Abstract
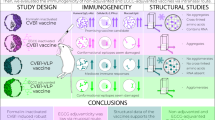
Cystic echinococcosis (CE) is a global zoonotic parasitic disease that represents a significant public health challenge. Although vaccination is considered an ideal strategy for controlling CE, no effective vaccines are currently available for dogs. Herein, bioinformatic approaches were employed to identify vaccine candidates. The selected proteins, including Echinococcus granulosus enolase (EgENO), severin (EgSev), cyclophilin (EgCyc), fatty acid-binding protein 1 (EgFABP1), calmodulin (EgCaM), and serine protease inhibitor 1 (EgSrp1), were expressed in Escherichia coli. These proteins were grouped into cocktail vaccines: rEgENO&rEgSev&rEgCyc and rEgFABP1&rEgCaM&rEgSrp1, and were combined with the Quil-A adjuvant to evaluate vaccine efficacy in beagles. After two subcutaneous immunizations, the rEgENO&rEgSev&rEgCyc and rEgFABP1&rEgCaM&rEgSrp1 vaccines reduced the parasite burden by 80.58% (p < 0.01) and 47.92% (p < 0.01), respectively. Additionally, Ligilactobacillus, Fusobacterium, and Streptococcus correlated significantly with immunoprotection. This study demonstrated bioinformatically screened antigens were effective vaccine candidates, and vaccine-microbiota interactions provided a potential strategy to improve vaccine efficacy.
Similar content being viewed by others
Introduction
Echinococcus granulosus sensu lato (E. granulosus s.l.), a cestode residing in the intestinal tract of canids, is the causative agent of cystic echinococcosis (CE), a globally prevalent zoonotic disease characterized by the establishment of its metacestodes in the organs of livestock (e.g., sheep, goats, cattle, yaks, pigs, horses, camels) and humans1. This disease poses significant threats to both public health and livestock economies, leading to considerable socioeconomic burdens2. Recognizing its impact, the World Health Organization (WHO) has included CE in its 2021–2030 strategic plan for control or elimination3,4. As definitive hosts, dogs serve as the primary source for CE infection, facilitating transmission to humans and animals primarily through the ingestion of food or water contaminated with feces containing E. granulosus eggs5. The current strategy to control E. granulosus infection in dogs relies on monthly praziquantel-based deworming programs6. However, this approach is hindered by significant challenges, including high resource demands and the emergence of anthelmintic resistance caused by prolonged drug use, ultimately undermining intervention efficacy7,8.
With further advances in our understanding of dogs’ mucosal immune responses following E. granulosus infection, the development of effective vaccines for dogs is increasingly regarded as a cornerstone strategy to mitigate CE transmission9,10. Compared with intermediate hosts such as sheep, cattle, and yak, the relatively smaller canine population makes dog vaccination a potentially more cost-effective strategy11,12. Current research on dog vaccination against E. granulosus primarily focuses on screening protective antigens. Recently, studies have demonstrated that dog vaccination provides protection by reducing the parasite burden and suppressing its developmental progression10,13,14. Candidate vaccines, such as those comprising EgM family proteins15,16, tropomyosin/paramyosin17, 3-hydroxyacyl-CoA dehydrogenase18, and the EgTIM-EgANXB3 (triosephosphate isomerase and annexin B3) cocktail vaccine19, have exhibited variable protective efficacy. To date, no vaccines have been licensed for use in dogs. A key challenge to commercialization is the inter-individual variability in protection rates, which compromises herd immunity and reduces the efficacy of population-level control13,14.
The inter-individual variability in vaccine-induced protective efficacy is modulated by multiple intrinsic and extrinsic factors, including host genetic background, innate immune status, and nutritional conditions20. Recent studies have revealed significant correlations between the gut microbiota composition and vaccine efficacy21,22,23. Specifically, a higher abundance of Actinobacteria is positively associated with robust vaccine responses, whereas Bacteroidetes is linked to reduced immunogenicity. The association between Firmicutes and Proteobacteria abundance and vaccine responsiveness appears to be taxon-specific24,25,26,27,28. Nevertheless, research exploring vaccine-induced microbiota modulation remains limited, particularly in the context of parenteral immunization20. Notably, helminth infections lead to substantial microbiota remodeling29,30, suggesting a potential interplay between parasite-induced dysbiosis and vaccine-mediated microbiota regulation. Determining this interaction could provide new insights into the mechanisms underlying vaccine-induced immunoprotection20,31. A systematic investigation into the correlation between vaccine-induced protection and alterations in fecal microbiota composition would provide valuable insights to develop next-generation vaccine adjuvants and delivery strategies.
In this study, we utilized bioinformatics to identify E. granulosus vaccine candidate antigens, followed by vaccination trials to systematically assess their efficacy and investigate the correlations between immunoprotection efficacy and fecal microbiota dynamics. Our findings provide a theoretical basis for controlling echinococcosis and offer novel insights into optimizing vaccine design.
Results
Bioinformatics-assisted identification of vaccine antigens
Proteomic profiling of exosomes identified 1916 high-confidence proteins. To establish a complete excretory/secretory (ES) proteome of adult E. granulosus, we integrated prior ES proteomic datasets, resulting in a catalog of 1940 proteins (Supplementary Table 1).
Through the multi-step antigen discovery pipeline, we identified 13 potential vaccine candidates: (1) Transcriptomic profiling of developmental stages revealed 439 genes that were significantly upregulated during parasite maturation (Supplementary Fig. 1a–c). (2) Comparative proteomic analysis via the Venn diagram intersection of protein datasets (Supplementary Table 1) identified 92 shared antigen candidates (Supplementary Fig. 2 and Supplementary Table 2). (3) Functional annotation based on literature mining further refined the selection to 15 well-characterized proteins. In silico antigenicity prediction using ANTIGENpro (cutoff ≥0.5) further identified 13 antigenic candidates (Table 1). Among them, triosephosphate isomerase (TIM), annexin B3 (ANXB3), and fatty acid-binding protein 2 (FABP2) have been validated as vaccine candidates with protective efficacy by our previous study19. Consequently, six proteins (rEgENO, rEgSev, rEgCyc, rEgFABP1, rEgCaM, rEgSrp1) were prioritized for further vaccine development.
