Abstract
Pluripotency, the ability to generate all body cell types, emerges in a disorganized embryonic cell mass. After implantation, these cells form a columnar epithelium and initiate lumenogenesis. During gastrulation, some undergo epithelial-to-mesenchymal transition to form the primitive streak (PS). The signals controlling these events in humans are largely unknown. Here, to study them, we developed a chemically defined 3D model where conventional pluripotent stem cells self-organize into a columnar epithelium with a lumen, from which PS-like cells emerge. We show that early TGFβ family inhibition prevents epithelial identity, also in murine 3D embryo models and in embryos. ZNF398 acts downstream of TGFβ1, activating the epithelial master regulator ESRP1 while repressing mesenchymal factors CDH2 and ZEB2. After epithelium formation, TGFβ1 stimulation is dispensable for its maintenance. However, treatment via ACTIVIN—a distinct TGFβ family ligand—induces PS efficiently. Thus, signalling of the TGFβ family dynamically governs pluripotent epiblast epithelial identity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Sequencing data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE248567. Previously published human48 and mouse41 embryo data that were reanalysed here are available under accession codes GSE136447 (human), GSE121650 and GSE133725 (mouse). The processed human embryo reference scRNA-seq dataset40 used only for Fig. 1 was retrieved from https://petropoulos-lanner-labs.clintec.ki.se/dataset.download.html. The previously published ZNF398 ChIP-seq dataset13 that was reanalysed here is available under accession code GSE133630. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Boroviak, T. & Nichols, J. The birth of embryonic pluripotency. Philos. Trans. R. Soc. Lond. B 369, 20130541 (2014.
Zhu, M. & Zernicka-Goetz, M. Principles of self-organization of the mammalian embryo. Cell 183, 1467–1478 (2020).
Pera, M. F. & Rossant, J. The exploration of pluripotency space: charting cell state transitions in peri-implantation development. Cell Stem Cell 28, 1896–1906 (2021).
Rossant, J. & Tam, P. P. L. Early human embryonic development: blastocyst formation to gastrulation. Dev. Cell 57, 152–165 (2022).
Sheng, G., Martinez Arias, A. & Sutherland, A. The primitive streak and cellular principles of building an amniote body through gastrulation. Science 374, abg1727 (2021).
Bedzhov, I. & Zernicka-Goetz, M. Self-organizing properties of mouse pluripotent cells initiate morphogenesis upon implantation. Cell 156, 1032–1044 (2014).
Kyprianou, C. et al. Basement membrane remodelling regulates mouse embryogenesis. Nature 582, 253–258 (2020).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science https://doi.org/10.1126/science.282.5391.1145 (1998).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007).
Tyser, R. C. V. et al. Single-cell transcriptomic characterization of a gastrulating human embryo. Nature 600, 285–289 (2021).
Wang, Z., Oron, E., Nelson, B., Razis, S. & Ivanova, N. Distinct lineage specification roles for NANOG, OCT4, and SOX2 in human embryonic stem cells. Cell Stem Cell 10, 440–454 (2012).
Zorzan, I. et al. The transcriptional regulator ZNF398 mediates pluripotency and epithelial character downstream of TGF-beta in human PSCs. Nat. Commun. 11, 2364 (2020).
Chia, N. Y. et al. A genome-wide RNAi screen reveals determinants of human embryonic stem cell identity. Nature 468, 316–320 (2010).
Naxerova, K. et al. Integrated loss- and gain-of-function screens define a core network governing human embryonic stem cell behavior. Genes Dev. 35, 1527–1547 (2021).
Eiselleova, L. et al. A complex role for FGF-2 in self-renewal, survival, and adhesion of human embryonic stem cells. Stem Cells 27, 1847–1857 (2009).
Greber, B. et al. Conserved and divergent roles of FGF signaling in mouse epiblast stem cells and human embryonic stem cells. Cell Stem Cell 6, 215–226 (2010).
Martello, G. & Smith, A. The nature of embryonic stem cells. Annu. Rev. Cell Dev. Biol. 30, 647–675 (2014).
