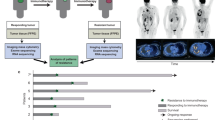
Abstract
Patients with impaired tumour-specific major histocompatibility complex class I (tsMHC-Iimpaired) often fail to respond to immune checkpoint blockade (ICB), presenting a major clinical challenge. However, through our multicentre investigation, we observed that a subset of patients with tsMHC-Iimpaired remains responsive to ICB, a phenomenon that has not been fully explained. Here we identify a COTL1high natural killer (NK) subset that mediates ICB responsiveness in these patients. Mechanistically, PD-L1+ macrophages coexpress GITRL and engage GITR on COTL1high NK cells, whereas PD-L1 blockade relieves the PD-1-mediated inhibition of GITR signalling and promotes NK cell activation. Activated COTL1high NK cells enhance immunological synapse stability and IFN-γ production via a metabolic–H3K27ac–RBPJ axis, thereby upregulating tsMHC-I expression and reinforcing adaptive anti-tumour immunity. Notably, GITR activation significantly enhances the sensitivity to anti-PD-L1 therapy in tsMHC-Iimpaired models. Our findings identify COTL1high NK cells as key determinants of ICB responsiveness and highlight the GITRL–GITR axis as a promising therapeutic target for tsMHC-Iimpaired tumours.
Similar content being viewed by others
Main
Immune checkpoint blockade (ICB), particularly therapy targeting PD-1/PD-L1 (αPD-1/PD-L1), has reshaped treatment paradigms for multiple solid tumours1,2,3. However, only a subset of patients (approximately 30%) derives clinical benefit4,5. ICB primarily acts by reinvigorating exhausted CD8+ T cells6,7, whose anti-tumour function depends on the recognition of tumour antigens presented by major histocompatibility complex class I (MHC-I) molecules8. Consequently, the deficiency or downregulation of tumour-specific MHC-I (tsMHC-I) is a key mechanism of immune evasion, often resulting in a ‘cold’ tumour microenvironment (TME) and poor response to ICB9,10. Intriguingly, a proportion of patients with tsMHC-Iimpaired tumours still respond to ICB, despite limited CD8+ T cell infiltration, though the underlying mechanisms remain unclear11. Unravelling these mechanisms is crucial for improving immunotherapy outcomes in this patient population.
In cancers such as hepatocellular carcinoma (HCC), classic genetic alterations causing MHC-I loss (for example, B2M mutations) are rare, and tumour mutation burden is typically low12,13,14 (Supplementary Fig. 1). This implies that non-genetic mechanisms probably underlie the tsMHC-Iimpaired phenotype, where tumours exhibit a heterogeneous mix of MHC-Ihigh and MHC-Ilow tumour cells rather than uniform loss. Thus, the restoration of tsMHC-I through ICB may explain the favourable responses in several patients with impaired tsMHC-I expression15. Type I and II interferons (IFNs) are pivotal for inducing tsMHC-I expression and fostering immune cell crosstalk within the TME16. Natural killer (NK) cells, which are abundant in the liver, are key players in this context17. They can recognize and eliminate tumour cells with impaired MHC-I and produce IFN-γ to enhance immune responses18. Understanding how to harness NK cell function represents a promising avenue for enhancing ICB efficacy in tsMHC-Iimpaired cancers.
Effective NK cell function, including target cell killing and productive cell–cell communication, relies on the formation of a stable immunological synapse (IS)19,20. The stability of the IS and the efficiency of signal transduction depend on F-actin regulation21,22. In this study, we identify a COTL1high NK cell subset that is enriched in ICB-responsive, tsMHC-Iimpaired tumours. COTL1 is an actin-binding protein critical for F-actin polymerization, a process essential for IS stability23. We demonstrate that these COTL1high NK cells establish stable synaptic interactions with PD-L1+ macrophages. Mechanistically, αPD-L1/PD-1 therapy unleashes a functional crosstalk via the GITRL–GITR axis between these macrophages and NK cells. GITR triggering induces metabolic and epigenetic reprogramming in COTL1high NK cells, leading to RBPJ upregulation and a feedback loop that potentiates IFNG and COTL1 expression. This loop sustains stable IS formation and continuous IFN-γ production, which in turn enhances tsMHC-I expression and promotes adaptive CD8+ T cell responses. Importantly, combining GITR agonism with αPD-L1 treatment elicits robust anti-tumour activity in multiple tsMHC-Iimpaired models.
In summary, our findings delineate a critical role for COTL1high NK cells in mediating the ICB response of tsMHC-Iimpaired tumours and propose the therapeutic reactivation of the GITRL–GITR signalling, as a promising strategy to overcome ICB resistance in this challenging context.
Results
COTL1high NK cell infiltration level predicts immunotherapy response in tumours with impaired tsMHC-I expression
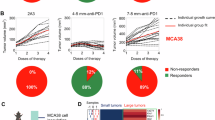
The loss or degradation of tumour MHC-I impairs CD8+ T cell recognition and is a recognized mechanism of immunotherapy resistance9. Intriguingly, an analysis of 56 pretreatment tumours from three independent centres revealed a subset of patients who responded to αPD-1/PD-L1 therapy (alone or in combination with kinase/VEGFR inhibitors), despite exhibiting both impaired tsMHC-I and limited CD8+ T cell infiltration (Fig. 1a,b and Supplementary Table 1). To investigate the underlying mechanisms, we analysed bulk RNA sequencing (RNA-seq) data from patients treated with atezolizumab ± bevacizumab (EGAD00001008128)24. Using deconvolution analysis25, we classified 289 pretreatment tumours into tsMHC-Ihigh and tsMHC-Iimpaired groups (Fig. 1c). Although high tsMHC-I correlated with increased CD8+ T cell infiltration and better response (Fig. 1c and Extended Data Fig. 1a), 26.2% of patients with the tsMHC-Iimpaired phenotype still responded to ICB (Fig. 1c). The subsequent immune subset analysis (Supplementary Table 2) identified NK cells as significantly enriched in these responders (Fig. 1d and Extended Data Fig. 1b,c). On the basis of HCC tsMHC-I expression patterns (Supplementary Fig. 2), we validated this finding in 24 tsMHC-Iimpaired tumours (9 responders, 15 non-responders), defined by a tsMHC-I rate <80% and fewer than 200 CD8+ T cells per field of view (FOV), using multiplex immunohistochemistry (mIHC) or imaging mass cytometry. Responders demonstrated a significantly higher density of tumour-infiltrating NK cells (Fig. 1e,f and Extended Data Fig. 1d). Although γδ T cells also possess MHC-I-independent cytotoxic potential, their infiltration and functional profiles showed no significant difference between responders and non-responders in this context (Extended Data Fig. 1e). The functional necessity of NK cells was further confirmed in an orthotopic, tsMHC-Iimpaired H22 model. NK cell depletion not only accelerated tumour progression in untreated mice but also completely abrogated the efficacy of αPD-L1 therapy (Fig. 1g,h and Extended Data Fig. 1f,g). This establishes NK cells as essential mediators of the ICB response under tsMHC-Iimpaired conditions. Consistent with this, MHC-I modulation models confirmed that reduced tsMHC-I expression enhanced NK cell infiltration while diminishing CD8+ T cell presence (Extended Data Fig. 2a–l). Further in vivo depletion studies in the orthotopic Hep-53.4 model revealed that the loss of either NK or CD8+ T cells significantly impaired αPD-L1 efficacy (Extended Data Fig. 2m,n). Notably, the comparable effects of combined or individual depletion of NK and CD8+ T cells highlight NK cells as critical effectors of anti-tumour immunity under tsMHC-Iimpaired conditions.
a, The representative mIHC staining for Pan-CK, HLA-ABC and CD8 for responders with high tsMHC-I expression (left) and CD8+ T cell infiltration or with impaired tsMHC-I expression and CD8+ T cell infiltration (right). n = 56 patients. Scale bars, 50 μm. b, Bar plots depicting the average counts of CD8+ T cells and tumour MHC-I expression levels across 56 pretreated tumour samples from the JSPH, WXPH and RJH centres, as measured by mIHC. Patients were treated with durvalumab or camrelizumab or tislelizumab or sintilimab treatment for at least three cycles. Grey and red arrows are used to indicate tsMHC-Iimpaired non-responders (n = 10) and responders (n = 9), respectively. c, A heat map showing the T cell, CD8+ T cell and neoantigen reactive CD8+ T cells (NeoTCR8) signature expression in 289 pretreated tumours from GO30140 and IMbrave150 cohorts. tsMHC-I expression was measured by the Bisque package. d, A lollipop chart showing the correlation between therapy response and different immune cell signature score in 144 pretreatment tumours with impaired MHC-I expression and low CD8+ T cell score from GO30140 and IMbrave150 cohorts (EGAD00001008128). Odds ratio (OR) values were determined by logistic regression. e,f, Representative magnetic resonance imaging images (left) and the paired haematoxylin and eosin (H&E) and mIHC staining of CD8 and CD56 (right) in responders (n = 9) and non-responders (n = 15) with impaired tsMHC-I and limited CD8+ T cells (e); and the statistical results of mean NK cell numbers in ten random fields (1,000 μm × 1,000 μm) per HCC tumour are shown (f) (P = 0.0013). The red arrows mark the tumour regions. The blue and red arrows in the mIHC images mark NK and CD8+ T cells, respectively. Scale bars, 200 μm and 20 μm. g,h, Representative images (g) and statistical results (h) of whole-liver morphology from isotype control or αPD-L1 or anti-asialo GM1 or combination-treated H22-bearing orthotopic HCC models (each n = 5). The dashed white lines mark the tumour boundary. Scale bars, 1 cm. Isotype versus anti-asialo GM1 P = 0.0238, αPD-L1 P = 0.0397; αPD-L1+anti-asialo GM1 versus αPD-L1 P = 0.0079, anti-asialo GM1 P > 0.9999. i, The UMAP plot of 30,241 sorted CD56+ NK cells from four patients with variable tsMHC-I expression, demonstrating the formation of six distinct clusters. j, The UMAP view of cell density displaying tumour-infiltrated NK cell distribution across tsMHC-Ihigh and tsMHC-Iimpaired groups. High relative cell density is shown as bright magma. k, The UMAP plots showing the expression of COTL1, GZMK, ITGA1 and ITGAE in the tumour-derived NK cells. l, Left: the representative H&E and mIHC staining depicting CD56+COTL1+ NK cells and CD3+ T cells of HCC specimens before treatment are shown. Scale bar, 50 μm. Right: the statistical analysis of the distribution of average CD56+ COTL1+ NK cells per patient, comparing responders (n = 9) and non-responders (n = 15) with a tsMHC-I+ rate below 80% and CD8+ T cell counts under 200 (P < 0.0001). Each patient, ten FOVs (1,000 μm × 1,000 μm) for analysis. m, An analysis of MHC-I, NeoTCR8 and COTL1high NK signature distribution in two pretreated HCC tissues (one responder and one non-responder) based on the Stereo-seq data (https://doi.org/10.6084/m9.figshare.22332352). n, The COTL1high NK cell signature score was analysed in 144 pretreated tumours with impaired tsMHC-I expression and low CD8+ T cell score from GO30140 and IMbrave150 cohorts (P = 0.01; responders, n = 31; non-responders, n = 113). P values were determined by unpaired two-sided Mann–Whitney test in f, h, l and n. Data are shown as the mean ± s.e.m. In the box plots, the centre line is the median, box limits are the first and third quartiles and whiskers are 1.5× interquartile range. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; n.s., not significant.
Guided by these findings and prior studies linking specific NK subsets to immunotherapy outcomes26,27, we performed single-cell RNA-seq (scRNA-seq) on NK cells from four treatment-naive patients with HCC with heterogeneous tsMHC-I expression. This analysis identified six distinct NK cell states (Fig. 1i and Supplementary Fig. 3a). Among these, a COTL1high subset was notably enriched in tsMHC-Iimpaired tumours (Fig. 1j). This subset coexpressed markers associated with tissue residency (ITGA1, ITGAE, ZNF683, CXCR6) and adaptive NK cells (KLRC2, CD52, IL32) (Fig. 1k and Supplementary Fig. 3b,c). The spatial analysis confirmed that tumours with impaired tsMHC-I, particularly those from responders, were enriched for COTL1+CD56+ NK cells compared with tsMHC-Ihigh tumours (Fig. 1l–n and Extended Data Fig. 2o,p). Collectively, these results identify COTL1high NK cells as a key cellular correlate of ICB response in tsMHC-Iimpaired HCC and suggest their potential role in rescuing anti-tumour immunity in this context.
Tumour-infiltrating COTL1high NK cells display both activated and dysfunctional states
Although NK cells are generally reduced in HCC tumours compared with adjacent normal tissue (Extended Data Fig. 3a,b) and are known to become dysfunctional in the TME28, we observed that COTL1high NK cells were more abundant and functionally relevant in tsMHC-Iimpaired tumours. The transcriptional profiling of this subset revealed a unique signature, coexpressing markers associated with dysfunction (NR4A2, TOX, TIGIT) and effector or activation (CD44, GZMK, GZMA) genes (Fig. 2a). Although sharing effector molecules such as GZMA and IFNG with conventional NK cells, COTL1high NK cells were distinguished by the elevated expression of exhaustion markers (PDCD1, HAVCR2, LAG3) and specific receptors (KIR2DL4, KLRC2, KLRC1) (Fig. 2b and Extended Data Fig. 3c,d).
a, Scatter plots showing the DEGs between COTL1high NK cells and other NK populations in the tumours. b, The frequencies of positive cells for selected functional genes across six NK clusters from each HCC tumour (n = 4) analysed using scRNA-seq. Data are presented as median ± interquartile range, with individual samples (dots). KIRs, killer cell immunoglobulin-like receptors. c, The t-distributed stochastic neighbour embedding map plots showing the seven NK clusters defined by 25 protein markers. The black arow marks the COTL1high NK cluster. cNK, conventional NK cells; lrNK, liver-resident NK cells. d, A heat map showing the protein expression used in the clustering across seven NK subtypes defined by CyTOF data. e, Bar plots showing the positive rate of CD11b, CD27, granzyme B and IFN-γ in the four tumour-enriched subtypes. f–h, A cytometry analysis of NK cells from tumour tissue of patients with HCC (n = 14) expressing PD-1 (f) (P = 0.0034), IFN-γ (g) (P = 0.0001) and granzyme K (h) (P = 0.0012). i, A violin plot showing the expression distribution of ten-gene IFN-γ-related signature score in COTL1high NK cells from pretreated and post-treated data (GSE235863). j, The RNA-seq data analysis showing the average MHC-I gene expression of paired pretreated and post-treated tumours (EGAD00001010132, n = 5 versus 11; responder, pre versus post, P = 0.0079; non-responder, pre versus post, P = 0.3248). k, The enrichment of IFN-γ response pathways in matched samples with or without MHC-I increase (EGAD00001010132). (with MHC-I increase, pre versus post, P = 0.0009; without MHC-I increase, pre versus post, P = 0.4518). l, The enrichment of IFN-γ downstream JAK–STAT signalling pathway in matched samples with or without MHC-I increase (EGAD00001010132) (with MHC-I increase, pre versus post, P = 0.0001; without MHC-I increase, pre versus post, P = 0.8383). m, Representative mIHC staining (left) and statistical results (right) depicting the tumour MHC-I expression of paired pretreated and post-treated specimens (n = 5 versus 5). Scale bar, 250 μm. (responder, pre versus post, P = 0.0021; non-responder, pre versus post, P = 0.7990). n, The spatial transcriptomics analysis of post-treated samples showing the MHC-I expression, COTL1high NK cell positive spot and IFN-γ response score in responders and non-responders (GSE238264). P values were determined by paired two-sided Mann–Whitney test in f–h and j–m. P values were determined by unpaired two-sided Mann–Whitney test in i. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; n.s., not significant.
