Health, NIH USA, guidelines). The protocols performed at UNSAM were also approved by the Institutional Committee for the use and care of experimental animals (CICUAE) from National University of San Martin (UNSAM) (26/2024).
Animals and immunizations
Eight-week-old female BALB/c mice were obtained from IIB-UNSAM animal facility. In prime immunization experiments, animals were intramuscularly (i.m) inoculated at days 0 and 14 with (i) Omicron XBB.1.5 RBD (10 µg) + Alum (100 µg) (N = 6), (ii) Omicron BA.4/5 RBD (10 µg) + Alum (100 µg) (N = 6), (iii) Gamma RBD (5 µg) + Omicron BA.4/5 RBD (5 µg) + Alum (100 µg) (N = 6), (iv) Gamma RBD (5 µg) + Omicron XBB.1.5 RBD (5 µg) + Alum (100 µg) (N = 6) and (vi) placebo (Alum alone, N = 6). In other experiment, animals were intramuscularly (i.m) inoculated at days 0 and 14 with (i) JN.1 RBD (10 µg) + Alum (100 µg) (N = 6), (ii) XBB.1.5 RBD (10 µg) + Alum (100 µg) (N = 6), (iii) Gamma RBD (5 µg) + Omicron BA.4/5 RBD (5 µg) + Alum (100 µg) (N = 6), (iv) Gamma RBD (5 µg) + JN.1 RBD (5 µg) + Alum (100 µg) (N = 6) and (vi) placebo (Alum alone, N = 6). Blood was collected on days 28 and 42 post prime immunization. Intramuscular injections and blood collection via the submandibular route were performed by trained personnel using a precise and rapid technique without anesthesia. Animals were sacrificed at day 42 post prime immunization and spleens or lungs were harvested. Euthanasia was performed using Ketamine + Xylazine (80 mg/kg ketamine + 16 mg/kg Xylazine) administered by intraperitoneal route followed by cervical dislocation to minimize animal suffering.
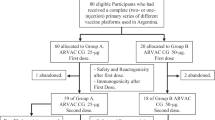
In booster experiments, 8-week-old female BALB/c mice were i.m inoculated on days 0 and 14 with mRNA bivalent Wuhan/Omicron BA.5 or with monovalent XBB.1.5 vaccines (Spikevax). Two months later a booster dose was administered with ARVAC adapted vaccines (N = 6) or Spikevax (bivalent Wuhan/Omicron BA.5 or with monovalent XBB.1.5, N = 6). Blood was collected on day 75 post prime immunization. Intramuscular injections and blood collection via the submandibular route were performed by trained personnel using a precise and rapid technique without anesthesia. Euthanasia was performed using Ketamine + Xylazine (80 mg/kg ketamine + 16 mg/kg Xylazine) administered by intraperitoneal route followed by cervical dislocation to minimize animal suffering.
Viruses
Ancestral SARS-CoV-2 (B.1, GISAID Accession ID EPI_ISL_16290469), Gamma (P.1, GISAID Accession ID EPI_ISL_15807444), Omicron BA.5 (GISAID Accession ID EPI_ISL_16297058), Omicron XBB.1.18 (GISAID ID EPI_ISL_18746184), Omicron XBB.1.5 (GISAID ID EPI_ISL_19035679), JN.1 (GISAID Accession ID: EPI_ISL_19037293), KP.2.3 (GISAID ID EPI_ISL_19364512), KP.3.1.1 (GISAID ID EPI_ISL_19364512) and XEC.15.11 (GISAID EPI_ISL_19506555) were isolated from nasopharyngeal specimens at the Instituto de Investigaciones Biotecnológicas (IIB-UNSAM) and adapted in Vero E6 cultures. Nasopharyngeal swab samples of XBB.1.18 were kindly provided by Dario Alvarez from Centro Rossi (Buenos Aires City, Argentina). Nasopharyngeal swab samples of JN.1, KP.2.3 and KP.3.1.1 were kindly provided by Eugenia Ibañez from Hospital Alemán (Buenos Aires City, Argentina) and Dr. Viegas (Public Health Laboratory, National University of La Plata, Buenos Aires, Argentina).
