Abstract
The “Nitrite Bottleneck” in nitrite-based nitrogen removal processes undermines the efficiency of partial nitrification-anammox (PN/A). This perspective proposes leveraging bacteriophages for precise microbial community engineering in nitrogen removal: selectively lysing nitrite-oxidizing bacteria (NOB) via targeted lysis, enhancing ammonia-oxidizing and anammox bacteria through auxiliary metabolic genes, and facilitating nutrient redistribution via the viral shunt. We explore the feasibility, technical challenges, and potential biosafety risks, offering a roadmap for phage-based advancements in wastewater treatment systems.
Similar content being viewed by others
Introduction
The rapid growth of the global population and urbanization has substantially elevated nitrogen pollution levels1,2, exceeding planetary boundaries and driving demand for sustainable wastewater treatment technologies. Traditional nitrification-denitrification processes, although widely implemented, are increasingly inadequate to meet the dual challenges of stringent effluent standards and carbon neutrality goals due to their high energy consumption, substantial carbon source requirements, and significant greenhouse gas emissions. The discovery of nitrite accumulation during wastewater treatment in 1975 laid the foundation for revolutionary nitrogen removal technologies3. This breakthrough led to the development of nitrite-based novel nitrogen removal processes, fundamentally represented by partial nitrification (NH₄⁺ to NO₂⁻) coupled with anaerobic ammonium oxidation (anammox NH₄⁺ and NO₂⁻ to N₂). These approaches represent a major shift, offering remarkable advantages: 60% reduction in aeration energy, 80% decrease in sludge production, and 83% lower N₂O emissions compared to conventional processes4. However, the process efficiency, long-term stability, and scalability of these revolutionary technologies are consistently plagued by the “Nitrite Bottleneck”, a persistent challenge in maintaining optimal nitrite levels, predominantly driven by the robust ecological resilience and competitive activity of nitrite-oxidizing bacteria (NOB)5,6. These NOB either rapidly consume nitrite intended for desired pathways, such as anammox, or outcompete ammonia-oxidizing bacteria (AOB) for essential resources. Despite numerous attempts to suppress NOB through conventional strategies encompassing wastewater treatment operational parameter control (e.g., adjustments in dissolved oxygen levels, sludge retention time)7,8, and chemical inhibitor addition9, these methods often lack the requisite precision, selectivity, and long-term efficacy in sustaining nitrite levels and system stability10,11. Thus, there is an urgent need for a disruptive technology that can fundamentally reshape the paradigm of microbial community regulation to decisively overcome the Nitrite Bottleneck.
Bacteriophages, with their exquisite host specificity and diverse life strategies12, offer unique opportunities for precisely maintaining the nitrite bottleneck through targeted microbial community engineering in innovative nitrite-based nitrogen removal processes. Unlike conventional physicochemical approaches that rely on broad environmental perturbations, phage-mediated regulation allows targeted lysis and functional enhancement at the single-species level13,14. Theoretically, lysis uses phages’ lytic cycle (receptor binding, replication, host rupture), while enhancement involves temperate phages transferring AMGs via horizontal gene transfer. Yet current related research in nitrite-based nitrogen removal remains exploratory15,16. Cutting-edge research in environmental virology has uncovered the extensive presence and activity of bacteriophages in wastewater treatment systems17,18. Critically, phage-mediated interventions have demonstrated success in related applications: targeted lysis of problematic bacterial populations enables specific community restructuring without compromising overall system functionality19,20, and Phage-encoded auxiliary metabolic genes (AMGs) associated with ammonia oxidation (e.g., amoABC) and nitrite reduction (e.g., nirK) enhance nitrogen transformation efficiency in activated sludge systems by modulating host metabolic adaptability21,22,23. These findings indicate that phages have the potential to serve as precision tools for sustaining the nitrite bottleneck by selectively suppressing nitrite-oxidizing bacteria (NOB) while enhancing functional bacteria in nitrite-driven processes.
Despite their great potential, bacteriophages have not been extensively investigated for maintaining the nitrite bottleneck in innovative nitrite-based nitrogen removal processes. A systematic analysis is urgently required to tackle the complexity of phage-host dynamics, potential ecological risks, and hurdles in practical implementation. This perspective presents an innovative solution to surmount this bottleneck by using bacteriophages as precise tools for engineering microbial communities. We put forward a conceptual framework: lytic phages can selectively target and eliminate nitrite-oxidizing bacteria (NOB). Meanwhile, temperate phages can enhance the metabolic performance of ammonia-oxidizing and anammox bacteria through the delivery of auxiliary metabolic genes. Additionally, phage-mediated lysis promotes nutrient recycling within the microbial community via the viral shunt mechanism. Leveraging recent advancements in environmental virology, synthetic biology, and multi-omics technologies, we rigorously assess the feasibility, technical challenges, and biosafety implications of implementing phage-based regulation in engineered wastewater systems. In conclusion, we propose a research roadmap to integrate intelligent, phage-based control systems into next-generation treatment processes. This approach reimagines the Nitrite Bottleneck not as a limiting factor, but as a manageable ecological checkpoint, thus promoting resilient and sustainable nitrogen removal in wastewater treatment.
The enduring challenge: why the nitrite bottleneck persists
Sustaining the nitrite bottleneck, the central prerequisite for innovative nitrogen removal strategies based on partial nitrification, is the cornerstone of these advanced processes. However, ensuring its stability continues to be one of the most persistent challenges in the field24. The reason for this persistence lies not in insufficient efforts but in a fundamental mismatch: the sophisticated ecological strategies of NOB continuously outmaneuver the comparatively blunt instruments of conventional process control.