Singh, A. M. et al. Signaling network crosstalk in human pluripotent cells: a Smad2/3-regulated switch that controls the balance between self-renewal and differentiation. Cell Stem Cell 10, 312–326 (2012).
Vallier, L. Activin/Nodal and FGF pathways cooperate to maintain pluripotency of human embryonic stem cells. J. Cell Sci. 118, 4495–4509 (2005).
Vallier, L. et al. Activin/Nodal signalling maintains pluripotency by controlling Nanog expression. Development 136, 1339–1349 (2009).
Wamaitha, S. E. et al. IGF1-mediated human embryonic stem cell self-renewal recapitulates the embryonic niche. Nat. Commun. 11, 764 (2020).
Massagué, J. & Sheppard, D. TGFβ signaling in health and disease. Cell 186, 4007–4037 (2023).
Brumm, A. S. et al. Initiation and maintenance of the pluripotent epiblast in pre-implantation human development is independent of NODAL signaling. Dev. Cell 60, 174–185 (2025).
Ross, S. & Hill, C. S. How the Smads regulate transcription. Int. J. Biochem. Cell Biol. 40, 383–408 (2008).
Yoney, A. et al. WNT signaling memory is required for ACTIVIN to function as a morphogen in human gastruloids. eLife 7, e38279 (2018).
Osnato, A. et al. TGFβ signalling is required to maintain pluripotency of human naïve pluripotent stem cells. eLife 10, e67259 (2021).
Ai, Z. et al. Dissecting peri-implantation development using cultured human embryos and embryo-like assembloids. Cell Res. 33, 661–678 (2023).
Hislop, J. et al. Modelling post-implantation human development to yolk sac blood emergence. Nature 626, 367–376 (2024).
Liu, L. et al. Modeling post-implantation stages of human development into early organogenesis with stem-cell-derived peri-gastruloids. Cell 186, 3776–3792.e16 (2023).
Moris, N. et al. An in vitro model of early anteroposterior organization during human development. Nature 582, 410–415 (2020).
Oldak, B. et al. Complete human day 14 post-implantation embryo models from naive ES cells. Nature 622, 562–573 (2023).
Pedroza, M. et al. Self-patterning of human stem cells into post-implantation lineages. Nature https://doi.org/10.1038/s41586-023-06354-4 (2023).
Simunovic, M., Siggia, E. D. & Brivanlou, A. H. In vitro attachment and symmetry breaking of a human embryo model assembled from primed embryonic stem cells. Cell Stem Cell 29, 962–972 (2022).
Simunovic, M. et al. A 3D model of a human epiblast reveals BMP4-driven symmetry breaking. Nat. Cell Biol. 21, 900 (2019).
Sozen, B. et al. Reconstructing aspects of human embryogenesis with pluripotent stem cells. Nat. Commun. 12, 5550 (2021).
Weatherbee, B. A. T. et al. A model of the post-implantation human embryo derived from pluripotent stem cells. Nature https://doi.org/10.1038/s41586-023-06368-y (2023).
Zheng, Y. et al. Controlled modelling of human epiblast and amnion development using stem cells. Nature https://doi.org/10.1038/s41586-019-1535-2 (2019).
Taniguchi, K. et al. Lumen formation is an intrinsic property of isolated human pluripotent stem cells. Stem Cell Rep. 5, 954–962 (2015).
Zhao, C. et al. A comprehensive human embryo reference tool using single-cell RNA-sequencing data. Nat. Methods 22, 193–206 (2025).
Argelaguet, R. et al. Multi-omics profiling of mouse gastrulation at single-cell resolution. Nature 576, 487–491 (2019).
Shao, Y. et al. Self-organized amniogenesis by human pluripotent stem cells in a biomimetic implantation-like niche. Nat. Mater. 16, 419–427 (2017).
Baillie-Benson, P., Moris, N. & Martinez Arias, A. Pluripotent stem cell models of early mammalian development. Curr. Opin. Cell Biol. 66, 89–96 (2020).