SARS-CoV-2 neutralization assay
Serum samples were heat-inactivated at 56 °C for 30 min. Serial dilutions were performed and then incubated for 1 h at 37 °C in the presence of 300 TCID50 of SARS-CoV-2 in DMEM 2% FBS. One hundred μl of the mixtures were then added onto Vero cells monolayers. After 72 h at 37 °C and 5% CO2, cells were fixed with PFA 4% (4 °C overnight) and stained with crystal violet solution in methanol. The cytopathic effect (CPE) of the virus on the cell monolayer was assessed by surface scanning at 585 nm in a microplate reader (FilterMAx F5 Microplate reader, Molecular Devices, San Jose, CA, USA). Average optical density (OD) of each well was used for the calculation of the percentage of neutralization of viral CPE for each sample as: Neutralization% = 100 × (DOsample − DOvirus control) / (DOcell control − DOvirus control). Non-linear curves of Neutralization (%) vs. Log 1/sera dilution were fitted to obtain the titer corresponding to the 50% of neutralization (NT50).
Antigenic cartography
The R package Racmacs (https://acorg.github.io/Racmacs/index.html) was used to create antigen cartography maps from serum neutralization titers against the SARS-CoV-2 live viruses (Gamma, Omicron BA.5, Omicron XBB.1.8 and JN.1 variants and the ancestral strain Wuhan). Antigenic distances were measured in antigenic units (AU). One AU corresponds to a two-fold dilution of the antibody in the neutralization assay. Each square in the map indicates 1 AU. The antigenic distance is measured in any direction of the map.
SARS-CoV-1 and SARS-CoV-2 pseudotyped lentivirus neutralization assay
SARS-CoV-1 and SARS-CoV-2 neutralization assay was performed using lentiviral particles expressing SARS-CoV-1 or SARS-CoV-2 spike protein on their surface and containing firefly luciferase reporter gene for detection of infection as previously described29,30. Briefly, two-fold serial dilutions of plasma samples, positive control and negative control were incubated with the Spike SARS-CoV-1 or SARS-CoV-2 pseudotypes (1 × 108 relative luminescence units (RLU)/well). Human IgG1 anti SARS-CoV-2 Spike CR3022 (The Native Antigen Company, code: MAB12453-100) was used as positive control for SARS-CoV-1 PSVNA tests. The second World Health Organization (WHO) International Standard for anti-SARS-CoV-2 immunoglobulin 2022 (NIBSC, code: 21/340) was used as positive control in SARS-CoV-2 PSVNA assays. Serum minus IgA/IgM/IgG human (Sigma-Merck, code: S5393) was used as negative control of the PSVNA assays.
The plates containing the pseudotype-serum mixture were incubated at 37 °C with 5% CO2 for 1 h. At the end of the incubation, HEK293T-ACE2-TMPRSS2 stable cell line (NIBSC, code: 101008) (1 × 105 cells/well) were added to the plate. Pseudotype-only and cell-only wells were included as controls. After 72 h, 50 μL of Bright-GloTM Luciferase Assay System reagent were added to each well and the plate was incubated for 5 min at 400 rpm in the dark (Fisherbrand, code: 88861024). The plates were read using a luminometer (Promega, Glomax Discover).
The GraphPad Prism version 8.4 package (GraphPad Software, GraphPad, 2365 Northside Dr, Suite 560, San Diego, CA 92108, USA) was used to analyze the pseudotype-based neutralization assays. Titers were first normalized based on the pseudotype-only mean and the cell-only mean, and inhibitory concentration 50% (IC50) values were calculated by non-linear regression model analysis (log (inhibitor) vs. normalized response variable slope). Titers were then expressed as the dilution at which the IC50 value fell. Samples with IC50 > 20 were considered positive and ≤20 as negative31.
