Abstract
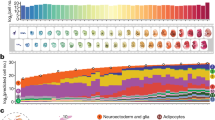
International efforts have yielded extensive single-cell time-series atlas datasets, such as those on mouse embryogenesis, providing a reference for mapping disease models across biomedical research. However, effectively using such data for temporal analysis of individual datasets is challenging due to the intricate nature of cell states and the tight coupling between time stamps and experimental batches. Here we introduce TemporalVAE, a deep generative model in a dual-objective setting that infers the biological time of each cell from a compressed latent space, even in a zero-shot setting. With a mouse development atlas, we demonstrated its scalability with millions of cells, accuracy in atlas-based cell staging across platforms and interpretability by identifying temporally sensitive genes with in silico perturbation. TemporalVAE effectively stages cells during human peri-implantation under both in vivo and in vitro conditions, and supports cross-primate comparisons among human, cynomolgus and marmoset embryos, highlighting its potential for broad biomedical applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All datasets used in this study are previously published datasets. The three small datasets used in Fig. 2 were downloaded from Psupertime, with the Acinar dataset directly downloaded via the Psupertime R package. The Embryonic beta cells and Human germline F datasets were accessed via GEO (GSE87375 and GSE86146). The mouse embryo development atlas was downloaded from https://omg.gs.washington.edu and can also be accessed via GEO (GSE228590). The mouse Stereo-seq dataset was downloaded from https://db.cngb.org/stomics/mosta/ and can be accessed via CNGB (CNP0001543). The Cynomolgus and Marmoset datasets were introduced by ref. 32 and are available via GitHub at https://github.com/Boroviak-Lab/SpatialModelling/tree/master/Data. For human peri-implantation, the eight datasets were downloaded from GEO or ArrayExpress: Xiang et al.28 (GSE136447), Petropoulos et al.26 (E-MTAB-3929), Molè et al.25 (E-MTAB-8060), Zhou et al.27 (GSE109555), Liu et al.24 (samples GSM3901995-GSM3902021, GSM3902030-GSM3902031 and GSM4058370-GSM4058375 from GSE133200), Tyser et al.29 (E-MTAB-9388), Xiao et al.31 (https://cs8.3dembryo.com/#/download) and Cui et al.30 (https://cs7.3dembryo.com/#/download). Moreover, all raw and preprocessed matrices and our trained model are available via Zenodo at https://doi.org/10.5281/zenodo.15339078 (ref. 53).
Code availability
TemporalVAE is freely available as a Python package via GitHub at https://github.com/StatBiomed/TemporalVAE. Detailed Jupyter notebooks to reproduce figures and results in this Report are also available in this repository. A step-by-step protocol can also be found at https://doi.org/10.17504/protocols.io.eq2ly47yplx9/v1 (ref. 54).
References
Ding, J., Sharon, N. & Bar-Joseph, Z. Temporal modelling using single-cell transcriptomics. Nat. Rev. Genet. 23, 355–368 (2022).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Gao, M., Qiao, C. & Huang, Y. UniTVelo: temporally unified RNA velocity reinforces single-cell trajectory inference. Nat. Commun. 13, 6586 (2022).
Schiebinger, G. et al. Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell 176, 928–943 (2019).
Lönnberg, T. et al. Single-cell RNA-seq and computational analysis using temporal mixture modeling resolves TH1/TFH fate bifurcation in malaria. Sci. Immunol. 2, eaal2192 (2017).
Lin, C. & Bar-Joseph, Z. Continuous-state HMMs for modeling time-series single-cell RNA-Seq data. Bioinformatics 35, 4707–4715 (2019).
Tran, T. N. & Bader, G. D. Tempora: cell trajectory inference using time-series single-cell RNA sequencing data. PLoS Comput. Biol. 16, e1008205 (2020).
Fischer, D. S. et al. Inferring population dynamics from single-cell RNA-sequencing time series data. Nat. Biotechnol. 37, 461–468 (2019).
Yeo, G. H. T., Saksena, S. D. & Gifford, D. K. Generative modeling of single-cell time series with PRESCIENT enables prediction of cell trajectories with interventions. Nat. Commun. 12, 3222 (2021).
Gao, M. et al. CLADES: a hybrid NeuralODE-Gillespie approach for unveiling clonal cell fate and differentiation dynamics. Nat. Commun. 16, 8174 (2025).
Macnair, W., Gupta, R. & Claassen, M. Psupertime: supervised pseudotime analysis for time-series single-cell RNA-seq data. Bioinformatics 38, i290–i298 (2022).
