Abstract
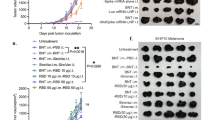
It is challenging for cancer vaccines to identify immunogenic antigens that are specifically and uniformly expressed on heterogeneous solid tumours and that can elicit production of T cells to lyse antigen-positive tumour cells and expand within the immunosuppressive tumour microenvironment. In contrast, microbial antigens are well-defined and robustly immunogenic and can activate specific memory T cells to eliminate microbes within the tumour microenvironment. Inspired by this, we developed a hepatitis B surface antigen (HBsAg)-tagged tumour vaccine system (H-TVAC). H-TVAC leverages HBsAg-specific memory T cells from a HBsAg mRNA vaccine to target and lyse HBsAg-tagged tumour cells using the vaccinia virus. This approach also elicits a tumour-specific immune response through epitope spreading by recruiting dendritic cells, thereby eliminating heterogeneous solid tumours. In various preclinical murine models, including the B16-OVA, B16F10, MC38, CT26, 4T1 and H22 hepatocellular carcinoma, as well as a B16F10 bilateral tumour model, H-TVAC demonstrates anti-tumour immune responses, improved survival rates and reduced metastasis and recurrence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. Raw data for sequencing results are available via the National Institutes of Health at https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1300966 (ref. 70). Source data are provided with this paper.
References
Hu, Z., Ott, P. A. & Wu, C. J. Towards personalized, tumour-specific, therapeutic vaccines for cancer. Nat. Rev. Immunol. 18, 168–182 (2018).
Blass, E. & Ott, P. A. Advances in the development of personalized neoantigen-based therapeutic cancer vaccines. Nat. Rev. Clin. Oncol. 18, 215–229 (2021).
Lopez, J. S. et al. A phase Ib study to evaluate RO7198457, an individualized neoantigen specific immunotherapy (iNeST),in combination with atezolizumab in patients with locally advanced or metastatic solid tumors. Cancer Res. 80, CT301 (2020).
Fang, Y. et al. A pan-cancer clinical study of personalized neoantigen vaccine monotherapy in treating patients with various types of advanced solid tumors. Clin. Cancer Res. 26, 4511–4520 (2020).
Sellars, M. C., Wu, C. J. & Fritsch, E. F. Cancer vaccines: building a bridge over troubled waters. Cell 185, 2770–2788 (2022).
Lang, F., Schrors, B., Lower, M., Tureci, O. & Sahin, U. Identification of neoantigens for individualized therapeutic cancer vaccines. Nat. Rev. Drug Discov. 21, 261–282 (2022).
Caushi, J. X., et al. Transcriptional programs of neoantigen-specific TIL in anti-PD-1-treated lung cancers. Nature 596, 126–132 (2021).
Rosenthal, R. et al. Neoantigen-directed immune escape in lung cancer evolution. Nature 567, 479–485 (2019).
Kubli, S. P., Berger, T., Araujo, D. V., Siu, L. L. & Mak, T. W. Beyond immune checkpoint blockade: emerging immunological strategies. Nat. Rev. Drug Discov. 20, 899–919 (2021).
O’Donnell, J. S., Teng, M. W. L. & Smyth, M. J. Cancer immunoediting and resistance to T cell-based immunotherapy. Nat. Rev. Clin. Oncol. 16, 151–167 (2019).
Kraehenbuehl, L., Weng, C. H., Eghbali, S., Wolchok, J. D. & Merghoub, T. Enhancing immunotherapy in cancer by targeting emerging immunomodulatory pathways. Nat. Rev. Clin. Oncol. 19, 37–50 (2022).
Katsikis, P. D., Ishii, K. J. & Schliehe, C. Challenges in developing personalized neoantigen cancer vaccines. Nat. Rev. Immunol. 24, 213–227 (2024).
Rudloff, M. W. et al. Hallmarks of CD8+ T cell dysfunction are established within hours of tumor antigen encounter before cell division. Nat. Immunol. 24, 1527–1539 (2023).