Mechanisms of NOB resilience and adaptability
The challenge persists because NOB are not a monolithic target, but rather a highly adaptable and resilient consortium25, rendering their selective inhibition a complex scientific and engineering problem. The instability of the bottleneck is mainly due to the resilient ecology of NOB, which uses a multi-faceted approach to survive and dominate in the competitive wastewater environment. This resilience stems from their extraordinary adaptability. NOB populations exhibit substantial functional diversity, enabling them to thrive across a broad range of operating conditions. Different evolutionary lineages have adapted to specific ecological niches: Lineage I dominates in high-nitrite environments (>0.5 mM), while Lineage II excels in low-nitrite conditions (<0.1 mM) with fluctuating dissolved oxygen (DO) levels, which are common in systems attempting to suppress them26. This ecological differentiation enables the NOB consortium to shift its activity to alternative opportunities when one closes, recovering up to 78% of its activity after 60 days of system operation21. This adaptability is underpinned by immense genetic diversity, with up to 121 species-level operational taxonomic units (OTUs) of the nxrB gene identified in activated sludge26. This metabolic adaptability also includes denitrification facilitated by genes like narG and napA under low-oxygen conditions, with expression levels increasing by 2.1 to 3.2 times21, thereby maintaining energy metabolism through nitrite reduction. A more profound challenge is the evolution of complete ammonia oxidation (comammox) in some Nitrospira strains, enabling them to oxidize ammonia directly to nitrate27,28. These organisms exhibit substrate affinity 10 to 30 times higher than conventional AOB (Km ~0.84 μM vs 6 to 27 μM)21. Comammox undermines the core concept of partial nitrification by competing for ammonia right at the beginning, rendering nitrite accumulation nearly impossible. Furthermore, NOB can establish a competitive advantage over anammox bacteria, as the latter are more susceptible to inhibition when nitrogen level fluctuates6,28. NOB also exhibits strong defensive traits, such as the ability to form biofilms and generate extracellular polymeric substances that create diffusion barriers, shielding them from external stressors29.
Limitations of current NOB suppression strategies
Current engineering strategies consistently fall short against such a highly adaptable opponent. The most common strategy involves adjusting DO levels to exploit the different oxygen affinities of AOB (oxygen half-saturation coefficient 0.3 to 1.2 mg/L) and NOB (1.2 to 2.5 mg/L)30. Although this approach can initially achieve nitrite accumulation rates (NAR) over 90%, the success is often short-lived: as low-DO-adapted NOB strains proliferate over time, system performance declines steadily, leading to a drop in NAR to 63% after 120 days of operation6,31. Similarly, intermittent aeration, which alternates between anoxic/aerobic phases, is an inconsistent strategy, as its effectiveness largely depends on the initial microbial community structure32,33. While some studies report high efficiency, achieving NAR values of up to 92.5%34, others using different sludge sources report a complete failure, with the NAR dropping to 0%33. Physical controls, such as adjusting sludge retention time (SRT), have a limited operational range; the strategy works well at high temperatures (30 °C)35 but fails at the standard 20 to 25 °C36,37 range where AOB and NOB growth rates align. The use of chemical inhibitors and advanced biofilm systems also reveals fundamental limitations. The addition of inhibitors like hydroxylamine can sharply reduce nitrate production by 89% within 24 hours38. However, this effect is transient; nitrate levels rebound to 92% of their initial concentration within 48 hours of stopping the treatment39. This approach not only incurs high operational costs but also carries the risk of promoting resistant strains. Ironically, biofilm and granular sludge systems, designed to enhance microbial processes, can actually provide a safe harbor for NOB. Due to microscale redox gradients resulting from diffusion limitation, microbial communities become stratified40, creating protected inner zones where NOB with high oxygen affinity thrive, evading external control efforts41. In summary, the nitrite bottleneck endures because we are using static, indiscriminate, and unsustainable strategies to combat a sophisticated, adaptable, and multifaceted biological system. This ongoing challenge clearly highlights the need for more than just environmental measures and to develop advanced regulatory tools that provide targeted, sustained, and intelligent species-level control to effectively manage the resilience of these vital microorganisms.
Harnessing phages for precision control of the nitrite bottleneck
To overcome the entrenched resilience of nitrite-oxidizing bacteria (NOB), we need a fundamental shift in strategy-replacing indiscriminate system-level controls with targeted, phage-enabled precision engineering. This approach re-imagines bacteriophages not as passive environmental entities, but as a programmable arsenal of biological tools capable of reshaping microbial communities at the species level. Through the strategic use of various phage-mediated mechanisms, the competitive dominance of NOB can be dismantled while simultaneously fortifying the desired functional guilds, thereby transforming the nitrite bottleneck from an intractable problem into a controllable node.
Direct control through selective lysis of NOB
One of the most effective ways to mitigate NOB dominance is through selective elimination via targeted lysis. This approach is based on the lytic cycle of bacteriophages, where a phage recognizes and injects its genome into a targeted host, takes over the cell’s metabolic machinery to produce viral progeny, and ultimately ruptures the cell wall using lytic enzymes like endolysins to release new virions42,43. This mechanism provides the foundation for a Kill the Winner (KtW) strategy. In this model, phages selectively attack the most abundant or fastest-growing members of a community, maintaining balance by preventing any single species from monopolizing resources44. This dynamic has been observed in aquatic systems, where viral genotypes and their susceptible host strains exhibit cyclical fluctuations, maintaining ecological balance. This principle can be designed to precisely target the nitrite bottleneck. Although no lytic phages infecting canonical nitrite-oxidizing bacteria (NOB) such as Nitrospira or Nitrobacter have been experimentally isolated to date, several lines of robust peer-reviewed evidence strongly indicate that phages actively interact with nitrifying microorganisms and can be harnessed for targeted biocontrol in engineered systems. For example, Lee et al.45 employed a differential-inhibition enrichment strategy to identify 61 high-quality viral genomes actively infecting soil nitrifiers, including viruses linked to Nitrospira-like NOB, indicating that NOB and phage interactions are ecologically widespread and detectable in situ45. Likewise, multiple phage-responsive genetic elements (proviruses, CRISPR spacers, defense islands) have been identified within cultivated Nitrospira and Nitrobacter genomes, further supporting the long-term co-evolution between NOB and associated viruses.
Importantly, while bona fide NOB-specific phages have not yet been isolated, closely related nitrifier-infecting phages have been successfully cultured and experimentally validated, providing strong precedents for isolating NOB-targeting viruses. Quirós et al. isolated φNF-1, the first virulent phage infecting Nitrosomonas europaea, N. communis, and N. nitrosa. φNF-1 strongly inhibited host growth and reduced NH₄⁺ oxidation rates to levels comparable to cell-free controls, demonstrating that phage-mediated suppression of nitrification is feasible in controlled systems46. Similarly, Choi showed that multiple environmental stressors induce prophage activation in Nitrosospira multiformis, causing substantial viral particle release (up to 2.3 × 10¹⁰ VLPs within 5 h at pH 5) and leading to measurable declines in ammonia-oxidation activity47. These studies collectively confirm that nitrifiers-including taxa phylogenetically adjacent to NOB, are susceptible to phage-mediated lysis.
