Abstract
Whether a distinct subset of cancer stem cells (CSCs) is exclusively responsible for metastasis and how this process occurs remain unresolved. Through multi-omics, pan-cancer analysis and multiple tumour-bearing models, we identify THY1⁺ CSCs as the key drivers of metastasis and uncover a previously unrecognized ‘pseudohypoxic’ state (independent of classical hypoxia) as a central regulatory factor. The self-renewal of THY1⁺ CSCs is maintained by IL-6–MYC signalling. Upon encountering neutrophils, THY1⁺ CSCs activate the THY1–Mac1 axis, triggering the Src–Akt/Erk pathway, Rac1 activation and a migrasome-dependent process that induces neutrophils to expel reactive oxygen species-enriched damaged mitochondria. THY1 signalling further enhances macropinocytosis, enabling CSCs to internalize these mitochondria and adopt a pseudohypoxic state, thereby facilitating CSC metastasis. Notably, targeting the IL-6–Myc, THY1–Mac1 or Src–Akt/Erk signalling pathways effectively suppresses pseudohypoxia-driven CSC metastasis. These findings unveil previously unexplored mechanisms by which CSCs undergo metastasis, offering potential strategies to combat tumour metastasis and improve cancer prognosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All the expression data that support the findings of this study can be obtained from the Gene Expression Omnibus, Mendeley Data or NGDC (GSA for human), and the selected studies are listed below. The RNA sequencing dataset presented in Fig. 3f and Extended Data Fig. 4f, g has been deposited in the Genome Sequence Archive under the accession numer CRA036043. Bulk RNA-seq from the TCGA dataset were obtained from Genomic Data Commons at https://portal.gdc.cancer.gov/. Previously published scRNA-seq data reanalysed here are available under accession codes (1) pan-HCC scRNA-seq data, GSE149614 (ref. 62), PRJCA007744 (ref. 63) and skrx2fz79n (ref. 51); (2) pan-cancer CSC analysis, PRJCA007744 (HCC and intrahepatic cholangiocarcinoma) (ref. 63), GSE131907 (lung cancer) (ref. 64), GSE132465 (colon cancer) (ref. 65), E-MTAB-6149 (lung cancer) (ref. 66), E-MTAB-6653 (lung cancer) (ref. 66), GSE267718 (bladder cancer) (ref. 67), GSE176078 (breast cancer) (ref. 68), GSE154778 (pancreatic cancer) (ref. 69), GSE167297 (gastric cancer) (ref. 70), GSE188711 (colon cancer) (ref. 71) and GSE139829 (melanoma) (ref. 72); (3) pan-cancer neutrophil analysis, PRJCA007744 (HCC and intrahepatic cholangiocarcinoma) (ref. 63), GSE267718 (bladder cancer) (ref. 67), GSE171145 (ref. 73)/GSE127465 (ref. 74) (lung cancer), OEP003254 (pancreatic cancer) (ref. 75) and PRJCA020880 (gastric cancer and colon cancer) (ref. 57); (4) spatial transcriptome data, HRA000437 (ref. 52) and skrx2fz79n (Mendeley Data) (ref. 51); (5) pan-cancer analysis of primary and matched metastatic lesions, GSE149614 (HCC) (ref. 62), GSE197177 (ref. 76)/GSE263733 (ref. 77) (pancreatic cancer), GSE225857 (colon cancer) (ref. 78) and GSE131907 (lung cancer) (ref. 64). All data reported in this paper will be shared by the lead contact upon request. The detailed information of publicly available datasets used in this study is listed in Supplementary Table 1. Source data are provided with this paper.
Code availability
All analysis and figures were generated using publicly available software packages. No custom code was used in this study.
References
de Visser, K. E. & Joyce, J. A. The evolving tumor microenvironment: from cancer initiation to metastatic outgrowth. Cancer Cell 41, 374–403 (2023).
Boumahdi, S. & de Sauvage, F. J. The great escape: tumour cell plasticity in resistance to targeted therapy. Nat. Rev. Drug Discov. 19, 39–56 (2020).
Loh, J. J. & Ma, S. Hallmarks of cancer stemness. Cell Stem Cell 31, 617–639 (2024).
Lee, T. K., Guan, X. Y. & Ma, S. Cancer stem cells in hepatocellular carcinoma–from origin to clinical implications. Nat. Rev. Gastroenterol. Hepatol. 19, 26–44 (2022).