Shahbazi, M. N. et al. Self-organization of the human embryo in the absence of maternal tissues. Nat. Cell Biol. 18, 700–708 (2016).
Ludwig, T. E. et al. Derivation of human embryonic stem cells in defined conditions. Nat. Biotechnol. 24, 185–187 (2006).
Chen, G. et al. Chemically defined conditions for human iPS cell derivation and culture. Nat. Methods 8, 424–429 (2011).
Deglincerti, A. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016).
Xiang, L. et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos. Nature 577, 537–542 (2020).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Molè, M. A. et al. A single cell characterisation of human embryogenesis identifies pluripotency transitions and putative anterior hypoblast centre. Nat. Commun. 12, 3679 (2021).
Stirparo, G. G. et al. Integrated analysis of single-cell embryo data yields a unified transcriptome signature for the human pre-implantation epiblast. Development 145, dev158501 (2018).
Nakanoh, S. et al. Human surface ectoderm and amniotic ectoderm are sequentially specified according to cellular density. Sci. Adv. 10, eadh7748 (2024).
Rostovskaya, M., Andrews, S., Reik, W. & Rugg-Gunn, P. J. Amniogenesis occurs in two independent waves in primates. Cell Stem Cell 29, 744–759.e6 (2022).
Matthews, K. R. W., Wagner, D. S. & Warmflash, A. Stem cell-based models of embryos: the need for improved naming conventions. Stem Cell Rep. https://doi.org/10.1016/j.stemcr.2021.02.018 (2021).
Zhang, Y. E. Non-Smad pathways in TGFβ signaling. Cell Res. 19, 128–139 (2009).
Tang, W. W. C. et al. Sequential enhancer state remodelling defines human germline competence and specification. Nat. Cell Biol. 24, 448–460 (2022).
Brennan, J. et al. Nodal signalling in the epiblast patterns the early mouse embryo. Nature 411, 965–969 (2001).
Shahbazi, M. N. et al. Pluripotent state transitions coordinate morphogenesis in mouse and human embryos. Nature 552, 239–243 (2017).
Carbognin, E. et al. Esrrb guides naive pluripotent cells through the formative transcriptional programme. Nat. Cell Biol. 25, 643–657 (2023).
Papanayotou, C. et al. A novel nodal enhancer dependent on pluripotency factors and smad2/3 signaling conditions a regulatory switch during epiblast maturation. PLoS Biol. 12, e1001890 (2014).
Amadei, G. et al. Inducible stem-cell-derived embryos capture mouse morphogenetic events in vitro. Dev. Cell 56, 366–382.e9 (2021).
Amadei, G. et al. Embryo model completes gastrulation to neurulation and organogenesis. Nature 610, 143–153 (2022).
Kim, Y. S. et al. Deciphering epiblast lumenogenesis reveals proamniotic cavity control of embryo growth and patterning. Sci. Adv. 7, eabe1640 (2021).
Ma, H. et al. In vitro culture of cynomolgus monkey embryos beyond early gastrulation. Science 366, eaax7890 (2019).
Hu, C. et al. CellMarker 2.0: an updated database of manually curated cell markers in human/mouse and web tools based on scRNA-seq data. Nucleic Acids Res. 51, D870–D876 (2023).
Warzecha, C. C. et al. An ESRP-regulated splicing programme is abrogated during the epithelial–mesenchymal transition. EMBO J. 29, 3286–3300 (2010).
Otani, T. & Furuse, M. Tight Junction Structure and Function Revisited. Trends Cell Biol. 30, 805–817 (2020).
Davis, R. P. et al. Targeting a GFP reporter gene to the MIXL1 locus of human embryonic stem cells identifies human primitive streak–like cells and enables isolation of primitive hematopoietic precursors. Blood 111, 1876–1884 (2008).
Bernardo, A. S. et al. BRACHYURY and CDX2 mediate BMP-induced differentiation of human and mouse pluripotent stem cells into embryonic and extraembryonic lineages. Cell Stem Cell 9, 144–155 (2011).