To validate these features at the protein level, we leveraged our scRNA-seq data, which showed that the COTL1high subset coexpresses COTL1, TNFRSF18 (encoding GITR) and ITGA1 (encoding CD49a) (Fig. 1j). The cytometry by time of flight (CyTOF) analysis of 109,930 NK cells confirmed that CD49a+CD103+ (COTL1high) NK cells expressed NKG2A, PD-1 and IFN-γ (Fig. 2c,d, Extended Data Fig. 3e,f and Supplementary Table 3). Furthermore, these cells predominantly exhibited a CD11b+CD27− effector phenotype (Fig. 2e)29. Flow cytometry corroborated that tumour-infiltrating CD49a+ NK cells expressed higher levels of PD-1, IFN-γ and granzyme K compared with their CD49a− counterparts (Fig. 2f–h, Extended Data Fig. 3g,h, Supplementary Fig. 3d and Supplementary Table 4).
We next evaluated the functional role of these cells in ICB therapy. Following αPD-1 treatment, the IFN-γ signature score30 of COTL1high NK cells increased significantly (Fig. 2i). Accordingly, bulk RNA-seq analyses revealed enhanced IFN‑γ pathway activity and MHC-I gene expression in responders (Fig. 2j–l). This pattern was corroborated at the tissue level by mIHC and spatial transcriptomics, which demonstrated that ICB led to increased tsMHC‑I expression and greater infiltration of both COTL1high NK and CD8+ T cells specifically in responders (Fig. 2m,n and Extended Data Fig. 3i,j). Together, these data indicate that COTL1high NK cells drive IFN‑γ-mediated tsMHC‑I upregulation, which promotes adaptive anti‑tumour immunity under ICB.
PD-1/PD-L1 blockade boosts COTL1high NK cells producing IFN-γ to upregulate tsMHC-I expression
To investigate whether the ICB response in tsMHC-Iimpaired tumours depends on IFN-γ from COTL1high NK cells, we turned to preclinical models. By integrating transcriptomic data from four published datasets across three murine HCC models and single-cell cross-species mapping analysis, we identified a strong transcriptional concordance between human COTL1high NK cells and murine CD49a+CD49b− NK cells (also referred to as ILC1s) (Extended Data Fig. 4a–d). At epigenetic levels, they also shared similar chromatin accessibility profiling (Extended Data Fig. 4e). These data support the conservation between human and mice. We therefore characterized CD49a+CD49b− NK cells in Hep‑53.4 and H22 orthotopic HCC models, both of which exhibit impaired MHC‑I expression (Extended Data Figs. 1f and 2a). In these models as well as in the Diethylnitrosamine–CCl4-induced HCC model, CD49a+CD49b− NK cells expressed higher levels of PD‑1 and IFN‑γ compared with CD49a−CD49b+ NK cells, recapitulating the phenotype of human COTL1high NK cells (Extended Data Fig. 4f–j).
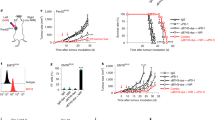
In the liver, Zfp683+ NK cells coincided with CD49a+ NK cells31 (Fig. 3a and Extended Data Fig. 4c,k). Although Zfp683 deficiency has been reported to cause a selective loss of liver-resident CD49a+ NK cells and tissue-resident memory T (Trm) cells32, it did not significantly affect either the proportion or absolute number of tumour-infiltrating CD8+ Trm cells under tsMHC-Iimpaired conditions (Extended Data Fig. 4l). We therefore used Zfp683−/− mice bearing orthotopic Hep-53.4 tumours (Fig. 3b), which are resistant to αPD-L1 treatment33, to assess the functional requirement of this NK cell subset in ICB. Compared with wild-type (WT) controls, Zfp683 deficiency not only accelerated baseline tumour growth but also completely abolished the anti-tumour efficacy of αPD-L1 therapy (Fig. 3c). Before treatment, Zfp683 deletion did not alter the proportion or effector function of CD4+ or CD8+ T cells (Fig. 3d,e and Extended Data Fig. 4m–o). By contrast, following αPD-L1 administration, intratumoural CD8+ T cells in Zfp683−/− mice failed to elicit enhanced effector functions (Fig. 3d,e). Moreover, Zfp683−/− mice displayed an impaired upregulation of tumour MHC-I in response to αPD-L1 treatment (Fig. 3f). These results underscore the essential role of IFN-γ derived from COTL1high NK cells in driving adaptive anti-tumour immunity, particularly in tsMHC-I-impaired settings.
a, The relative abundance of NK cells stained with CD49a and CD49b in Hep-53.4 tumours from WT and Zfp683−/− mice demonstrating CD49a+CD49b− NK cell depletion. b, A schematic illustrating the process of αPD-L1 treatment in orthotopic HCC tumours. Ab, antibody. c, The representative images and statistical results of whole-liver morphology from isotype control- or αPD-L1-treated Hep-53.4-bearing orthotopic HCC models on Zfp683+/+ and Zfp683−/− mice (each n = 5; Zfp683+/+–isotype versus Zfp683−/−–isotype, P = 0.0238, Zfp683+/+–αPD-L1, P = 0.0079; Zfp683−/−–αPD-L1 versus Zfp683−/−–isotype, P = 0.0635, Zfp683+/+–αPD-L1, P = 0.0397). Scale bars, 1 cm. The dashed white lines mark the tumour boundary. d, The flow cytometry analysis and statistical results of IFN-γ expression in tumour-infiltrating CD8+ T cells from isotype control- or αPD-L1-treated Hep-53.4-bearing orthotopic HCC models on Zfp683+/+ and Zfp683−/− mice (each n = 5; Zfp683+/+–isotype versus Zfp683−/−–isotype, P = 0.2540, Zfp683+/+–αPD-L1, P = 0.0317; Zfp683−/−–αPD-L1 versus Zfp683−/−–isotype, P = 0.5476, Zfp683+/+–αPD-L1, P = 0.0079). e, The flow cytometry analysis and statistical results of granzyme B expression in tumour-infiltrating CD8+ T cells from isotype control- or αPD-L1-treated Hep-53.4-bearing orthotopic HCC models on Zfp683+/+ and Zfp683−/− mice (each n = 5; Zfp683+/+–isotype versus Zfp683−/−–isotype, P = 0.6905, Zfp683+/+–αPD-L1, P = 0.0159; Zfp683−/−–αPD-L1 versus Zfp683−/−–isotype, P = 0.8413, Zfp683+/+–αPD-L1, P = 0.0238). f, The mean fluorescence intensity of MHC-I in Hep-53.4-implanted tumours from WT and Zfp683−/− mice determined by mean values of five random FOVs (each n = 4; Zfp683+/+–isotype versus Zfp683−/−–isotype, P = 0.2000, Zfp683+/+–αPD-L1, P = 0.0286; Zfp683−/−–αPD-L1 versus Zfp683−/−–isotype, P = 0.3429, Zfp683+/+–αPD-L1, P = 0.0286). g, The flow cytometry histogram showing the protein expression of MHC-I (H-2Kb/H-2Db) on sgCtrl and sgIfngr1 Hep-53.4 cells. h, A schematic illustrating the process of αPD-L1 treatment in orthotopic sgCtrl and sgIfngr1 Hep-53.4 tumours. i, Representative images and statistical results of whole-liver morphology from αPD-L1 treated WT Hep-53.4 and Ifngr1-knockout (KO) Hep-53.4-bearing orthotopic HCC models (each n = 5; sgCtrl–isotype versus sgCtrl–αPD-L1, P = 0.0238, sgIfngr1–isotype, P = 0.2143; sgIfngr1–αPD-L1 versus sgIfngr1–isotype, P = 0.1190, sgCtrl–αPD-L1, P = 0.0079). Scale bars, 1 cm. The dashed white lines mark the tumour boundary. j, The mean fluorescence intensity of MHC-I in sgCtrl and sgIfngr1 Hep-53.4-implanted tumours from WT mice determined by mean values of five random FOVs (each n = 4; sgCtrl–isotype versus sgCtrl–αPD-L1, P = 0.0286, sgIfngr1–isotype, P = 0.0286; sgIfngr1–αPD-L1 versus sgIfngr1–isotype, P = 0.3429, sgCtrl–αPD-L1, P = 0.0286). k, The flow cytometry analysis and statistical results of CD8+ T cells frequency from WT and Ifngr1-KO Hep-53.4 tumours (each n = 5; sgCtrl–isotype versus sgCtrl–αPD-L1, P = 0.0159, sgIfngr1–isotype, P = 0.5476; sgIfngr1–αPD-L1 versus sgIfngr1–isotype, P = 0.1111, sgCtrl–αPD-L1, P = 0.0079). l, Flow cytometry analysis and statistical results of CD8+IFN-γ+ T cells frequency from WT and Ifngr1-KO Hep-53.4 tumours (each n = 5; sgCtrl–isotype versus sgCtrl–αPD-L1, P = 0.0079, sgIfngr1–isotype, P = 0.7381; sgIfngr1–αPD-L1 versus sgIfngr1–isotype, P = 0.4524, sgCtrl–αPD-L1, P = 0.0079). P values were determined by unpaired two-sided Mann–Whitney test in c–f and i–l. Axes represent relative fluorescence intensity in arbitrary units. Data are shown as the mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; n.s., not significant.
As IFN‑γ‑mediated MHC‑I upregulation depends on tumour‑intrinsic IFN-γR signalling, we inoculated mice with sgIfngr1 Hep-53.4 cells to test the contribution of this pathway to tsMHC‑I induction and αPD‑L1 efficacy (Fig. 3g,h). The loss of IFN-γR in tumours markedly attenuated the therapeutic effect of αPD‑L1 and failed to increase tsMHC‑I expression (Fig. 3i,j). Moreover, PD‑L1 blockade could not rescue CD8+ T cell infiltration or enhance their effector function in IFN-γR-deficient tumours (Fig. 3k,l). Thus, tumour-intrinsic IFN-γR signalling is essential not only for MHC-I induction but also for CD8+ T cell recruitment and effector activation. In conclusion, these findings demonstrate that IFN-γ from COTL1high NK cells is crucial for enhancing tsMHC-I expression and sustaining CD8+ T cell function during PD-1/PD-L1 blockade.
αPD-1/PD-L1 treatment eliminates the suppression on GITRL–GITR axis-mediated IFN-γ production in COTL1high NK cells
We next investigated how the HCC microenvironment sustains COTL1high NK cell function. A cell–cell interaction analysis indicated that COTL1high NK cells interact most strongly with macrophages (Fig. 4a), a finding corroborated by their spatial colocalization in tumour sections (Fig. 4b and Extended Data Fig. 5a). Given our earlier work identifying macrophages as a major source of PD‑L1 in HCC34 (Extended Data Fig. 5b), we cocultured sorted COTL1high NK cells with PD‑L1+ macrophages. The αPD-L1 treatment significantly boosted IFN-γ production only when cells were in direct contact, pointing to a requirement for membrane-bound signals in NK cell activation (Fig. 4c). Among the surface receptors upregulated in COTL1high NK cells, TNFRSF18 (GITR) stood out for its established immunomodulatory roles35 (Fig. 4d and Extended Data Fig. 5c). The flow cytometric analysis further confirmed the presence of an NK cell subset coexpressing COTL1, GITR and CD49a (Extended Data Fig. 5d). Notably, GITR expression was higher in CD49a+ NK cells than in CD49a− NK cells or other immune cell subsets (Fig. 4e and Supplementary Fig. 4). Its ligand, GITRL (TNFSF18), was highly expressed on PD‑L1+ macrophages (Fig. 4f,g and Extended Data Fig. 5e–g). Although dendritic cells (DCs) also expressed GITRL, the spatial transcriptomic analysis revealed that PD-L1+ macrophages were consistently positioned in closer proximity to COTL1high NK cells within HCC tissues (Extended Data Fig. 5h–j). Notably, this spatial preference was not restricted to HCC; similar patterns were observed in nasopharyngeal carcinoma and pancreatic cancer (Extended Data Fig. 5k,l), where PD-L1+ macrophages were likewise the immune population most closely associated with COTL1high NK cells. We further validated this spatial organization by mIHC analysis (Extended Data Fig. 5m–o). Functionally, the small interfering RNA (siRNA)-mediated GITR knockdown or antibody-mediated GITR blockade in vitro-derived COTL1high NK cells significantly reduced IFN-γ production (Fig. 4h,i). The treatment of patient-derived COTL1high NK cells with recombinant GITRL (rGITRL) enhanced IFN-γ secretion, an effect reversed by cotreatment with recombinant PD-L1 (rPD-L1) (Fig. 4j). In the coculture with macrophages, αPD-L1 boosted IFN-γ production, but this enhancement was abolished when GITRL was concurrently blocked (Fig. 4k), indicating that GITR signalling is required to sustain IFN-γ production, whereas PD-1/PD-L1 engagement functions as a negative regulator. Furthermore, PD-L1+ macrophage-cocultured COTL1high NK cells increased CD8+ T cell granzyme B expression ex vivo in the presence of tumour cells after PD-L1 blockade, and this effect was diminished by IFN-γ neutralization (Fig. 4l). Clinically, the high colocalization of COTL1high NK cells and PD-L1+ macrophages in tsMHC-Iimpaired responders (Fig. 4m,n) and the positive association with survival in ICB-treated patients (Fig. 4o) underscore the clinical relevance of this axis. Collectively, these findings indicate that PD-1/PD-L1 blockade engages a GITRL–GITR circuit that empowers COTL1high NK cells and boosts downstream CD8+ T cell activity. This circuit is especially important in tsMHC-Iimpaired tumours, where COTL1high NK cells help compensate for defective antigen presentation.