Cheng, Y. et al. Non-terminally exhausted tumor-resident memory HBV-specific T cell responses correlate with relapse-free survival in hepatocellular carcinoma. Immunity 54, 1825–1840.e7 (2021).
Sobao, Y. et al. The role of hepatitis B virus-specific memory CD8 T cells in the control of viral replication. J. Hepatol. 36, 105–115 (2002).
Wang, W. G. et al. Systemic immune responses to irradiated tumours via the transport of antigens to the tumour periphery by injected flagellate bacteria. Nat. Biomed. Eng. 6, 44–53 (2022).
Wang, W. et al. Perfluorocarbon regulates the intratumoural environment to enhance hypoxia-based agent efficacy. Nat. Commun. 10, 1580 (2019).
Tang, X. et al. Adenovirus-mediated specific tumor tagging facilitates CAR-T therapy against antigen-mismatched solid tumors. Cancer Lett. 487, 1–9 (2020).
Zuo, S. et al. An engineered oncolytic vaccinia virus encoding a single-chain variable fragment against TIGIT induces effective antitumor immunity and synergizes with PD-1 or LAG-3 blockade. J. Immunother. Cancer 9, e002843 (2021).
Gnant, M. F. X. et al. Tumor-specific gene delivery using recombinant vaccinia virus in a rabbit model of liver metastases. J. Natl Cancer Inst. 91, 1744–1750 (1999).
Nakano, S. et al. Recent advances in immunotherapy for hepatocellular carcinoma. Cancers 12, 775 (2020).
Macedo, N., Miller, D. M., Haq, R. & Kaufman, H. L. Clinical landscape of oncolytic virus research in 2020. J. Immunother. Cancer 8, e001486 (2020).
Ma, L. et al. Vaccine-boosted CAR T crosstalk with host immunity to reject tumors with antigen heterogeneity. Cell 186, 3148–3165 (2023).
Tai, W. et al. An mRNA-based T-cell-inducing antigen strengthens COVID-19 vaccine against SARS-CoV-2 variants. Nat. Commun. 14, 2962 (2023).
Schmidt, T. et al. Cellular immunity predominates over humoral immunity after homologous and heterologous mRNA and vector-based COVID-19 vaccine regimens in solid organ transplant recipients. Am. J. Transpl. 21, 3990–4002 (2021).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Liu, Q., Sun, Z. & Chen, L. Memory T cells: strategies for optimizing tumor immunotherapy. Protein Cell 11, 549–564 (2020).
Guo, M. et al. Durable and enhanced immunity against SARS-CoV-2 elicited by manganese nanoadjuvant formulated subunit vaccine. Signal Transduct. Target. Ther. 8, 462 (2023).
Shalhout, S. Z., Miller, D. M., Emerick, K. S. & Kaufman, H. L. Therapy with oncolytic viruses: progress and challenges. Nat. Rev. Clin. Oncol. 20, 160–177 (2023).
Decker, T. & Lohmann-Matthes, M. L. A quick and simple method for the quantitation of lactate dehydrogenase release in measurements of cellular cytotoxicity and tumor necrosis factor (TNF) activity. J. Immunol. Methods 115, 61–69 (1988).
Curran, M. A., Montalvo, W., Yagita, H. & Allison, J. P. PD-1 and CTLA-4 combination blockade expands infiltrating T cells and reduces regulatory T and myeloid cells within B16 melanoma tumors. Proc. Natl Acad. Sci. USA 107, 4275–4280 (2010).
Roy, D. G. et al. Adjuvant oncolytic virotherapy for personalized anti-cancer vaccination. Nat. Commun. 12, 2626 (2021).
Brossart, P. The role of antigen spreading in the efficacy of immunotherapies. Clin. Cancer Res. 26, 4442–4447 (2020).
Mimura, K. et al. Combined inhibition of PD-1/PD-L1, Lag-3, and Tim-3 axes augments antitumor immunity in gastric cancer-T cell coculture models. Gastric Cancer 24, 611–623 (2021).