McLean, A. B. et al. Activin a efficiently specifies definitive endoderm from human embryonic stem cells only when phosphatidylinositol 3-kinase signaling is suppressed. Stem Cells 25, 29–38 (2007).
Sturgeon, C. M., Ditadi, A., Awong, G., Kennedy, M. & Keller, G. Wnt signaling controls the specification of definitive and primitive hematopoiesis from human pluripotent stem cells. Nat. Biotechnol. 32, 554–561 (2014).
D’Amour, K. A. et al. Efficient differentiation of human embryonic stem cells to definitive endoderm. Nat. Biotechnol. 23, 1534–1541 (2005).
Funa, N. S. et al. β-Catenin regulates primitive streak induction through collaborative interactions with SMAD2/SMAD3 and OCT4. Cell Stem Cell 16, 639–652 (2015).
Gadue, P., Huber, T. L., Paddison, P. J. & Keller, G. M. Wnt and TGF-beta signaling are required for the induction of an in vitro model of primitive streak formation using embryonic stem cells. Proc. Natl Acad. Sci. USA 103, 16806–16811 (2006).
Winnier, G., Blessing, M., Labosky, P. A. & Hogan, B. L. M. Bone morphogenetic protein-4 is required for mesoderm formation and patterning in the mouse. Genes Dev. 9, 2105–2116 (1995).
Heldin, C.-H., Landström, M. & Moustakas, A. Mechanism of TGF-beta signaling to growth arrest, apoptosis, and epithelial–mesenchymal transition. Curr. Opin. Cell Biol. 21, 166–176 (2009).
Warmflash, A., Sorre, B., Etoc, F., Siggia, E. D. & Brivanlou, A. H. A method to recapitulate early embryonic spatial patterning in human embryonic stem cells. Nat. Methods 11, 847–854 (2014).
Krammer, T. et al. Mouse neural tube organoids self-organize floorplate through BMP-mediated cluster competition. Dev. Cell 59, 1940–1953.e10 (2024).
Boroviak, T. et al. Single cell transcriptome analysis of human, marmoset and mouse embryos reveals common and divergent features of preimplantation development. Development 145, dev167833 (2018).
Clark, A. T. et al. Stem cell-based embryo models: the 2021 ISSCR stem cell guidelines revisited. Stem Cell Rep. 20, 102514 (2025).
Takashima, Y. et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 158, 1254–1269 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based Analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Proserpio, V., Duval, C., Falvo, V., Donati, G. & Oliviero, S. in Immune Receptors: Methods and Protocols (eds Rast, J. & Buckley, K.) 217–229 (Springer, 2022); https://doi.org/10.1007/978-1-0716-1944-5_15
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Amezquita, R. A. et al. Orchestrating single-cell analysis with Bioconductor. Nat. Methods 17, 137–145 (2020).
L. Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Acknowledgements
We thank all the members of the Martello Laboratory and Oliviero Laboratory for discussions and suggestions. In particular, we thank M. Chieregato for technical support. We thank S. Rapelli, I. Laurence Polignano and F. Mirzadeh for helping with the pilot experiment of scRNA-seq. We also thank H. T. Stuart for helping with immunostaining protocol in the 3D structures. G.M.’s laboratory is supported by grants from the Giovanni Armenise–Harvard Foundation (Dissecting the Human Pluripotency Network), the Telethon Foundation (GJC21157), an ERC Starting Grant (MetEpiStem), the Progetti di Rilevante Interesse Nazionale PRIN 2022 (Dissecting genetic, epigenetic and metabolic alterations caused by reprogramming of somatic cells to pluripotency), the Microsoft Research Ltd Grant (The Pluripotency Program in Human Embryonic Stem Cells), the HUMANIZE Project la CAIXA Foundation (Generation of humanized organs from human iPS cells) and the European Union – Next Generation EU, Mission 4 Component 1, CUP C93C22002780006, Spoke n.3 (AAV-delivered MTF1 to suppress suppressors of polyQ toxicity), S.O. is supported by the Associazione Italiana per la Ricerca sul Cancro (AIRC) IG IG 2022 ID 27155, PRIN 2018, and IIGM institutional funds. G.M. and S.O. laboratories were supported by CN3 funding-National Center for Gene Therapy and Drugs based on RNA Technology. G.A.’s laboratory is supported by a STARS Starting Grant of the University of Padova, a PRID 2024 grant from the Department of Biology and a Federation of European Biochemical Societies Excellence Award.