a, The cell–cell interactions between COTL1high NK cells and other cell types indicated by the scPLC data (HRA001748). The line thickness represents the number of significant ligand–receptor pairs. b, The mIHC staining for COTL1, CD56, CD68 and CD3 reveals the colocalization of COTL1high NK cells and macrophages. The white arrows marked marophages and COTL1+ NK cells accordingly. Scale bars, 50 μm and 10 μm. c, An analysis of intratumoural COTL1high NK cells in medium or with PD-L1+ macrophages directly or in a Transwell chamber (each n = 4; medium versus Transwell, P = 0.2000, direct, P = 0.0286; direct versus Transwell, P = 0.0286). d, A Venn diagram showing the upregulated receptors in COTL1high NK cells across two scRNA-seq datasets. e, The flow cytometry analysis of GITR expression in CD49a+ NK and CD49a− NK cells from human HCC tumours (n = 19, P < 0.0001). FMO, fluorescence minus one. f, A dot plot showing the expression level of TNFSF18 (GITRL) in cell subtypes from HCC tumours using the scPLC data (HRA001748). g, The CD274 expression between GITRL− and GITRL+ macrophages in the scPLC data. h,i, Blocking GITRL–GITR using antibodies (Abs) (h) (each n = 3; medium versus GITR Ab, P = 0.0001, GITRL Ab, P = 0.0004) and siRNA (i) (each n = 4; si-NC versus si-GITR 1, P = 0.0286, si-GITR 2, P = 0.0286) in the coculture system abrogated macrophage-mediated effector function of in vitro-induced COTL1high NK cells. j, The tumour-infiltrating COTL1high NK cells were cultured in medium alone or treated by recombinant PD-L1 and GITRL (n = 11). The levels of IFN-γ production were detected by flow cytometry (untreated versus rGITRL, P = 0.0010, rGITRL + rPD-L1, P = 0.3652; rGITRL versus rGITRL + rPD-L1, P = 0.0010). k, The tumour-infiltrating COTL1high NK cells were cultured in medium alone or cocultured with autologous PD-L1+ macrophages in the presence or absence of isotype Ab, αPD-L1 Ab or GITR-shielding Ab (n = 11). The levels of IFN-γ production were detected by flow cytometry (PD-L1 Ab versus isotype, P = 0.0010, GITR Ab, P = 0.0010; GITR Ab versus GITR Ab + PD-L1 Ab, P = 0.4512). l, The tumour-derived CD8+ T cells isolated from HCC tumours were treated with PD-L1 Ab and cultured in medium alone or cocultured with PD-L1+ macrophages cocultured autologous COTL1high NK cells in the presence or absence of isotype Ab and IFN-γ blocking Ab (n = 11). The expression of granzyme B was detected by flow cytometry (isotype versus IFN-γ Ab, P = 0.0010). m, A schematic of the distance calculation between two spots in the spatial transcriptomics dataset. n, The spatial colocalization between GITRL+ macrophages and COTL1high NK cells in patients with MHC-Iimpaired HCC who respond or not respond to αPD-1 therapy (GSE238264 and skrx2fz79n). o, Kaplan–Meier survival curves for overall survival showing the synergistic positive effect of COTL1high NK cells and GITRL+ macrophages in GO30140 and IMbrave150 cohorts (EGAD00001008128). P values were determined by paired or unpaired two-sided Mann–Whitney test in c, e, f and i–l. Data are shown as the mean ± s.e.m. P values were determined by unpaired two-sided Student’s t-test in h. The P values for survival analysis were determined by the log-rank test. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; n.s., not significant.
GITRL–GITR axis potentiated by αPD-1/PD-L1 induces positive feedback between COTL1high NK cells and PD-L1+ macrophages
COTL1, an F-actin-binding protein that stabilizes actin structures by antagonizing cofilin23, plays an important role in organizing the IS. In NK cells, a stable IS is essential not only for target cell killing but also for productive cell–cell communication20.
To structurally assess the role of COTL1 in IS formation, we examined CD49a-sorted COTL1high and COTL1low NK cells from HCC tumours using transmission electron microscopy (TEM). The tumour-derived COTL1high NK cells displayed more surface protrusions than their COTL1low counterparts (Fig. 5a and Extended Data Fig. 6a). Moreover, these protrusions were frequently polarized towards PD-L1+ macrophages (Fig. 5b). Consistent with these observations, immunofluorescence confocal imaging revealed the stronger colocalization of F-actin with COTL1 at the contact interface between COTL1high NK cells and PD-L1+ macrophages, indicating a stabilized IS morphology (Fig. 5c). Accordingly, the siRNA‑mediated knockdown of COTL1 impaired IS formation between COTL1high NK cells and macrophages (Extended Data Fig. 6b). We next sought to understand how IS formation translates into sustained effector function. We hypothesized that direct cell–cell interactions at the IS engage transcriptional programmes that reinforce NK cell function. To investigate downstream signalling, we performed SCENIC analysis and identified RBPJ, a key transcription factor in the NOTCH pathway, as a prominent regulon enriched in COTL1high NK cells (Extended Data Fig. 6c,d). Consistent with this finding, cleavage under targets and release using nuclease (CUT&RUN) sequencing showed higher RBPJ binding in CD49a+ NK cells compared with CD49a− NK cells (Fig. 5d). An integrated analysis of promoter binding, chromatin accessibility and gene expression data identified candidate RBPJ targets (Fig. 5e), including direct binding at the IFNG and COTL1 promoters (Fig. 5f). Notably, the RBPJ expression strongly correlated with GITR levels (Fig. 5g and Extended Data Fig. 6e), and GITR agonism further enhanced RBPJ binding at these loci (Fig. 5h). The truncation of the binding region abolished the repression of the IFNG and COTL1 promoters by RBPJ according to the results of the luciferase assay (Fig. 5i). Therefore, these data suggest the αPD-L1-rescued GITRL–GITR axis links COTL1-mediated IS architecture to RBPJ-driven transcriptional reinforcement of NK cell effector programmes.
a, The TEM showing the membrane protrusions of COTL1high and COTL1low NK cells sorted from resected HCC tumours on the basis of CD49a staining (left). The mean numbers of membrane protrusions are shown (COTL1high versus COTL1low, P < 0.0001). b, The confocal imaging shows NK cells and autologous macrophages were coincubated, fixed and then stained with COTL1 (green) and with Actin-Tracker Red-594 (red) to mark the location of F-actin. Scale bar, 5 μm. c, The TEM showing the interface of COTL1high/COTL1low NK cells and PD-L1+ macrophages. NK cells and PD-L1+macrophages were freshly purified from HCC tumours. The large cells are macrophages; the small cells are NK cells. Scale bar, 1 μm. An analysis of the fluorescence intensity demonstrates an obvious increase in the amount of F-actin at the synapse. d, A heat map of RBPJ peaks of CD49a+ and CD49a− NK cells from human HCC tissues using CUT&RUN (each n = 3). e, A Venn diagram showing genes with upregulated expression, open chromatin and RBPJ binding at promoter regions in CD49a+ NK cells. f, The genome track view of the IFNG and COTL1 locus showing ATAC-seq and RBPJ CUT&RUN peaks between CD49a+ and CD49a− NK cells. g, The Pearson correlation between RBPJ and GITR mean fluorescence intensity (MFI) in COTL1high NK cells from seven HCC tumours. The error bands represent the 95% confidence interval for the correlation. h, The CUT&RUN quantitative PCR assay showing the level of RBPJ binding to the IFNG and COTL1 promoter region in f by GITR activation and its control (each n = 3; IFNG promoter, P = 0.0008; COTL1 promoter, P = 0.0059). i, The luciferase reporter assay showing IFNG and COTL1 promoter-mediated expression in 293T cells overexpressing empty vector (oeVec) and RBPJ (oeRBPJ) and the full-length or binding site truncations of the IFNG or COTL1 promoter (each n = 3 times independent experiments, IFNG, basic, P = 0.653572, full length, P = 0.028122, Δbinding sites, P = 0.197995; COTL1, basic, P = 0.181410, full length, P = 0.009578, Δbinding sites, P = 0.487092). j, The real-time OCR between GITR agonist treated COTL1high NK cells and control (n = 3; P = 0.0262). k, The MFI (right) and representative histogram (left) of MitoTracker Green (MG) staining of live in vitro-induced COTL1high NK cells treated with GITR agonist and its control (each n = 4, P = 0.0286). l, The MFI (right) and representative histogram (left) of MitoTracker Deep Red (MDR) staining of live in vitro-induced COTL1high NK cells treated with GITR agonist and its control (each n = 4, P = 0.0286). m, The MFI (right) and representative histogram (left) of pSTAT5 staining of live in vitro-induced COTL1high NK cells treated with GITR agonist and its control (each n = 3, P = 0.0073). FMO, fluorescence minus one. n, CUT&RUN quantitative PCR assay showing the level of STAT5 binding to the DLAT promoter region by GITR activation and its control (each n = 3, P = 0.0110). o, The level of acetyl-CoA after GITR activation was detected by ELISA (each n = 4, P = 0.0286). p, The percentage of IFN-γ+ COTL1high NK cells treated by an acetyl-CoA inhibitor (ACSS2-IN-2) and its control (each n = 4, P = 0.0286). q, The genome track view of the RBPJ promoter locus showing H3K27ac CUT&Tag modification peaks on COTL1high NK cells after GITR activation and its control (each n = 3). r, The p-SHP-1 MFI of COTL1high NK cells treated with rPD-L1 and its control (each n = 3, P = 0.0048). s, The p-STAT5 MFI of COTL1high NK cells cultured in medium alone or treated by recombinant PD-L1, GITRL and SHP-1 inhibitor TPI-1 (each n = 3, medium versus rGITRL, P = 0.0004, rGITRL + rPD-L1, P = 0.4646, rGITRL + rPD-L1 + TPI-1, P = 0.0026). P values were determined by unpaired two-sided Mann–Whitney test in a, j–l, o and p. P values were determined by unpaired two-tailed Student’s t-test in h, i, m, n, r and s. Data are shown as the mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; n.s., not significant. FITC, fluorescein isothiocyanate.
Given that RBPJ can function both dependently and independently of NOTCH signalling36, we assessed the contribution of NOTCH activity to IFN-γ production. The inhibition of NOTCH with a γ-secretase inhibitor did not reduce IFN-γ expression in COTL1high NK cells (Extended Data Fig. 7a), indicating NOTCH-independent RBPJ regulation. We next explored how GITR signalling enhances IFNG expression via RBPJ. Because cellular metabolism critically regulates NK cell activity19, we analysed metabolic pathways and found COTL1high NK cells enriched for oxidative phosphorylation (Extended Data Fig. 7b,c). Indeed, the GITR agonist treatment significantly increased the mitochondrial oxygen consumption rate (OCR) (Fig. 5j and Extended Data Fig. 7d) and improved mitochondrial function, as evidenced by elevated mitochondrial mass, membrane potential and Ca2+ levels (Fig. 5k,l and Extended Data Fig. 7e,f). AUCell analysis further revealed enhanced complex V activity in COTL1high NK cells, suggesting increased ATP synthesis (Extended Data Fig. 7g). The inhibition of complex V with oligomycin or mitochondrial uncoupling by carbonyl cyanide m-chlorophenylhydrazone (CCCP)/carbonyl cyanide 4-(trifluoromethoxy) phenylhydrazone (FCCP) reduced the IFN-γ level of COTL1high NK cells, confirming the dependence of IFN-γ production on mitochondrial respiration (Extended Data Fig. 7h). In parallel, the JAK–STAT pathway was prominently active in COTL1high NK cells, with STAT5 emerging as the most enriched STAT family member (Extended Data Fig. 7i,j). GITR activation selectively increased phosphorylated STAT5 (p-STAT5) but not p-STAT1 or p-STAT3 (Fig. 5m and Extended Data Fig. 7k,l). Given that JAK–STAT5 regulates catabolic processes37,38, we examined downstream mechanisms and found that p-STAT5 translocates to the nucleus and interacts with PDC-E2 (encoded by DLAT), a core component of the pyruvate dehydrogenase complex required for acetyl-CoA production (Fig. 5n). Consistently, GITR stimulation increased intracellular acetyl-CoA levels (Fig. 5o), whereas the inhibition of acetyl-CoA synthesis reduced IFN-γ production (Fig. 5p). As nuclear acetyl-CoA primarily fuels histone acetylation39,40, the cleavage under targets and tagmentation (CUT&Tag) profiling revealed increased H3K27ac at the RBPJ promoter in GITR-activated COTL1high NK cells (Fig. 5q). These findings delineate a coherent signalling cascade in which GITR engagement boosts mitochondrial metabolism and acetyl-CoA production, thereby promoting histone acetylation and RBPJ-mediated transcription of IFNG and COTL1.
Given that PD‑L1/PD‑1 engagement recruits SHP‑1/2 phosphatases to suppress CD8+ T cells by negatively regulating JAK–STAT signalling41,42,43, we investigated whether PD‑L1 similarly inhibits GITR downstream signalling in COTL1high NK cells. The PD-L1/PD-1 axis significantly increased p-SHP-1 levels s in COTL1high NK cells (Fig. 5r). Conversely, SHP-1 inhibition restored STAT5 phosphorylation, indicating that PD-L1/PD-1 suppresses GITR-mediated p-STAT5 through p-SHP-1 activation (Fig. 5s and Extended Data Fig. 7n). In summary, our results indicate that the PD-1/PD-L1 blockade initiates a GITRL–GITR-driven positive feedback loop between COTL1high NK cells and PD-L1+ macrophages, thereby enhancing immunotherapy responses in patients with tsMHC-Iimpaired (Extended Data Fig. 7o).