Recombinant protein expression
Six E. granulosus genes were successfully cloned and expressed. PCR amplification using cDNA templates derived from PSCs and strobilated worms yielded distinct bands corresponding to EgENO (1302 bp), EgSev (1101 bp), EgCyc (489 bp), EgFABP1 (402 bp), EgCaM (450 bp), and EgSrp1 (1104 bp) (Supplementary Fig. 3a). Six recombinant proteins (rEgENO, rEgSev, rEgCyc, rEgFABP1, rEgCaM, and rEgSrp1) were confirmed by the SDS-PAGE analysis. The molecular weights of the recombinant proteins, including the pET-32a (+) vector-derived tag, were consistent with their theoretically approximate values: rEgENO (65.7 kDa), rEgSev (58.4 kDa), rEgCyc (35.9 kDa), rEgFABP1 (32.7 kDa), rEgCaM (34.5 kDa), and rEgSrp1 (58.5 kDa) (Supplementary Fig. 3b, c).
Vaccine-induced protection against E. granulosus in dogs
At 28 days post-challenge, necropsy was performed, which demonstrated varying levels of protection efficacy among the groups (Fig. 1 and Supplementary Table 3). In the control group, the worm burden in each dog reached 100,451 ± 10,340, with a mean worm length of 1.199 ± 0.014 mm and width of 0.199 ± 0.003 mm. The rEgFABP1&rEgCaM&rEgSrp1 vaccine group exhibited a 47.92% reduction in worm burden compared with that of the control (p < 0.01), with 52,313 ± 8754 worms. Worm length showed a marked reduction by 15.51% (p < 0.05), whereas width remained comparable (Fig. 2 and Supplementary Table 4). Notably, the rEgENO&rEgSev&rEgCyc vaccine demonstrated superior efficacy, achieving an 80.58% reduction in worm burden (p < 0.01), with 19,511 ± 6621 worms. Additionally, worms exhibited 0.86 fewer proglottids, and 24.85% (p < 0.01) and 22.11% (p < 0.01) reductions in body length and width, respectively (Fig. 2 and Supplementary Table 4).
Schematic summary of the study design, including the vaccination regimen and sample collection across three experimental groups. The timeline illustrates the major experimental procedures, including vaccination, parasite challenge, and designated sampling points (marked above the timeline). Stool and blood samples were obtained at specified time points (indicated by specific labels) for 16S rRNA sequencing and assessment of serum IgG levels and cytokine dynamics. Sample labels: A_a: Group A, time point Day0; A_b: Group A, Day21; A_c: Group A, Day35; A_d: Group A, Day49. B_a: Group B, Day0; B_b: Group B, Day21; B_c: Group B, Day35; B_d: Group B, Day49. C_a: Group C, Day0; C_b: Group C, Day21; C_c: Group C, Day35; C_d: Group C, Day49.
Quantitative evaluation of parasite burden and morphology, including a worm burden reduction, b worm length, and c worm width at 49 days post-infection across the experimental groups. Representative micrographs of 28-day worms from d the control group, e the rEgFABP1&rEgCaM&rEgSrp1 group, and f the rEgENO&rEgSev&rEgCyc group. Scale bar: 500 μm. Data are presented as mean ± SEM; *p < 0.05, **p < 0.01, ***p < 0.001.
Histopathological evaluation
Histopathological analysis indicated some degree of vaccine-associated protection against intestinal damage caused by E. granulosus infection (Fig. 3a–c). Gross examination of control dogs showed diffuse mucosal edema and hemorrhagic exudates, whereas vaccinated groups generally exhibited better-preserved mucosal integrity. H&E staining demonstrated notable villus architectural disruption in control dogs, including immune cell infiltration at parasite attachment sites (Fig. 3a). Vaccinated dogs maintained well-preserved villus morphology, exhibiting fewer parasite adhesion sites (Fig. 3b, c). PAS staining showed strong mucin production across all groups. Notably, goblet cell density was elevated in the vaccinated dogs, consistent with parasite-driven mucus hypersecretion.
Small intestines collected from dogs in the control (a) and vaccinated groups (b, c) (rEgFABP1&rEgCaM&rEgSrp1 and rEgENO&rEgSev&rEgCyc) at 4 weeks post-PSC challenge and processed for H&E and PAS staining. Parasite attachment sites, hemorrhagic exudates, and goblet cells were indicated by black frames, black arrows, and yellow arrows, respectively. Scale bar: 200 μm.
Serum IgG and cytokine dynamics
Serum IgG profiling (Fig. 4a–f) revealed distinct antibody kinetics among the vaccine groups. The rEgFABP1&rEgCaM&rEgSrp1 group achieved high IgG titers at D21, but the levels had declined by D49, with greater inter-individual variation. Additionally, the rEgENO&rEgSev&rEgCyc group exhibited a robust IgG response, peaking at D21 and persisting at significantly higher levels than the control group (p < 0.05) until D49. Serum IgG longevity in dogs vaccinated with the rEgENO&rEgSev&rEgCyc vaccine suggested that antibody responses could be maintained for at least 6 months, potentially contributing to sustained immunoprotection (Fig. 4g–i).
Serum-specific IgG levels against rEgFABP1 (a), rEgCaM (b), rEgSrp1 (c), rEgENO (d), rEgSev (e), and rEgCyc (f) in the experimental groups. Serum IgG longevity. Serum-specific IgG levels against rEgENO (g), rEgSev (h), rEgCyc (i) over 6 months. j–l Cytokine response patterns. j In the control group, IL-2 levels increased significantly at day 21, whereas IL-4 levels decreased, indicating a Th1-skewed response. k In the rEgFABP1&rEgCaM&rEgSrp1 group, IL-2, IL-4, and IFN-γ levels increased significantly at day 21. l In dogs vaccinated with rEgENO, rEgSev, and rEgCyc, no notable cytokine fluctuations were observed at day 21. Post PSCs challenge, the IL-2 levels decreased, whereas the IL-4 and IL-10 levels increased, favoring a Th2-biased response. *p < 0.05, **p < 0.01, ***p < 0.001.
Cytokine profiling (Fig. 4k) revealed a mixed Th1/Th2 response in the rEgFABP1&rEgCaM&rEgSrp1 group at D21 after the second immunization, characterized by concurrent IL-2, IL-4, and IFN-γ upregulation (p < 0.05). Following parasite challenge (Day 35), levels of IL-2, IL-4, and IFN-γ significantly decreased (p < 0.05), suggesting a suppression of host immune responses. Interestingly, the rEgENO&rEgSev&rEgCyc group displayed a unique immune response (Fig. 4l). While cytokine levels remained unchanged at D21, a pronounced Th2-skewed immunity occurred at D35 post-challenge, characterized by significant IL-4 and IL-10 upregulation (p < 0.05) and IL-2 suppression below the detection limit. Furthermore, IFN-γ levels rose significantly at D35 (p < 0.05), indicating IFN-γ pathway activation.