Zhang, J. et al. Osr2 functions as a biomechanical checkpoint to aggravate CD8+ T cell exhaustion in tumor. Cell 187, 3409–3426.e24 (2024).
Lu, S. X. et al. Pharmacologic modulation of RNA splicing enhances anti-tumor immunity. Cell 184, 4032–4047 (2021).
Chen, Y. E. et al. Engineered skin bacteria induce antitumor T cell responses against melanoma. Science 380, 203–210 (2023).
Yadav, M. et al. Predicting immunogenic tumour mutations by combining mass spectrometry and exome sequencing. Nature 515, 572–576 (2014).
Symons, J. A., Tscharke, D. C., Price, N. & Smith, G. L. A study of the vaccinia virus interferon-gamma receptor and its contribution to virus virulence. J. Gen. Virol. 83, 1953–1964 (2002).
Kaech, S. M. & Cui, W. G. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12, 749–761 (2012).
Li, X., Sun, X., Wang, B., Li, Y. & Tong, J. Oncolytic virus-based hepatocellular carcinoma treatment: current status, intravenous delivery strategies, and emerging combination therapeutic solutions. Asian J. Pharm. Sci. 18, 100771 (2023).
Lybaert, L. et al. Challenges in neoantigen-directed therapeutics. Cancer Cell 41, 15–40 (2023).
Ji, D. et al. An engineered influenza virus to deliver antigens for lung cancer vaccination. Nat. Biotechnol. 42, 518–528 (2024).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Stronen, E. et al. Targeting of cancer neoantigens with donor-derived T cell receptor repertoires. Science 352, 1337–1341 (2016).
Chen, X. et al. An oncolytic virus delivering tumor-irrelevant bystander T cell epitopes induces anti-tumor immunity and potentiates cancer immunotherapy. Nat. Cancer 5, 1063–1081 (2024).
Rosato, P. C. et al. Virus-specific memory T cells populate tumors and can be repurposed for tumor immunotherapy. Nat. Commun. 10, 567 (2019).
Meier, S. L., Satpathy, A. T. & Wells, D. K. Bystander T cells in cancer immunology and therapy. Nat. Cancer 3, 143–155 (2022).
Simoni, Y. et al. Bystander CD8+ T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557, 575–579 (2018).
Martin, J. D., Cabral, H., Stylianopoulos, T. & Jain, R. K. Improving cancer immunotherapy using nanomedicines: progress, opportunities and challenges. Nat. Rev. Clin. Oncol. 17, 251–266 (2020).
Robert, C. et al. Immunotherapy discontinuation - how, and when? Data from melanoma as a paradigm. Nat. Rev. Clin. Oncol. 17, 707–715 (2020).
Zhao, H. et al. A therapeutic hepatitis B mRNA vaccine with strong immunogenicity and persistent virological suppression. npj Vaccines 9, 22 (2024).
Hou, A. J., Chen, L. C. & Chen, Y. Y. Navigating CAR-T cells through the solid-tumour microenvironment. Nat. Rev. Drug Discov. 20, 531–550 (2021).
Vincent, R. L. et al. Probiotic-guided CAR-T cells for solid tumor targeting. Science 382, 211–218 (2023).
Zhang, A. Q. et al. Universal redirection of CAR T cells against solid tumours via membrane-inserted ligands for the CAR. Nat. Biomed. Eng. 7, 1113–1128 (2023).
Park, A. K. et al. Effective combination immunotherapy using oncolytic viruses to deliver CAR targets to solid tumors. Sci. Transl. Med. 12, eaaz1863 (2020).
Caushi, J. X. et al. Transcriptional programs of neoantigen-specific TIL in anti-PD-1-treated lung cancers. Nature 596, 126–132 (2021).
Ngo, W. et al. Why nanoparticles prefer liver macrophage cell uptake in vivo. Adv. Drug Deliv. Rev. 185, 114238 (2022).