GITR activation synergizes with PD-1/PD-L1 blockade against tsMHC-Iimpaired HCC
To evaluate whether GITR triggering activates COTL1high NK cells and enhances immunotherapy in Hep-53.4 tumours, mice were treated with the GITR agonist DTA-1, αPD-L1 or their combination (Fig. 6a). The combination therapy was significantly more effective at controlling tumour growth than either monotherapy (Fig. 6b). An analysis of tumour-infiltrating immune cells revealed that the combination therapy synergistically increased the proportions of IFN-γ+CD49a+CD49b− NK cells and IFN-γ+CD8+ T cells, while reducing the frequency and suppressive capacity of regulatory T cells (Treg), without broadly affecting other immune populations (Fig. 6c–e, Extended Data Fig. 8a–f and Supplementary Fig. 5). Furthermore, tumour MHC-I expression was significantly upregulated by the combination treatment (Fig. 6f). Consistent results were obtained in H22 tumour models (Extended Data Fig. 9a–j and Supplementary Fig. 6). To confirm that the role of GITR agonism is dependent on COTL1high NK cells, we treated αPD-L1 or combination with DTA-1 on Zfp683+/+ and Zfp683−/− mice bearing orthotopic Hep-53.4 tumours (Fig. 6g). The anti-tumour efficacy of both αPD-L1 monotherapy and the combination was abrogated in Zfp683−/− mice (Fig. 6h,i). Although COTL1high NK cells express PD-1, αPD-L1 monotherapy only induced a trend towards reduced tumour growth in WT mice but not significant compared with Zfp683−/− mice (Fig. 6i, P = 0.0635). By contrast, the combination therapy potently enhanced CD8+ T cell effector function in WT but not Zfp683−/− mice (Fig. 6j). In addition, the depletion of CD4+ T cells neither enhanced nor impaired the therapeutic efficacy of PD-L1 blockade, indicating that GITR signalling in CD4+ T cells, including Treg and helper T cells, is not a major determinant of treatment response in tsMHC-Iimpaired tumours (Extended Data Fig. 8g). To directly assess the requirement for NK cell-intrinsic GITR signalling in mediating the therapeutic synergy, E4bp4−/− mice (NK-deficient mice) with established Hep-53.4 tumours received a single intratumoural injection of sgTnfrsf18 or sgControl (sgCtrl) NK cells before PD-L1 blockade (Fig. 6k and Extended Data Fig. 9k). A better tumour control was observed in NK-deficient mice receiving sgCtrl NK cells compared with those receiving sgTnfrsf18 NK cells, supporting a requirement for GITR signalling in NK cells for optimal tumour control during PD-L1 blockade (Fig. 6l). To extend these findings to human HCC, we next sought to validate these findings using an ex vivo organ culture (EVOC) system (Fig. 6o). We found that the combination of αPD-L1 and a GITR agonist significantly reduced Ki-67, increased cleaved caspase 3 and upregulated tsMHC-I expression, whereas αPD-L1 alone had no effect (Fig. 6p–s and Supplementary Table 6). These findings suggest that combining GITR agonists with αPD-L1 therapy could overcome immunotherapy resistance in patients with tsMHC-Iimpaired HCC.
a, A schematic illustration showing Hep-53.4-bearing mice treated with DTA-1, αPD-L1, combination or isotype control. b, The representative images and statistical results of whole-liver morphology from Hep-53.4-bearing orthotopic HCC model (each n = 5; isotype versus αPD-L1, P = 0.0079, DTA-1, P = 0.0079, DTA-1 + αPD-L1, P = 0.0079; αPD-L1 versus DTA-1, P = 0.4365, DTA-1 + αPD-L1, P = 0.0238). The dashed white lines mark the tumour boundary. c, The flow cytometry analysis of IFN-γ production in CD49a+CD49b− NK cells (each n = 5; isotype versus αPD-L1, P = 0.0079, DTA-1, P = 0.0079, DTA-1 + αPD-L1, P = 0.0079; αPD-L1 versus DTA-1, P = 0.0556, DTA-1 + αPD-L1, P = 0.0079). d, The flow cytometry analysis of IFN-γ production in CD8+ T cells (each n = 5; isotype versus αPD-L1, P = 0.0079, DTA-1, P = 0.0317, DTA-1 + αPD-L1, P = 0.0079; αPD-L1 versus DTA-1, P = 0.0952, DTA-1 + αPD-L1, P = 0.1508). e, The flow cytometry analysis of Treg frequencies (each n = 5; isotype versus αPD-L1, P = 0.0556, DTA-1, P = 0.0079, DTA-1 + αPD-L1, P = 0.0079; DTA-1 + αPD-L1 versus αPD-L1, P = 0.1508, DTA-1, P = 0.0079). f, The mean fluorescence intensity of MHC-I in Hep-53.4-implanted tumours from WT mice determined by mean values of five random FOVs (each n = 4, isotype versus αPD-L1, P = 0.0286, DTA-1, P = 0.0286, DTA-1 + αPD-L1, P = 0.0286; DTA-1 + αPD-L1 versus αPD-L1, P = 0.0286, DTA-1, P = 0.0286; αPD-L1 versus DTA-1, P = 0.6857). g, A schematic illustration showing Hep-53.4-bearing Zfp683+/+ or Zfp683−/− mice treated with αPD-L1 or combination with DTA-1. h, Representative images of whole-liver morphology from WT or Zfp683−/− mice treated with αPD-L1 or combination with DTA-1. The dashed white lines mark the tumour boundary. i, The statistical results of maximum tumour diameter from WT or Zfp683−/− mice treated with αPD-L1 or combination with DTA-1 (each n = 5, Zfp683+/+–αPD-L1 versus Zfp683+/+–αPD-L1 + DTA-1, P = 0.0476, Zfp683−/−–αPD-L1, P = 0.0635; Zfp683−/−–αPD-L1 + DTA-1 versus Zfp683−/−–αPD-L1, P = 0.8492, Zfp683+/+–αPD-L1 + DTA-1, P = 0.0079). j, The representative images and statistical results showing the average number of CD8+Granzyme B+ cells of five FOVs (500 μm × 500 μm) in Hep-53.4 hepatoma (each n = 4, each mouse five FOVs; Zfp683+/+–αPD-L1 versus Zfp683+/+–αPD-L1 + DTA-1, P = 0.0286, Zfp683−/−–αPD-L1, P = 0.0286; Zfp683−/−–αPD-L1 + DTA-1 versus Zfp683−/−–αPD-L1, P = 0.4857, Zfp683+/+–αPD-L1 + DTA-1, P = 0.0286). k, A schematic illustration showing NK-deficient mice receiving intratumoural injection of sgCtrl or sgTnfrsf18 NK cells. l, Representative images and statistical results of whole-liver morphology from NK-deficient mice receiving an intratumoural injection of sgCtrl or sgTnfrsf18 NK cells (each n = 6; P = 0.0043). Scale bar, 1 cm. The dashed white lines mark the tumour boundary. m, The tumour tissues from five patients with MHC-Iimpaired HCC were collected and rapidly dissected into clusters that were treated with or without autologous PBMC-induced COTL1high NK cells plus atezolizumab or ragifilimab for 3 days (Methods). n, The representative immunostaining of EVOCs using cleaved caspase 3 and Ki-67 staining was performed. Scale bars, 100 μm. o, The quantitation results are shown (n = 4 FOVs for each five patients; Ki-67, NC versus αPDL1 + COTL1high NK, P = 0.4291, GITR agonist + COTL1high NK, P = 0.0024, αPDL1 + GITR agonist + COTL1high NK, P < 0.0001; cleaved caspase 3, NC versus αPDL1 + COTL1high NK, P = 0.4020, GITR agonist + COTL1high NK, P = 0.4612, αPDL1 + GITR agonist + COTL1high NK, P = 0.0026). p, The representative immunostaining of EVOCs using Pan-CK and HLA-ABC staining was performed. Scale bars, 150 μm. The quantitation results are shown (each n = 5, NC versus αPDL1 + GITR agonist + COTL1high NK, P = 0.0079). P values were determined by unpaired two-sided Mann–Whitney test in c–f, i, j, l, o and p. Data are shown as the mean ± s.e.m. In the box plots, the centre line is the median, box limits are the first and third quartiles and whiskers are 1.5× interquartile range. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; n.s., not significant. Ab, antibody. NC, negative control. Icons in m created in BioRender; You, W. https://biorender.com/7uda7v9 (2026).
COTL1high NK cells signature predicts better survival and ICB response in several tsMHC-Iimpaired solid tumours
Our data so far indicate a high degree of correlation between COTL1high NK cells and ICB efficiency. COTL1high NK cell-like phenotypes are present in various solid tumours, including lung adenocarcinoma (LUAD), intrahepatic cholangiocarcinoma and head and neck squamous cell carcinoma (Fig. 7a–c). The infiltration of this type of NK cells is associated with favourable survival in most solid tumours (Fig. 7d). To explore the relationship between COTL1high NK cells and ICB responses in other MHC-Iimpaired tumours, we analysed patients with LUAD from the the OAK and POPLAR trials (n = 196)44 and patients with metastatic melanoma from PRJEB23709 (n = 72) (Extended Data Fig. 10a, b). In patients with tsMHC-Iimpaired, responders had significantly higher COTL1high signature scores (Fig. 7e,f). In an independent spatial transcriptomics cohort of nasopharyngeal carcinoma patients treated with αPD-L1, we observed the notable infiltration of COTL1high NK cells and spatial co-occurrence with GITRL+ macrophages in responders with impaired MHC-I expression but not in non-responders (Fig. 7g and Extended Data Fig. 10c). To extend our analysis at single-cell level, we incorporated public scRNA-seq data from ICB-treated patients with non-small-cell lung cancer (NSCLC), colorectal cancer (CRC), clear cell renal cell carcinoma (ccRCC) and Merkel cell carcinoma (MCC) (Fig. 7h). We also found a subset of patients with tsMHC-I+ <80% respond to ICB therapy. The further analysis of the NK cell populations of these patients showed that COTL1high NK cells are enriched in responders (Fig. 7i). These data underscore the critical role of GITR–GITRL interaction in promoting effective anti-tumour responses and sustaining ICB efficacy in MHC-Iimpaired tumours (Extended Data Fig. 10d).
a, An integrated analysis of NK cells from scRNA-seq data among six cancer types. b, The COTL1high NK cell score of COTL1high NK cell signatures in NK cells from six cancer types. The dashed black lines mark the COTL1high NK subset. c, A pie plot showing the distribution of COTL1high NK cells among six cancer types. d, The survival analysis of COTL1high NK cell signature in TCGA database. e, A box plot showing the COTL1high NK cell signature score in CD8+ Tlow/MHC-Iimpaired patients with LUAD with response or no response to atezolizumab (αPD-L1) treatment in EGAS00001005013 (n = 124, P = 0.02). f, A box plot showing the COTL1high NK cell signature score in patients with CD8+ Tlow/MHC-Iimpaired metastatic melanoma with response or no response to nivolumab or pembrolizumab (αPD-1) treatment (n = 44, P = 0.01). g, The tumour-regional MHC-I expression and COTL1high NK cell positive spot assessed by COTL1high NK cell signature score in responders and non-responders from patients with nasopharyngeal carcinoma who received toripalimab (αPD-1) treatment (GSE206245). h, The total scRNA-seq of tumours treated with ICB from four cancer types were collected. NSCLC data were downloaded from SRP485503; MCC data were downloaded from GSE235093; CRC data were downloaded from GSE205506; and ccRCC data were downloaded from SRZ190804. i, The tumour MHC-I analysis and NK cell clustering showed an enrichment of COTL1high NK cells in responders with impaired tsMHC-I. High relative cell density is shown as bright magma. P values were determined by unpaired two-sided Mann–Whitney test in e and f. In the box plots, the centre line is the median, box limits are the first and third quartiles and whiskers are 1.5× interquartile range. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. HNSCC, head and neck squamous cell carcinoma; ICCA, intrahepatic cholangiocarcinoma; RCC, renal cell carcinoma; GC, gastric cancer; Tfh, T follicular helper cells. Icons in h created in BioRender; You, W. https://biorender.com/7uda7v9 (2026).
Discussion
Immunotherapy, particularly αPD-1/PD-L1 therapy, has transformed cancer treatment, yet its efficacy is limited in tumours with impaired tsMHC-I expression9,45. Recent studies have linked tsMHC-I loss to immunotherapy resistance, yet a subset of patients with tsMHC-Iimpaired tumours still respond to ICB46,47. The underlying mechanisms enabling immunotherapy responsiveness in these tsMHC-Iimpaired tumours remain poorly understood. Here, through single-cell and spatial analysis, we identify COTL1high NK cells as a key cellular correlate of ICB responsiveness in tsMHC-Iimpaired tumours. These NK cells, characterized by high IFN-γ production and PD-1 expression, compensate for defective antigen presentation by sustaining IFN-γ production and promoting secondary CD8+ T cell activation. The disruption of the PD-1/PD-L1 axis releases GITRL-mediated costimulatory signalling from PD-L1+ macrophages, establishing a self-reinforcing circuit that maintains NK cell effector function. In contrast to mismatch repair deficient colon tumours, where γδ T cells contribute to ICB responses11,48, their numbers and functions showed no differences between responders and non-responders, suggesting γδ T cells do not dominantly mediate response to immunotherapy in tsMHC-Iimpaired HCC. Thus, boosting COTL1high NK cells may benefit these patients. Although CD49a-expressing NK subsets, a defining feature of COTL1high NK cells, have been reported in various solid tumours, their functional role in anti‑tumour immunity remains context‑dependent49,50,51. In several settings, CD49a+ NK cells exhibit tissue-resident characteristics with limited cytotoxicity, whereas in others, they retain robust effector functions. Our data reveal that in tsMHC-Iimpaired HCC, CD49a marks a functionally specialized NK cell subset characterized by high COTL1 expression, preserved effector capacity and robust IFN-γ production. To interrogate the molecular mechanisms sustaining this functional state in vivo, we sought a genetic model that selectively targets the COTL1high NK cells. Considering that COTL1 is highly, but not uniquely, expressed in this NK subset and that we did not identify a marker gene that unequivocally distinguishes these cells from all other immune populations, in vivo antibody-based depletion or NK cell-specific gene knockout strategies were not feasible. Cross-species single-cell mapping and chromatin accessibility profiling show that human COTL1high NK cells correspond to the Cd49a+Cd49b− liver-resident NK subset in mice, supporting their evolutionary conservation and enabling genetically tractable mechanistic studies of their functional maintenance. Zfp683 (also known as Hobit) is a key transcription factor controlling the development, tissue residency and functional adaptation of Cd49a+Cd49b− NK cells and CD8+ Trm cells32. However, Zfp683 knockout did not significantly alter the abundance of CD8+ Trm cells in our models. Importantly, Zfp683−/− mice lost the therapeutic benefit of PD-L1 blockade, either alone or in combination with GITR agonism, establishing a causal requirement for this NK subset.
A central insight from our ligand–receptor, spatial and functional analysis is that PD-L1+ macrophages, not DCs or other myeloid populations, constitute the dominant and therapeutically relevant source of GITRL engaging COTL1high NK cells. In HCC, PD-L1 is preferentially expressed by macrophages34, whereas tsMHC-Iimpaired tumour cells upregulate IGSF8, a checkpoint molecule that directly suppresses NK cell cytotoxicity52. Thus, although PD-L1+ macrophages provide the essential GITRL signal to activate COTL1high NK cells, tumour cells themselves deploy a parallel inhibitory mechanism via IGSF8 to counteract NK activation. Furthermore, αPD-1/PD-L1 therapy amplifies GITRL–GITR interactions by stabilizing ISs, thereby reinforcing NK cell activation. Tumour-derived metabolic suppression may counteract this effect53, suggesting that strategies stabilizing immune synapses or targeting tumour metabolism could further enhance ICB efficacy.
Importantly, GITR activation has been shown to reduce Treg number and suppressive function, highlighting its dual roles54. However, our genetic and functional data demonstrate that in tsMHC-Iimpaired HCC, the therapeutic benefit of GITR agonism critically depends on COTL1high NK cells rather than on Treg or other GITR+ populations. Consistently, PD-L1+ macrophages expressing GITRL adopt immunostimulatory phenotypes in HCC and other tumours55, supporting a model in which GITRL signalling from macrophages sustains NK cell effector function.