Dirkse, A. et al. Stem cell-associated heterogeneity in glioblastoma results from intrinsic tumor plasticity shaped by the microenvironment. Nat. Commun. 10, 1787 (2019).
Phan, T. G. & Croucher, P. I. The dormant cancer cell life cycle. Nat. Rev. Cancer 20, 398–411 (2020).
Hu, J. et al. STING inhibits the reactivation of dormant metastasis in lung adenocarcinoma. Nature 616, 806–813 (2023).
Dalla, E. et al. Lung-resident alveolar macrophages regulate the timing of breast cancer metastasis. Cell 187, 6631–6648 (2024).
Zuo, H. et al. Differential regulation of breast cancer bone metastasis by PARP1 and PARP2. Nat. Commun. 11, 1578 (2020).
Liang, H. et al. Host STING-dependent MDSC mobilization drives extrinsic radiation resistance. Nat. Commun. 8, 1736 (2017).
Jaillon, S. et al. Neutrophil diversity and plasticity in tumour progression and therapy. Nat. Rev. Cancer 20, 485–503 (2020).
Mantovani, A., Marchesi, F., Malesci, A., Laghi, L. & Allavena, P. Tumour-associated macrophages as treatment targets in oncology. Nat. Rev. Clin. Oncol. 14, 399–416 (2017).
Xia, H. et al. Autophagic adaptation to oxidative stress alters peritoneal residential macrophage survival and ovarian cancer metastasis. JCI Insight 5, e141115 (2020).
Godet, I. et al. Hypoxia induces ROS-resistant memory upon reoxygenation in vivo promoting metastasis in part via MUC1-C. Nat. Commun. 15, 8416 (2024).
Wei, X. et al. Mechanisms of vasculogenic mimicry in hypoxic tumor microenvironments. Mol. Cancer 20, 7 (2021).
Mantovani, A. & Locati, M. Macrophage metabolism shapes angiogenesis in tumors. Cell Metab. 24, 887–888 (2016).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Teschendorff, A. E. & Enver, T. Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome. Nat. Commun. 8, 15599 (2017).
Chanvorachote, P., Sriratanasak, N. & Nonpanya, N. C-myc contributes to malignancy of lung cancer: a potential anticancer drug target. Anticancer Res. 40, 609–618 (2020).
Sies, H. & Jones, D. P. Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat. Rev. Mol. Cell Biol. 21, 363–383 (2020).
Hu, P., Leyton, L., Hagood, J. S. & Barker, T. H. Thy-1-integrin interactions in cis and trans mediate distinctive signaling. Front. Cell. Dev. Biol. 10, 928510 (2022).
Jiao, H. et al. Mitocytosis, a migrasome-mediated mitochondrial quality-control process. Cell 184, 2896–2910 (2021).
Xu, W. et al. Integrin-induced PIP5K1C kinase polarization regulates neutrophil polarization, directionality, and in vivo infiltration. Immunity 33, 340–350 (2010).
Liu, J., Chen, Y. & O’Neill, L. Phospholipid metabolism in innate immunity and inflammation: from basic to clinic. Immun. Inflamm. 1, 6 (2025).
Colin, M. et al. Dysregulation of macropinocytosis processes in glioblastomas may be exploited to increase intracellular anti-cancer drug levels the example of temozolomide. Cancers 11, 411 (2019).
Ding, X., Yao, T., Liu, X., Fan, Z. & Liu, Y. A macropinocytosis-related gene signature predicts the prognosis and immune microenvironment in hepatocellular carcinoma. Front. Oncol. 13, 1143013 (2023).
Bayona-Bafaluy, M. P. et al. Revisiting the mouse mitochondrial DNA sequence. Nucleic Acids Res. 31, 5349–5355 (2003).
Ludwig, L. S. et al. Lineage tracing in humans enabled by mitochondrial mutations and single-cell genomics. Cell 176, 1325–1339 (2019).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Zhang, H. et al. Systematic investigation of mitochondrial transfer between cancer cells and T cells at single-cell resolution. Cancer Cell 41, 1788–1802 (2023).
Tian, Y. et al. An IL-6-induced STAT3-to-PI3K signaling switch potently drives PD-L1 transcription in cancer stem cells of colorectal cancer. Sci. Bull. https://doi.org/10.1016/j.scib.2025.07.013 (2025).