Suh, S. et al. Nanoscale bacteria-enabled autonomous drug delivery system (NanoBEADS) enhances intratumoral transport of nanomedicine. Adv. Sci. (Weinh.) 6, 1801309 (2019).
Gögenur, M. et al. Intratumoral influenza vaccine in early colorectal cancer. J. Immunother. Cancer 10, A575–A575 (2022).
Townsend, J. P., Hassler, H. B., Sah, P., Galvani, A. P. & Dornburg, A. The durability of natural infection and vaccine-induced immunity against future infection by SARS-CoV-2. Proc. Natl Acad. Sci. USA 119, e2204336119 (2022).
Ferdinands, J. M. et al. Intraseason waning of influenza vaccine protection: evidence from the US Influenza Vaccine Effectiveness Network, 2011-12 through 2014-15. Clin. Infect. Dis. 64, 544–550 (2017).
Palache, A., Oriol-Mathieu, V., Fino, M. Xydia-Charmanta, M. & Influenza Vaccine Supply task force (IFPMA IVS). Seasonal influenza vaccine dose distribution in 195 countries (2004–2013): little progress in estimated global vaccination coverage. Vaccine 33, 5598–5605 (2015).
Harrington, K., Freeman, D. J., Kelly, B., Harper, J. & Soria, J. C. Optimizing oncolytic virotherapy in cancer treatment. Nat. Rev. Drug Discov. 18, 689–706 (2019).
Atasheva, S. et al. ystemic cancer therapy with engineered adenovirus that evades innate immunity. Sci. Transl. Med. 12, eabc6659 (2020).
Palmer, C. D. et al. Individualized, heterologous chimpanzee adenovirus and self-amplifying mRNA neoantigen vaccine for advanced metastatic solid tumors: phase 1 trial interim results. Nat. Med. 28, 1619–1629 (2022).
Papachristofilou, A. et al. Phase Ib evaluation of a self-adjuvanted protamine formulated mRNA-based active cancer immunotherapy, BI1361849 (CV9202), combined with local radiation treatment in patients with stage IV non-small cell lung cancer. J. Immunother. Cancer 7, 38 (2019).
Mackensen, A. et al. CLDN6-specific CAR-T cells plus amplifying RNA vaccine in relapsed or refractory solid tumors: the phase 1 BNT211-01 trial. Nat. Med. 29, 2844–2853 (2023).
Lee, J. M. et al. The phase 3 INTerpath-002 study design: individualized neoantigen therapy (INT) V940 (mRNA-4157) plus pembrolizumab vs placebo plus pembrolizumab for resected early-stage non-small-cell lung cancer (NSCLC). J. Clin. Oncol. 42, TPS8116 (2024).
Wang W. et al. HBsAg-tagged tumour vaccine system eliminates solid tumours through virus-specific memory T cells. NCBI https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1300966 (2025).
Acknowledgements
This study was supported by grants from the National Natural Science Foundation of China (grant no. 82304428 to W.W.) and the Fundamental Research Funds for the Central Universities (grant no. 2632025ZD03 to W.W.), the Anti-tumor New Drug Rapid Transformation Public Service Platform of Jiangsu Province (grant no. BM2023002 to Y.Y.) and the Health Science and Technology Innovation Joint Project of Hainan Province, China (grant no. WSJK2024MS231 to Y.Y.). Additional financial support was provided through the Research Start-Up Funds from China Pharmaceutical University (grant no. 3150120054 to W.W.) as well as the Qingyun Project Research Fund (grant no. PB-QY012106 to W.W.) from Polaris Biology (Shanghai, China). We sincerely thank A. Lin for his guidance on the construction of the mRNA-LNP vaccine, and thank Y. Wang, J. Qiu and X. Wang from Polaris Biology for the comprehensive technical support for this project.