Alpha diversity and beta diversity analysis of the fecal microbiota composition
To investigate the interplay between the fecal microbiota and vaccine-induced immune responses, we performed 16S rRNA gene sequencing to profile fecal microbiota composition. Comparative analysis of the three experimental groups (Fig. 1) demonstrated dynamic shifts in microbial diversity at both the phylum and genus levels throughout the vaccination process and E. granulosus infection (Supplementary Fig. 4).
Analysis of longitudinal α-diversity (Fig. 5a–f) revealed distinct microbial community shifts in dogs in the rEgENO&rEgSev&rEgCyc vaccine group (Fig. 5e, f). Shannon and Chao1 indices exhibited significant increases at 7 days post-secondary immunization (D21, p < 0.05) relative to baseline (D0), indicating a vaccine-mediated expansion of microbial diversity. The sustained elevation of the Shannon index at 14 days post-challenge (D35, p < 0.05) suggested that microbiota remodeling might contribute to enhanced host immune defense. In contrast, the dogs in the rEgFABP1&rEgCaM&rEgSrp1 group exhibited microbiota dysregulation (Fig. 5c, d), with a transient increase in the Shannon index at D21, followed by a sustained decline post parasite challenge, leading to a significant decrease at D49 (p < 0.05). Meanwhile, the Chao1 indices remained statistically unchanged. To assess vaccine-specific effects on α-diversity dynamics, control group analyses were conducted. The control group exhibited a progressive decline in both Shannon and Chao1 indices throughout the study, with a significant reduction at D35 post-challenge (p < 0.05, Fig. 5a, b), confirming the parasite’s ability to suppress microbiota diversity. The divergence between vaccinated and control groups suggested that vaccine antigens might counteract parasite-driven microbial adaptations by modulating the fecal microbiota composition.
a–f Temporal α-diversity analysis (Shannon and Chao1 indices). In the control group, both indices exhibited a continuous decline, with a significant reduction observed at day 35 (p < 0.05) (a, b). In the rEgFABP1&rEgCaM&rEgSrp1 group, the Shannon index showed a transient increase at day 21, followed by a progressive decline that became significant at day 49 (p < 0.05), whereas the Chao1 index remained unchanged (c, d). In the rEgENO&rEgSev&rEgCyc group, both the Shannon and Chao1 indices increased significantly at day 21 and day 35 (p < 0.05) (e, f). g–i β-diversity analysis based on Bray–Curtis distance. Principal coordinate analysis (PCoA) revealed distinct clustering of fecal microbiota from the control (g), rEgFABP1&rEgCaM&rEgSrp1 (h), and rEgENO&rEgSev&rEgCyc (i) groups. Sample labels: A_a: Group A, Day0; A_b: Group A, Day21; A_c: Group A, Day35; A_d: Group A, Day49. B_a: Group B, Day0; B_b: Group B, Day21; B_c: Group B, Day35; B_d: Group B, Day49. C_a: Group C, Day0; C_b: Group C, Day21; C_c: Group C, Day35; C_d: Group C, Day49.
Analysis of β-diversity (Fig. 5g–i) demonstrated distinct vaccine-mediated clustering of rEgENO&rEgSev&rEgCyc samples along the PC1 and PC2 coordinates (ANOSIM: R = 0.7636, p = 0.001). Principal coordinate analysis (PCoA) based on Bray–Curtis distances revealed that PC1 (32.36%) and PC2 (19.39%) explained 51.75% of the total variance, suggesting dynamic, vaccine-induced shifts in the fecal microbiota composition. In contrast, the control group exhibited substantial post-parasite challenge displacement along PC1 (27.32%) and PC2 (16.64%) (ANOSIM: R = 0.5876, p = 0.001), forming a distinct clustering pattern relative to the vaccinated groups.
Variability of the fecal microbiota in vaccinated or E. granulosus-infected dogs
To investigate the impact of E. granulosus infection and vaccination on fecal microbiota regulation, longitudinal dynamic analysis was performed, revealing intervention-specific taxonomic alterations. By analyzing the top 10 core bacterial genera (Figs. 6a, 7a), combined with LEfSe analysis (LDA > 3.5, FDR-corrected p < 0.05 via Kruskal–Wallis test; Figs. 6b, 7b), identified three key taxa—Ligilactobacillus (phylum Firmicutes), Fusobacterium (phylum Fusobacteria), and Streptococcus (phylum Firmicutes)—exhibiting distinct responses to vaccination and E. granulosus infection.
a Heatmap illustrating relative abundance changes in the top 10 bacterial genera. b LEfSe analysis (LDA score >3.5, p < 0.05) identifying key discriminative taxa. c Fecal microbiota composition changes at the phylum level. d Genus-specific relative abundance shifts. Sample labels: A_a: Group A, Day0; A_c: Group A, Day35.
a Heatmap illustrating relative abundance changes in the top 10 bacterial genera before and after vaccination. b LEfSe analysis (LDA score >3.5, p < 0.05) identifying key discriminatory taxa. c Phylum-level compositional alterations. d Genus-specific relative abundance variations post-vaccination. Sample labels: C_a: Group C, Day0; C_b: Group C, Day21.
In the control group, following E. granulosus colonization (D35), the relative abundance of Ligilactobacillus increased by 33.98% (p = 0.008) and that of Streptococcus increased by 7.87% (p = 0.014), whereas Fusobacterium exhibited a 9.46% decline (p = 0.014) (Fig. 6d). Conversely, the rEgENO&rEgSev&rEgCyc vaccine group exhibited an inverse regulatory trend (Fig. 7a–d), with Ligilactobacillus and Streptococcus abundances decreasing by 13.03% (p = 0.008) and 27.28% (p = 0.008), respectively, while Fusobacterium levels increased by 5.76% (p = 0.007) (Fig. 7d).
LEfSe analysis further confirmed Ligilactobacillus (LDA = 5.23/4.79), Fusobacterium (LDA = 4.78/4.43), and Streptococcus (LDA = 4.66/5.15) as key discriminative taxa across both experimental groups (Wilcoxon test, FDR-adjusted p < 0.05) (Figs. 6b and 7b). These findings suggested that vaccination might counteract E. granulosus-induced microbiota dysbiosis by modulating the relative abundances of key bacterial taxa.