Determination of T cell immune responses
In prime immunization experiments, 4 weeks after the second dose mice were sacrificed to study cellular immune responses. Splenocytes and lung cells were obtained and cultured for 3 or 5 days in the presence of complete medium or recombinant RBD antigen (Omicron BA.4/5 or JN.1 as detailed in the legends). Then, supernatants were collected and cytokines (TNF-α, IFN-γ and IL-5) were measured by ELISA (Biolegend. San Diego, CA). For intracellular cytokine determination: splenocytes were cultured (4×106 cells/well) in the presence of stimulus medium (complete medium supplemented with anti-CD28 and anti-CD49d) or Ag stimuli (stimulus medium + Omicron BA.4/5 or JN.1 RBD-peptides pool (JPT Peptide Technologies GmbH, Berlin) + recombinant Omicron BA.4/5 or JN.1 RBD protein as detailed) for 18 h. Next, brefeldin A was added for 5 h to the samples. Cells were then washed, fixed, permeabilized, stained, and analyzed by flow cytometry. The cells were stained with Viability dye (Zombie Acqua), anti-mouse-CD8a Alexa Fluor 488, anti-mouse-CD4 Alexa Fluor 647, anti-TNF-α PeCy7 and anti-IFN-γ PE (Biolegend, San Diego, CA). BD Fortessa LSR-X20 cytometer with DIVA Software were used for analysis.
Statistical analysis
Statistical analysis and plotting were performed using GraphPad Prism 8 software (GraphPad Software, San Diego, CA). In experiments with more than two groups, data were analyzed using one-way ANOVA with Kruskal–Wallis or Bonferroni post-test. When necessary, a logarithmic transformation was applied prior to the analysis to obtain data with a normal distribution. In experiments with two groups, an unpaired t test or Mann–Whitney U test was used. A p value < 0.05 was considered significant. When exact p values are not shown we used the following reference: *p < 0.05, **p < 0.01 and ***p < 0.001. When bars were plotted, results were expressed as means ± SEM or GMT ± SD for each group.
Data availability
The data that support the findings of this study are not openly available due to reasons of sensitivity, but may be obtained from the corresponding author upon reasonable request. The data are stored in controlled-access facilities at the Instituto de Investigaciones Biotecnológicas, Universidad de San Martín.
References
Wang, Q. et al. Alarming antibody evasion properties of rising SARS-CoV-2 BQ and XBB subvariants. Cell 186, 279–286.e278 (2023).
Bennett, C. et al. Immunogenicity and safety of a bivalent (Omicron BA.5 plus ancestral) SARS-CoV-2 recombinant spike protein vaccine as a heterologous booster dose: interim analysis of a phase 3, non-inferiority, randomised, clinical trial. Lancet Infect. Dis. 24, 581–593 (2024).
WHO. Statement on the antigen composition of COVID-19 vaccines. https://www.who.int/news/item/26-04-2024-statement-on-the-antigen-composition-of-covid-19-vaccines (2023).
Patel, N. et al. XBB.1.5 spike protein COVID-19 vaccine induces broadly neutralizing and cellular immune responses against EG.5.1 and emerging XBB variants. Sci. Rep. 13, https://doi.org/10.1038/s41598-023-46025-y (2023).
Modjarrad, K. et al. Preclinical characterization of the Omicron XBB.1.5-adapted BNT162b2 COVID-19 vaccine. NPJ Vaccines 9, 229 (2024).
Tartof, S. Y. et al. Effectiveness of BNT162b2 XBB vaccine against XBB and JN.1 sublineages. Open Forum Infect. Dis. 11, ofae370 (2024).
Ma, K. C. et al. Effectiveness of updated 2023-2024 (Monovalent XBB.1.5) COVID-19 vaccination against SARS-CoV-2 Omicron XBB and BA.2.86/JN.1 lineage hospitalization and a comparison of clinical severity-IVY network, 26 Hospitals, October 18, 2023-March 9, 2024. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciae405 (2024).