Li, G. et al. Sceptic: pseudotime analysis for time-series single-cell sequencing and imaging data. Genome Biol. 26, 209 (2025).
Rood, J. E. et al. The Human Cell Atlas from a cell census to a unified foundation model. Nature 637, 1065–1071 (2024).
Qiu, C. et al. A single-cell time-lapse of mouse prenatal development from gastrula to birth. Nature 626, 1084–1093 (2024).
Zhao, C. et al. A comprehensive human embryo reference tool using single-cell RNA-sequencing data. Nat. Methods 22, 193–206 (2024).
Eze, U. C., Bhaduri, A., Haeussler, M., Nowakowski, T. J. & Kriegstein, A. R. Single-cell atlas of early human brain development highlights heterogeneity of human neuroepithelial cells and early radial glia. Nat. Neurosci. 24, 584–594 (2021).
Karlsson, K. et al. Deterministic evolution and stringent selection during preneoplasia. Nature 618, 383–393 (2023).
Kingma, D. P. & Welling, M. Auto-encoding variational Bayes. In Proc. 2nd International Conference on Learning Representations (ICLR) (2014).
Calderon, D. et al. The continuum of drosophila embryonic development at single-cell resolution. Science 377, eabn5800 (2022).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using dna nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Liu, D. et al. Primary specification of blastocyst trophectoderm by scRNA-seq: new insights into embryo implantation. Sci. Adv. 8, eabj3725 (2022).
Molè, M. A. et al. A single cell characterisation of human embryogenesis identifies pluripotency transitions and putative anterior hypoblast centre. Nat. Commun. 12, 3679 (2021).
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Zhou, F. et al. Reconstituting the transcriptome and dna methylome landscapes of human implantation. Nature 572, 660–664 (2019).
Xiang, L. et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos. Nature 577, 537–542 (2020).
Tyser, R. C. et al. Single-cell transcriptomic characterization of a gastrulating human embryo. Nature 600, 285–289 (2021).
Cui, L. et al. Spatial transcriptomic characterization of a Carnegie stage 7 human embryo. Nat. Cell Biol. 27, 360–369 (2025).
Xiao, Z. et al. 3D reconstruction of a gastrulating human embryo. Cell 187, 2855–2874 (2024).
Bergmann, S. et al. Spatial profiling of early primate gastrulation in utero. Nature 609, 136–143 (2022).
Sierra, C. et al. The lncRNA Snhg11, a new candidate contributing to neurogenesis, plasticity, and memory deficits in Down syndrome. Mol. Psychiatry 29, 2117–2134 (2024).
Maguire, S. et al. Targeting of Slc25a21 is associated with orofacial defects and otitis media due to disrupted expression of a neighbouring gene. PLoS ONE 9, e91807 (2014).
Fogel, B. L. et al. RBFOX1 regulates both splicing and transcriptional networks in human neuronal development. Hum. Mol. Genet. 21, 4171–4186 (2012).
Mahony, C. B. et al. Hapln1b, a central organizer of the ECM, modulates kit signaling to control developmental hematopoiesis in zebrafish. Blood Adv. 5, 4935–4948 (2021).
Wang, R. N. et al. Bone morphogenetic protein (BMP) signaling in development and human diseases. Genes Dis. 1, 87–105 (2014).
Hadas, R. et al. Temporal BMP4 effects on mouse embryonic and extraembryonic development. Nature 634, 652–661 (2024).
Sapkota, D. et al. Onecut1 and onecut2 redundantly regulate early retinal cell fates during development. Proc. Natl Acad. Sci. USA 111, E4086–E4095 (2014).
Fajner, V. et al. Hecw controls oogenesis and neuronal homeostasis by promoting the liquid state of ribonucleoprotein particles. Nat. Commun. 12, 5488 (2021).
Werren, E. A. et al. Biallelic variants in CSMD1 are implicated in a neurodevelopmental disorder with intellectual disability and variable cortical malformations. Cell Death Dis. 15, 379 (2024).
Baum, M. L. et al. CSMD1 regulates brain complement activity and circuit development. Brain Behav. Immun. 119, 317–332 (2024).
Homma, S., Shimada, T., Hikake, T. & Yaginuma, H. Expression pattern of LRR and Ig domain-containing protein (LRRIG protein) in the early mouse embryo. Gene Expr. Patterns 9, 1–26 (2009).
Williams, S. M. et al. An integrative analysis of non-coding regulatory DNA variations associated with autism spectrum disorder. Mol. Psychiatry 24, 1707–1719 (2019).