Links between vaccine-induced immunity and fecal microbiota alterations
Building on previous evidence linking host immunity with the gut microbiota32, we employed a multidimensional integrative approach to determine the relationship between vaccine-induced immune responses and microbiota alterations. Environmental factor analysis revealed significant dynamic covariation between the relative abundances of Ligilactobacillus, Fusobacterium, and Streptococcus and the host Th1/Th2 cytokine profiles (Fig. 8a).
a Heatmap of Pearson’s correlation analysis between key bacterial genera and cytokine levels. b Mantel test and RDA of cytokine-microbiota interactions. c RDA ordination plot illustrating the contributions of cytokines to microbiota variation. The first two constrained axes explained 24.42% of total microbiota variability (RDA1: 17.6%, RDA2: 6.82%, p < 0.001). Sample labels: C_a: Group C, Day0; C_b: Group C, Day21; C_c: Group C, Day35; C_d: Group C, Day49.
Mantel tests and redundancy analysis (RDA) identified IFN-γ as the principal factor of microbiota restructuring following vaccination (Mantel’s r = 0.79, p = 0.01). IFN-γ exhibited a strong positive correlation with IL-4 and IL-10, while showing a negative association with IL-2 (Fig. 8b). These findings suggested that the Th1/Th2 equilibrium plays a pivotal role in orchestrating temporal microbial shifts. Notably, RDA constrained 24.42% of microbiota variance (RDA1: 17.60%, RDA2: 6.82%; p < 0.001) (Fig. 8c), with IFN-γ (r = 0.79, p = 0.01, Mantel test) accounting for the highest proportion of constrained variation. These findings highlighted vaccine-induced host immune shifts as a key determinant of microbiota remodeling.
Discussion
Cystic echinococcosis (CE), caused by E. granulosus, remains a global public health challenge, imposing significant burdens on both human health and livestock productivity3,33. Given the chronic and complex nature of CE, control strategies should prioritize simple, cost-effective, and efficient approaches6,7,14. Vaccination, widely regarded as the most cost-effective approach of infectious disease control34, holds great promise for CE prevention. Moreover, mathematical models of E. granulosus transmission dynamics have further demonstrated that vaccinating dogs can reduce infection pressure on intermediate hosts; if herd immunity reaches 75%, parasite elimination may be achievable35. However, the lack of effective commercial vaccines for dogs—the primary definitive hosts and reservoirs of E. granulosus—remains a critical barrier in CE eradication10,13,14.
The complex life cycle of E. granulosus, involving multiple host transitions5, facilitates its sophisticated immune evasion strategies36,37,38. This complexity poses a significant challenge in identifying antigens with high protective efficacy, constituting a major bottleneck in vaccine development. Traditional antigen screening approaches, which rely on animal infection models, are hindered by lengthy experimental timelines, high costs, and biosafety concerns, severely restricting progress in vaccine research9,14. The integration of multi-omics data with bioinformatics provides an innovative strategy to identify novel vaccine antigens39. Based on the developmental biology of E. granulosus, key biological processes, such as PSC growth, adaptation to the intestinal environment, and immune evasion, are mediated by specific functional proteins40. These proteins, critical for parasite survival and host-parasite interactions, represent promising candidate targets for dog vaccines.
This study firstly established a comprehensive proteomic database of EVs from 28-day E. granulosus, integrating three functional protein subsets based on E. granulosus biological processes. Bioinformatic screening identified thirteen annotated proteins, including three previously validated vaccine candidates19. Based on this selection framework, six of the remaining ten proteins were prioritized for vaccine efficacy assessment. These proteins were tegument-localized or E. granulosus secreted proteins likely to interact directly with the host intestinal mucosa, thereby triggering antigen-specific immune responses41,42. Notably, EgENO functions not only as a glycolytic enzyme essential for parasite energy metabolism, but also as a major exosome component in adult worms, playing a role in host immune modulation43. Similarly, the tegument-localized EgSev protein is presumed to facilitate parasite motility via mechanisms similar to those of EgA3144,45. Bioinformatic profiling of EgCyc identified highly immunogenic epitopes, reinforcing its potential as a vaccine candidate46. EgFABP1, a lipid trafficking protein crucial for parasite survival16, mediates host-derived lipid acquisition essential for growth and development40, while exhibiting strong immunogenic properties for vaccination47. Functional analyses of EgCaM48,49 and EgSrp150,51 further delineated their roles in parasite maturation and adaptation to the gut microenvironment. Collectively, these findings confirm the reliability of our bioinformatics-driven vaccine antigen discovery framework. Additionally, antigen delivery strategies significantly influence vaccine efficacy. The antigenic complexity of multicellular helminths such as E. granulosus means that single-antigen vaccines might be insufficient to provide immunoprotection because of epitope diversity52. Cocktail vaccines provide a viable approach to enhance protective efficacy and durability, representing a promising direction in antiparasitic vaccine development53,54. Our cocktail vaccines were designed to align with this approach.
Building upon these foundational studies, our cocktail vaccine (rEgENO&rEgSev&rEgCyc) achieved an 80.58% reduction in the worm burden following a two-dose immunization regimen, representing an approximately 10% improvement over existing two-dose vaccines19. Comparative analysis highlighted three key advantages: (1) The two-dose subcutaneous regimen confers enhanced protection against high-dose PSC challenge, while maintaining operational feasibility; (2) sustained antibody levels for 6 months post-immunization support a biannual vaccination strategy to prevent parasite establishment; (3) high sequence homology (>98%) with Echinococcus multilocularis homologous antigens (Table 1) suggests potential cross-protection against both cestode species10. Dogs also serve as definitive hosts of E. multilocularis, and in many CE-endemic areas, alveolar echinococcosis coexists as a major concern in Central Asia and western China, where E. multilocularis and E. granulosus exhibit high endemicity1,10,14. Given the absence of licensed vaccines against E. multilocularis55, the cross-protective potential of the vaccine developed in the present study holds substantial socioeconomic and public health significance for the simultaneous control of CE and alveolar echinococcosis within endemic regions.
Although existing candidate vaccines against E. granulosus induce variable protective responses in dogs, substantial inter-individual differences in vaccine efficacy have been reported14. A substantial proportion of low responders might hinder herd immunity development, thereby reducing overall disease control effectiveness56. Thus, unraveling the complex host-parasite interactions and immune protection mechanisms of E. granulosus is essential to optimize vaccination strategies. Recent studies highlighted the gut microbiota composition as a pivotal determinant of vaccine efficacy56,57. However, the relationships between E. granulosus infection, microbiome dynamics, and vaccine efficacy have yet to be explored in canine models.