European Medical Association Emergency Task Force: EMA confirms its recommendation to update the antigenic composition of authorised COVID-19 vaccines for 2024-2025. https://www.ema.europa.eu/en/documents/other/ema-confirms-its-recommendation-updateantigenic-composition-authorised-covid-19-vaccines-2024-2025_en.pdf (2024).
Coria, L. M. et al. A gamma-adapted subunit vaccine induces broadly neutralizing antibodies against SARS-CoV-2 variants and protects mice from infection. Nat. Commun. 15, https://doi.org/10.1038/s41467-024-45180-8 (2024).
Dai, L. et al. Efficacy and safety of the RBD-dimer-based Covid-19 vaccine ZF2001 in adults. N. Engl. J. Med. 386, 2097–2111 (2022).
Toledo-Romani, M. E. et al. Safety and efficacy of the two doses conjugated protein-based SOBERANA-02 COVID-19 vaccine and of a heterologous three-dose combination with SOBERANA-Plus: a double-blind, randomised, placebo-controlled phase 3 clinical trial. Lancet Reg. Health Am. 18, 100423 (2023).
Borralleras, C. et al. The PHH-1V HIPRA vaccine: a new tool in the vaccination strategy against COVID-19. Rev. Esp. Quimioter. 36, 507–515 (2023).
Heidary, M. et al. A comprehensive review of the protein subunit vaccines against COVID-19. Front. Microbiol. 13, 927306 (2022).
Pasquevich, K. A. et al. Safety and immunogenicity of a SARS-CoV-2 Gamma variant RBD-based protein adjuvanted vaccine used as booster in healthy adults. Nat. Commun. 14, 4551 (2023).
WHO. Statement on the antigen composition of COVID-19 vaccines. https://www.who.int/news/item/26-04-2024-statement-on-the-antigen-composition-of-covid-19-vaccines (2024).
Perez Marc, G. et al. Immunogenicity and safety of monovalent and bivalent SARS-CoV-2 variant adapted RBD-based protein booster vaccines in adults previously immunized with different vaccine platforms: a phase II/III, randomized clinical trial. Vaccine 54, 127045 (2025).
Perez-Marc, G. et al. Immunogenicity and Safety of Gamma, Omicron BA.4/5 and Bivalent SARS-CoV-2 RBD-based Protein Booster Vaccines in Adults Previously Immunized with Different Vaccine Platforms: A Phase II/III, Randomized, Clinical Trial (Cold Spring Harbor Laboratory, 2024).
Lee, D. W. et al. mRNA-1273 vaccines adapted to JN.1 or KP.2 elicit cross-neutralizing responses against the JN.1 sublineages of SARS-CoV-2 in mice. Vaccine 54, 126961 (2025).
Sun, Z. et al. The role of cellular immunity in the protective efficacy of the SARS-CoV-2 vaccines. Vaccines 10, https://doi.org/10.3390/vaccines10071103 (2022).
Wang, L. et al. T cell immune memory after Covid-19 and vaccination. BMJ Med. 2, e000468 (2023).
Geers, D. et al. Profiling the SARS-CoV-2-specific T-cell response. Lancet Infect. Dis. 24, e477–e478 (2024).
Sette, A., Sidney, J. & Crotty, S. T cell responses to SARS-CoV-2. Annu. Rev. Immunol. 41, 343–373 (2023).
Cankat, S., Demael, M. U. & Swadling, L. In search of a pan-coronavirus vaccine: next-generation vaccine design and immune mechanisms. Cell Mol. Immunol. 21, 103–118 (2024).
Lenart, K. et al. Three immunizations with Novavax’s protein vaccines increase antibody breadth and provide durable protection from SARS-CoV-2. NPJ Vaccines 9, 17 (2024).
Stankov, M. V. et al. Humoral and cellular immune responses following BNT162b2 XBB.1.5 vaccination. Lancet Infect. Dis. 24, e1–e3 (2024).
Martinez, D. R. et al. Vaccine-mediated protection against Merbecovirus and Sarbecovirus challenge in mice. Cell Rep. 42, 113248 (2023).