Author information
Authors and Affiliations
Contributions
W.W., Y.Y., Y.H. and S.Z. conceptualized and designed the research framework. Experiments were carried out and data analysis was performed by Y.C., L.Z., M.L., W.W., L.W., F.J., K.H., W.Z., L.C. and S.Q. The interpretation and discussion of the data were undertaken by W.W., Y.Y., W.Z., X.L. and S.Z. The drafting and preparation of the manuscript were managed by W.W., Y.C., L.Z., M.L., Y.Y., S.Z. and Y.H. All authors have meticulously read the final version of the paper and given their approval for its publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Luigi Buonaguro and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of the mRNA-H vaccine.
a, Characteristics of mRNA-LNPs, including their size, zeta potential and polydispersity index. b, Quantification of HBsAg secretion by 293T cells 24 h after mRNA-H transfection using an ELISA assay (n = 3 biologically independent samples). c, HBsAb titres in the serum were measured with an ELISA kit 7 days after the last of two immunizations with mRNA-H. The reference HepB-CHO served as a commercially available benchmark for immunization with recombinant hepatitis B (n = 6 biologically independent samples). These experiments were repeated twice independently with similar results. d, Flow cytometry gating strategy for the HBsAg-specific T cells (IFN-γ+, gate: CD3+CD8+ T cells) under the stimulation of HBsAg peptides in the spleen 30 days after immunization with the HBsAg vaccine (0.25 mg per mouse). e, Treatment scheme for the analysis of HBsAg-specific memory T cells on days 76 and 390 in the spleen from mice immunized by mRNA-H (0.25 mg per mouse). f, Flow cytometry gating strategy for the HBsAg-specific memory T cells (IFN-γ+CD44+, gate: CD3+CD8+ T cells) under the stimulation of HBsAg peptides from the spleen of the treated mice in (e). g,h, Representative flow cytometry plots (g) and quantitative analysis (h) (day 76 n = 5 and day 390 n = 3; biologically, independent samples) of HBsAg-specific memory T cells. i, Representative flow cytometry plots of eGFP expression during the preparation of B16-HBsAg tumour cells. eGFP and neomycin are the positive fluorescence and resistance markers used for the generation and screening of B16F10-HBsAg cells, respectively. The generated B16F10-HbsAg cells were continuously screened in neomycin-containing medium for 2 weeks before the experiments. j, qPCR analysis of the HBsAg mRNA level in B16-HBsAg cells compared with control cells (B16F10 and B16F10-pc) (n = 3 biologically independent samples). These experiments were repeated twice independently with similar results. For b, c, h and j, data are presented as mean ± s.e.m., with statistical significance assessed using one-way ANOVA (b, c and j) or two-tailed Student’s t-test (h).
Extended Data Fig. 2 Characterization of the intra-tumoural VV-H spreading and HBsAg expression at 24, 48 and 72 h post intra-tumoural injection.
a, Flow cytometry gating strategy of tumour cells infected with VV-H at 24, 48 and 72 h after intra-tumoural injection of VV-H (2 × 107 pfu per mouse). The VV-infected tumour cells were stained with anti-mouse CD45 antibody (PECy7) and anti-VV antibody (FITC), and identified as the CD45−VV+ population. b, Representative flow cytometry plots of the VV-H infected tumour cells. c, Ex vivo immunofluorescence images of B16F10 tumours infected with VV-H and intra-tumoural HBsAg expression at 24, 48 and 72 h after intra-tumoural injection of VV-H (2 × 107 pfu per mouse). Tumour sections were stained with an anti-VV antibody (yellow), an anti-HBsAg antibody (green) and DAPI (blue) to visualize VV-H, HBsAg and tumour nuclei, respectively. Scale bar, 50 μm. d, Quantitative analysis of tumour cells infected by VV-H and intra-tumoural HBsAg expression from the treated mice shown in (c) (n = 3 biologically independent samples). The proportion of infected cells and HBsAg expression levels was quantified by calculating the fluorescence area of VV-H or HBsAg relative to the DAPI-stained area. For d, data are presented as mean ± s.e.m.