Song, Y. et al. H19 promotes cholestatic liver fibrosis by preventing ZEB1-mediated inhibition of epithelial cell adhesion molecule. Hepatology 66, 1183–1196 (2017).
Larsen, J. E. et al. ZEB1 drives epithelial-to-mesenchymal transition in lung cancer. J. Clin. Invest. 126, 3219–3235 (2016).
Kumar, A., Bhanja, A., Bhattacharyya, J. & Jaganathan, B. G. Multiple roles of CD90 in cancer. Tumor Biol. 37, 11611–11622 (2016).
Lee, G. et al. Oxidative dimerization of PHD2 is responsible for its inactivation and contributes to metabolic reprogramming via HIF-1α activation. Sci. Rep. 6, 18928 (2016).
Taniguchi, C. M. et al. Cross-talk between hypoxia and insulin signaling through Phd3 regulates hepatic glucose and lipid metabolism and ameliorates diabetes. Nat. Med. 19, 1325–1330 (2013).
Yang, M., Su, H., Soga, T., Kranc, K. R. & Pollard, P. J. Prolyl hydroxylase domain enzymes: important regulators of cancer metabolism. Hypoxia 2, 127–142 (2014).
Yuan, X., Ruan, W., Bobrow, B., Carmeliet, P. & Eltzschig, H. K. Targeting hypoxia-inducible factors: therapeutic opportunities and challenges. Nat. Rev. Drug Discov. 23, 175–200 (2024).
Jiang, D. et al. Neutrophil-derived migrasomes are an essential part of the coagulation system. Nat. Cell Biol. 26, 1110–1123 (2024).
Jiao, H. & Yu, L. Migrasomes: biogenesis, physiological roles, and therapeutic potentials. J. Cell Biol. 223, e202403051 (2024).
Burgos-Ojeda, D. et al. A novel model for evaluating therapies targeting human tumor vasculature and human cancer stem-like cells. Cancer Res. 73, 3555–3565 (2013).
Mu, M. et al. Targeting ferroptosis-elicited inflammation suppresses hepatocellular carcinoma metastasis and enhances sorafenib efficacy. Cancer Res. 84, 841–854 (2024).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria (2021).
Korotkevich, G., Sukhov, V., Budin, N., Shpak, B., Artyomov, M. N. & Sergushichev, A. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2021).
Boyer, L. A. et al. Core Transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Menyhárt, O., Kothalawala, W. J. & Győrffy, B. A gene set enrichment analysis for the cancer hallmarks. J. Pharm. Anal. 15, 101065 (2025).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013).
Li, Q. scTour: a deep learning architecture for robust inference and accurate prediction of cellular dynamics. Genome Biol. 24, 149 (2023).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genom. 19, 477 (2018).
Liu, Y. et al. Identification of a tumour immune barrier in the HCC microenvironment that determines the efficacy of immunotherapy. J. Hepatol. 78, 770–782 (2023).
Wu, R. et al. Comprehensive analysis of spatial architecture in primary liver cancer. Sci. Adv. 7, eabg3750 (2021).
Kueckelhaus, J. et al. Inferring histology-associated gene expression gradients in spatial transcriptomic studies. Nat. Commun. 15, 7280 (2024).
Larsson, L., Franzén, L., Ståhl, P. L. & Lundeberg, J. Semla: a versatile toolkit for spatially resolved transcriptomics analysis and visualization. Bioinformatics 39, btad626 (2023).
Ma, Y. & Zhou, X. Spatially informed cell-type deconvolution for spatial transcriptomics. Nat. Biotechnol. 40, 1349–1359 (2022).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Wu, Y. et al. Neutrophil profiling illuminates anti-tumor antigen-presenting potency. Cell 187, 1422–1439 (2024).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 218 (2016).
Finotello, F. et al. Molecular and pharmacological modulators of the tumor immune contexture revealed by deconvolution of RNA-seq data. Genome Med. 11, 34 (2019).
Aran, D., Hu, Z. & Butte, A. J. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18, 220 (2017).
Lu, Y. et al. A single-cell atlas of the multicellular ecosystem of primary and metastatic hepatocellular carcinoma. Nat. Commun. 13, 4594 (2022).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147 (2022).
Kim, N. et al. Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma. Nat. Commun. 11, 2285 (2020).