In addition to evidence within nitrifiers, successful phage applications in full-scale and laboratory-scale wastewater systems provide robust proof-of-concept that targeted phage lysis can restructure activated-sludge communities without impairing treatment performance. Kotay isolated a Myoviridae phage infecting the filamentous bacterium Haliscomenobacter hydrossis and achieved a reduction of sludge volume index (SVI) from 155 to 105, corresponding to 32% improvement in settling, while maintaining 100% ammonia removal efficiency47. In other WWTP case studies, phage GTE7 effectively lysed multiple foaming-associated Gordonia and Nocardia species and completely prevented foam stabilization in activated sludge reactors48.
Collectively, these findings offer robust scientific support for proposing phage-mediated selective lysis as a viable strategy to suppress NOB. Although NOB-infecting phages have yet to be isolated, the demonstrated infectivity of phages toward closely related nitrifiers, combined with multiple successful phage-based interventions in wastewater treatment systems, provides a compelling precedent supporting the feasibility of developing targeted NOB-phage biocontrol in engineered nitrite-based nitrogen removal processes.
Lytic phages specifically infecting canonical NOB (e.g., Nitrospira, Nitrobacter) have not been isolated or experimentally validated to date. Existing evidence for NOB-phage associations is therefore limited to genome-based predictions and in silico analyses45. Several complete NOB genomes contain putative prophage regions identified using tools such as VirSorter2 and CheckV49, and numerous NOB-associated viral contigs have been deposited in the IMG/VR database50. For example, the viral contig IMGVR_UViG_2829791209_000001, derived from Nitrobacter vulgaris DSM 10236, is classified within Caudoviricetes with high confidence, suggesting the presence of prophage remnants rather than active lytic phages-akin to the stable lysogeny observed in other host-virus systems51. Likewise, environmental metagenomes contain additional Nitrospira-associated viral signatures, yet none have been experimentally confirmed as a functional lytic phage infecting NOB45. Thus, isolating bona fide NOB-infecting phages remains an important research frontier, and current NOB-phage discussions must be framed as hypothesis-generating rather than evidence-based. These knowledge gaps highlight the urgency of phage discovery in NOB-enriched engineered wastewater systems; isolating or engineering phages that exclusively target dominant NOB (e.g., Nitrospira) could directly weaken their competitive edge via continuous predation, stabilizing the partial nitrification pathway52. The feasibility of such targeted biocontrol in wastewater has already been demonstrated. For instance, targeted lysis of the filamentous bacterium Haliscomenobacter hydrossis key contributor to sludge bulking-led to a 32% reduction in the Sludge Volume Index (SVI) from 155 to 105, while maintaining 100% ammonia nitrogen removal efficiency47. This successful application serves as a powerful proof-of-concept, suggesting that a similar phage-based “precision strike” could be deployed to suppress NOB and resolve the nitrite bottleneck.
Indirect control via enhancement of functional guilds
A comprehensive strategy should not only limit competitors but also actively preserve and strengthen key functional bacteria like ammonia-oxidizing bacteria (AOB) and anaerobic ammonium oxidation (anammox) bacteria. According to the “Piggyback the Winner” (PtW) model, temperate (lysogenic) phages tend to integrate into the host genome as prophage rather than lysing the cell right away, particularly in high-host-density environments such as AOB-rich biofilms53,54. This lysogenic state confers a key advantage via a mechanism termed “superinfection exclusion”, wherein the resident prophage protects its host from infection and lysis by other (often related) phages53,55. This effectively transforms the phage into a guardian, shielding the dominant functional bacteria against viral threats and maintaining their population stability. In addition to providing passive protection, phages can actively enhance their hosts’ metabolic capabilities through the horizontal transfer of Auxiliary Metabolic Genes (AMGs)56,57,58. These phage-encoded genes have the potential to modulate the host’s metabolism, often benefiting both the host and the phage. For example, metagenomic and metatranscriptomic analyses of wastewater treatment plants have revealed that phage-borne ammonia monooxygenase subunit genes (amoC) are actively expressed at levels ~1.7 times higher than the host’s own amoC59. This transcriptional evidence suggests that phage-carried amoC might contribute to the host AOB’s ammonia oxidation process; however, direct experimental proof of an enhanced nitrogen metabolism in the host due to this phage gene has not yet been demonstrated. Similarly, some phages carry and express genes involved in queuosine synthesis (e.g., folE, queD, queE), which may improve host adaptability under stress by modulating RNA modification59. Phage-mediated gene transfer can also bolster host resistance to heavy metals, indirectly preserving the ecological niche of sensitive functional microbes like anammox bacteria60. This “metabolic hijacking” strategy thus serves as a powerful tool to improve the fitness and competitive advantage of AOB and anammox bacteria.
Ecosystem-level control by modulating nutrient fluxes
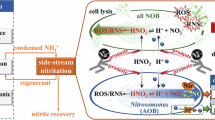
In addition to targeting individual cells, phages can regulate entire microbial ecosystems on a larger scale through a process known as the “viral shunt“61,62. Bacteriophages lyse host cells by rupturing their membrane and releasing the entire cellular contents-a rich soup of dissolved organic matter (DOM), carbon, nitrogen, and phosphorus-directly into the environment63. This quick release of nutrients diverts resources away from higher trophic levels and immediately supplies them to the surrounding microbial community64. This process occurs on a massive scale in marine ecosystems, viruses lyse approximately 20% of the bacterial biomass daily65, contributing an estimated 150 billion tons of carbon flux annually66. This constant redistribution of resources can serve as an effective tool for ecosystem engineering. In an innovative nitrogen removal system, the targeted lysis of abundant NOB would not only eliminate the competitor but also trigger a localized viral shunt. The nutrients released from the lysed NOB could then be readily absorbed by other bacteria in the system. This process could selectively favor slower-growing but desirable species like anammox bacteria, providing them with a pulse of readily available substrates. By selectively lysing NOB, phage-based therapy has the potential to reshape the flow of resources within the microbial food web, establishing a new ecological balance that inherently favors the desired nitrogen removal pathways (Fig.1).