Author information
Authors and Affiliations
Contributions
I.Z. and E.C. set up the 3D-hE-gastruloid protocol, designed and performed most of the experiments. I.Z. performed ZNF398 overexpression experiments. I.Z., I.P. and F.B. performed experiments with MIXL1-GFP reporter cells. I.Z. and A. Drusin performed molecular analyses. V.P., I.Z. and F.B. performed single-cell RNA-seq experiments. A.L. performed analysis of bulk and single-cell RNA-seq data, ChIP-seq data and statistical analyses. I.Z., E.C., M.S., D. Benvegnù and A.L. prepared figures. M.A. analysed bulk RNA-seq data. D. Benvegnù performed the mES cell rosette assay. D. Benvegnù, P.B., D. Bizzotto and G.A. performed experiments with mouse embryos. C.D. and G.A. designed and performed experiments with cysts composed of WT and Gata4-inducible ES cells. S.D.J.U. and G.A. designed and performed experiments with ETiX embryoids. A. Ditadi and R.S. generated ZNF398 KO cells. G.P. and E.P. generated CRISPRi cell lines. S.O., G.A. and G.M. supervised the study and secured fundings. G.M. wrote the manuscript with suggestions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Ligand and receptor expression analyses in human embryos and 3D models.
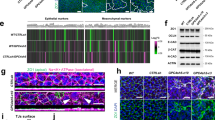
a, Box and dot plots showing the expression levels of receptors of the FGF, INSULIN and TGF-beta signalling pathways in the post-EPIs cell population of the embryo dataset48 (72 cells). Expression is reported as log-normalized counts. Horizontal line indicates median, box indicates the interquartile range (IQR) and whiskers denote the 1.5 × IQR. b, Box and dot plots showing the expression levels of ligands of the FGF, INSULIN and TGF-beta signalling pathways in the post-EPIs (72 cells), CTBs (159 cells) and Hypoblast (25 cells) cell population of the embryo dataset. Expression is reported as log-normalized counts. Horizontal line indicates median, box indicates the interquartile range (IQR) and whiskers denote the 1.5 × IQR. c, Optimised protocol to mimic epiblast self-organisation in vitro. Single hPSCs were seeded on a Matrigel layer with E8 medium + 5% of Matrigel (v/v). After 4 days, 3D structures were analysed as indicated. See also Method details. d, Bar plots showing gene expression levels measured by RNA-seq of KiPS cultured in the conventional 2D system or in 3D after 4 days of self-organisation for selected marker genes of pluripotency. Expression is reported as CPM after TMM normalization. Bars indicate the mean of 2 experiments, shown as dots. e, Representative image of immunostaining for SOX2 and TBXT of H9 after 8 days of self-organisation. On day 8, the 3D structures still express pluripotency markers, such as SOX2, but they also express the anterior primitive streak marker, T. Note that SOX2 and T markers are mutually exclusive (see merge SOX2/T). Images are representative of 2 experiments. Scale bars: 50 µm. f,g, Bar plots showing gene expression levels measured by RNA-seq of KiPS cultured in the conventional 2D system or in 3D after 4 days of self-organisation for selected marker genes of primitive streak (f) and Wnt/BMP signalling (g). Expression is reported as CPM after TMM normalization. Bars indicate the mean of 2 experiments, shown as dots.
Extended Data Fig. 2 Integrated single-cell transcriptomics analysis of human embryos and 3D-hE-gastruloids.
