Abstract
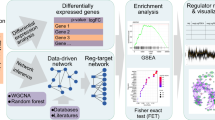
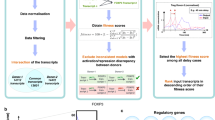
The regulation of gene expression is crucial for the functional integration of evolutionarily young genes, particularly those that emerge de novo. However, the regulatory programmes governing the expression of de novo genes remain unknown. To address this, we applied computational methods to single-cell RNA sequencing data, identifying key transcription factors probably instrumental in regulating de novo genes. We found that transcription factors do not have the same propensity for regulating de novo genes; some transcription factors regulate more de novo genes than others. Leveraging genetic and genomic tools in Drosophila, we further examined the role of two key transcription factors, achintya and vismay, and the regulatory architecture of new genes. Our findings identify key transcription factors associated with the expression of de novo genes and highlight how transcription factors, and possibly their duplications, are linked to the expressional regulation of de novo genes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
RNA-seq data are submitted to NCBI BioProject under accession number PRJNA1197503.
Code availability
Code and scripts for this study are available at https://github.com/LiZhaoLab/DeNovoGene_Transcriptional_Regulation and available via Zenodo at https://doi.org/10.5281/zenodo.15304293 (ref. 71).
References
Rebeiz, M., Patel, N. H. & Hinman, V. F. Unraveling the tangled skein: the evolution of transcriptional regulatory networks in development. Annu. Rev. Genom. Hum. Genet. 16, 103–131 (2015).
Zabidi, M. A. & Stark, A. Regulatory enhancer–core-promoter communication via transcription factors and cofactors. Trends Genet. 32, 801–814 (2016).
Zhao, L., Svetec, N. & Begun, D. J. De novo genes. Ann. Rev. Genet 58, 211–232 (2024).
Long, M., VanKuren, N. W., Chen, S. & Vibranovski, M. D. New gene evolution: little did we know. Annu. Rev. Genet. 47, 307–333 (2013).
Voordeckers, K., Pougach, K. & Verstrepen, K. J. How do regulatory networks evolve and expand throughout evolution? Curr. Opin. Biotechnol. 34, 180–188 (2015).
Pougach, K. et al. Duplication of a promiscuous transcription factor drives the emergence of a new regulatory network. Nat. Commun. 5, 4868 (2014).
Rich, A., Acar, O. & Carvunis, A.-R. Massively integrated coexpression analysis reveals transcriptional regulation, evolution and cellular implications of the yeast noncanonical translatome. Genome Biol. 25, 183 (2024).
Lee, B.-K. et al. Cell-type specific and combinatorial usage of diverse transcription factors revealed by genome-wide binding studies in multiple human cells. Genome Res. 22, 9–24 (2012).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Van Oss, S. B. & Carvunis, A.-R. De novo gene birth. PLoS Genet. 15, e1008160 (2019).
McLysaght, A. & Hurst, L. D. Open questions in the study of de novo genes: what, how and why. Nat. Rev. Genet. 17, 567–578 (2016).
Chen, S., Krinsky, B. H. & Long, M. New genes as drivers of phenotypic evolution. Nat. Rev. Genet. 14, 645–660 (2013).
Xia, S., Chen, J., Arsala, D., Emerson, J. J. & Long, M. Functional innovation through new genes as a general evolutionary process. Nat. Genet. 57, 295–309 (2025).
Begun, D. J., Lindfors, H. A., Thompson, M. E. & Holloway, A. K. Recently evolved genes identified from Drosophila yakuba and D. erecta accessory gland expressed sequence tags. Genetics 172, 1675–1681 (2006).
Kaessmann, H. Origins, evolution, and phenotypic impact of new genes. Genome Res. 20, 1313–1326 (2010).
Witt, E., Benjamin, S., Svetec, N. & Zhao, L. Testis single-cell RNA-seq reveals the dynamics of de novo gene transcription and germline mutational bias in Drosophila. eLife 8, e47138 (2019).