Lee, H. O. et al. Lineage-dependent gene expression programs influence the immune landscape of colorectal cancer. Nat. Genet. 52, 594–603 (2020).
Zhao, Y. et al. Integrative transcriptome analysis reveals the molecular events underlying impaired T-cell responses in EGFR-mutant lung cancer. Sci. Rep. 14, 18366 (2024).
Tran, M. A. et al. Urine scRNAseq reveals new insights into the bladder tumor immune microenvironment. J. Exp. Med. 221, e20240045 (2024).
Wu, S. Z. et al. A single-cell and spatially resolved atlas of human breast cancers. Nat. Genet. 53, 1334–1347 (2021).
Lin, W. et al. Single-cell transcriptome analysis of tumor and stromal compartments of pancreatic ductal adenocarcinoma primary tumors and metastatic lesions. Genome Med. 12, 80 (2020).
Jeong, H. Y. et al. Spatially distinct reprogramming of the tumor microenvironment based on tumor invasion in diffuse-type gastric cancers. Clin. Cancer Res. 27, 6529–6542 (2021).
Guo, W. et al. Resolving the difference between left-sided and right-sided colorectal cancer by single-cell thesequencing. JCI Insight 7, e152616 (2022).
Durante, M. A. et al. Single-cell analysis reveals new evolutionary complexity in uveal melanoma. Nat. Commun. 11, 496 (2020).
Lambrechts, D. et al. Phenotype molding of stromal cells in the lung tumor microenvironment. Nat. Med. 24, 1277–1289 (2018).
Zilionis, R. et al. Single-cell transcriptomics of human and mouse lung cancers reveals conserved myeloid populations across individuals and species. Immunity 50, 1317–1334 (2019).
Wang, L. et al. Single-cell RNA-seq analysis reveals BHLHE40-driven pro-tumour neutrophils with hyperactivated glycolysis in pancreatic tumour microenvironment. Gut 72, 958–971 (2023).
Zhang, S. et al. Single cell transcriptomic analyses implicate an immunosuppressive tumor microenvironment in pancreatic cancer liver metastasis. Nat. Commun. 14, 5123 (2023).
Park, J. K. et al. Single-cell transcriptome analysis reveals subtype-specific clonal evolution and microenvironmental changes in liver metastasis of pancreatic adenocarcinoma and their clinical implications. Mol. Cancer 23, 87 (2024).
Wang, F. et al. Single-cell and spatial transcriptome analysis reveals the cellular heterogeneity of liver metastatic colorectal cancer. Sci. Adv. 9, eadf5464 (2023).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (82341014, 32530037, U25C2022 and 82025016) and the Natural Science Foundation of Guangdong Province, China (2023A1515012466 and 2024A1515010549) awarded to D.-M.K., and by grants from the National Natural Science Foundation of China (82322051 and 82271773) awarded to Y.W.
Author information
Authors and Affiliations
Contributions
D.-M.K., Y.W. and X.-M.L. conceived the project. W.-H.W. and P.-L.L. designed experiments. W.-H.W., P.-L.L., W.-J.C., Z.-X.L. and Y.-Q.X. performed most of the experiments and analysed the results. M.C., L.Z. and J.-C.W. provided clinical samples and analysed the related clinical data. L.C. and L.L. made intellectual contributions. D.-M.K., Y.W. and X.-M.L. contributed to study design, supervised the study and contributed to writing the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Xue-Yan He and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identification of THY1+ cells as a metastatic-related cancer stem cells (CSCs) in HCC.
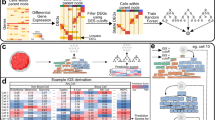
a–f, Annotations of CSCs in human HCC scRNA datasets. Schematic representation of a modified stemness-related signature approach to derive CSCs from HCC scRNA datasets (a). 3D co-embedding of stemness-related signature and cell entropy score in CSCs and non-CSCs (b). The reliability of the approach to derive CSCs was examined by CytoTRACE and cell entropy score (c). Heatmap displaying marker gene expression across CSC clusters. The top two marker genes of each CSC cluster were listed (d). Proliferation, self-renewal, multipotency, drug resistance, and anti-apoptotic score of CSC clusters (e). Canonical cancer stem cell surface marker expression in defined CSCs (f). g, Univariate (left) and multivariate (right) regression analyses of factors associated with patients’ recurrence in HCC (n = 86 patients). Points represent hazard ratios; error bars indicate 95% confidence intervals. h, Association of THY1⁺ CSC infiltration with patients’ survival utilizing TCGA datasets across nine cancer types. Statistical significance was determined using Cox regression analyses with two-sided tests (g), no adjustment for multiple comparisons was applied, as the analyses were predefined and hypothesis-driven, Kaplan–Meier analysis with log-rank test (h), or unpaired two-tailed Student’s t-test (c). Violin plots show data distribution with median and interquartile range (c). Panel a created with BioRender.com. AFP, α-fetoprotein; CI, confidence interval; HR, hazard ratio.