Proposed phage interventions: a Lytic phages execute targeted NOB lysis via “Kill-the-Winner” dynamics; b Temperate phages employ “Piggyback the Winner” (PtW) strategy-prioritizing infection of dominant AOB/anammox populations-to form lysogens (integrated phage genomes in host cells), which transfer auxiliary metabolic genes (AMGs) via horizontal gene transfer (HGT) to enhance their nitrite utilization and metabolic flexibility; c Viral shunt redistributes nutrients from lysed cells to functional bacteria, rewiring ecosystem nutrient flux. Adapted with modifications from Freepik.
From concept to reality: charting the roadmap for phage-based engineering
Although phage-based regulation holds great potential, translating this concept into a reliable and safe technology will require extensive research efforts. Translating the principle into practice requires addressing three key challenges: innate phage host specificity, the complexities of integrating them into dynamic engineered systems, and the need for stringent biosafety measures. A well-defined strategy to overcome these challenges is essential for harnessing the full potential of phages as advanced microbial control solutions in wastewater treatment.
The host range challenge from phage cocktails to synthetic biology
The main challenge in effective phage therapy is its narrow host range; a single phage strain often infects only a specific bacterial strain, such as Nitrospira multiformis. To tackle this issue, a tiered approach is necessary to enhance treatment effectiveness67. A straightforward approach involves using “phage cocktails”-blends of multiple phage strains-to effectively target the diverse NOB community. At a more advanced level, combining machine learning and bioinformatics with metagenomic data can accelerate the screening and discovery of novel, naturally occurring broad-spectrum phages while improving predictions of phage-host interactions68. The most promising long-term solution, though, involves synthetic biology and gene-editing tools. With technologies like CRISPR-Cas, scientists could modify phage tail fiber proteins, essentially “programming” their host range to create bespoke phages capable of targeting the most problematic NOB strains within a specific system69,70 (Fig. 2).
Addressing narrow host range requires tiered approaches: Phage cocktails broaden NOB targeting; Machine learning predicts phage-host interactions via genomic signatures; Synthetic biology enables CRISPR-guided tail fiber engineering to reprogram infectivity against resilient NOB strains (e.g., Nitrospira). Created with BioGDP.com88.
The implementation challenge: ensuring efficacy and stability in engineered systems
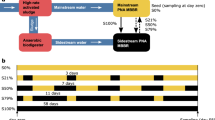
Beyond host range, ensuring phage effectiveness and persistence within complex engineered systems (e.g., wastewater reactors) poses a major implementation challenge. To operationalize such phage-based strategies, it is essential to delineate the physicochemical boundaries under which phages remain active and to establish quantitative metrics that can reliably indicate successful control of target guilds such as NOB. Recent studies on phages in engineered and terrestrial environments provide relevant empirical constraints on the physicochemical windows that support productive infection. Various studies quantified ionic strength thresholds for infection of E. coli by dsDNA and ssRNA phages, demonstrating that while many alkaline earth and alkali metals facilitate infection, phage-specific EC50 values can exceed those typically observed in freshwater systems71. In activated sludge, a Myoviridae phage infecting Haliscomenobacter hydrossis retained considerable infectivity across pH 5-8 and remained structurally stable even under 42 °C exposure47. Likewise, in ammonia-oxidizing cultures, the Podoviridae-like phage ΦNF-1 preserved infectivity over 7 days at pH ≈8 and achieved titers up to 10⁸ PFU mL⁻¹, failing only below ≈10² PFU mL⁻¹46. Complementary work examined the SRB-associated phage SRB7757 in corrosion-relevant reactors, tracking phage titers, host cell concentrations, and sulfide production over 48h72. Collectively, these studies provide experimentally grounded bounds on temperature, pH, ionic strength, and redox chemistry that can guide preliminary specification of operational windows for NOB-targeting phage application in PN/A systems.
Equally important are methodological precedents that establish how to quantify successful phage-mediated control. In the ΦNF-1-Nitrosomonas system, qPCR targeting the amoB gene was used to track host population decline while simultaneously quantifying phage genome copies, confirming active phage replication71. Functional inhibition was assessed through ammonium measurements using the salicylate method, where infected cultures exhibited markedly reduced NH₄⁺ consumption relative to controls46. In activated sludge, classical plaque assays, Sludge Volume Index (SVI) measurements, and epifluorescence microscopy to track virus-like particles (VLPs) demonstrated a 32% SVI reduction (155 to 105 mL g⁻¹) without impairing COD or nutrient removal47. For SRB7757, phage performance was quantified by monitoring sulfide concentrations, phage titers via double-layer plating, and host abundance through OD₆₀₀ measurements73. Together, these multi-layered indicators-functional gene dynamics, nitrogen species trends, settling performance, phage titers, and VLP counts-provide a transferable framework for defining success criteria for future NOB-targeting phage interventions.