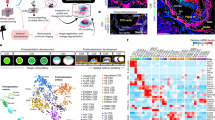
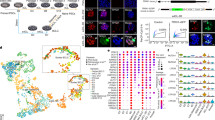
a, t-SNE embedding of 394 single-cell transcriptomes showing the results of clustering analysis for the identification of cell populations in the 3D epiblast model dataset. Cells are coloured by their differentiation day (left) and inferred cell type (right). b, Barplots showing the relative percentage of cells by differentiation day in each cluster/inferred cell population. c, Box and dot plots showing the expression levels by cell type of selected marker genes for pluripotency, epithelial, polarity, primitive streak and amnion cell identities (total number of cells = 394). Expression is reported as log-normalized counts. Horizontal line indicates median, box indicates the interquartile range (IQR) and whiskers denote the 1.5 × IQR. Coloured dots indicate the cell cluster in which the gene was found as a marker. d,e,f, t-SNE embedding of 892 single-cell transcriptomes coloured according to the relative (Z-score) expression levels of selected marker genes for primitive streak (CDH2, EOMES, VIM), ICM (ESRRB), Pre-implantation epiblast (KLF17, TFCP2L1), general pluripotency (PRDM14, SOX2) and epithelial/polarity (CLDN6) cell identities. g, UMAP embedding of 337 single-cell transcriptomes showing the results of pseudo-temporal ordering by reverse graph embedding using Monocle3. The line plot on the leftmost UMAP represents the embedded trajectory graph. Cells are coloured according to pseudotime (left), differentiation day (middle) and inferred cell type (right).
Extended Data Fig. 3 Validation and characterisation of TGF-beta signalling inhibition in 3D-hE-gastruloids.
a, Top: Representative images of 3D structures generated from H9 hPSCs treated with different concentrations of SB43. Scale bar: 50 µm. Bottom left: Bar plots showing the fraction of 3D structures in the indicated conditions. Bars indicate the mean of 2 experiments, shown as individual dots. Bottom right: Bar plots showing gene expression analysis by qPCR of LEFTY1 in the indicated conditions. Bars indicate the mean of 2 experiments, shown as individual dots. b, Top: Representative images of 3D structures generated from H9 hPSCs treated with different concentrations of A83. Scale bar: 50 µm. Bottom left: Bar plots showing the fraction of 3D structures in the indicated conditions. Bars indicate the mean of 2 experiments, shown as individual dots. Bottom right: Bar plots showing gene expression analysis by qPCR of LEFTY1 in the indicated conditions. Bars indicate the mean of 3 experiments (except for 2D, which is based on 2 experiments), shown as individual dots. c, Representative images of immunostaining for phalloidin (F-Actin) and Hoechst (nuclei) in 3D-hE- Gastruloids generated from KiPS cell line cultured in E8, E8 + 10 µM SB43, and E8 + 10 µM A83. Images are representative of 3 experiments. Scale bar: 50 µm. d, Representative brightfield and fluorescence images of 2D SMAD2_CRISPRi and parental cell lines cultured without and with DOX/TMP for 14 days. Images are representative of 2 experiments. Scale bar: 100 µm. e, Bar plots showing gene expression analysis by qPCR of SMAD2 and LEFTY1 genes on 2D SMAD2_CRISPRi and parental cell lines cultured without and with DOX/TMP for 14 days. N = 2 experiments. f, Schematic representation of the experimental strategy used to generate SMAD2_CRISPRi and parental 3D-hE-Gastruloids. Structures were cultured for 4 days and subsequently analyzed by immunofluorescence, qPCR, and morphological assessment. g, Bar plots showing fraction of organised (red bars), not organised (grey bars) and no cavity (orange bars) SMAD2_CRISPRi-derived structures with and without DOX/TMP after 4 days of self-organisation. N = 2 experiments. h, Barplot showing the number of structures per field in SMAD2_CRISPRi cell line with and without DOX and TMP. N = 10 technical replicates, shown as dots. i, Bar plots showing gene expression analysis by qPCR of NANOG and OCT4 genes on SMAD2_CRISPRi and parental 3D-hE-Gastruloids without and with DOX/TMP after 4 days of self-organisation. N = 2 experiments. j, Schematic representation of the experimental strategy used for TGF-beta family inhibition at different days. Inhibition was achieved by adding 1 µM A83 or 1 µM SB43 to the E8 medium, while DMSO was used as a control. After 4 days, the structures were analyzed by morphological assessment and qPCR. k, Representative images of KiPS-derived structures following a 4 days time-course of signalling inhibition with 1 µM SB43. Images are representative of 4 experiments. Scale bar: 50 µm. l, Bar plots showing fraction of organised (red bars), not organised (grey bars) and no cavity (orange bars) KiPS-derived structures in a 4 days time-course of signalling inhibition with 1 µM SB43. Bars indicate the mean +/- s.e.m. of 4 experiments, shown as dots. Unpaired two-tailed Welch’s t-test. m, Bar plots showing gene expression analysis by qPCR of pluripotency marker genes performed in KiPS-derived structures following a 4 days time-course of signalling inhibition with 1 µM SB43. Bars indicate the mean +/- s.e.m. of 3 experiments, shown as dots. Unpaired two-tailed Welch’s t-test.