This study provides the first comprehensive characterization of the interactions between E. granulosus infection, fecal microbiota alterations, and vaccine-mediated immunity, offering novel insights into vaccine response variability. Experimental validation confirmed that E. granulosus infection drives distinct fecal microbiota remodeling in dogs. Unlike typical helminth infections that promote α-diversity29, E. granulosus infection significantly decreased the Shannon index (p < 0.05) while increasing Ligilactobacillus and Streptococcus abundance and suppressing Fusobacterium. These microbial alterations might enhance parasite survival through three mechanisms: (1) Ligilactobacillus-derived lactate might disrupt mucosal immune surveillance by modulating dendritic cell-T cell interactions, thereby facilitating immune evasion58; (2) Streptococcus enrichment might sustain parasite-favorable microenvironments by modulating metabolic reprogramming59. Notably, the depletion of Fusobacterium mirrors patterns observed in enterobiasis60, suggesting the presence of a conserved helminth-driven immunoregulatory mechanism. Collectively, these findings suggest that E. granulosus strategically manipulates its ecological niche through targeted microbiota modulation.
The rEgENO&rEgSev&rEgCyc vaccine, which exhibited the highest efficacy in reducing E. granulosus burden in this study, comprises ES antigens from adult worms that are potentially involved in host-parasite interactions42,61. To investigate the microbiota-immunity interplay underpinning anthelmintic protection, comparative analyses between the high-efficacy vaccine groups and controls revealed inverse abundance patterns of Ligilactobacillus, Fusobacterium, and Streptococcus. Vaccine-induced microbiota remodeling was observed to influence E. granulosus colonization, thereby contributing mechanistically to protective immunity. Notably, longitudinal variations in Ligilactobacillus, Fusobacterium, and Streptococcus abundances correlated positively with IL-4 concentrations in vaccinated dogs, supporting a Th2-skewed immune response that promoted parasite clearance and mitigated intestinal pathology62. Simultaneously, IFN-γ fluctuations exhibited microbiota-dependent associations, suggesting that vaccine-induced IFN-γ production might sustain long-term protection through interactions with microbial communities63. These findings provide novel insights to optimize vaccine design to enhance efficacy. However, this study is limited in its ability to identify specific bacterial species and to elucidate the mechanisms underlying microbiota-mediated modulation of host immunity. Future studies employing metagenomic sequencing and mechanistic approaches are warranted to further clarify the roles of key microbial taxa in shaping vaccine-induced protection.
Methods
Animals and parasites
A total of 22 6-month-old beagles (11 males and 11 females) were obtained from the Beagle Breeding Center at the Sichuan Institute of Musk Deer Breeding (Sichuan, China). Eighteen dogs were used for vaccine efficacy assessment and fecal microbiome dynamics assessment. Four were used to assess serum IgG longevity in vaccinated dogs. All dogs were housed under specific-pathogen-free conditions in a controlled quarantine facility and fed a standardized diet of commercial dog food with free access to tap water.
Hydatid cysts were harvested from the livers of sheep naturally infected with CE and slaughtered at an abattoir in Sichuan Province, China. Protoscoleces (PSCs) were isolated from these cysts following the methods described in our previous study19. The viability of the PSCs exceeded 95% were used for vaccine trials. Species identification was performed via PCR amplification and sequencing, confirming the E. granulosus sensu stricto G1 genotype64.
Bioinformatics-assisted screening of vaccine candidates
Based on our previously acquired mass spectrometry proteomics dataset of adult E. granulosus exosomes, raw spectral data were analyzed using Mascot v2.3.0265 against the UniProt database under rigorous quality control criteria (false discovery rate [FDR] ≤1%). This study employed a multi-step vaccine antigen screening pipeline by integrating our 28-day E. granulosus worm-derived exosome proteomic profile with publicly available E. granulosus proteomic and transcriptomic datasets40,41,42,61,66,67: (1) Comparative transcriptomics identified genes functionally upregulated in adult worms relative to PSCs (log2fold-change >2, p < 0.05) through volcano plot analysis; (2) gene selection was refined using threshold-optimized multi-group volcano plots and GO clustering, focusing on host-parasite interaction and immune evasion genes; (3) Four protein sets were constructed based on the multi-omics data, with the selection rationale detailed in Supplementary Table 1. Notably, cross-species interaction proteins were identified through Western blot analysis of the E. granulosus proteome using sera from infected dogs. Then, Venn diagram analysis was used to identify proteins shared among at least two sets, prioritizing those essential proteins for vaccine candidates. Antigenicity was predicted using ANTIGENpro (http://scratch.proteomics.ics.uci.edu/) with a cut-off value of ≥0.5, while cross-species protection potential was evaluated through BLAST (https://www.uniprot.org/blast) alignment against Echinococcus multilocularis orthologs.
Protein expression
The six recombinant E. granulosus proteins (rEgENO (enolase), rEgSev (severin), rEgCyc (cyclophilin), rEgFABP1 (fatty acid-binding protein 1), rEgCaM (calmodulin) and rEgSrp1 (serine protease inhibitor 1) were expressed as follows. Primers with homologous arms were designed using Primer Premier 5.0 (Premier Biosoft, San Francisco, CA, USA) based on target sequences obtained from GenBank (Supplementary Table 5). The target genes were amplified by PCR, and the amplicons were verified via 1% agarose gel electrophoresis before being inserted into the pET-32a (+) expression vector (Invitrogen, USA) using a Seamless Cloning Kit (Genesand Biotech, China). Recombinant plasmids were subsequently transformed into Escherichia coli BL21 (DE3) competent cells (Tiangen, China) and induced with 1 mM Isopropyl β-D-1-thiogalactopyranoside (IPTG) at 16 °C for 6 h. The his-tagged recombinant proteins were purified using an Ni²⁺-NTA affinity chromatography column (Bio-Rad, USA), and the final protein concentrations were quantified using a bicinchoninic acid (BCA) protein assay kit (Beyotime, China).
Vaccination and parasite challenge
Eighteen beagles (n = 6 per group, half male and half female) were randomly assigned into three experimental groups, following the design outlined in Fig. 1. Each dog received two doses of a subcutaneous injection of the designated cocktail vaccine formulations rEgENO&rEgSev&rEgCyc and rEgFABP1&rEgCaM&rEgSrp1 (detailed in Fig. 1), with a primary immunization at Day 0 (D0) and a booster at D14. On D21, each dog was orally challenged with 150,000 PSCs. At D49, animals were first sedated with an intramuscular injection of xylazine hydrochloride (2 mg/kg) to induce anesthesia. After confirming the absence of reflexes, euthanasia was performed by intravenous administration of sodium pentobarbital at a dose of 100 mg/kg body weight. This method was consistent with the guidelines of the American Veterinary Medical Association. The entire small intestine was excised aseptically, and the intestinal contents were systematically collected for microscopic enumeration of the total worm burden. In each dog, 50 randomly selected worms were measured for body length and maximal width. Intestinal tissue samples were fixed in 4% paraformaldehyde for subsequent histopathological assessment.