Phage activity is influenced by environmental factors (e.g., temperature) that may inhibit lysis or accelerate host demise, requiring adaptive optimization of process parameters74. A key research focus is the development of effective dosing strategies, such as analyzing the effects of pulsed versus continuous dosing on microbial community structure using advanced multi-omics approaches75. Furthermore, to maintain a stable and active phage population, innovative approaches such as phage immobilization or integrating them directly with biofilm and granular sludge systems could be explored to establish resilient “phage-reactive layers.” Finally, a key strategy will be to explore the synergistic effects of phage treatment alongside existing process controls. For instance, integrating phage treatment with suboptimal conditions like low temperatures or controlled sludge retention times could significantly enhance NOB suppression, resulting in a synergistic effect that exceeds individual measures60. A critical challenge for phage-based suppression of NOB lies in the rapid evolutionary responses of bacterial hosts. Extensive experimental evidence shows that bacteria often acquire phage resistance through receptor modification, surface-structure alteration, or CRISPR-Cas immunity, with measurable fitness trade-offs in receptor-modified mutants76. Such resistance can emerge quickly: for example, in controlled evolution experiments, more than 50% of E. coli populations became resistant to an untrained phage within 3 days, and resistance reached 100% by day 10, whereas resistance against a coevolved, host-trained phage did not appear until approximately 15-30 days and required multiple mutations, occurring at a ~100-fold lower frequency77. Phage cocktails can further delay resistance: combining two or three complementary phages prevented detectable resistant mutants for 24–48 hours in vitro78, whereas resistance emerged substantially earlier under single-phage treatments. These findings highlight the necessity of incorporating predictive eco-evolutionary models to anticipate arms-race dynamics between NOB and their associated phages and to design intervention strategies that remain effective under rapid coevolution. Nevertheless, the practical application of these strategies still requires consideration of ecological risks. Compared with the aforementioned conventional strategies, phage engineering strategies based on genetic modification offer distinct advantages in overcoming bacterial resistance, with core mechanisms divided into two directions: precise targeting of resistance genes and optimization of host recognition ability. Against bacterial resistance mediated by CRISPR-Cas immunity, the scarless Cas9-assisted Recombineering (no-SCAR) system enables efficient editing of phage genomes to modify tail fiber proteins and lysozyme genes79. The engineered phages can simultaneously overcome both “receptor mutation resistance” and “biofilm resistance” of Pseudomonas aeruginosa, resulting in a 3-log reduction in target bacterial load within 72 hours in vitro compared to the original phage group, with a 40% increase in bactericidal efficiency. Directed mutagenesis of the host recognition region in phage tail fiber proteins is also an effective strategy; Yehl obtained T7 phage tail fiber mutants through saturation mutagenesis80, which delayed the emergence of resistance by 24 to 48 hours in Escherichia coli populations that originally developed 100% resistance within 10 days, and reduced the final proportion of resistant strains to less than 15%. More groundbreaking is the dual-functional engineering strategy of “CRISPR-Cas loading + tail fiber modification”: integrating a CRISPR-Cas system targeting core metabolic genes (e.g., nxrB) of drug-resistant bacteria into the phage genome while modifying tail fiber proteins to recognize novel receptors. This engineered phage achieved a 92% clearance rate in a mouse model infected with carbapenem-resistant Enterobacteriaceae (CRE), and no recurrence of resistant strains was detected during a 14-day continuous observation period81. Similarly, adaptive mutant phages obtained through 30-day phage-bacteria coevolution screening exhibited a 75% 72-hour inhibition rate against extensively drug-resistant (XDR) Klebsiella pneumoniae, which was significantly superior to that of the original phages (27% to 42%)82. Additionally, CRISPR-Cas-assisted personalized phage engineering technology allows customization of engineered phages based on the resistance gene characteristics of specific drug-resistant bacteria, further enhancing the targeting of resistance83. These data confirm that engineered phages can provide an efficient solution to the resistance issue in NOB inhibition through the dual mechanism of “resistance evasion and precise killing”, complementing conventional strategies and opening up new avenues for the long-term application of phages in NOB inhibition.
Beyond evolutionary constraints, phage-mediated lysis can generate broader ecological consequences within complex wastewater microbiomes. Lysis releases dissolved organic matter, intracellular nitrogen, and micronutrients that may stimulate opportunistic or undesirable taxa. In a controlled sponge-microbiome experiment, selective phage disturbance triggered a bloom of opportunistic Vibrio spp., increasing from nearly 0% to approximately 13% relative abundance within 72 hours84. Similar risks exist in wastewater systems, where phages are abundant and tightly associated with flocs and granules; nutrient release from phage lysis may unintentionally promote filamentous bacteria or pathogens85, potentially counteracting intended NOB suppression and destabilizing functional guilds essential for PN/A performance. Furthermore, community context itself can modulate phage-host interactions, with multispecies assemblages shown to suppress both phage and host densities and even constrain coevolutionary trajectories. Taken together, these insights underscore the need for phage-based NOB control strategies to integrate evolutionary forecasting, dynamic phage-cocktail design, and ecosystem-level modeling to minimize unintended ecological cascades.
The biosafety challenge: mitigating unintended ecological consequences
A major issue regarding the use of phages is biosafety, particularly the risk of unintended ecological consequences. A primary hazard is phage-mediated horizontal gene transfer, especially the potential dissemination of antibiotic resistance genes (ARGs)86 or other virulence factors. Studies reveal that ARGs, including sul1 and tetM, are frequently found in the viral fraction of wastewater, accounting for 35% and 22% of detected genes87, respectively. To reduce this risk, a comprehensive, multi-layered safety strategy is essential. First, every potential phage candidate must undergo rigorous genomic screening to select suitable “chassis” strains that lack known virulence or antibiotic resistance genes. Second, synthetic biology can be employed to integrate safety switches into the phage genome, such as “self-destruction” mechanisms that activate under certain conditions or limit the phage’s ability to transfer genetic material. Finally, comprehensive and long-term pilot-scale studies are crucial before any large-scale application. These studies, conducted in reactors such as sequencing batch reactors (SBRs) that mimic real-world conditions, should employ multiple omics approaches, such as viral metagenomics, transcriptomics, and metabolomics, to thoroughly assess the ecological risks and monitor the long-term effects of phage application on the indigenous microbial community (Fig. 3)
Phage-based strategies for NOB control in PN/A systems (contextualized with implementation & biosafety considerations) a Phage AMGs Regulate Nitrogen Metabolism: Phages deliver auxiliary metabolic genes (AMGs) to modulate key nitrogen transformation steps (e.g., NH₄⁺-NO₂⁻-N₂ conversion), aligning with efforts to reshape microbial nitrogen pathways in phage-mediated metabolic regulation; b Biofilm Phage Lysis: Challenges & Process Synergy: Phages target biofilm-forming bacteria (mitigating antibiotic resistance gene dissemination, a biosafety concern) and exert host-specific NOB inhibition under conditions (35°C, SRT 15 d) matching PN/A operational stability windows; c Synbio Phage Engineering via Metagenomics & CRISPR: Phage genomes from samples are characterized by viral metagenomics (to screen safe chassis) then edited via CRISPR/Cas (e.g., tail fiber modification, a strategy against NOB resistance) to construct functional engineered phages.
Concluding perspective and outlook
The long-standing battle against the ecological resilience of nitrite-oxidizing bacteria (NOB) has revealed the fundamental limitations of our existing control strategies. Relying on macroscale environmental manipulations (e.g., dissolved oxygen regulation, sludge retention time adjustment) is analogous to using a blunt tool when fine precision is required. This Perspective suggests the need for fundamental change in our mindset: advancing from broad environmental control to the precise, targeted manipulation of microbial species. In this new framework, the “Nitrite Bottleneck” is no longer seen as an intractable problem, but is redefined as a controllable ecological node, a critical juncture in the nitrogen cycle that can be precisely regulated with the appropriate tools.