Extended Data Fig. 2 THY1 signalling regulates the metastasis of CSCs.
a, Percentage of THY1+ cells in Hepa1-6 and H22 cells was analysed by flow cytometry (n = 5). b, Gating strategies for FACS of THY1+ and EpCAM+ CSCs from tumour cells. c,d, Extreme limiting dilution analysis of unsorted, THY1+, or EpCAM+ Hepa1-6 cells (n = 5). e, Survival curves of mice bearing tumours from unsorted, THY1⁺, or EpCAM⁺ Hepa1-6 cells (n = 8). f, Migration potential of unsorted, THY1+, EpCAM+ Hepa1-6 cultured in vitro (n = 5; scale bar, 60 µm). g,h, Indicated B16 cells were intradermally injected at the right flank of C57BL/6 mice. Lung metastases were assessed (n = 5; scale bar, 250 µm). i, Indicated 4T1 cells were inoculated in mammary fat pads of BALB/c mice. Lung metastases were assessed (n = 5; scale bar, 250 µm). j, Indicated CT26 cells were intradermally injected at the right flank of BALB/c mice. Lung metastases were assessed (n = 5; scale bar, 250 µm). k, Efficiency of Thy1-Knockout (sgThy1) in Hepa1-6 cells (n = 5). l, Effects of the αTHY1 antibody on lung metastasis in mice bearing THY1⁺ Hepa1-6 hepatomas (n = 5; scale bar, 250 µm). m, Effects of sgThy1 on survival in mice bearing THY1⁺ Hepa1-6 hepatomas (n = 8). n,o, WT or Epcam-overexpressing (Epcam-OE) THY1+ Hepa1-6 cells were inoculated into mouse livers. Efficiency of Epcam-OE in Hepa1-6 cells (n; n = 5). Lung metastases were analysed (o; n = 5). Data are from three (d,e,m) or five (k,n) independent experiments. Data are presented as mean ± s.e.m. of three (h–j,l,o) or five (a,f) independent experiments. Statistical significance was determined using two-sided likelihood ratio tests (d), one-way ANOVA with Tukey’s post hoc test (f,h–j), unpaired two-tailed Student’s t-test (l,o), or the log-rank (Mantel–Cox) test (e,m). Panel b,c,g,i,j,l,o created with BioRender.com.
Extended Data Fig. 3 IL-6-Myc signalling is crucial for the self-renewal of THY1+ CSCs.
a, FACS-sorted THY1− human and murine hepatoma cell lines were cultured for different durations in vitro. Percentage of THY1+ cells regenerated from THY1− cells over time (n = 5). b,c, Efficiency of shNANOG, shMYC (b) or MYC-OE (c) in Huh-7 cells (each n = 5). d, Percentage of THY1+ cells generated from unsorted, wild-type THY1− (cultured with control medium), or MYC-OE THY1− (cultured with doxycycline) Huh-7 cells over time (n = 5). The induced group was compared with their corresponding uninduced group. e, GSEA of NANOG or c-Myc signatures (right: M5926) on pseudotime-ordered developmental trajectory from CSC.c4 to CSC.c3 populations. f, Schematic illustration of c-Myc binding sites in the THY1 promoter region. TSS, transcription start site. g, Sequence analysis identifying c-Myc binding sites within the promoter regions of the THY1 gene. h, Schematic diagram illustrating the truncated 5′-flanking regions of the THY1 gene. i,j, Effect of signalling inhibitors treatment (i) or shIL6R (j) on c-Myc expression in Huh-7 cells (each n = 5). k, Effects of IL-6 family cytokines on c-Myc expression in Huh-7 cells. (n = 5). l–n, Wild-type or Il6ra-knockdown EpCAM+ Hepa1-6 cells were inoculated into mouse livers. The efficiency of Il6ra-knockdown was examined (l; n = 5). Gating strategy for FACS analysis of GFP+ tumour cells was shown (m). Tumour volume was measured (n; n = 5; scale bar, 1cm). o, GSEA of c-Myc signatures (left: M5926) or IL-6–STAT3 pathway (right: M5897) in THY1+ versus THY1− CSCs from pan-cancer scRNA datasets. Data are from five (b,c,i,j,k) independent experiments. Data are presented as mean ± s.e.m. of three (l,n) or five (a,d) independent experiments. Statistical significance was determined using two-way ANOVA with Tukey’s post hoc test (d), a one-sided, permutation-based test (e,o), or unpaired two-tailed Student’s t-test (l,n).