Extended Data Fig. 3 Representative images of B16-OVA tumour-bearing mice and flow cytometric analysis of T-cell populations.
a, Images of B16-OVA tumour-bearing mice at the therapeutic endpoint. b, Flow cytometry gating strategy for intra-tumoural CD3+CD8+ T cells. c,d, Representative flow cytometry plots of intra-tumoural CD3+CD8+ T (c) and CD3+CD4+ T cells (d).
Extended Data Fig. 4 Analysis of T-cell and NK cell populations following H-TVAC administration with or without targeted depletion.
a, Treatment scheme for B16-OVA tumour-bearing mice with or without depletion of CD4+ T cells, CD8+ T cells and NK cells with H-TVAC as indicated. Depletion was achieved through the intraperitoneal administration of blocking antibodies targeting CD4+ T cells (10 mg kg−1), CD8+ T cells (10 mg kg−1) and NK cells (10 mg kg−1) on days 11, 13 and 15, respectively. b, Representative flow cytometry gating strategy for intra-tumoural CD8+ T cells (gate: CD45+CD3+), CD4+ T cells and NK cells (gate: CD45+). c,d, Representative flow cytometry plots of intra-tumoural CD8+ T cells, CD4+ T cells (c) and NK cells (d). e–g, Quantification of CD8+ T cells (e), CD4+ T cells (f) and NK cells (g) in the blood 3 days after various treatments (n = 3 biologically independent samples). For e–g, data are presented as mean ± s.e.m., with statistical significance assessed using one-way ANOVA (e–g) followed by Tukey’s multiple comparison test.
Extended Data Fig. 5 H-TVAC triggers a robust immune response in B16F10 tumour-bearing mice.
a, Treatment scheme for B16F10 tumour-bearing mice. b, Averaged tumour growth curves for B16F10 tumour-bearing mice (n = 8 biologically independent animals). c,d, Quantitation of intra-tumoural CD8+ T cells (c) and CD4+ T cells (d) at the therapeutic endpoint by flow cytometry (mRNA-H+VV-C and H-TVAC n = 7 and others n = 6; biologically independent samples). e, Representative flow cytometry plots of intra-tumoural CD3+CD8+ T and CD3+CD4+ T cells (gate: CD45+ cells). For b–d, data are presented as mean ± s.e.m., with statistical significance assessed using two-way ANOVA (b) and one-way ANOVA (c and d) followed by Tukey’s multiple comparison test.
Extended Data Fig. 6 Characterization of intra-tumoural HBsAg-specific memory T cells.
a, Treatment regimen for the analysis of intra-tumoural HBsAg specific memory T cells (mRNA-H, 0.25 mg kg−1; VV-H, 2 × 107 pfu per mouse). b, Representative flow cytometry plots of intra-tumoural HBsAg specific memory T cells. c, Flow cytometry gating strategy for the intra-tumoural HBsAg specific memory T cells (IFN-γ+CD44+, gate: CD3+CD8+ T cells) under the stimulation of HBsAg peptides. d, Flow cytometry gating strategy for exhausted (PD-1+TIM-3+, PD-1+LAG-3+ and TIM-3+LAG-3+) and functional phenotype (IFN-γ+TNF+) analysis of intra-tumoural HBsAg-specific memory T cells (gate: IFN-γ+CD44+; the gate for saline group, CD44+). e–h, Representative flow cytometry plots of the exhausted and functional phenotype analysis of intra-tumoural HBsAg-specific memory T cells. i, Quantification of IFN-γ+ TNF+ T cells in (h). For i, data are presented as mean ± s.e.m. (n = 6 biologically independent animals), with statistical significance assessed using a two-tailed Student’s t-test.