Witt, E., Langer, C. B., Svetec, N. & Zhao, L. Transcriptional and mutational signatures of the Drosophila ageing germline. Nat. Ecol. Evol. 7, 440–449 (2023).
Zhao, L., Saelao, P., Jones, C. D. & Begun, D. J. Origin and spread of de novo genes in Drosophila melanogaster populations. Science 343, 769–772 (2014).
Cridland, J. M., Majane, A. C., Zhao, L. & Begun, D. J. Population biology of accessory gland-expressed de novo genes in Drosophila melanogaster. Genetics 220, iyab207 (2022).
Raz, A. A. et al. Emergent dynamics of adult stem cell lineages from single nucleus and single cell RNA-seq of Drosophila testes. eLife 12, e82201 (2023).
Witt, E., Svetec, N., Benjamin, S. & Zhao, L. Transcription factors drive opposite relationships between gene age and tissue specificity in male and female Drosophila gonads. Mol. Biol. Evol. 38, 2104–2115 (2021).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Li, H. et al. Fly Cell Atlas: a single-nucleus transcriptomic atlas of the adult fruit fly. Science 375, eabk2432 (2022).
Peng, J. & Zhao, L. The origin and structural evolution of de novo genes in Drosophila. Nat. Commun. 15, 810 (2024).
Kryuchkova-Mostacci, N. & Robinson-Rechavi, M. Tissue-specific evolution of protein coding genes in human and mouse. PLoS ONE 10, e0131673 (2015).
Leader, D. P., Krause, S. A., Pandit, A., Davies, S. A. & Dow, J. A. T. FlyAtlas 2: a new version of the Drosophila melanogaster expression atlas with RNA-Seq, miRNA-Seq and sex-specific data. Nucleic Acids Res. 46, D809–D815 (2018).
Sonawane, A. R. et al. Understanding tissue-specific gene regulation. Cell Rep. 21, 1077–1088 (2017).
Luscombe, N. M. et al. Genomic analysis of regulatory network dynamics reveals large topological changes. Nature 431, 308–312 (2004).
Rauluseviciute, I. et al. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 52, D174–D182 (2024).
Perkins, K. K., Admon, A., Patel, N. & Tjian, R. The Drosophila Fos-related AP-1 protein is a developmentally regulated transcription factor. Genes Dev. 4, 822–834 (1990).
Soory, A. & Ratnaparkhi, G. S. SUMOylation of Jun fine-tunes the Drosophila gut immune response. PLoS Pathog. 18, e1010356 (2022).
Vogt, P. K. & Bos, T. J. in Advances in Cancer Research Vol. 55 (eds Vande Woude, G. F. & Klein, G.) 1–35 (Elsevier, 1990).
Ayyar, S., Jiang, J., Collu, A., White-Cooper, H. & White, R. A. H. Drosophila TGIF is essential for developmentally regulated transcription in spermatogenesis. Development 130, 2841–2852 (2003).
Wang, Z. & Mann, R. S. Requirement for two nearly identical TGIF-related homeobox genes in Drosophila spermatogenesis. Development 130, 2853–2865 (2003).
Zaborowski, A. B. & Walther, D. Determinants of correlated expression of transcription factors and their target genes. Nucleic Acids Res. 48, 11347–11369 (2020).
Wang, Y., Wang, L. & Wang, Z. Transgenic analyses of TGIF family proteins in Drosophila imply their role in cell growth. J. Genet. Genom. 35, 457–465 (2008).
Perezgasga, L. et al. Regulation of transcription of meiotic cell cycle and terminal differentiation genes by the testis-specific Zn-finger protein matotopetli. Development 131, 1691–1702 (2004).
Beall, E. L. et al. Discovery of tMAC: a Drosophila testis-specific meiotic arrest complex paralogous to Myb–Muv B. Genes Dev. 21, 904–919 (2007).