Extended Data Fig. 4 Pseudohypoxia drives the metastatic potential of THY1+ CSCs.
a, Schematic overview of datasets used in pan-cancer scRNA analysis. b–d, Metastasis score (b) and hypoxia signature score (c) of THY1+ CSCs in pan-cancer scRNA datasets. The correlation of average metastasis score and hypoxia signature score across different cancer types (d). e, Distribution of hypoxic area (stained by hypoxyprobe) and HIF1α+ cells in tumours of Thy1-OE Hepa1-6 hepatoma-bearing mice (n = 11; scale bar, 75 µm). The numbers of HIF1α+ cells in normoxic and hypoxic areas were calculated. f,g, Venn diagram showing genes related to pseudohypoxia in Hepa1-6 hepatoma, defined as those upregulated in Thy1-OE versus WT mouse hepatoma, as well as in Hif1a-competent versus Hif1a-knockdown Thy1-OE groups (f). Cancer hallmark enrichment analysis of these genes was performed (g) using bulk RNA-seq data (n = 3; CRA036043). The red line indicates adjusted P < 0.05. h, Comparative analysis of metastasis-related gene expression profiles among THY1− CSCs, HIF-signaturehigh THY1+ CSCs, and HIF-signaturelow THY1+ CSCs in human HCC. i,j, Expression of hypoxia-related proteins (i) and EMT-related proteins (j) of WT and Thy1-OE Hepa1-6 cells was analysed (each n = 5). k, Migration potential of WT and Thy1-OE Hepa1-6 cells cultured in vitro was estimated (n = 5; scale bar, 60 µm). l, Effects of Thy1-OE on survival in mice bearing Hepa1-6 hepatomas (n = 8). Data are from three (f,g) or five (i,j) independent experiments. Data are presented as mean ± s.e.m. of three (e,l) or five (k) independent experiments. Statistical significance was determined using linear regression with two-sided t-tests for regression coefficients (d), over-representation analysis based on the hypergeometric test, with false discovery rate correction applied for multiple comparisons (g,h), unpaired two-tailed Student’s t-test (e,k) or the log-rank (Mantel–Cox) test (l). Violin plots show data distribution with median and interquartile range (b,c). Panel a created with BioRender.com. ERI, enabling replicative immortality; REM, reprogramming energy metabolism; RCD, resisting cell death; TPI, tumour-promoting inflammation; TIM, tissue invasion and metastasis; IA, inducing angiogenesis; AID, avoiding immune destruction; EGS, evading growth suppressors; GI, genome instability and mutation; SPS, sustaining proliferative signalling.
Extended Data Fig. 5 Neutrophil-triggered pseudohypoxia is required for the metastasis property of THY1+ CSCs.
a, GO enrichment analysis of notably upregulated genes (log2FC > 1, p.adj < 0.01) in HIF-signaturelow THY1+ CSC versus HIF-signaturelow THY1− tumour cell-related regions. b, Heatmap of neutrophil infiltration indices in TCGA cancer tissues with low or high THY1+ CSC hypoxia signature score, calculated by GSVA using specific markers (log2FC > 1, min.pct > 0.2) defining THY1⁺ CSC-related hypoxic regions in the spatial transcriptome analysis. c, Efficiency of THY1-knockdown (shTHY1) or overexpression (THY1-OE) in Huh-7 cells (n = 5). d, Efficiency of HIF1A-GFP-overexpression (HIF1α-GFP) in Huh-7 cells (n = 3; scale bar, 10 µm), wild-type GFP was used as control. e, Metastasis score of THY1+ CSCs in patients with HCC with high versus low neutrophil infiltration (n = 94 patients). f, Gating strategies for FACS of neutrophils from mouse hepatoma tumours. g, Depleting efficiency of αLy6G antibody on neutrophils in mouse hepatoma model (n = 5). Data are from three (d) or five (c) independent experiments. Data are presented as mean ± s.e.m. of three (g) independent experiments. Statistical significance was determined using over-representation analysis based on the hypergeometric test, with false discovery rate correction applied for multiple comparisons (a), the Mann–Whitney U test (b,e), with false discovery rate correction applied for multiple comparisons (b), or two-way ANOVA with Tukey’s post hoc test (g). Violin plots show data distribution with median and interquartile range (e).