Extended Data Fig. 7 Characterization of all intra-tumoural T cells at the therapeutic endpoint.
a, Treatment regimen for the analysis of intra-tumoural T cells at the therapeutic endpoint (mRNA-H 0.25 mg kg−1 and VV-H 2 × 107 pfu per mouse). b, Flow cytometry gating strategy for intra-tumoural exhausted T cells (PD-1+TIM-3+, PD-1+LAG-3+ and TIM-3+LAG-3+). HBsAg specific memory T cells were determined as IFN-γ+CD44+ cell populations under the stimulation of HBsAg peptides (gate: CD3+CD8+). c–e. Quantification of intra-tumoural PD-1+LAG-3+(c), PD-1+TIM-3+(d) and TIM-3+LAG-3+ T cells (e) (n = 7 biologically independent samples). f–h. Representative flow cytometry plots of the exhausted T cells, including PD-1+TIM-3+(f), PD-1+LAG-3+(g) and TIM-3+LAG-3+ T cells (h). i,j, Immunohistochemical staining (i) and quantification (j) of intra-tumoural granzyme B secretion (n = 3 biologically independent samples). k,l, Immunohistochemical staining (k) and quantification (l) of intra-tumoural perforin secretion (n = 3 biologically independent samples). For c–e, j and l, data are presented as mean ± s.e.m., with statistical significance assessed using two-tailed Student’s t-test (c–e), and one-way ANOVA (j and l) followed by Tukey’s multiple comparison test.
Extended Data Fig. 8 Safety evaluation of the H-TVAC system.
a–o, Serum biochemical indexes (a–i) and haematological indexes (j–o) of CT26 tumour-bearing mice 14 days after treatment: AST (a), ALT (b), GGT (c), CKI (d), TGL (e), CHOL (f), BUN (g), TP (h), TBI (i), HCT (j), RBC (k), HGB (l), MCH (m), MCV (n) and MCHC (o) (saline and mRNA-H + saline n = 7 and others n = 8; biologically independent samples). p, images of HE staining in major organs (heart, liver, spleen, lung and kidney) 14 days post-treatment. For a–o, data are presented as mean ± s.e.m., with statistical significance assessed using two-way ANOVA (a–o) followed by Tukey’s multiple comparison test.
Extended Data Fig. 9 An overview of the H-TVAC platform for universal cancer vaccine and its advantages over traditional tumour neoantigen vaccines.
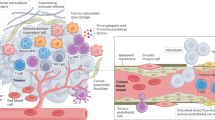
H-TVAC leverages microbial-specific memory T cells, induced by microbial vaccines, to lyse tumour cells tagged with microbial antigens. This approach facilitates the efficient elimination of tagged tumour cells via virus-specific memory T cells. Furthermore, tumour cell lysis, combined with DC recruitment mediated by the VV, enhances tumour-specific immune responses through epitope spreading, thereby promoting the eradication of heterogeneous solid tumours. Compared with traditional tumour neoantigen vaccines, H-TVAC therapy provides significant advantages. First, H-TVAC utilizes well-defined, conserved and highly immunogenic microbial antigens, effectively eliciting microbial-specific memory T cells. In contrast, tumour neoantigens are heterogeneous, highly mutated and require a complex screening process. Despite these efforts, their immunogenicity remains low. Second, unlike tumour-specific T cells that often become exhausted within the TME, microbe-specific memory T cells can resist this exhaustion and stimulate robust tumour-specific immune responses through epitope spreading.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–11, Methods, Tables 1 and 2 and References.
Source data
Source Data Fig. 1 (download XLSX )
Source data.
Source Data Fig. 2 (download XLSX )
Source data.
Source Data Fig. 3 (download XLSX )
Source data.
Source Data Fig. 4 (download XLSX )
Source data.
Source Data Fig. 5 (download XLSX )
Source data.
Source Data Fig. 6 (download XLSX )
Source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, W., Chu, Y., Zhao, L. et al. HBsAg-tagged tumour vaccine system eliminates solid tumours through virus-specific memory T cells. Nat. Biomed. Eng (2025). https://doi.org/10.1038/s41551-025-01555-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-025-01555-w
This article is cited by
-
Multi-strategy embedded framework for neoantigen vaccine maturation
Scientific Reports (2026)