We envision a future where wastewater treatment systems are equipped with ‘intelligent’ phage-based regulatory platforms capable of autonomous optimization. Such systems would integrate real-time multi-omics monitoring with artificial intelligence to detect incipient increases in problematic NOB populations and trigger the release of personalized synthetic phage cocktails. These are not merely natural viruses, but are purposefully engineered biological constructs. Through synthetic biology, their host range can be precisely programmed by modifying tail fiber proteins to target specific NOB strains. Additionally, their genomes can be edited to incorporate auxiliary metabolic genes, enhancing the activity of key functional groups like AOB or anammox bacteria. Furthermore, these engineered phages can be equipped with safety features, including self-destruction mechanisms, to address biosafety risks such as the unintended horizontal transfer of antibiotic resistance genes.
Translating this vision into reality is a bold endeavor that requires a collaborative, multidisciplinary research approach. To ensure the success and ecological sustainability of phage application, meticulous pilot-scale studies in controlled environments (e.g., sequencing batch reactors, SBRs) are essential. Although research on NOB-targeting phages is still in its early stages, the way forward is clear. It demands seamless integration of synthetic biology for phage design, artificial intelligence for predictive host-phage modeling, and process engineering for real-world applications. Despite ongoing challenges with host specificity and environmental stability, the potential to create resilient, high-efficiency nitrogen removal systems justifies the pursuit. Ultimately, harnessing the power of phages will not only resolve the nitrite bottleneck but also pave the way for a new era of precision eco-engineering in sustainable water management.
Data availability
No datasets were generated or analyzed during the current study.
References
Tian, X. et al. Will reaching the maximum achievable yield potential meet future global food demand?. J. Clean. Prod. 294, 126285 (2021).
Rockstrom, J. et al. A safe operating space for humanity. Nature 461, 472–475 (2009).
Voets, J., Vanstaen, H. & Verstraete, W. Removal of Nitrogen From Highly Nitrogenous Wastewaters. J. Water Pollut. Control Fed. 47, 394–398 (1975).
Kartal, B., Kuenen, J. G. & van Loosdrecht, M. C. M. Sewage treatment with Anammox. Science 328, 702–703 (2010).
Oshiki, M., Shimokawa, M., Fujii, N., Satoh, H. & Okabe, S. Physiological characteristics of the anaerobic ammonium-oxidizing bacterium ‘Candidatus Brocadia sinica’. Microbiology 157, 1706–1713 (2011).
Wang, Z. et al. Unravelling adaptation of nitrite-oxidizing bacteria in mainstream PN/A process: Mechanisms and counter-strategies. Water Res. 200, 117239 (2021).
Feng, L. et al. Intracellular electron competition in response to the oxygen pressure of the aerobic denitrification process in an O2-based membrane biofilm reactor (MBfR) for nitrate removal. Sci. Total Environ. 875, 162592 (2023).
Huang, R., Meng, T., Liu, G., Gao, S. & Tian, J. Simultaneous nitrification and denitrification in membrane bioreactor: Effect of dissolved oxygen. J. Environ. Manag. 323, 116183 (2022).
Zhang, X. et al. Enhancement of nitrite production via addition of hydroxylamine to partial denitrification (PD) biomass: Functional genes dynamics and enzymatic activities. Bioresour. Technol. 318, 124274 (2020).
Zhao, J. et al. Selective Enrichment of Comammox Nitrospira in a Moving Bed Biofilm Reactor with Sufficient Oxygen Supply. Environ. Sci. Technol. 56, 13338–13346 (2022).
Zheng, M. et al. Predominance of comammox bacteria among ammonia oxidizers under low dissolved oxygen condition. Chemosphere 308, 136436 (2022).
Teklemariam, A. D. et al. The Battle between Bacteria and Bacteriophages: A Conundrum to Their Immune System. Antibiotics 12, 381 (2023).
Saw, P. E. & Song, E.-W. Phage display screening of therapeutic peptide for cancer targeting and therapy. Protein Cell 10, 787–807 (2019).
Karimi, M. et al. Bacteriophages and phage-inspired nanocarriers for targeted delivery of therapeutic cargos. Adv. Drug Deliv. Rev. 106, 45–62 (2016).
Penades, J. R., Chen, J., Quiles-Puchalt, N., Carpena, N. & Novick, R. P. Bacteriophage-mediated spread of bacterial virulence genes. Curr. Opin. Microbiol. 23, 171–178 (2015).
Touchon, M., Moura de Sousa, J. A. & Rocha, E. P. Embracing the enemy: the diversification of microbial gene repertoires by phage-mediated horizontal gene transfer. Curr. Opin. Microbiol. 38, 66–73 (2017).
Mathieu, J., Yu, P., Zuo, P., Da Silva, M. L. B. & Alvarez, P. J. J. Going viral: emerging opportunities for phage-based bacterial control in water treatment and reuse. Acc. Chem. Res. 52, 849–857 (2019).
Sohail, H. A. et al. Bacteriophages: emerging applications in medicine, food, and biotechnology. Phage 1, 75–82 (2020).
Wang, D. et al. Distributions, interactions, and dynamics of prokaryotes and phages in a hybrid biological wastewater treatment system. Microbiome 12, 134 (2024).
Qi, H. et al. Symbiotic bacteriophages exhibit multiple adaptive strategies in activated sludge flocs and contribute to floc stability. Chem. Eng. J. 492, 152448 (2024).
Daims, H. et al. Complete nitrification by Nitrospira bacteria. Nature 528, 504 (2015).
Wang, Y. et al. Heterogeneity of soil bacterial and bacteriophage communities in three rice agroecosystems and potential impacts of bacteriophage on nutrient cycling. Environ. Microbiome 17, 17 (2022).
Jin, M., Cai, L., Lu, L., Yu, M. & Zhang, R. Combined metabolomic and genomic analyses reveal phage-specific and infection stage-specific alterations to marine Roseobacter metabolism. ISME Commun. 5, ycaf047 (2025).
Duan, H., Ye, L., Lu, X. & Yuan, Z. Overcoming nitrite oxidizing bacteria adaptation through alternating sludge treatment with free nitrous acid and free ammonia. Environ. Sci. Technol. 53, 1937–1946 (2019).
Yu, L. et al. The differential proliferation of AOB and NOB during natural nitrifier cultivation and acclimation with raw sewage as seed sludge. RSC Adv. 10, 28277–28286 (2020).