Chen, V. S. et al. Histology atlas of the developing prenatal and postnatal mouse central nervous system, with emphasis on prenatal days E7.5 to E18.5. Toxicol. Pathol. 45, 705–744 (2017).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. In Proc. 3rd International Conference on Learning Representations (ICLR) (2015).
Courty, N., Flamary, R., Habrard, A. & Rakotomamonjy, A. Joint distribution optimal transportation for domain adaptation. Adv. Neural Inf. Process. Syst. 30, 3733–3742 (2017).
Enge, M. et al. Single-cell analysis of human pancreas reveals transcriptional signatures of aging and somatic mutation patterns. Cell 171, 321–330 (2017).
Qiu, W.-L. et al. Deciphering pancreatic islet β cell and α cell maturation pathways and characteristic features at the single-cell level. Cell Metab. 25, 1194–1205 (2017).
Li, L. et al. Single-cell RNA-seq analysis maps development of human germline cells and gonadal niche interactions. Cell Stem Cell 20, 858–873 (2017).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 1–5 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Liu, Y.-J. Resource of “TemporalVAE: atlas-assisted temporal mapping of time-series single-cell transcriptomes during embryogenesis”. Zenodo https://doi.org/10.5281/zenodo.15339078 (2025).
Liu, Y. et al. A step-by-step protocol for using TemporalVAE on developmental timing prediction. protocols.io https://doi.org/10.17504/protocols.io.eq2ly47yplx9/v1 (2025).
Acknowledgements
We thank StatBiomed Lab members for being supportive throughout this study. This study was jointly supported by the National Natural Science Foundation of China (grant no. 62222217), the Research Grants Council of the Hong Kong SAR, China (grant no. T12-705-24-R), the InnoHK initiative of the Innovation and Technology Commission of the Hong Kong SAR Government and the University of Hong Kong through a startup fund and a seed fund (Y.H.), as well as the Guangdong Basic and Applied Basic Research Foundation, China (grant no. 2023A1515220177), the National Key Research and Development Program of China (grant nos. 2022YFC2702500 and 2022YFC2702503), Key R&D Program of the Ministry of Science and Technology, China (grant no. 2023YFF0905400) and National Natural Science Foundation of China through grants (grant no. U2341229).
Author information
Authors and Affiliations
Contributions
Y.H. conceived and supervised the project. Y.L. and Y.H. designed the study. Y.L. developed the whole model, implemented the software, and analysed the data and results, with support from F.C. and M.B. D.C. and Y.C. provided guidance and results interpretation. Y.L. and Y.H. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Magnus Rattray and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
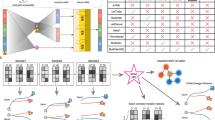
Extended Data Fig. 1 Detail architecture of TemporalVAE.
In this model, each layer is a fully connected neural network with the shape of (input dimension, output dimension).
Extended Data Fig. 2 Benchmark results on Human female germline datasets.
The results format is the same as main Fig. 2.
Extended Data Fig. 3 Model ablation experiments.
Ablation experiments of TemporalVAE (without the Time-predictor) on Acinar (A, B, C), Embryo beta (D, E, F), and Human female germline datasets (G, H, I). The Ablated-TemporalVAE has the same model structure as TemporalVAE but without the Time-Predictor. The Ablated-TemporalVAE uses the same training parameters as TemporalVAE to train the ablated model on the training data to obtain their low-dimensional representations, then a linear regression (LR) model is fitted on these representations and applied to predict the pseudo-time of test data. The Spearman results (A, D, G) highlight the role of the Time-Predictor in learning temporally meaningful latent features. The distributions of the predicted time by Ablated-TemporalVAE (B, E, H) and TemporalVAE (C, F, I) on three datasets demonstrate the Time-Predictor module is essential for pseudo-time prediction.
Extended Data Fig. 4 Violin plots of time prediction in the Xiang dataset.
Shown is the predicted time by TemporalVAE either (a) via pre-trained on human peri-implantation atlas (related to main Fig. 4b) or (b) via leave-one-stage-out cross-validation.
Extended Data Fig. 5 Results of in silico perturbation on five shared temporally sensitive genes.
Mean Δt (A) and expression pattern (B) of the five temporally sensitive genes shared among time clusters along the developmental timeline of mouse atlas data.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Y., Cai, F., Barile, M. et al. TemporalVAE: atlas-assisted temporal mapping of time-series single-cell transcriptomes during embryogenesis. Nat Cell Biol 27, 1982–1992 (2025). https://doi.org/10.1038/s41556-025-01787-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-025-01787-7