Mechanistically, COTL1high NK cells displayed the enhanced activation of STAT5 and RBPJ programmes downstream of GITR signalling. In the hypoxic TME, STAT5-driven metabolic fitness supports oxidative phosphorylation-dependent NK survival and cytotoxicity56,57. PD-L1/PD-1 blockade both amplifies macrophage-derived GITRL costimulation and relieves PD-1/SHP-1 mediated inhibition of STAT5, creating a dual-signal amplification loop. This metabolic–epigenetic regulation explains the heightened sensitivity of COTL1high NK cells to GITR agonism and the pronounced synergy between PD-L1 blockade and GITR activation in tsMHC-Iimpaired tumours.
Collectively, our study uncovers a previously unappreciated mechanism by which COTL1high NK cells orchestrate ICB responses in tsMHC-Iimpaired cancers through PD-L1+ macrophage-derived GITRL signalling. This axis not only explains why a subset of patients responds to ICB despite impaired antigen presentation but also provides a rationale for combining GITR agonists with PD-1/PD-L1 blockade to rescue anti-tumour immunity in otherwise resistant settings. Targeting this pathway may therefore expand the therapeutic benefit of immunotherapy in tsMHC-Iimpaired malignancies.
Methods
This study was conducted in full compliance with the applicable ethical guidelines. The HCC specimens were obtained from the First Affiliated Hospital of Nanjing Medical University (JSPH), the Affiliated Wuxi People’s Hospital of Nanjing Medical University (WXPH) and the Renji Hospital of Shanghai Jiaotong University School of Medicine (RJH). All patients were informed of the study and provided informed consent before specimen collection. All experiments were performed in compliance with government policies and the Declaration of Helsinki. The study was approved by the ethics committee of JSPH (2023-SR-191), the ethics committee of WXPH (KY22105) and the ethics committee of the RJH (KY2025-091-C). The detailed clinical and pathological information of the patients is presented in Supplementary Tables 1, 4 and 6. All animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC) at Nanjing Medical University in accordance with NIH and institutional guidelines (IACUC-2201004).
Animal experiments
The Zfp683−/− and NK-deficient (E4bp4−/−) mice were obtained from the Shanghai Model Organisms Center. WT male BALB/c or C57BL/6 mice (6–8 weeks old) were obtained from the Animal Core Facility of Nanjing Medical University. To induce mouse hepatoma models, 1 × 106 H22 (China Centre for Type Culture Collection, GDC0091), Hep-53.4 (CLS-Cell Lines Service, CVCL_5765) or Hepa1-6 (American Type Culture Collection, CRL-1830) tumour cells were inoculated into the liver of mice. All mice were maintained under specific pathogen-free conditions in the animal facilities of Nanjing Medical University. In several cases, the C57BL/6 mice were administrated with diethylnitrosamine and CCl4 to induce spontaneous hepatoma. The maximal tumour burden permitted by IACUC did not exceed 2,000-mm3 volume or 2-cm diameter in all animal experiments.
Tumour dissociation and single-cell library preparation
The liver tissues were surgically removed and kept in MACS Tissue Storage Solution (Miltenyi Biotec) until processing. The tissue samples were processed as described below. In brief, samples were first washed with phosphate-buffered saline (PBS), minced into small pieces (approximately 1 mm3) on ice and enzymatically digested with 50 U ml−1 collagenase I (Worthington) and 30 U ml−1 DNase I (Worthington) for 45 min at 37°C, with agitation. After digestion, samples were sieved through a 70-µm cell strainer and centrifuged at 300g for 5 min. After the supernatant was removed, the pelleted cells were suspended in red blood cell lysis buffer (Miltenyi Biotec) to lyse red blood cells. After washing with PBS containing 0.04% BSA, the cell pellets were resuspended in PBS containing 0.04% BSA and refiltered through a 35-μm cell strainer. Dissociated single cells were then stained for viability assessment using Calcein AM (Thermo Fisher Scientific) and Draq7 (BD Biosciences).
The BD Rhapsody system was used to profile single-cell transcriptomes. Single cells were randomly distributed across >200,000 microwells via limited dilution. Barcoded beads were added to saturation, ensuring one bead per microwell. After cell lysis, polyadenylated RNA transcripts hybridized to the beads, which were then collected for reverse transcription. During complementary DNA (cDNA) synthesis, each molecule was tagged at the 5′ end with a unique molecular identifier (UMI) and a cell barcode indicating its origin. Whole-transcriptome amplification was performed following the BD Rhapsody workflow: second-strand synthesis, WTA adaptor ligation and 18 cycles of PCR. Sequencing libraries were prepared by random priming PCR to enrich the 3′ ends of transcripts containing cell labels and UMIs. Libraries were quantified using a High-Sensitivity DNA chip (Agilent) on a Bioanalyzer 2200 and the Qubit High-Sensitivity DNA assay. Each sample was sequenced on an Illumina platform with 150 bp paired-end reads.
TEM
NK cells were sorted by flow cytometry and fixed with 2.5% glutaraldehyde for 48 h, rinsed with phosphate buffer and postfixed with 1% osmium tetroxide. Then, the samples were rinsed and dehydrated in ethanol and acetone. Finally, the samples were embedded in Spurr resin. Ultrathin sections were then stained with 2% uranyl acetate and lead citrate and visualized using an FEI Tecnai G2 Spirit Bio TWIN electron microscope.
Mass cytometry (CyTOF) staining
Antibodies used for CyTOF analysis are listed in Supplementary Table 3. In brief, cells were washed once with 1× PBS, then stained with 100 μl of 250 nM cisplatin (Fluidigm) for 5 min on ice to exclude dead cells and then incubated in Fc receptor blocking solution before being stained with the surface antibodies cocktail for 30 min on ice. Cells were washed twice with fluorescence-activated cell sorting (FACS) buffer (1× PBS + 0.5% BSA) and fixed in 200 μl of intercalation solution (Maxpar Fix and Perm Buffer containing 250 nM 191/193Ir, Fluidigm) overnight. After fixation, cells were washed once with FACS buffer and then perm buffer (eBioscience) and stained with the intracellular antibodies cocktail for 30 min on ice. Cells were washed and resuspended with deionized water, adding into 20% EQ beads (Fluidigm), acquired on a mass cytometer (Helios, Fluidigm).
Antibody staining and flow cytometry analysis
Following tumour excision or biopsy, a representative tumour fragment that was fresh and sterile was transferred to the laboratory for study. All surrounding macroscopic tissue, including any obvious tumour capsule, was removed from the tumour. Every sample was cut into small pieces (<1 mm in diameter) and then was dissociated through gentleMACS (programme: E_01) with Roswell Park Memorial Institute (RPMI) 1640 medium. Digested tumour pieces were teased through a 70-µm strainer and diluted. After centrifugation, the cell pellet was resuspended in a 40% Percoll solution (170891, GE Healthcare), and a phase of 80% Percoll was underlaid using a glass Pasteur pipette. The resulting gradient was centrifuged at 2,000 rpm for 30 min at room temperature without brakes. After the removal of the red blood cell-containing pellet on the bottom and excess buffer-containing cellular debris on the top, the cell population at the Percoll interphase enriched for TILs was washed twice.
Homogenized cells were labelled with monoclonal antibodies for 30 min at 4 °C in FACS buffer (2% FBS in Dulbecco’s PBS), washed twice in FACS buffer, then fixed in IC Fixation buffer (00-8222-49, Invitrogen) in Dulbecco’s PBS. Viable cells were revealed using the Zombie NIRTM Fixable Viability Dye (BioLegend). Multiparameter FACS data were acquired on the BD LSR Fortessa X-20 FACS instrument (BD Biosciences), and data were analysed using FlowJo software (version 10, Treestar). The antibodies used in this Article are listed in Supplementary Table 7.
scRNA-seq raw data processing and analysis
We used the BD Rhapsody WTA analysis pipeline (v1.9.1) to process raw FASTQ files, including filtering, alignment and cell calling. R1 reads were used to identify cell label sequences and UMIs. R2 reads were mapped to reference to determine mRNA origin. The reference genome GRCh38 was built using STAR software (version 2.5.2b). Further adjustment was performed with the recursive substitution error correction algorithm to remove artefact molecules owing to amplification bias in the molecule annotation step. Recursive substitution error correction counts were subsequently used for downstream analysis.
The raw count files were imported into R Statistical Software Environment (v4.1.0). The cells were removed that had either more than 2,000 expressed genes or more than 50% mitochondrial UMIs were removed. A deep neural network-based tool called Single-Cell Variational Inference was used to normalize, scale, train and remove data batch effects. The Louvain modularity optimization algorithm built-in Seurat (v3.2.3) was applied to iteratively group cells together into clusters. Cell clusters were annotated using highly expressed genes. Cells expressing IL7R or CD3 that were considered as innate lymphoid cells or T cells were removed before clustering analysis.
Analysis of scPLC dataset from Xue et al.
Raw fastq files were downloaded from HRA001748 (https://ngdc.cncb.ac.cn/gsa-human/browse/HRA001748). ScRNA-seq data were aligned to human reference genome GRCh38 and quantified using the CellRanger toolkit (v7.0.0). Empty droplets and doublets were removed using dropletUtils (v1.28.0) and scDblFinder (v1.21.2) R package. Cells with more than 50% of the percentage of mitochondrial genes and expressing more than 6,000 genes were removed before downstream analysis. Single-Cell Variational Inference was also used to normalize, scale, train and remove data batch effects. NK cells were annotated on the basis of the well-known genes, including NCAM1, NKG7, NCR1, NCR3 and CD7. Next, we performed a second round of clustering to further characterize subpopulations of NK cells.
Cell–cell communication analysis
To investigate cell–cell interactions among NK cells and macrophages, we analysed the ligand–receptor pairs using CellphoneDB (v.3.1.0), a public repository used to identify significant ligand–receptor interactions58. The interaction strength refers to the number of significant pairs between different cell types.
Spatial transcriptomics datasets analysis
The Stereo-seq data in Li’s research59 is available at the Figshare database (https://doi.org/10.6084/m9.figshare.22332655). We downloaded the annotated Seurat object file and visualized the NK cell in the spatial locations. For 10x Visium data, the processed data were downloaded from GSE238264, GSE206245, Mendeley Database (https://data.mendeley.com/datasets/skrx2fz79n/1) and http://lifeome.net/supp/livercancer-st/data.htm. To identify COTL1high NK-positive spots, AddMoculeScore function from Seurat was used to quantify signature gene score. The spots with a score more than the median value of all spots from one slice was considered as COTL1high NK-positive spots.
To define the tsMHC-Iimpaired at the spatial transcriptomics (ST) level, we used a comprehensive method. In brief, we utilized Cancer-Finder software60 to identify malignant spots. Subsequently, the MHC-I gene score was computed using the AddModuleScore function in the Seurat package. Tumour spots with an MHC-I score greater than 0 were classified as MHC-I+ spots. Finally, we calculated the MHC-I+ rate in tumour regions.
Single-cell regulatory network inference and clustering
Gene regulatory networks for six distinct NK cell groups were identified using pySCENIC (v0.11.2) GRNBoost2 mode61. The potential direct-binding targets (regulons) specificity score was calculated to find the regulators for NK cell groups.
Pathway signalling signatures analysis
We performed pathway analysis on NK subsets using PROGENy62. In brief, scores were computed using the run_aucell function from the decoupleR package (v2.2.2) based on PROGENy network model. The top 1,000 target genes of the progeny model were used, as recommended for single-cell data.
Bulk RNA-seq data analysis
The raw reads were aligned to GRCh38 genome using hisat2 software. FeatureCounts was used to determine the read counts per gene. Transcripts per million was calculated by the rnanorm python package. The immune cells score was estimated on the basis of transcripts per million values by the ssGSEA algorithm in the GSVA R package. The fraction of MHC-I+ and MHC-I− tumour cells was estimated by Bisque R package. In brief, MHC-I+ and MHC-I− tumour cells were identified in the scPLC (single-cell data of primary liver cancer) dataset. Then, we utilized scPLC data as a reference to deconvolute the relative percentage of MHC-I+ and MHC-I− tumour cells in bulk RNA-seq data.
Bulk ATAC-seq data analysis
The raw bulk assay for transposase-accessible chromatin using sequencing (ATAC-seq) data of CD49a+ and CD49a− NK cells from human liver were downloaded from SRP28522763. The raw bulk ATAC-seq data of CD49a+ and CD49a− NK cells from murine liver were downloaded from GSE19671664. Fastq-dump software (v3.0.0) was used to convert the sra-format file to raw fastq files. Next, we utilized the pipeline from Cebola Lab (https://github.com/CebolaLab/ATAC-seq) to analyse the ATAC-seq data. In brief, we first aligned the data to human GRCh38 or mouse GRCm38 reference genome using Bowtie2 (v2.4.1) software. ATAC-seq experiments commonly include a high proportion of mitochondrial reads. Then, mitochondrial reads and duplicates or low-quality alignments were removed from the alignments using MarkDuplicates from PicardTools (v.2.23.3). We also removed the ENCODE blacklist regions in the human or mouse before further analysis. Peaks were identified using MACS2 software (v2.2.7). Finally, normalized BigWig files were generated by DeepTools (v.3.5.1) and merged for visualization by transPlotR package (v0.0.4).
CUT&RUN or CUT&Tag data analysis
CUT&RUN or CUT&Tag sequencing data were analysed as described previously65, following the standard pipeline from Henikoff lab. In brief, paired-end 150-bp reads were aligned to GRCh38 human genome using Bowtie2 (v2.4.1) with the following options: --end-to-end --very-sensitive --no-mixed --no-discordant --phred33. Technical replicates were used to increase the power of peak calling. The duplicates or low-quality alignments were removed from the alignments using MarkDuplicates from PicardTools (v.2.23.3). HOMER software (v4.11.1) was used to call peaks from bam files and calculated the differential peaks between CD49a+ and CD49a− NK cells. The heat maps showing peak distribution for each datum of IgG and RBPJ were generated using the deepTools function ‘plotHeatmap’.
Gene enrichment analysis
To identify the biological function(s) of the DEGs in each cluster, we performed Gene Ontology enrichment analysis using ClusterProfiler R package (v3.18).
CyTOF data analysis
CD45+ immune cells were gated and exported from the generated.fcs files. The processed.fcs files were imported into R environment as the SingleCellExperiment object using flowCore package (v2.10.0). The R package cytofworkflow was used to complete quality control, clustering, cell annotation and visualization. We applied two runs to get final NK groups. Specifically, in the first run, we clustered all CD45+ immune cells on the basis of the expression of cell-specific lineage markers. In the second run, CD45+CD56+CD3−CD19−CD14−CD127− cell clusters were further clustered using NK-associated markers (SOM, Self-Organizing Map, 100; maxK, 35).