Lin, D. Y., Huang, S., Milinovich, A., Duggal, A. & Wang, X. Effectiveness of XBB.1.5 vaccines and antiviral drugs against severe outcomes of omicron infection in the USA. Lancet Infect. Dis. 24, e278–e280 (2024).
Van Werkhoven, C. H. et al. Early COVID-19 vaccine effectiveness of XBB.1.5 vaccine against hospitalisation and admission to intensive care, the Netherlands, 9 October to 5 December 2023. Eurosurveillance 29, https://doi.org/10.2807/1560-7917.es.2024.29.1.2300703 (2024).
Hyseni, I. et al. Characterisation of SARS-CoV-2 lentiviral pseudotypes and correlation between pseudotype-based neutralisation assays and live virus-based micro neutralisation assays. Viruses 12, https://doi.org/10.3390/v12091011 (2020).
Temperton, N. J. et al. Longitudinally profiling neutralizing antibody response to SARS coronavirus with pseudotypes. Emerg. Infect. Dis. 11, 411–416 (2005).
Ferrara, F. & Temperton, N. Pseudotype neutralization assays: from laboratory bench to data analysis. Methods Protoc. 1, https://doi.org/10.3390/mps1010008 (2018).
Acknowledgements
The authors are grateful to the technical staff at the I+D Biofármacos Laboratorio Pablo Cassará and to Fundación Pablo Cassará for their significant assistance in the production, preparation and control of the antigen pilot lots and the regulatory documentation: Valeria Krum, Francisco M. Zurvarra, Ignacio Drehe, Morgan Miller Blanco, Mariano Picchio, Jonathan A. Baqué, Mariana Li Causi, Analia V. De Nichilo, Rocío Rodriguez, Cristian J. Payes, Juan Ignacio Sagues, Matias Lorch, Brenda Heinrich, María Victoria Román, Maria Victoria Pepe and all technician and analyst of Laboratorio Pablo Cassará involved in this project. We thank Ministerio de Salud de la Provincia de Buenos Aires for providing mRNA Spikevax vaccines.
Author information
Authors and Affiliations
Contributions
L.M.C. was responsible of overall experiment design, conducted humoral and cellular experiments, collected data, performed data analysis, and wrote the manuscript. J.C. was responsible for overall experimental design and conceptualization, wrote the manuscript and received funding support. J.M.R., S.A.D.P., A.C.H.I. and I.G.K. generated high expression vectors, designed the antigen and was responsible of development of the vaccine formulation and downstream development. Coordination of antigen and vaccine production. K.A.P. and J.M.F. participated of experimental design, data analysis and reviewed the manuscript. L.A.B., C.G.F.C. performed virus neutralization studies. C.P.C., A.D., L.P., L.M.S. and F.P.C. conducted and analyzed humoral studies and conducted cellular studies. G.P., R.A. and G.L. conducted pseudotyped SARS-CoV-1 neutralization assay. N.T. provided the plasmid for SARS-CoV-1 pseudotype production. J.C.V. participated in the development of vaccine formulation, collaborates in the overall process of vaccine development. All authors revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.M.R. and A.C.H.I. are salaried employees of Fundacion Pablo Cassara. J.M.F., S.A.D.P., I.G.K. and J.C.V. are salaried employees of Laboratorio Pablo Cassara. L.M.C., L.A.B., C.P.C., A.D., L.P., C.G.F.C., L.M.S., F.P.C., J.C. and K.A.P. declare no competing interests relevant to this article. G.P., R.A. and G.L. are employees of VisMederi srl. N.T. declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bruno, L.A., Pueblas Castro, C., Demaría, A. et al. Development of bivalent RBD adapted COVID-19 vaccines for broad sarbecovirus immunity. npj Vaccines 10, 108 (2025). https://doi.org/10.1038/s41541-025-01156-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41541-025-01156-3
This article is cited by
-
Targeting ACE2 with a camelid antibody inhibits SARS-CoV-2 binding and has protective effects in vivo
Nature Communications (2025)