Extended Data Fig. 6 THY1 signalling promote ROS-enriched mitochondria extrusion by neutrophils.
a, THY1+ Huh-7 cells were left untreated, co-cultured with neutrophils directly or in a transwell system. HIF1α protein level was analysed (n = 5). b,c, THY1+ Huh-7 cells were left untreated, treated with Co-CM or THY1-CM in the absence (b,c) or presence (c) of the proteasome inhibitor MG132. mRNA (b) and protein (c) level of HIF1α were analysed (each n = 5). d, Schematic overview of achieving different Co-CM fractions. e, Effects of blocking antibodies on mitochondrial proteins in the 18,000g pellet from Co-CM derived from cocultures of THY1+ Huh-7 cells and neutrophils (n = 5). f, Effects of THY1-Fc on mitochondrial proteins in the 18,000g pellet from neutrophil-conditioned medium (n = 5). g,h, Mitochondrial content (g) and mitochondrial membrane potential (h) in untreated or THY1-Fc-treated neutrophils were analysed (each n = 5). i, Schematic representation of isolating neutrophil mitochondria. j, Intracellular mitochondria were extracted from fresh, 12-hour-cultured untreated or THY1-Fc-treated neutrophils. Mitochondrial membrane potential of these mitochondria was analysed (n = 5). k, Selection of stable mitochondrial content indicator genes for Fig. 4n, o. The selected genes are highlighted in red, with variation coefficient less than 1.0 in both blood and tumour samples from patients with HCC. l, Gating strategies for FACS of neutrophils from HCC tumours. m, Mitochondrial content in neutrophils from HCC tumour and paired blood was analysed in the original HCC cohort by flow cytometry (n = 17 patients). Data are from five (a,c,e,f,j) independent experiments. Data are presented as mean ± s.e.m. of five independent experiments (b,g,h,m). Statistical significance was determined using unpaired two-tailed Student’s t-test (g,h), paired two-tailed Student’s t-test (m), or one-way ANOVA with Tukey’s post hoc test (b). Panels d,i and l created with BioRender.com.
Extended Data Fig. 7 THY1 signalling facilitate neutrophil unhealthy mitochondria extrusion via migrasome.
a, Schematic representation of the protocol used for MEP sorting. Mitochondrial proteins from each indicated layer were analysed by immunoblotting (n = 5). b, Schematic overview of the isolation of different THY1-CM fractions. Marker proteins for migrasomes, exosomes, and apoptotic bodies in the purified THY1-CM fractions obtained from the same centrifugation process were analysed by immunoblotting. Samples were normalized to total protein concentration determined by the bicinchoninic acid (BCA) assay (n = 5). c, Time-lapse imaging of untreated neutrophils. The white dashed line outlined the cells. (n = 3; scale bar, 5 µm). The cell membrane was labelled with DIL. d, Membrane potential in excreted and intracellular mitochondria from untreated or THY1-Fc -treated neutrophils was analysed using Mitotracker Deep Red staining. (n = 3; scale bar, 5 µm). Representative images of THY1-Fc-treated neutrophils were shown. CFSE staining was employed to outline the cells. e, Lysosome-, endoplasmic reticulum-, and Golgi-related proteins in migrasomes from THY1-Fc-treated neutrophils were analysed (n = 5). f, Dot plot of ITGAM expression in each cell type, with dot size representing the fraction of expressing cells and colour indicating mean expression intensity, based on a publicly available scRNA dataset (skrx2fz79n). g, Monocytes were left untreated or treated with THY1-Fc. Migrasomal and mitochondrial proteins in MEP derived from CM were determined (n = 5). Data are from five (a,b,e,g) independent experiments. Data are presented as mean ± s.e.m. of over 100 cells (c,d) across three independent experiments. Statistical significance was determined using unpaired two-tailed Student’s t-test (d) or the two-sided Wilcoxon rank-sum test, and false discovery rate correction was applied for multiple comparisons across cell types (f). Violin plots show data distribution with median and interquartile range (d). Panel b created with BioRender.com.