Luciferase reporter assay
For the luciferase reporter assay, the full-length promoters of IFNG and COTL1, or their key truncated regions containing RBPJ-binding sites, were cloned into the pGL3 vector. The cDNA encoding the transcription factor RBPJ was constructed into the pcDNA4 plasmid for overexpression. T 293T cells were seeded in 24-well plates 1 day before transfection. For each well, 100 ng of pGL3 reporter plasmid, 1 ng of pRL-TK internal control plasmid and 500 ng of pcDNA4-RBPJ overexpression plasmid were transfected. At 24 h after transfection, cells were transferred to 96-well assay plates at a density of 10,000 cells per well. After cell attachment, luminescence signals were measured using a Luciferase Assay Kit (Vazyme, DL101-01) according to the manufacturer’s instructions.
Single-cell mapping analysis
We applied Symphony (v0.1.1) to map NK cells from Xue et al. to our dataset. For mapping, we used the reference built by our scRNA-seq data66. The mapQuery function was performed using the gene expression data and cell metadata. We further applied the knnPredict (k = 30) function to transfer the cell types from the reference dataset to the clusters identified in our data.
For the similar analysis of human and murine NK cells, cell label transfer was performed using Seurat FindTransferAnchors() and MapQuery() functions. A reference Uniform Manifold Approximation and Projection (UMAP) was generated from our dataset by setting return.model = TRUE and rerunning RunUMAP(). Transfer anchors between both datasets were computed and projected onto the reference UMAP via the MapQuery() function.
Inducing COTL1high NK cells in vitro
As previously described, using cytokines TGF-β1 and IL-15, peripheral NK cells could be induced to be a COTL1high phenotype shared with hepatic COTL1high NK cells67,68. In brief, NK cells were firstly purified from peripheral blood mononuclear cells (PBMC) via negative selection with a MojoSortTM Human NK Cell Isolation Kit (BioLegend). The NK cells were cultured in RPMI 1640 media supplemented with recombinant human TGF-β1 (5 ng ml−1), IL-15 (10 ng ml−1) and induced for approximately 2 days at 37 °C in a humid atmosphere with 5% CO2. On day 3, COTL1high NK cells were purified using EasySepTM PE Positive Selection Kit (StemCell), and follow-up assays were collected.
Seahorse metabolic assays
The in vitro-induced COTL1high NK cells from healthy donors’ blood were stimulated with ragifilimab for 2 h. Then, the OCR was measured using the XF96 extracellular flux analyser following the manufacturer’s instructions. In the seahorse assays, the COTL1high NK cells were treated with oligomycin (1 μM), FCCP (1.5 μM), rotenone (0.5 μM) and antimycin A (0.5 μM). Each condition was performed with three replicates.
Immunohistochemistry and mIHC staining and analysis
Paraffin-embedded human or mouse HCC tissues were cut into 5-μm sections to perform immunohistochemistry and immunofluorescence staining. The sections were stained with antibodies against human or mouse PanCK (1:500, Abcam), HLA-ABC (1:400, Abcam), CD8 (1:500, Abcam), CD56 (1:300, Abcam), CD49a (1:4,000, Novus Biologicals), NKp46 (1:200,BioLegend), GITR (1:500, Abcam), CD68 (1:300, Abcam) and CD3 (1:100, Abcam) followed by horseradish peroxidase (HRP)-conjugated anti-rat or anti-rabbit IgG (Thermo Fisher). For immunofluorescence staining, the sections staining was performed on PANO 7-plex IHC kit (Panovue). The slides were detected by digital pathology slide scanner (KF-FL-020, kfbiopathology) and analysed by Qupath or HALO software.
To quantify the tsMHC-I levels in HCC tumours, we initially performed mIHC analysis of Pan-CK and HLA-ABC in a tissue microarray cohort. The results revealed a distinct bimodal distribution of tsMHC-I expression, with a significant patient subset exhibiting high expression levels at 80–95% range. On the basis of this biologically anchored distribution, we established the following classification threshold: tumours demonstrating the positive percentage of HLA-ABC in Pan-CK+ tumour cells <80% were defined as tsMHC-Iimpaired tumours.
Mitochondrial functional and phenotypic characterization
MitoTracker Deep Red (Beyotime, C1034), MitoTracker Green (Beyotime, C1048), TMRE (Beyotime, C2001S) and Fluo-3 (Sigma, 73881-1MG) were suspended with NK cells at room temperature for 30 min and washed using DPBS. Cells were further analysed using flow cytometry (BD LSR Fortessa X-20). Each condition was performed with three to four replicates.
ELISA
Concentrations of acetyl-CoA from the lysate of COTL1high NK cells were detected using enzyme-linked immunosorbent assay (ELISA) kits according to the manufacturer’s instructions (Byabscience).
EVOC experiment
Human HCC tumours were washed with cold PBS and cutted into fragments of about 750-μm diameter in ice-cold RPMI 1640 supplemented with 8% foetal calf serum, followed by washing and resuspension in RPMI 1640 supplemented with 8% foetal bovine serum, 2% autologous human serum, IL2 (300 IU ml−1), 1% transferrin–insulin–selenium mix (Gibco), 1% glutamine (Gibco), 1% penicillin–streptomycin (Gibco), 0.5% HEPES (Gibco), 0.6% nonessential amino acids (Gibco), 2.5 μg ml−1 amphotericin (Gibco) and 50 μg ml−1 gentamycin (Gibco). The resuspended EVOC was divided into 500 μl well−1 in 24-well plates and cultured at 37 °C and 5% CO2. After culturing for 8 h for EVOC initiation, atezolizumab (αPD-L1, STARTER) and ragifilimab (GITR agonist, Selleck) was added at a finial concentration of 35 μg ml−1 for additional culturing for 3 days.
CRISPR–Cas9-knockout of Ifngr1 and B2m
The CRISPR–Cas9 system was used for genetic knockouts. CRISPR guide RNA were designed to target two exons in GRCm38 genome of Ifngr1 (g1: TGGAGCTTTGACGAGCACTG, g2: AGCTGGCAGGATGATTCTGC) and B2m (g1: GACAAGCACCAGAAAGACCA, g2: GGATTTCAATGTGAGGCGGG). For lentivirus construction, the Cas9 gene cDNA cloned by PCR was inserted into EF1-CAS9-P2A-PUR lentiviral vectors (Hanbio). The recombinant lentivirus with Cas9 gene coding sequence was produced by cotransfection of 293T cells with plasmids PSPAX2 and PMD2G with LipoFiterTM (Hanbio). Lentivirus-containing supernatant were collected 48 h after transfection and filtered through 0.22-μm cellulose acetate filters (Millipore). Recombinant lentiviruses were concentrated by ultracentrifugation (2 h at 50,000g). To establish stable Cas9 gene-overexpressing cell lines, Hep-53.4 cells were transduced with lentiviral vector (HBLV-cas9-PURO) at a multiplicity of infection of approximately five in the presence of 5 μg ml−1 polybrene. After 24 h, culture medium was removed and fresh medium was added to the tumour cells (Hep-53.4 or Hepa1-6). At 72 h after cell transduction, 5 μg ml−1 puromycin (Hanbio) was added to the culture medium to screen for cell lines stably transduced with the Cas9 gene.
Then, Cas9 cells were transduced with lentiviral vector at a multiplicity of infection of approximately five in the presence of 5 μg ml−1 polybrene. After 24 h, culture medium was removed, and fresh medium was added to the Cas9 cells. Hep-53.4 cells incubated with Ifngr1-directed guides were stained with anti-mouse Ifngr1. Hepa1-6 cells incubated with B2m-directed guides were stained with anti-mouse H2-Kb. Ifngr1/H2-Kb-negative and GFP-positive cells were sorted into single cells using BD FACS Vantage SE (BD Biosciences) and cultured for 2–3 weeks.
Construction of B2m-overexpressed cell lines
B2m-overexpressing lentiviruses (Hanbio) were transduced into Hep-53.4 cells at 40–50% confluency. The viral particles and transfection reagent were diluted in medium, incubated for 5 min, combined and incubated for a further 20 min before addition to target cells. After 48 h, the viral supernatant was replaced with fresh medium. Stable lines were selected with 3 μg ml−1 puromycin (48–72 h).
CRISPR gene editing in mouse primary NK cells
CRISPR–Cas9-mediated gene editing targeting GITR (sgTnfrsf18: CAACTACACTCGGCTGACCT) and the control locus AAVS1 was performed by electroporation of the Cas9–gRNA ribonucleoprotein (RNP) complexes. RNPs were assembled by incubating recombinant Cas9 protein with synthetic sgRNAs at room temperature before electroporation. For each reaction, 5 × 10⁵ mouse primary NK cells were resuspended in 9 μl of Buffer R (Thermo Fisher Scientific, Neon Transfection System) and mixed with 1 μl of the preassembled RNP complex. The resulting 10 μl cell–RNP mixture was loaded into a Neon Pipette tip without introducing air bubbles and electroporated using the following parameters: 1,700 V, 10 ms and a single pulse. Cells were immediately transferred into prewarmed complete culture medium (IMMOCELL, IMP-M204-2) for recovery and downstream analyses.
In vivo immune cell depletion and blockade using antibodies
NK cell and CD8+ T cell depletion was performed by the intraperitoneal injection of anti-Asialo-GM1 antibody (FUJIFILM Wako, 986-10001, 20 μl each for a single dose per mouse) and anti-CD8a antibody (BioXCell, BE0004-1, 100 μg each for a single dose per mouse) 24 h before tumour challenge, and after the challenge, the antibody was injected every 3 days. For in vivo treatments, mice were treated every 3 days by intraperitoneal injection of 100 μg anti-GITR agonist DTA-1 (BioXCell, BE0063), αPD-L1 (BioXCell, BE0101) and IgG control.
Statistics and reproducibility
Statistical analysis was performed using R (v4.1.0), Qupath (v0.5.0) and GraphPad Prism (v10) software. We used the Kruskal–Wallis or one-way analysis of variance test to assess overall differences among multiple groups. Then, Mann–Whitney U exact or approximate test or Student’s t-test was used for pairwise comparisons. All P values reported were two-sided and statistical significance was defined as P less than 0.05. The univariate and multivariate analyses were performed with the Cox proportional hazards model. No statistical methods were used to predetermine sample sizes. All collected data were included in subsequent analyses without exclusion. For both in vitro and in vivo experimental paradigms, sample or animal allocation to treatment groups used simple randomization protocols. Researchers were not blinded to group allocation during experimental procedures or the outcome evaluation.
Reporting summary
Further information on the research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The scRNA-seq and CUT&RUN data that support the findings of this study have been deposited in the Genome Sequence Archive for Human database under accession number HRA010073. The human sequencing data are available under restricted access because of data privacy and supervision. Researchers can send data requests for scientific purposes via the website (https://ngdc.cncb.ac.cn/gsa-human/) after registration. Data access requests will be processed within 1 month by following the guidelines for Genome Sequence Archive for noncommercial use. Once access has been approved, the data will be available for 1 month. Previously published data that were re-analysed here are available under accession code GSE238264, GSE206245, GSE196716, GSE158485, GSE235863, scPLC (GSA for human: https://ngdc.cncb.ac.cn/gsa-human/browse/HRA001748), EGAD00001008128, EGAD00001010132, EGAS00001005013, PRJEB23709, skrx2fz79n, SRP485503, GSE235093, SRZ190804, GSE205506, GSE206245 and GSE278687. The human pancancer RNA-seq data were derived via the TCGA Research Network at http://cancergenome.nih.gov/. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Code availability
The R or python code used for analysis in this study is publicly available via GitHub at https://github.com/youwh1996/HCC_NK_PAPER.
References
Kim, H. D. et al. Regorafenib plus nivolumab in unresectable hepatocellular carcinoma: the phase 2 RENOBATE trial. Nat. Med. 30, 699–707 (2024).
Li, C. et al. Pathologic response of phase III study: perioperative camrelizumab plus rivoceranib and chemotherapy versus chemotherapy for locally advanced gastric cancer (DRAGON IV/CAP 05). J. Clin. Oncol. 43, 464–474 (2025).
Fukuoka, S. et al. Regorafenib plus nivolumab in patients with advanced gastric or colorectal cancer: an open-label, dose-escalation, and dose-expansion phase Ib trial (REGONIVO, EPOC1603). J. Clin. Oncol. 38, 2053–2061 (2020).
D’Alessio, A. et al. Pathological response following neoadjuvant immune checkpoint inhibitors in patients with hepatocellular carcinoma: a cross-trial, patient-level analysis. Lancet Oncol. 25, 1465–1475 (2024).
Valero, C. et al. Clinical-genomic determinants of immune checkpoint blockade response in head and neck squamous cell carcinoma. J. Clin. Invest. 133, e169823 (2023).
Dammeijer, F. et al. The PD-1/PD-L1-checkpoint restrains T cell immunity in tumor-draining lymph nodes. Cancer Cell 38, 685–700 (2020).
Hashimoto, M. et al. PD-1 combination therapy with IL-2 modifies CD8+ T cell exhaustion program. Nature 610, 173–181 (2022).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Gu, S. S. et al. Therapeutically increasing MHC-I expression potentiates immune checkpoint blockade. Cancer Discov. 11, 1524–1541 (2021).
Chen, X. et al. A membrane-associated MHC-I inhibitory axis for cancer immune evasion. Cell 186, 3903–3920 (2023).
de Vries, N. L. et al. γδ T cells are effectors of immunotherapy in cancers with HLA class I defects. Nature 613, 743–750 (2023).
Chen, L. et al. Deep whole-genome analysis of 494 hepatocellular carcinomas. Nature 627, 586–593 (2024).
Pyke, R. M. et al. A machine learning algorithm with subclonal sensitivity reveals widespread pan-cancer human leukocyte antigen loss of heterozygosity. Nat. Commun. 13, 1925 (2022).
Teo, J. M. N. et al. Tumor-associated neutrophils attenuate the immunosensitivity of hepatocellular carcinoma. J. Exp. Med. 222, e20241442 (2025).
Li, Z. et al. Neoadjuvant tislelizumab plus stereotactic body radiotherapy and adjuvant tislelizumab in early-stage resectable hepatocellular carcinoma: the Notable-HCC phase 1b trial. Nat. Commun. 15, 3260 (2024).
Benci, J. L. et al. Opposing functions of interferon coordinate adaptive and innate immune responses to cancer immune checkpoint blockade. Cell 178, 933–948 (2019).
Berrien-Elliott, M. M., Jacobs, M. T. & Fehniger, T. A. Allogeneic natural killer cell therapy. Blood 141, 856–868 (2023).
Dubrot, J. et al. In vivo CRISPR screens reveal the landscape of immune evasion pathways across cancer. Nat. Immunol. 23, 1495–1506 (2022).
Zheng, X. et al. Tumors evade immune cytotoxicity by altering the surface topology of NK cells. Nat. Immunol. 24, 802–813 (2023).
Oh, S. C. et al. NgR1 is an NK cell inhibitory receptor that destabilizes the immunological synapse. Nat. Immunol. 24, 463–473 (2023).
Paillon, N. et al. PD-1 inhibits T cell actin remodeling at the immunological synapse independently of its signaling motifs. Sci. Signal. 16, eadh2456 (2023).