Serum IgG detection
Serum antigen-specific IgG levels were quantified using an indirect enzyme-linked immunosorbent assay (ELISA). Recombinant proteins (rEgENO, rEgSev, rEgCyc, rEgFABP1, rEgCaM, and rEgSrp1) were coated in 96-well plates at 5 μg/mL in 0.05 M carbonate-bicarbonate buffer (pH 9.6) (100 μL/well) and incubated at 4 °C for 16 h. After three washes with PBST (0.05% Tween-20 in phosphate-buffered saline (PBS); 200 μL/well, 5 min per wash), the wells were blocked using 5% (w/v) skim milk in PBS at 37 °C for 2 h. Serum samples (diluted 1:100 in PBST) were then added and incubated at 37 °C for 1 h, followed by incubation with horseradish peroxidase (HRP)-conjugated rabbit anti-dog IgG (1:3000, Solarbio, China) at 37 °C for 1.5 h. The 3,3′,5,5′-Tetramethylbenzidine (TMB; Tiangen) substrate was added for 20 min in the dark, followed by reaction termination using 100 μL of 2 M H2SO4. The optical density (OD) was then measured at 450 nm.
Serum IgG longevity in dogs vaccinated with the rEgENO&rEgSev&rEgCyc vaccine was assessed using an indirect ELISA following the above protocol, with blood sera collected over 6 months.
Serum cytokine profiling
Serum levels of interleukin (IL)-2, IL-4, IL-5, IL-10, and interferon gamma (IFN-γ) were quantified using commercial ELISA kits (Solarbio, China) according to the manufacturer’s instructions. Before assay execution, all kit components were equilibrated to room temperature. Standard curves were generated using recombinant cytokine calibrators provided in each kit.
Histopathological analysis
Dog small intestinal tissue samples were processed using standardized histopathological protocols. Freshly excised small intestinal segments (1 cm³) were immediately fixed in 4% paraformaldehyde (0.1 M PBS, pH 7.4) at 4 °C for 24 h, with one solution replacement. Fixed tissues were subsequently dehydrated through a graded ethanol series, cleared in xylene, and embedded in paraffin at 56–58 °C. Serial 5 μm sections were stained with hematoxylin and eosin (H&E) and periodic acid-Schiff (PAS), followed by mounting with neutral resin. Three consecutive sections per sample were prepared.
Dog fecal microbiome profiling
Total genomic DNA was extracted from canine fecal samples using the E.Z.N.A.® DNA Kit (Omega Bio-tek, USA) following the manufacturer’s instructions. The V3-V4 hypervariable regions of the 16S rRNA gene were amplified using primers 338 F (5′-ACTCCTACGGGAGGCAGCAG-3′) and 806 R (5′-GGACTACHVGGGTWTCTAAT-3′)68. Amplicon sequencing was performed on the Illumina NextSeq 2000 platform (Majorbio, China), and raw reads were deposited in the NCBI SRA database (Accession: PRJNA1244299).
Quality filtering and denoising of raw reads were conducted using DADA269 within QIIME270 under default parameters. Sequences were subsampled to the minimum sequencing depth (Good’s coverage >99.09%) before α- and β-diversity analyses. Amplicon sequence variants (ASVs) were taxonomically classified against the SILVA 16S rRNA database (v138) using QIIME2’s Naïve Bayes classifier.
Bioinformatic analyses were conducted on the Majorbio Cloud Platform (https://cloud.majorbio.com). α-Diversity indices (Chao1, Shannon) were computed using mothur71, with intergroup differences assessed via the Wilcoxon rank-sum test. β-Diversity was evaluated via Bray–Curtis-based principal coordinate analysis (PCoA) with ANOSIM (999 permutations). Linear discriminant analysis effect size (LEfSe)72 was used to identify differentially abundant taxa (linear discriminant analysis (LDA) > 3.5, p < 0.05) from phylum to genus levels. Redundancy analysis (RDA) was employed to explore cytokine-microbiome associations, while Spearman correlation networks (p < 0.05) were constructed to visualize key species interactions.
Statistical analysis
All parasitological parameters (total worm count, body length, and maximal width) were analyzed using SPSS 20.0 (IBM Corp., USA) and GraphPad Prism 8.0 Software (GraphPad Inc., USA). Protection efficacy was calculated as: (Mean worm burden in the control group−Mean worm burden in the vaccinated group)/Mean worm burden in the control group × 100%. The Mann–Whitney U test was used for statistical analysis, and differences with a p value less than 0.05 were considered statistically significant.
Ethics
The animal study was reviewed and approved by the Animal Care and Use Committee of Sichuan Agricultural University (SYXK2019-187). All animal procedures used in this study were carried out in accordance with the Guide for the Care and Use of Laboratory Animals (National Research Council, Bethesda, MD, USA) and recommendations of the ARRIVE guidelines (https://www.nc3rs.org.uk/arrive-guidelines). All methods were carried out in accordance with relevant guidelines and regulations.
Data availability
All data generated or analyzed during this study are included in this published article and its Supplementary Information files. The 16S rRNA gene sequence data generated in this study have been deposited in the NCBI Sequence Read Archive (accession number PRJNA1244299). The dataset is publicly accessible at: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1244299/.
References
Wen, H. et al. Echinococcosis: advances in the 21st century. Clin. Microbiol. Rev. 32, e00075-18 (2019).
Zhou, H., Wang, X., Han, S. & Xiao, N. Advances and challenges in the prevention, control and research of echinococcosis in China. Decoding Infect. Transm. 3, 100041 (2025).
Noguera, Z. L. P., Charypkhan, D., Hartnack, S., Torgerson, P. R. & Rüegg, S. R. The dual burden of animal and human zoonoses: a systematic review. PLoS Negl. Trop. Dis. 16, e0010540 (2022).
Widdicombe, J. et al. The economic evaluation of Cystic echinococcosis control strategies focused on zoonotic hosts: a scoping review. PLoS Negl. Trop. Dis. 16, e0010568 (2022).