Extended Data Fig. 4 Morphometric analyses of ETiX-embryoids.
a, Representative confocal images of ETiX obtained after 120 h, with or without A83 inhibitor. The structures were stained for PODXL. Nuclei were stained with Hoechst, F-Actin with phalloidin. The staining was performed to assess the presence of the cavity in the structures, as indicated by the dashed line. Scale bar: 50 µm. b. Violin plot showing the aspect ratio of ETiX after 120 h of culture, with or without treatment with 5 µM A83 inhibitor. Each dot represents a single structure and the dot’s color represents an independent experiment, for a total of 5 experiments (Control, n = 134; 5 μM A83, n = 222). Paired two tailed t-test. c, Representative confocal images of ETiX after 24 h, 72 h and 120 h with or without 5 µM A83 inhibitor treatment. F-actin is stained with phalloidin and nuclei with Hoechst. The white dashed line indicates an example of the region where intensity was tracked and plotted in d. Scale bar: 50 µm. d, Quantification of phalloidin and Hoechst intensity along a radial line from the center to the edge of a structure (see Methods for details). Number of independent experiments: at 24 h, N = 3; at 72 h, Control, N = 4; 5 µM, N = 3; at 120 h, N = 5. For each experiment, n = 5 structures were analyzed, with 5 randomly selected radii quantified per structure. Solid lines indicate the mean, and shaded areas represent the s.e.m.
Extended Data Fig. 5 Characterisation of ZNF398 genetic perturbations in both 2D and 3D hPSC models.
a, Gene expression analysis by qPCR of KiPS stably expressing the empty vector control (Empty) without (blue) or with SB43 (yellow), or expressing ZNF398 without SB43 (lilac) or with SB43 (orange) after 4 days of self-organisation. Bars indicate the mean +/- s.e.m. of 4 experiments, shown as dots. Expression was normalised to the mean of Empty DMSO samples. Unpaired two-tailed Welch’s t-test. b, Heatmap showing gene expression analysis by qPCR for markers of pluripotency. Human PSCs, either expressing ZNF398 or an empty vector, were analysed when cultured under conventional conditions (2D) and after 4 days of self-organisation (3D) in presence of SB43 or DMSO. Values are expressed as fold-change relative to the mean of Empty 2D samples from 4 experiments and are scaled by rows (genes). Red and purple indicate high and low expression, respectively. c, Representative chromatograms of the ZNF398 locus in a WT (top panel) and a heterozygous clone. The matching part of the guide (before the cut) is highlighted in light blue. Table summarizing the ZNF398 sequence analyses in 365 clones. d, Representative brightfield and fluorescence images of 2D ZNF398_CRISPRi and parental cell lines cultured without and with DOX/TMP for 7 days. Scale bar: 200 µm. e, Bar plots showing gene expression analysis by qPCR of pluripotency and epithelial genes on 2D ZNF398_CRISPRi cell line cultured without and with DOX/TMP for 7 days. Bars indicate the mean of 2 experiments, shown as dots. f, Bar plots showing the fraction of organised (red), non-organised (gray), and no-cavity (orange) ZNF398_CRISPRi-derived structures, with or without DOX and TMP. Each bar represents the average of 2 experiments. g, Bar plots showing gene expression analysis by qPCR of pluripotency and epithelial genes on ZNF398_CRISPRi 3D-hE-Gastruloids without and with DOX/TMP after 4 days of self-organisation. N = 2 independent experiments. h, PCA of the ZNF398_CRISPRi 3D-hE-Gastruloids RNA-seq experiment, computed using the 2000 most variable genes. Colors indicate the analyzed conditions, while shape indicates whether the culture is 2D or 3D.