Mace, E. M. & Orange, J. S. Lytic immune synapse function requires filamentous actin deconstruction by coronin 1A. Proc. Natl Acad. Sci. USA 111, 6708–6713 (2014).
Kim, J. et al. Coactosin-like 1 antagonizes cofilin to promote lamellipodial protrusion at the immune synapse. PLoS ONE 9, e85090 (2014).
Cappuyns, S. et al. PD-1− CD45RA+ effector-memory CD8 T cells and CXCL10+ macrophages are associated with response to atezolizumab plus bevacizumab in advanced hepatocellular carcinoma. Nat. Commun. 14, 7825 (2023).
Jew, B. et al. Accurate estimation of cell composition in bulk expression through robust integration of single-cell information. Nat. Commun. 11, 1971 (2020).
Tang, F. et al. A pan-cancer single-cell panorama of human natural killer cells. Cell 186, 4235–4251 (2023).
Xu, L. et al. A comprehensive single-cell breast tumor atlas defines epithelial and immune heterogeneity and interactions predicting anti-PD-1 therapy response. Cell Rep. Med. 5, 101511 (2024).
Pouxvielh, K. et al. Tumor-induced natural killer cell dysfunction is a rapid and reversible process uncoupled from the expression of immune checkpoints. Sci. Adv. 10, eadn0164 (2024).
Fu, B. et al. CD11b and CD27 reflect distinct population and functional specialization in human natural killer cells. Immunology 133, 350–359 (2011).
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Weizman, O. E. et al. ILC1 confer early host protection at initial sites of viral infection. Cell 171, 795–808 (2017).
Mackay, L. et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue-residency in lymphocytes. Eur. J. Immunol. 46, 753–753 (2016).
Cappuyns, S. et al. Single-cell RNA sequencing-derived signatures define response patterns to atezolizumab + bevacizumab in advanced hepatocellular carcinoma. J. Hepatol. 82, 1036–1049 (2025).
Lu, L. G. et al. PD-L1 blockade liberates intrinsic antitumourigenic properties of glycolytic macrophages in hepatocellular carcinoma. Gut 71, 2551–2560 (2022).
Croft, M. The role of TNF superfamily members in T-cell function and diseases. Nat. Rev. Immunol. 9, 271–285 (2009).
Friedrich, T. et al. Notch-dependent and -independent functions of transcription factor RBPJ. Nucleic Acids Res. 50, 7925–7937 (2022).
Lin, Y. et al. Pancreatic STAT5 activation promotes KrasG12D-induced and inflammation-induced acinar-to-ductal metaplasia and pancreatic cancer. Gut 73, 1831–1843 (2024).
Jones, N. et al. Akt and STAT5 mediate naive human CD4+ T-cell early metabolic response to TCR stimulation. Nat. Commun. 10, 2042 (2019).
Guertin, D. A. & Wellen, K. E. Acetyl-CoA metabolism in cancer. Nat. Rev. Cancer 23, 156–172 (2023).
Young, C. M. et al. Metabolic dependencies of metastasis-initiating cells in female breast cancer. Nat. Commun. 14, 7076 (2023).
Latchman, Y. et al. PD-L2, a novel B7 homologue, is a second ligand for PD-1 and inhibits T cell activation. FASEB J. 15, A345 (2001).
Yokosuka, T. et al. Programmed cell death 1 forms negative costimulatory microclusters that directly inhibit T cell receptor signaling by recruiting phosphatase SHP2. J. Exp. Med. 209, 1201–1217 (2012).
Hu, X. Y., Li, J., Fu, M. R., Zhao, X. & Wang, W. The JAK/STAT signaling pathway: from bench to clinic. Signal Transduct. Target. Ther. 6, 402 (2021).
Patil, N. S. et al. Intratumoral plasma cells predict outcomes to PD-L1 blockade in non-small cell lung cancer. Cancer Cell 40, 289–300 (2022).
Zhu, A. X. et al. Ramucirumab after sorafenib in patients with advanced hepatocellular carcinoma and increased alpha-fetoprotein concentrations (REACH-2): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 20, 282–296 (2019).
Huang, J. et al. FASN inhibition decreases MHC-I degradation and synergizes with PD-L1 checkpoint blockade in hepatocellular carcinoma. Cancer Res. 84, 855–871 (2024).
Liu, T. et al. High-affinity neoantigens correlate with better prognosis and trigger potent antihepatocellular carcinoma (HCC) activity by activating CD39+CD8+ T cells. Gut 70, 1965–1977 (2021).
Hayday, A., Dechanet-Merville, J., Rossjohn, J. & Silva-Santos, B. Cancer immunotherapy by γδ T cells. Science 386, eabq7248 (2024).
Kirchhammer, N. et al. NK cells with tissue-resident traits shape response to immunotherapy by inducing adaptive antitumor immunity. Sci. Transl. Med. 14, eabm9043 (2022).
Zhang, Y. et al. CD49a targeting enhances NK cell function and antitumor immunity. Cancer Immunol. Res. 13, 139–151 (2025).
Vermeer, M. et al. Tissue localization of natural killer cells dictates surveillance of lung metastasis. Nat. Commun. 16, 9464 (2025).
Li, Y. L. et al. IGSF8 is an innate immune checkpoint and cancer immunotherapy target. Cell 187, 2703–2716 (2024).
Li, Y. et al. Tumor cells impair immunological synapse formation via central nervous system-enriched metabolite. Cancer Cell 42, 985–1002 (2024).
Coe, D. et al. Depletion of regulatory T cells by anti-GITR mAb as a novel mechanism for cancer immunotherapy. Cancer Immunol. Immunother. 59, 1367–1377 (2010).
Wang, L. et al. PD-L1-expressing tumor-associated macrophages are immunostimulatory and associate with good clinical outcome in human breast cancer. Cell Rep. Med. 5, 101420 (2024).
Delconte, R. B. et al. Fasting reshapes tissue-specific niches to improve NK cell-mediated anti-tumor immunity. Immunity 57, 1923–1938 (2024).
Zhou, Y. et al. The application of interleukin-2 family cytokines in tumor immunotherapy research. Front. Immunol. 14, 1090311 (2023).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Li, Z. et al. Presence of onco-fetal neighborhoods in hepatocellular carcinoma is associated with relapse and response to immunotherapy. Nat. Cancer 5, 167–186 (2024).
Zhong, Z. X. et al. Domain generalization enables general cancer cell annotation in single-cell and spatial transcriptomics. Nat. Commun. 15, 1929 (2024).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Holland, C. H. et al. Robustness and applicability of transcription factor and pathway analysis tools on single-cell RNA-seq data. Genome Biol. 21, 36 (2020).
Stary, V. et al. A discrete subset of epigenetically primed human NK cells mediates antigen-specific immune responses. Sci. Immunol. 5, eaba6232 (2020).
Nixon, B. G. et al. Cytotoxic granzyme C-expressing ILC1s contribute to antitumor immunity and neonatal autoimmunity. Sci. Immunol. 7, eabi8642 (2022).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Kang, J. B. et al. Efficient and precise single-cell reference atlas mapping with Symphony. Nat. Commun. 12, 5890 (2021).
Du, X. et al. Human-induced CD49a+ NK cells promote fetal growth. Front. Immunol. 13, 821542 (2022).
Moreno-Nieves, U. Y. et al. Landscape of innate lymphoid cells in human head and neck cancer reveals divergent NK cell states in the tumor microenvironment. Proc. Natl Acad. Sci. USA 118, e2101169118 (2021).
Acknowledgements
We are grateful for the scPLC data of patients with HCC provided by R. Xue from Peking University and the bulk RNA-seq data provided by W. Zhai from the Institute of Zoology, Chinese Academy of Sciences. This work was supported by grants from the National Natural Science Foundation of China (grant nos. 82230059 and U25A20111), the Jiangsu Provincial Key Research Development Program of China (grant no. BE2022770), Major Program of Wuxi Medical Centre, Nanjing Medical University (grant no. WMCM202308) awarded to Y.C. and grants from the National Natural Science Foundation of China (grant no. 82573130), Postdoctoral Fellowship Program of China Postdoctoral Science Foundation (grant no. GZB20250491), the Natural Science Excellent Youth Foundation of Jiangsu Province (grant no. BK20250161), Jiangsu Funding Program for Excellent Postdoctoral Talent (grant no. 2025ZB138) and the Jiangsu Province Young Science and Technology Talent Support Project awarded to C.H.
Author information
Authors and Affiliations
Contributions
Y.C. and W.Y. conceived the project and designed the experiments. W.Y., C.H., Y.Z., Y. Huang, C.W., M.Z., Y.Ha., J.Y., D.K. and R.S. performed the experiments. M.C. performed experiments of CRISPR gene editing in mouse primary NK cells. W.Y., C.H., Y.Z., C.W. and Y.C. analysed the data. W.Y., Y. Han, Y.Z., J.Y., D.K., S.L., Y.Z., Q.L. and R.S. were involved in patient inclusion and sample acquisition. J.L. helped us in the flow cytometry experiment. X.-F.Y. and B.Y. helped us in the response to the reviewers. W.Y. and Y.C. wrote the manuscript. Q.X., W.C., D.-M.K. and Y.C. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Zhongjun Dong, Nicholas Huntington and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 COTL1high NK cells are associated with αPD-1/PD-L1 efficacy in tsMHC-Iimpaired HCC.
a, The correlation between MHC-I+ tumor cells estimated by Bisque R package and CD8+ T cell scores in EGAD00001008128 dataset. The error bands represent the 95% confidence interval for the correlation. b, Volcano plot showing the differential expression genes between responders and non-responders among 144 patients with low CD8+ T cells. c, GSEA analysis of genes enrolled in ‘GSE45365_NK_CELL_VS_CD8_TCELL_UP’ from msigDB database. d, Imaging mass cytometry analysis of NK cells infiltration between tsMHC-Iimpaired responders and non-responders (P < 0.0001). Scale bar, 150 μm. e, Representative mIHC staining depicting TCRγδ+ cells (P = 0.2810) and TCRγδ+ Granzyme B+ cells (P = 0.6943) of HCC specimens before treatment are shown (left). Scale bar, 100 and 20 μm. The statistical results of mean γδ T and Granzyme B+ γδ T numbers in 10 random fields (1000 μm* 1000μm) per HCC tumor were shown (responder, n = 7; non-responder, n = 8). f, MHC-I expression in H22 cells was evaluated by flow cytometry. g, Representative images showing the efficiency of depleting NK cells. R, responder; NR, non-responder. P values were determined by unpaired two-sided Mann-Whitney tests (d, e). Data are shown as the mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 2 The variation of tumor MHC class I reshapes immune infiltration of HCC.
a, Flow cytometry histogram showing the protein expression of MHC-I (H-2Kb/H-2Db) on vector and B2mOE Hep-53.4 cells. b, Schematic representing the construction of a mouse model bearing vector and B2mOE Hep-53.4 hepatoma. c, Representative images and statistical results of whole-liver morphology from vector and B2mOE Hep-53.4-bearing orthotopic HCC models (each n = 5, P = 0.0159). Scale bars, 1 cm. d, Flow cytometry analysis of tumor-infiltrating T and NK cells in vector and B2mOE Hep-53.4-bearing orthotopic HCC tumors (each n = 5; T cells, P = 0.0079; NK cells, P = 0.0159). e, Representative mIHC staining and statistical results showing the number of NKp46+ and CD8+ cells per FOV (500 μm * 500 μm) in vector and B2mOE Hep-53.4 hepatoma (each n = 5, each mouse five FOVs, T cells, P < 0.0001; NK cells, P < 0.0001). Scale bars, 50 μm. f, Flow cytometry histogram showing the protein expression of MHC-I (H-2Kb/H-2Db) on Hepa1-6 cells. g, Flow cytometry histogram showing the protein expression of MHC-I (H-2Kb/H-2Db) on sgCtrl and sgB2m Hepa1-6 cells. h, Schematic representing the construction of a mouse model bearing Hepa1-6 hepatoma with variable MHC-I expression backgrounds. i, Representative images and statistical results of whole-liver morphology from Hepa1-6-bearing orthotopic HCC models with variable MHC-I expression (each n = 6; sgCtrl: sgB2m 1:0 versus 1:1, P = 0.6190, 0:1, P = 0.0087; 1:1 versus 0:1, P = 0.0130). Scale bars, 1 cm. j, Representative mIHC staining of NKp46 and CD8 in WT, WT/B2m KO mixed (1:1) and B2m KO Hepa1-6 bearing orthotopic HCC tumors (CD8+ T: sgCtrl: sgB2m 1:0 versus 1:1, P = 0.6834, 0:1, P = 0.0728; 1:1 versus 0:1, P = 0.0005; NK cell: sgCtrl: sgB2m 1:0 versus 1:1, P = 0.0016, 0:1, P = 0.2236; 1:1 versus 0:1, P = 0.0051). Scale bars, 50 μm. k, Statistical results showing the number of NKp46+ and CD8+ cells per FOV (650 μm * 650 μm) in CD8 in WT, WT/B2m KO mixed (1:1) and B2m KO Hepa1-6 bearing orthotopic HCC tumors (each n = 5 and each mouse four FOVs; NK cell: sgCtrl: sgB2m 1:0 versus 1:1, P = 0.0152, 0:1, P = 0.3095; 1:1 versus 0:1, P = 0.0043; T cell: sgCtrl: sgB2m 1:0 versus 1:1, P > 0.9999, 0:1, P = 0.0022; 1:1 versus 0:1, P = 0.0022). l, Representative flow cytometry images of depleting NK, CD8+ T and both in orthotopic Hep-53.4 tumors. m, Representative images of whole-liver morphology from Hep-53.4-bearing orthotopic HCC models treated with isotype antibody, anti-asialo GM1, anti-CD8 and combination. Scale bars, 1 cm. n, Statistical results of maximum tumor diameter in Hep-53.4-bearing orthotopic HCC models treated with isotype antibody, anti-asialo GM1, anti-CD8 and combination (each n = 5; Isotype versus anti-asialo GM1, P = 0.0079, anti-CD8, P = 0.0079, anti-asialo GM1+ anti-CD8, P = 0.0079; anti-asialo GM1 versus anti-CD8, P = 0.4365, anti-asialo GM1+ anti-CD8, P = 0.5714). o, Flow cytometry analysis of representative cases of CD49+ (COTL1high) NK cells in human HCC tumors between over 80% tsMHC-I+ fraction and less than 80% tsMHC-I+ fraction. Frequencies of CD49+ (COTL1high) NK cells in relative to CD45+ immune cells (n = 11 vs 13; P = 0.0030). p, Integration analysis of 101 HCC tumors showed the distribution of COTL1high NK cells between over 80% tsMHC-I+ fraction and less than 80% tsMHC-I+ fraction (P = 0.0380). q, Label transfer analysis of our data and GSE235863 (ICB-treated patients) by symphony R package showing COTL1high NK cells have a higher fraction in responders. P values were determined by unpaired two-sided Mann-Whitney tests (c-e, i-k and n-p). Data are shown as the mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 3 The phenotype of COTL1high NK cells in the treatment-naive HCC TME.