Pal, M., Alemu, H. H., Marami, L. M., Garedo, D. R. & Bodena, E. B. Cystic echincoccoosis: a comprehensive review on life cycle, epidemiology, pathogenesis, clinical spectrum, diagnosis, public health and economic implications, treatment, and control. Int. J. Clin. Exp. Med. Res. 6, 131–141 (2022).
Rong, X., Fan, M., Zhu, H. & Zheng, Y. Dynamic modeling and optimal control of cystic echinococcosis. Infect. Dis. Poverty 10, 1–13 (2021).
Larrieu, E., Gavidia, C. M. & Lightowlers, M. W. Control of cystic echinococcosis: background and prospects. Zoonoses Public Health 66, 889–899 (2019).
Coffeng, L. E., Stolk, W. A. & de Vlas, S. J. Predicting the risk and speed of drug resistance emerging in soil-transmitted helminths during preventive chemotherapy. Nat. Commun. 15, 1099 (2024).
Pourseif, M. M. et al. Current status and future prospective of vaccine development against Echinococcus granulosus. Biologicals 51, 1–11 (2018).
Zhang, W. & McManus, D. P. Vaccination of dogs against Echinococcus granulosus: a means to control hydatid disease?Trends Parasitol. 24, 419–424 (2008).
Craig, P. S., Giraudoux, P., Wang, Z. H. & Wang, Q. Echinococcosis transmission on the Tibetan Plateau. Adv. Parasitol. 104, 165–246 (2019).
Craig, P., Hegglin, D., Lightowlers, M., Torgerson, P. R. & Wang, Q. Echinococcosis: control and prevention. Adv. Parasitol. 96, 55–158 (2017).
Torgerson, P. R. Dogs, vaccines and Echinococcus. Trends Parasitol. 25, 57–58 (2009).
Borhani, M. et al. Echinococcus granulosus sensu lato control measures: a specific focus on vaccines for both definitive and intermediate hosts. Parasit. Vectors 17, 533 (2024).
Wang, N. et al. Evaluation of protective immune responses induced by DNA vaccines encoding Echinococcus granulosus EgM123 protein in Beagle dogs. Front. Vet. Sci. 11, 1444741 (2024).
Zhang, W. et al. Vaccination of dogs against Echinococcus granulosus the cause of cystic hydatid disease in humans. J. Infect. Dis. 194, 966–974 (2006).
Petavy, A. F. et al. An oral recombinant vaccine in dogs against Echinococcus granulosus, the causative agent of human hydatid disease: a pilot study. PLoS Negl. Trop. Dis. 2, e125 (2008).
Xian, J. et al. Molecular characterization and immune protection of the 3-hydroxyacyl-CoA dehydrogenase gene in Echinococcus granulosus. Parasit. Vectors 14, 1–11 (2021).
Shao, G. et al. Protective efficacy of six recombinant proteins as vaccine candidates against Echinococcus granulosus in dogs. PLoS Negl. Trop. Dis. 17, e0011709 (2023).
Zimmermann, P. The immunological interplay between vaccination and the intestinal microbiota. npj Vaccines 8, 24 (2023).
Zimmermann, P. & Curtis, N. The influence of the intestinal microbiome on vaccine responses. Vaccine 36, 4433–4439 (2018).
Jordan, A., Carding, S. R. & Hall, L. J. The early-life gut microbiome and vaccine efficacy. Lancet Microbe 3, e787–e794 (2022).
Lynn, D. J., Benson, S. C., Lynn, M. A. & Pulendran, B. Modulation of immune responses to vaccination by the microbiota: implications and potential mechanisms. Nat. Rev. Immunol. 22, 33–46 (2022).
Huda, M. N. et al. Stool microbiota and vaccine responses of infants. Pediatrics 134, e362–e372 (2014).
Mullié, C. et al. Increased poliovirus-specific intestinal antibody response coincides with promotion of Bifidobacterium longum-infantis and Bifidobacterium breve in infants: a randomized, double-blind, placebo-controlled trial. Pediatr. Res. 56, 791–795 (2004).
Harris, V. et al. Rotavirus vaccine response correlates with the infant gut microbiota composition in Pakistan. Gut Microbes 9, 93–101 (2018).
Zhao, T. et al. Influence of gut microbiota on mucosal IgA antibody response to the polio vaccine. npj Vaccines 5, 47 (2020).
Harris, V. C. et al. Significant correlation between the infant gut microbiome and rotavirus vaccine response in rural Ghana. J. Infect. Dis. 215, 34–41 (2017).
Walusimbi, B. et al. The effects of helminth infections on the human gut microbiome: a systematic review and meta-analysis. Front. Microbiomes 2, 1174034 (2023).
Sieng, S., Chen, P., Wang, N., Xu, J.-Y. & Han, Q. Toxocara canis-induced changes in host intestinal microbial communities. Parasit. Vectors 16, 462 (2023).
Paz, E. A. et al. Bacterial communities in the gastrointestinal tract segments of helminth-resistant and helminth-susceptible sheep. Anim. Microbiome 4, 23 (2022).
Su, C. et al. Helminth-induced alterations of the gut microbiota exacerbate bacterial colitis. Mucosal Immunol. 11, 144–157 (2018).
Hua, R. Q. et al. Genetic diversity of Echinococcus granulosus sensu lato in China: epidemiological studies and systematic review. Transbound. Emerg. Dis. 69, e1382–e1392 (2022).
Jorge, S. & Dellagostin, O. A. The development of veterinary vaccines: a review of traditional methods and modern biotechnology approaches. Biotechnol. Res Innov. 1, 6–13 (2017).
Torgerson, P. Canid immunity to Echinococcus spp.: impact on transmission. Parasite Immunol. 28, 295–303 (2006).
Wang, Y. et al. Impairment of dendritic cell function and induction of CD4+ CD25+ Foxp3+ T cells by excretory-secretory products: a potential mechanism of immune evasion adopted by Echinococcus granulosus. BMC Immunol. 16, 1–10 (2015).
Cao, S. et al. Arginase promotes immune evasion of Echinococcus granulosus in mice. Parasit. Vectors 13, 1–12 (2020).
Siracusano, A. et al. Immunomodulatory mechanisms during Echinococcus granulosus infection. Exp. Parasitol. 119, 483–489 (2008).
Ammar, A., Elleboudy, N. & Mohammed, S. The contribution of in silico studies in parasitology: a multifaceted approach. Parasitol. U. J. 16, 1–11 (2023).
Zheng, H. et al. The genome of the hydatid tapeworm Echinococcus granulosus. Nat. Genet. 45, 1168–1175 (2013).