Extended Data Fig. 6 Analysis of MIXL1-GFP and TGF-beta signalling components signal in 3D-hE-gastruloids.
a, Representative images and fluorescence images of MIXL1-GFP reporter for the indicated conditions are shown. Images are representative of 3 experiments. Scale bar: 50 µm. b, Gating strategy used for the quantification of MIXL1-GFP positive cells from 3D-hE-gastruloids. c, Box and dot plots showing the expression levels by cell type of the receptors of the TGF-beta family in 3D-hE-gastruloids. The left plot represents 3D-hE-gastruloids derived from H9 hPSCs (total number of cells = 337), while the right plot represents 3D-hE-gastruloids derived from HES3-MIXL1-GFP hESCs (total number of cells = 694). Expression is reported as log-normalized counts. Horizontal line indicates median, box indicates the interquartile range (IQR) and whiskers denote the 1.5 × IQR. Boxes and dots are colored according to cell types. d, Representative confocal images of KiPS cells. The cells were not treated (DMSO), treated overnight with 10 µM SB43 (SB34), and treated overnight with 10 µM SB43 followed by 1 h of 2 ng/ml TGFB1 treatment (SB43 + TGFB1). The cells were stained for p-SMAD3 and nuclei labeled using Hoechst. Images are representative of 2 experiments. Scale bar: 50 µm. e, Representative confocal images of KiPS (left) and H9 (right) 3D-hE-gastruloids stained for p-SMAD3 cultured in E8 and E6+Activin A. Nuclei were stained with Hoechst and F-actin with phalloidin. Images are representative of 2 experiments. Scale bar: 50 µm.
Extended Data Fig. 7 BMP signalling pathway analysis in 3D-hE-gastruloids.
a, Box and dot plots showing the expression levels by cell type of the ligands (top) and receptors (bottom) of BMP signalling pathway, in 3D-hE-gastruloids. The left plot represents 3D-hE-gastruloids derived from H9 hPSCs (total number of cells = 337), while the right plot represents 3D-hE-gastruloids derived from HES3-MIXL1-GFP hESCs (total number of cells = 694). Expression is reported as log-normalized expression counts. Horizontal line indicates median, box indicates the interquartile range (IQR) and whiskers denote the 1.5 × IQR. Boxes and dots are colored according to cell types. b, Representative confocal images of KiPS cells. The cells were not treated (DMSO), treated overnight with 0.1 µM LDN (LDN), and treated overnight with 0.1 µM LDN followed by 1 h of 100 ng/ml BMP4 treatment (LDN + BMP4). The cells were stained for p-SMAD1/5 and nuclei labeled using Hoechst. Images are representative of 2 experiments. Scale bar: 50 µm. c, Representative confocal images of KiPS (top) and H9 (bottom) 3D-hE-gastruloids stained for p-SMAD1/5 cultured in E8 and E6 + Activin A. Nuclei were stained with Hoechst and F-actin with phalloidin. Images are representative of 2 experiments. Scale bar: 50 µm.
Supplementary information
Supplementary Tables 1–3 (download XLSX )
Supplementary Table 1. List of antibodies used in this study. Supplementary Table 2. List of primers used in this study. Supplementary Table 3. Light microscopy reporting table.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zorzan, I., Carbognin, E., Lauria, A. et al. A human epiblast model reveals dynamic TGFβ-mediated control of epithelial identity during mammalian epiblast development. Nat Cell Biol 28, 49–65 (2026). https://doi.org/10.1038/s41556-025-01831-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-025-01831-6