a, Flow cytometry analysis of representative cases of CD45+CD3−CD56+ NK cells in human HCC tumors and adjacent liver tissues. Frequencies of total NK cells in paired relative to the CD45+ lymphocytes (n = 16, P = 0.016). b, The ration of NK cells among tumors, tumor edges and normal liver tissues assessed by a public CyTOF data (https://data.mendeley.com/datasets/jxsz3hdsyg/2; Normal versus edge, P = 0.0007, tumor, P = 0.0012; tumor versus edge, P = 0.0942). c, UMAP plots showing the distribution of PRF1, IFNG, GZMB, HAVCR2 and LAG3 expression between tumors and adjacent liver tissues. d, Violin plots showing the expression of inhibitory receptor genes (left) and activation receptor genes (right) in NK cell clusters. e, Gating strategy for the identification of NK cells used in CyTOF clustering. f, The tSNE plot showing the tissue distribution of NK cells in CyTOF data. g, h, Flow cytometry analysis of NK cells from tumor tissue of patients with HCC expressing Tim-3 (g, n = 12, P = 0.064) and Perforin (h, n = 14, P = 0.0022). i, Analysis of tsMHC-I expression in pre- and post-treated HCC tumors using spatial transcriptome data (Zenodo database: 15396298). j, Representative mIHC staining (left) and statistical results (right) depicting the number of CD8+ T cells and CD56+COTL1+ NK cells in paired pre- and post-treated specimens from responders and non-responders (n = 5 vs 5; CD8+ T, responder, P = 0.0625; non-responder, P = 0.3125; COTL1+ NK, responder, P = 0.0625; non-responder, P = 0.1250). Scale bar, 100 μm. R, responder; NR, non-responder. P values were determined by paired two-sided Mann-Whitney tests (a, b, g, h and j). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 4 The phenotypes of NK cells in murine HCC models.
a, Integrated analysis of single CD3−NK1.1+ cells from four public datasets. b, Transfer labeling analysis of human COTL1high NK cells and murine NK cells. Heatmap showing jaccard similar index between human and murine NK cells. c, UMAP plots showing the expression of Itga1, Itga2, Rbpj, Tnfrsf18, Zfp683 and Ifng which we observed highly expressed in human HCC. d, Violin plot showing the expression of Cotl1 and Tnfrsf18 between CD49a+CD49b− and CD49a−CD49b+ NK cells. e, Chromatin accessibility analysis showed similar epigenetic characteristics between human and murine CD49a+ NK cells. f, Analysis of IFN-γ (P = 0.0156), KLRG1 (P = 0.0156), and PD-1 (P = 0.0156) expression on NK cell subsets (CD49a+CD49b− NK and CD49a−CD49b+ NK) by flow cytometry (each n = 7) on Hep-53.4 bearing hepatoma. g Schematic illustration showing constructing the mice model bearing H22 hepatoma. h Flow cytometry analysis of Cd49a and Cd49b in NK cells from H22 tumors. i Analysis of IFN-γ (P = 0.0312), KLRG1 (P = 0.0312), and PD-1 (P = 0.0312) expression on NK cell subsets (CD49a+CD49b− NK and CD49a−CD49b+ NK) by flow cytometry (each n = 7) on H22-bearing hepatoma. j, Analysis of IFN-γ (P = 0.0312) and PD-1 (P = 0.0312) expression on NK cell subsets from DEN+CCl4 induced hepatocarcinogenesis by flow cytometry (n = 7). k, The fraction of CD49a+CD49b− NK cells in tumors from Zfp683+/+ and Zfp683−/− mice (each n = 5, P = 0.0079). l, Analysis of the fraction and the number of CD8+CD103+ Trm cells in Hep53.4-bearing hepatoma (each n = 5; fraction, P = 0.4603; number, P = 0.6905). m, n Flow cytometry analysis (m) and statistical results (n) of CD8+ and CD4+ T cells fraction (each n = 5) in tumors from Zfp683+/+ and Zfp683−/− mice treated with isotype control or αPD-L1 antibody (CD8+ T cells: Zfp683+/+ - isotype versus Zfp683−/− - isotype, P > 0.9999, Zfp683+/+ - αPD-L1, P = 0.0079, Zfp683−/− - αPD-L1, P = 0.6905; Zfp683−/− - αPD-L1 versus Zfp683+/+ - αPD-L1, P = 0.0079; CD4+ T cells: Zfp683+/+ - isotype versus Zfp683−/− - isotype, P = 0.2222, Zfp683+/+ - αPD-L1, P = 0.0952, Zfp683−/− - αPD-L1, P = 0.0556; Zfp683−/− - αPD-L1 versus Zfp683+/+ - αPD-L1, P > 0.9999). o, Analysis of IFN-γ+ CD4+ T cells fraction (each n = 5) in tumors from Zfp683+/+ and Zfp683−/− mice treated with isotype control or αPD-L1 antibody (Zfp683+/+ - isotype versus Zfp683−/− - isotype, P = 0.1508, Zfp683+/+ - αPD-L1, P = 0.4206, Zfp683−/− - αPD-L1, P = 0.8413; Zfp683−/− - αPD-L1 versus Zfp683+/+ - αPD-L1, P = 0.7381). P values were determined by unpaired two-sided Mann-Whitney tests (e and h-k). Data are shown as the mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 5 Crosstalk between PD-L1+ macrophages and COTL1high NK cells.
a, The colocalization of CD49a+ (COTL1high) NK cells and macrophages in an HCC tumor sample detected by MERFISH (https://info.vizgen.com/ffpe-showcase?submissionGuid=6d5172c8-49af-44c2-a90b-376059fd4268). A representative tissue section (HumanLiverCancerPatient2) is shown. The number of macrophages and other myeloid cells within the distance to COTL1high NK cells < 50μm was measured (P < 0.0001). b, mIHC staining of CD68 and PD-L1 in HCC tumors. c, The UAMP plot shows the expression of TNFRSF18 across NK cells. d, Flow cytometry-based dimensionality reduction showing the expression of CD49a, GITR and COTL1 in NK cells from 13 HCC tumors. e, UMAP plot showing the cell types identified in scPLC dataset. f, UMAP plot showing the expression of TNFSF18 and TNFRSF18 in scPLC dataset. g, Flow cytometry analysis of GITRL expression in macrophages and DCs from human HCC (P = 0.0556). h, The distance of PD-L1+ macrophages or DCs to COTL1high NK cells spots were measured in HCC 10x Visium data. i, The distance of PD-L1+ macrophages or DCs to COTL1high NK cells spots were measured in stereo-seq data. j, MERFISH analysis showing the transcript of COTL1high NK cells and macrophages. k, The distance of PD-L1+ macrophages or DCs to COTL1high NK cells spots were measured in 10x Visium data from nasopharyngeal carcinoma (GSE206245). Each point means average distance in one sample (P = 0.0286). l, The distance of PD-L1+ macrophages or DCs to COTL1high NK cells spots were measured in 10x Visium data from pancreatic cancer (GSE278687). Each point means average distance in one sample (P = 0. 0093). m, The diagram f HALO Proximity analysis. n, Differential spatial centroid distances between COTL1high NK cells and PD-L1+ macrophages versus DCs (P = 0.0286). n = 4 HCC samples, scale bar: 50μm. o, mIHC staining showing the colocalization between GITR+COTL1+ NK cells and PD-L1+ macrophages. P values were determined by unpaired two-sided Mann-Whitney tests (a, g, k, l, n). Data are shown as the mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 6 CUT&RUN sequencing analysis between CD49a+ and CD49a− NK cells sorted from human HCC tissues.
a, TEM showing the membrane protrusions of multiple cells in one image using low-magnification microscopy (n = 3 patients). b, Confocal laser scanning microscopy showing the F-actin staining between si-COTL1-treated COTL1high NK cells and macrophages. Scale bar, 5 μm. c, Bar plots showing the fraction of RBPJ-expressed cells across 6 NK clusters. d, Scatter plot showing the Pearson correlation between RBPJ and TNFRSF18 expression level in NK cells from HCC tumors. The significance was tested using a two-tailed t-test, with p-values derived from the t-distribution with n-2 degrees of freedom.
Extended Data Fig. 7 OXPHOS was markedly up-regulated in RBPJ+ COTL1high NK cluster.
a, MFI (left) and statistical results (right) of IFN-γ levels of live in vitro-induced COTL1high NK cells treated with DAPT (a γ-Secretase inhibitor which inhibits the activation of Notch signalling) and its’ control (each n = 4; medium versus DAPT, P = 0.3677; DMSO versus DAPT, P = 0.4968). b, The volcano plot represents the differentially metabolic activity between COTL1high NK cells and other NK clusters. c, GSEA analysis of 50 hallmark pathways between RBPJ+ and RBPJ− COTL1high NK cells. d, Basal respiration were quantified (P = 0.0565). e, f, MFI (right) and representative histogram (left) of Fluo-3 AM (e, P = 0.0286) and TMRE (f, P = 0.0286) staining of live in vitro-induced COTL1high NK cells treated with GITR agonist and its’ control (each n = 4). g, The box plot represents the mitochondrial respiration complex signature score based on scRNA-seq data of human HCC NK cells. The signature was from the wikipathways database and assessed by AUCell. h, Flow cytometry analysis (left) and statistical results (right) of IFN-γ expression of COTL1high NK cells upon mitochondrial respiration complex inhibition including carbonyl cyanide m-chlorophenylhydrazone (CCCP), carbonyl cyanide 4-(trifluoromethoxy) phenylhydrazone (FCCP) and oligomycin (each n = 4; Control versus CCCP, P = 0.0286, FCCP, P = 0.0286, Oligomycin, P = 0.0286). i, Heatmap showing the relative PROGENy pathway scores in each NK cluster. j, Transcription factor ranks, which are required for top 100 DEGs from scRNA-seq data analyzed by ChEA3 database. k, l, MFI (right) and representative histogram (left) of p-STAT1 (k, P > 0.9999) and p-STAT3 (l, P = 0.2000) staining of live in vitro-induced COTL1high NK cells treated with GITR agonist and its’ control (each n = 3). m, p-SHP-2 levels of COTL1high NK cells were measured by flow cytometry (each n = 3, p-SHP-2, P = 0.1187). n, p-SHP1 MFI of COTL1high NK cells cultured in medium alone or treated by rPD-L1, rGITRL and SHP-1 inhibitor TPI-1 (each n = 3, Medium versus rGITRL, P = 0.5586, rGITRL+rPD-L1, P < 0.0001, rGITRL+rPD-L1 + TPI-1, P = 0.9630). o, The mechanisms behind the functional interaction between PD-L1⁺ macrophages and COTL1high NK cells in the context of GITR activation and PD-L1 blockade. MFI, mean fluorescence intensity. pSTAT1/3/5, phosphorylated STAT1/3/5; pSHP-2, phosphorylated SHP-2; Fluo-3 AM, Fluo-3-pentaacetoxymethyl ester; TMRE, Tetramethylrhodamine ethyl ester perchlorate. P values were determined by unpaired two-sided Mann-Whitney tests (a, d-f, h, k and l). P values were determined by unpaired two-sided Students’ t-test (m and n). Data are shown as the mean ± s.e.m. In the box plots, the centre line is the median, box limits are the first and third quartiles and whiskers are 1.5× interquartile range. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 8 Combination therapy of αPD-L1 and GITR activation enhances CD8+ T cell and COTL1high NK cell responses in MHC-Iimpaired HCC.
(a-e) Statistical results showing the percentage of macrophages (a), monocytes (b), DCs (c), NKT cells (d) and IFN-γ+ CD49a−CD49b+ NK cells (e) among Hep-53.4-bearing mice treated with DTA-1, αPD-L1, combination or isotype control. f, The mean number of TGF-β+ Tregs in Hep-53.4 tumors treated with DTA-1, αPD-L1, combination or isotype control. The values were determined by mean values of 5 random FOVs (each n = 4; Isotype versus DTA-1, P = 0.0286, αPD-L1, P = 0.0286, αPD-L1 + DTA-1, P = 0.0286; DTA-1 versus αPD-L1, P = 0.0286, αPD-L1 + DTA-1, P = 0.3429; αPD-L1 versus αPD-L1 + DTA-1, P = 0.0286). g, Depletion of CD4+ T cells do not influence the αPD-L1 efficacy in Hep-53.4 HCC model (each n = 5, P = 0.7302). P values were determined by unpaired two-sided Mann-Whitney tests (a-g). Data are shown as the mean ± s.e.m.
Extended Data Fig. 9 The role of GITR signaling in H22-bearing orthotopic HCC models.
a, Schematic illustration showing H22-bearing mice treated with DTA-1, αPD-L1, combination or isotype control. b, Representative images and statistical results of whole-liver morphology from H22-bearing orthotopic HCC models (each n = 6). c-j Flow cytometry analysis of tumor-infiltrating immune cells from orthotopic HCC tumors (each n = 6). k, Flow cytometry histogram showing the protein expression of GITR on sgCtrl and sgTnfrsf18 murine primary NK cells. P values were determined by unpaired two-sided Mann-Whitney tests (b-j). Data are shown as the mean ± s.e.m.
Extended Data Fig. 10 The predictive value of COTL1high NK cells in ICB treatment of other MHC-Iimpaired solid cancers.
a, b, Tumor subtype classification based on CD8+ T cell gene expression of ICB cohorts: OAK & POLAR (lung adenocarcinoma, LUAD) cohort (a) and melanoma cohort (b). The correlation between MHC-I gene expression and CD8+ T cell infiltration was calculated. c, The process of identifying the expression ratio of tsMHC-I in spatial transcriptome data. d, A summary diagram of the main findings from this study.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods and Figs. 1–6.
Supplementary Tables (download XLSX )
Supplementary Table 1. The clinicopathological features in 56 ICB-treated patients with HCC, related to Figs. 1 and 2. Supplementary Table 2. Immune cell marker gene sets used in Fig. 1. Supplementary Table 3. List of CyTOF antibodies staining in this study, related to Fig. 2. Supplementary Table 4. The clinicopathological features in 90 treatment-naive patients with HCC, related to Figs. 2–4. Supplementary Table 5. Gene signatures of COTL1high NK cells and GITRL+ macrophages related to Fig. 4. Supplementary Table 6. The clinicopathological features in five patients for EVOC. Supplementary Table 7. Antibodies and chemicals used in this paper.
Supplementary Data 1 (download XLSX )
Statistical source data.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
You, W., Hu, C., Zhang, Y. et al. Reinvigorating COTL1high NK cells via GITR signalling overcomes immune checkpoint blockade resistance in tsMHC-I-impaired tumours. Nat Cell Biol (2026). https://doi.org/10.1038/s41556-026-01925-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41556-026-01925-9