Gruber-Dorninger, C. et al. Functionally relevant diversity of closely related Nitrospira in activated sludge. ISME J. 9, 643–655 (2015).
Ghimire-Kafle, S., Weaver, M. E. & Bollmann, A. Ecophysiological and genomic characterization of the freshwater complete ammonia oxidizer Nitrospira sp. Strain BO4. Appl. Environ. Microbiol. 89, (2023).
Wei, Y., Chen, Y., Xia, W., Ye, M. & Li, Y.-Y. Dynamic pulse approach to enhancing mainstream Anammox process stability: Integrating sidestream support and tackling nitrite-oxidizing bacteria challenges. Bioresour. Technol. 395, 130327 (2024).
Duan, H. et al. Nitrite oxidizing bacteria (NOB) contained in influent deteriorate mainstream NOB suppression by sidestream inactivation. Water Res. 162, 331–338 (2019).
Ali, M. et al. Physiological characterization of anaerobic ammonium oxidizing bacterium ‘Candidatus Jettenia caeni’. Environ. Microbiol. 17, 2172–2189 (2015).
Cui, B. et al. The effect of dissolved oxygen concentration on long-term stability of partial nitrification process. J. Environ. Sci. 90, 343–351 (2020).
Al-Hazmi, H. E., Yin, Z., Grubba, D., Majtacz, J. B. & Makinia, J. Comparison of the Efficiency of Deammonification under Different DO Concentrations in a Laboratory-Scale Sequencing Batch Reactor. Water 14, 368 (2022).
Yuan, Y., Xie, Y., Xu, P. & Li, X. Verification of inhibition effects of anoxic/aerobic alternation on NOB in nitrosation system under mainstream conditions. J. Water Process Eng. 45, 102479 (2022).
Qiu, J., Li, X., Peng, Y. & Jiang, H. Advanced nitrogen removal from landfill leachate via a two-stage combined process of partial nitrification-Anammox (PNA) and partial denitrification-Anammox (PDA). Sci. Total Environ. 810, 151186 (2022).
Hubaux, N., Wells, G. & Morgenroth, E. Impact of coexistence of flocs and biofilm on performance of combined nitritation-anammox granular sludge reactors. Water Res. 68, 127–139 (2015).
Dong, W., Lu, G., Yan, L., Zhang, Z. & Zhang, Y. Characteristics of pellets with immobilized activated sludge and its performance in increasing nitrification in sequencing batch reactors at low temperatures. J. Environ. Sci. 42, 202–209 (2016).
Seuntjens, D. et al. Mainstream partial nitritation/anammox with integrated fixed-film activated sludge: Combined aeration and floc retention time control strategies limit nitrate production. Bioresour. Technol. 314, 123711 (2020).
Li, J. et al. A critical review of one-stage anammox processes for treating industrial wastewater: Optimization strategies based on key functional microorganisms. Bioresour. Technol. 265, 498–505 (2018).
Wang, Z. et al. Nitrite accumulation in comammox-dominated nitrification-denitrification reactors: Effects of DO concentration and hydroxylamine addition. J. Hazard. Mater. 384, 121375 (2020).
Wu, W. et al. Contrasting the relative importance of species sorting and dispersal limitation in shaping marine bacterial versus protist communities. ISME J. 12, 485–494 (2018).
Winkler, M.-K. H. et al. An integrative review of granular sludge for the biological removal of nutrients and recalcitrant organic matter from wastewater. Chem. Eng. J. 336, 489–502 (2018).
Chamblee, J. et al. Endolysin Regulation in Phage Mu Lysis. mBio 13, (2022).
Nair, G. & Jain, V. An intramolecular cross-talk in D29 mycobacteriophage endolysin governs the lytic cycle and phage-host population dynamics. Sci. Adv. 10, eadh9812 (2024).
Voigt, E., Rall, B. C., Chatzinotas, A., Brose, U. & Rosenbaum, B. Phage strategies facilitate bacterial coexistence under environmental variability. PeerJ 9, e12194 (2021).
Lee, S., Hazard, C. & Nicol, G. W. Activity of novel virus families infecting soil nitrifiers is concomitant with host niche differentiation. ISME J. 18, wrae205 (2024).
Quiros, P. et al. Identification of a virulent phage infecting species of Nitrosomonas. ISME J. 17, 645–648 (2023).
Kotay, S. M., Datta, T., Choi, J. & Goel, R. Biocontrol of biomass bulking caused by Haliscomenobacter hydrossis using a newly isolated lytic bacteriophage. Water Res. 45, 694–704 (2011).
Petrovski, S., Seviour, R. J. & Tillett, D. Prevention of Gordonia and Nocardia Stabilized Foam Formation by Using Bacteriophage GTE7. Appl. Environ. Microbiol. 77, 7864–7867 (2011).
Guo, J. et al. VirSorter2: a multi-classifier, expert-guided approach to detect diverse DNA and RNA viruses. Microbiome 9, 37 (2021).
Paez-Espino, D. et al. IMG/VR v.2.0: an integrated data management and analysis system for cultivated and environmental viral genomes. Nucleic Acids Res. 47, D678–D686 (2019).
Baquero, D. P. et al. Stable coexistence between an archaeal virus and the dominant methanogen of the human gut. Nat. Commun. 15, 7702 (2024).
Rodriguez-Brito, B. et al. Viral and microbial community dynamics in four aquatic environments. ISME J. 4, 739–751 (2010).
Knowles, B. et al. Lytic to temperate switching of viral communities. Nature 531, 466–470 (2016).
Silveira, C. B. & Rohwer, F. L. Piggyback-the-Winner in host-associated microbial communities. npj Biofilms Microbiomes 2, 16010 (2016).
Silveira, C. B. & Rohwer, F. L. Piggyback-the-Winner in host-associated microbial communities. npj Biofilms Microbomes 2, 16010 (2016).
Mara, P. et al. Viral elements and their potential influence on microbial processes along the permanently stratified Cariaco Basin redoxcline. ISME J. 14, 3079–3092 (2020).
Hurwitz, B. L. & U’Ren, J. M. Viral metabolic reprogramming in marine ecosystems. Curr. Opin. Microbiol. 31, 161–168 (2016).
Sun, M., Yuan, S., Xia, R., Ye, M. & Balcázar, J. L. Underexplored viral auxiliary metabolic genes in soil: Diversity and eco-evolutionary significance. Environ. Microbiol. 25, 800–810 (2023).