Li, Z. J. & Wei, Z. Analysis of protoscoleces-specific antigens from Echinococcus granulosus with proteomics combined with Western blot. Biomed. Environ. Sci. 25, 718–723 (2012).
Miles, S. et al. Combining proteomics and bioinformatics to explore novel tegumental antigens as vaccine candidates against Echinococcus granulosus infection. J. Cell Biochem. 120, 15320–15336 (2019).
Pourseif, M. M., Yousefpour, M., Aminianfar, M., Moghaddam, G. & Nematollahi, A. A multi-method and structure-based in silico vaccine designing against Echinococcus granulosus through investigating enolase protein. Bioimpacts 9, 131 (2019).
Miles, S., Velasco-de-Andrés, M., Lozano, F. & Mourglia-Ettlin, G. Interactome analysis of CD5 and CD6 ectodomains with tegumental antigens from the helminth parasite Echinococcus granulosus sensu lato. Int. J. Biol. Macromol. 164, 3718–3728 (2020).
Zhang, X. et al. EgSeverin and Eg14-3-3zeta from Echinococcus granulosus are potential antigens for serological diagnosis of echinococcosis in dogs and sheep. Micro. Pathog. 179, 106110 (2023).
Khazaei, S. & Moghadamizad, Z. Echinococcus granulosus cyclophilin: Immunoinformatics analysis to provide insights into the biochemical properties and immunogenic epitopes. Inf. Med. Unlocked 30, 100925 (2022).
Chabalgoity, J. A. et al. Expression and immunogenicity of an Echinococcus granulosus fatty acid-binding protein in live attenuated Salmonella vaccine strains. Infect. Immun. 65, 2402–2412 (1997).
Mousavi, S. M. et al. Calmodulin-specific small interfering RNA induces consistent expression suppression and morphological changes in Echinococcus granulosus. Sci. Rep. 9, 3894 (2019).
Wang, N. et al. Molecular and biochemical characterization of calmodulin from Echinococcus granulosus. Parasit. Vectors 10, 1–9 (2017).
Guo, G., Tian, M. X., Jiao, H. J., Li, J. & Zhang, W. B. Comparison and analysis of structure and function of serine protease inhibitors from two strains of Echinococcus. Chin. J. Biol. 34, 928–935 (2021).
Li, X. et al. Echinococcus multilocularis serpin regulates macrophage polarization and reduces gut dysbiosis in colitis. Infect. Immun. 92, e00232-24 (2024).
Rivero, F. D. et al. Disruption of antigenic variation is crucial for effective parasite vaccine. Nat. Med. 16, 551–557 (2010).
Lightowlers, M. et al. Vaccination against cestode parasites: anti-helminth vaccines that work and why. Vet. Parasitol. 115, 83–123 (2003).
Manjunathachar, H. V. et al. Cocktail vaccine for the management of Hyalomma anatolicum and Rhipicephalus microplus. Front. Immunol. 15, 1471317 (2024).
Woolsey, I. D. & Miller, A. L. Echinococcus granulosus sensu lato and Echinococcus multilocularis: a review. Res. Vet. Sci. 135, 517–522 (2021).
Borey, M. et al. Links between fecal microbiota and the response to vaccination against influenza A virus in pigs. npj Vaccines 6, 92 (2021).
Ciabattini, A., Olivieri, R., Lazzeri, E. & Medaglini, D. Role of the microbiota in the modulation of vaccine immune responses. Front. Microbiol. 10, 1305 (2019).
Sanmarco, L. M. et al. Lactate limits CNS autoimmunity by stabilizing HIF-1α in dendritic cells. Nature 620, 881–889 (2023).
Beyhan, Y. E. & Yıldız, M. R. Microbiota and parasite relationship. Diagn. Microbiol. Infect. Dis. 106, 115954 (2023).
Yang, C. A. et al. Impact of Enterobius vermicularis infection and mebendazole treatment on intestinal microbiota and host immune response. PLoS Negl. Trop. Dis. 11, e0005963 (2017).
Wang, Y. et al. Proteomic analysis of the excretory/secretory products and antigenic proteins of Echinococcus granulosus adult worms from infected dogs. BMC Vet. Res. 11, 1–7 (2015).
Cortés, A., Muñoz-Antoli, C., Esteban, J. G. & Toledo, R. Th2 and Th1 responses: clear and hidden sides of immunity against intestinal helminths. Trends Parasitol. 33, 678–693 (2017).
McFarlane, A. J. et al. Enteric helminth-induced type I interferon signaling protects against pulmonary virus infection through interaction with the microbiota. J. Allergy Clin. Immunol. 140, 1068–1078 (2017).
Zhan, J. et al. Molecular and functional characterization of inhibitor of apoptosis proteins (IAP, BIRP) in Echinococcus granulosus. Front. Microbiol 11, 729 (2020).
Perkins, D. N., Pappin, D. J., Creasy, D. M. & Cottrell, J. S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Virginio, V. G. et al. Excretory/secretory products from in vitro-cultured Echinococcus granulosus protoscoleces. Mol. Biochem. Parasitol. 183, 15–22 (2012).
Nicolao, M. C., Rodriguez Rodrigues, C. & Cumino, A. C. Extracellular vesicles from Echinococcus granulosus larval stage: isolation, characterization and uptake by dendritic cells. PLoS Negl. Trop. Dis. 13, e0007032 (2019).
Liu, C. et al. Denitrifying sulfide removal process on high-salinity wastewaters in the presence of Halomonas sp. Appl. Microbiol. Biotechnol. 100, 1421–1426 (2016).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Barberán, A., Bates, S. T., Casamayor, E. O. & Fierer, N. Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J. 6, 343–351 (2012).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, 1–18 (2011).
Acknowledgements
The authors gratefully acknowledge all the participants involved in the dog vaccine development for their invaluable contributions. We also extend our sincere appreciation to the staff of the Beagles Breeding Center for their dedicated assistance with animal management. This work was supported by a grant from the Key Technology Research and Development Program of Sichuan Province in China (grant number 2022YFN0013).
Author information
Authors and Affiliations
Contributions
G.Y. and A.Y. conceived and designed the study, contributed to funding acquisition and data verification. G.S., R.H. and X.Z. were responsible for conducting experiments, performing data analysis and drafting the manuscript. Y.C. and Z.L. contributed to project administration and data collection. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shao, G., Zhu, X., Hua, R. et al. Cocktail vaccine induces immunoprotection and modulates the fecal microbiota in dogs against Echinococcus granulosus infection. npj Vaccines 10, 214 (2025). https://doi.org/10.1038/s41541-025-01275-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41541-025-01275-x