Extended Data Fig. 8 THY1 signalling enhanced macropinocytosis to initiate tumour pseudohypoxia.
a, Three-dimensional reconstruction and positional mapping of TOM20-GFP-loaded migrasomes in a THY1⁺ Huh-7 cell (n = 30 cells from three independent experiments; Scale bar, 10 µm). The cell membrane was labelled with DIL. b, Macropinocytosis of dextran in Carmil1 wild-type (Carmil1-WT) and Carmil1 mutant (Carmil1-AA) Thy1-overexpressing (Thy1-OE) Hepa1-6 cells (n = 3; scale bar, 10 µm). c, Carmil1-WT and Carmil1-AA Thy1-OE Hepa1-6 cells were inoculated into mice livers. The total number of metastatic foci of 30 lung sections were assessed (n = 5; scale bar, 250 µm). d, GSEA of the macropinocytosis signature in THY1⁺ CSCs versus THY1⁻ cancer cells in HCC single-cell transcriptomic datasets. e, Uptake of migrasomes from TOM20-GFP–expressing HL60 cells by THY1-overexpressing Huh-7 cells in the presence of recombinant Mac1-Fc protein (n = 3; scale bar, 10 µm). f, Effect of THY1-overexpression (THY1-OE) on signalling pathway activation in Huh-7 cells (n = 5). g,h, Effects of pathway inhibitors on migrasome uptake (g; n = 3) and HIF1α expression (h; n = 5) in THY1⁺ Huh-7 cells. i, Specificity of TthIII digestion on C57BL/6-specific mitochondrial DNA fraction was assessed (n = 5). j, UMAP visualization of the cell clusters, patient annotations, and NMT states of tumour cell from 11 patients with HCC with marked NMT. Data are from three (a) or five (f,h,i) independent experiments. Data are presented as mean ± s.e.m. of five mice (c) or over 100 cells (b,e,g) across three independent experiments. Statistical significance was determined using unpaired two-tailed Student’s t-test (b,c,e), a one-sided, permutation-based test (d), one-way ANOVA with Tukey’s post hoc test (g). Violin plots show data distribution with median and interquartile range (b,e,g). Panel i created with BioRender.com.
Supplementary information
Supplementary Table 1 (download XLSX )
Information on pan-cancer samples from publicly available datasets.
Supplementary Table 2 (download XLSX )
Clinical characteristics of the 135 patients with HCC.
Supplementary Table 3 (download XLSX )
Clinical characteristics of 22 patients with untreated HCC with paired PT and PVTT tissues.
Supplementary Table 4 (download XLSX )
Clinical characteristics of 86 patients with untreated HCC.
Supplementary Table 5 (download XLSX )
Clinical characteristics of ten patients with untreated HCC with fresh blood.
Supplementary Table 6 (download XLSX )
List of CSC transcription factors and ChIP-seq analysis of transcription factors binding the THY1 gene.
Supplementary Table 7 (download XLSX )
Clinical characteristics of 17 patients with untreated HCC with fresh tissues and blood.
Supplementary Table 8 (download XLSX )
Summary of cell lines.
Supplementary Table 9 (download XLSX )
Summary of primers.
Supplementary Table 10 (download XLSX )
Summary of plasmids.
Supplementary Table 11 (download XLSX )
Summary of shRNA, guideRNA.
Supplementary Table 12 (download XLSX )
Summary of antibodies.
Supplementary Table 13 (download XLSX )
Summary of critical kits.
Supplementary Table 14 (download XLSX )
Gene sets used in GSEA and GSVA.
Source data
Source Data (download PDF )
Unprocessed western blots and gels.
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wan, WH., Li, PL., Cao, WJ. et al. THY1+ cancer stem cells drive metastasis through a pseudohypoxic state shaped by neutrophil-derived mitochondria. Nat Cell Biol 28, 596–607 (2026). https://doi.org/10.1038/s41556-026-01876-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-026-01876-1