Yuan, L. & Ju, F. Potential auxiliary metabolic capabilities and activities reveal biochemical impacts of viruses in municipal wastewater treatment plants. Environ. Sci. Technol. 57, 5485–5498 (2023).
Wang, D. et al. Niche differentiation and symbiotic association among ammonia/nitrite oxidizers in a full-scale rotating biological contactor. Water Res. 225, 119137 (2022).
Fuhrman, J. A. Marine viruses and their biogeochemical and ecological effects. Nature 399, 541–548 (1999).
Weitz, J. S. & Wilhelm, S. W. Ocean viruses and their effects on microbial communities and biogeochemical cycles. F1000 Biol. Rep. 4, 17 (2012).
Weitz, J. S. et al. A multitrophic model to quantify the effects of marine viruses on microbial food webs and ecosystem processes. ISME J. 9, 1352–1364 (2015).
Shelford, E. J., Middelboe, M., Møller, E. F. & Suttle, C. A. Virus-driven nitrogen cycling enhances phytoplankton growth. Aquat. Microb. Ecol. 66, 41–46 (2012).
Pourtois, J., Tarnita, C. E. & Bonachela, J. A. Impact of Lytic Phages on Phosphorus- vs. Nitrogen-Limited Marine Microbes. Front. Microbiol. 11, (2020).
Suttle, C. A. Viruses in the sea. Nature 437, 356–361 (2005).
Bolsan, A. C. et al. Bacteriophages in wastewater treatment: can they be an approach to optimize biological treatment processes?. Environ. Sci. Pollut. Res. Int 29, 89889–89898 (2022).
Lood, C. et al. Digital phagograms: predicting phage infectivity through a multilayer machine learning approach. Curr. Opin. Virol. 52, 174–181 (2022).
Doudna, J. A. & Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Shalem, O., Sanjana, N. E. & Zhang, F. High-throughput functional genomics using CRISPR-Cas9. Nat. Rev. Genet. 16, 299–311 (2015).
Carlson, H. K. et al. Geochemical constraints on bacteriophage infectivity in terrestrial environments. ISME Commun. 3, 78 (2023).
Bacteriophage application in inhibiting corrosion- producing bacteria | BMC Microbiology | Full Text. https://bmcmicrobiol.biomedcentral.com/articles/10.1186/s12866-025-03952-2.
Zhang, G. et al. Bacteriophage application in inhibiting corrosion-producing bacteria. BMC Microbiol. 25, 241 (2025).
Huang, D. et al. Adaptive strategies and ecological roles of phages in habitats under physicochemical stress. Trends Microbiol. 32, 902–916 (2024).
Letten, A. D. & Ludington, W. B. Pulsed, continuous or somewhere in between? Resource dynamics matter in the optimisation of microbial communities. ISME J. 17, 641–644 (2023).
Chen, L., Zhao, X., Wongso, S., Lin, Z. & Wang, S. Trade-offs between receptor modification and fitness drive host-bacteriophage co-evolution leading to phage extinction or co-existence. ISME J. 18, wrae214 (2024).
Borin, J. M., Avrani, S., Barrick, J. E., Petrie, K. L. & Meyer, J. R. Coevolutionary phage training leads to greater bacterial suppression and delays the evolution of phage resistance. Proc. Natl. Acad. Sci. 118, e2104592118 (2021).
Costa, P., Pereira, C., Romalde, J. L. & Almeida, A. A game of resistance: War between bacteria and phages and how phage cocktails can be the solution. Virology 599, 110209 (2024).
Reisch, C. R. & Prather, K. L. J. The no-SCAR (Scarless Cas9 Assisted Recombineering) system for genome editing in Escherichia coli. Sci. Rep. 5, 15096 (2015).
Yehl, K. et al. Engineering Phage Host-Range and Suppressing Bacterial Resistance through Phage Tail Fiber Mutagenesis. Cell 179, 459 (2019).
Meile, S., Du, J., Dunne, M., Kilcher, S. & Loessner, M. J. Engineering therapeutic phages for enhanced antibacterial efficacy. Curr. Opin. Virol. 52, 182–191 (2022).
Peng, H., Chen, I. A. & Qimron, U. Engineering phages to fight multidrug-resistant bacteria. Chem. Rev. 125, 933–971 (2024).
Chaudhary, N. et al. CRISPR-Cas-assisted phage engineering for personalized antibacterial treatments. Indian J. Med. Microbiol. 53, 100771 (2025).
Steiner, L. X. et al. Phage-induced disturbance of a marine sponge microbiome. Environ. Microbiome 19, 97 (2024).
Balleste, E. et al. Bacteriophages in sewage: abundance, roles, and applications. FEMS Microbes 3, xtac009 (2022).
Pfeifer, E., Bonnin, R. A. & Rocha, E. P. C. Phage-plasmids spread antibiotic resistance genes through infection and lysogenic conversion. mBio 13, (2022).
Shi, L.-D. et al. A mixed blessing of viruses in wastewater treatment plants. Water Res. 215, 118237 (2022).
Jiang, S. et al. Generic Diagramming Platform (GDP): a comprehensive database of high-quality biomedical graphics. Nucleic Acids Res. 53, D1670–D1676 (2024).
Acknowledgements
The authors would like to thank the contributors for their valuable support. We would like to express our gratitude to Dr. Linchen He from Peking University for his valuable contributions in revising and enhancing the language throughout the manuscript. This work was supported by the National Natural Science Foundation of China (Grant Nos. 42407161, 42230411, and 42177099) and the Jing-Jin-Ji Regional Integrated Environmental Improvement-National Science and Technology Major Project (Grant No. 2025ZD1204700).
Author information
Authors and Affiliations
Contributions
J.P. conceived and designed the study, led the writing of the original draft, and was responsible for visualization. C.J. and X.L. defined the key information and provided funding support. X.W. contributed to the development of the content structure and assisted in reviewing and editing. S.X., A.S., and X.Z. supervised the study. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Peng, J., Jiang, C., Wang, X. et al. The nitrite bottleneck revisited: exploring the potential of phage-mediated microbial regulation for advancing novel nitrogen removal strategies. npj Clean Water 9, 11 (2026). https://doi.org/10.1038/s41545-025-00543-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41545-025-00543-6