Hiller, M. et al. Testis-specific TAF homologs collaborate to control a tissue-specific transcription program. Development 131, 5297–5308 (2004).
Vilstrup, A. P. et al. A germline PAF1 paralog complex ensures cell type-specific gene expression. Genes Dev. 38, 866–886 (2024).
Theofel, I. et al. tBRD-1 and tBRD-2 regulate expression of genes necessary for spermatid differentiation. Biol. Open https://doi.org/10.1242/bio.022467 (2017).
Gönczy, P., Matunis, E. & DiNardo, S. bag-of-marbles and benign gonial cell neoplasm act in the germline to restrict proliferation during Drosophila spermatogenesis. Development 124, 4361–4371 (1997).
Flaherty, M. S. et al. chinmo is a functional effector of the JAK/STAT pathway that regulates eye development, tumor formation, and stem cell self-renewal in Drosophila. Dev. Cell 18, 556–568 (2010).
Leatherman, J. L. & DiNardo, S. Germline self-renewal requires cyst stem cells and stat regulates niche adhesion in Drosophila testes. Nat. Cell Biol. 12, 806–811 (2010).
Schlötterer, C. Genes from scratch—the evolutionary fate of de novo genes. Trends Genet. 31, 215–219 (2015).
Carvunis, A.-R. et al. Proto-genes and de novo gene birth. Nature 487, 370–374 (2012).
Lee, U., Mozeika, S. M. & Zhao, L. A synergistic, cultivator model of de novo gene origination. Genome Biol. Evol. 16, evae103 (2024).
Van De Sande, B. et al. A scalable SCENIC workflow for single-cell gene regulatory network analysis. Nat. Protoc. 15, 2247–2276 (2020).
McLysaght, A. & Guerzoni, D. New genes from non-coding sequence: the role of de novo protein-coding genes in eukaryotic evolutionary innovation. Phil. Trans. R. Soc. B 370, 20140332 (2015).
Yona, A. H., Alm, E. J. & Gore, J. Random sequences rapidly evolve into de novo promoters. Nat. Commun. 9, 1530 (2018).
Galupa, R. et al. Enhancer architecture and chromatin accessibility constrain phenotypic space during Drosophila development. Dev. Cell 58, 51–62 (2023).
Khodursky, S. et al. The evolution and mutational robustness of chromatin accessibility in Drosophila. Genome Biol. 24, 232 (2023).
Lu, D., Sin, H.-S., Lu, C. & Fuller, M. T. Developmental regulation of cell type-specific transcription by novel promoter-proximal sequence elements. Genes Dev. 34, 663–677 (2020).
Bravo González-Blas, C. et al. SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks. Nat. Methods 20, 1355–1367 (2023).
Zhou, Q. et al. On the origin of new genes in Drosophila. Genome Res. 18, 1446–1455 (2008).
Xia, S. et al. Genomic analyses of new genes and their phenotypic effects reveal rapid evolution of essential functions in Drosophila development. PLoS Genet. 17, e1009654 (2021).
Andersen, P. R., Tirian, L., Vunjak, M. & Brennecke, J. A heterochromatin-dependent transcription machinery drives piRNA expression. Nature 549, 54–59 (2017).
Kryuchkova-Mostacci, N. & Robinson-Rechavi, M. A benchmark of gene expression tissue-specificity metrics. Brief. Bioinform. https://doi.org/10.1093/bib/bbw008 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 52, D33–D43 (2024).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Peng, J., Wang, B., Svetec, N. & Zhao, L. De novo gene transcriptional regulation. Zenodo https://doi.org/10.5281/zenodo.15304293 (2025).
Acknowledgements
We thank members of L.Z.'s laboratory for helpful discussions. Stocks obtained from the Bloomington Drosophila Stock Center (NIH P40OD018537) were used in this study. We thank The Rockefeller University High Performance Computing Center for the support in computation. This work was supported by National Institutes of Health MIRA R35GM133780, the Robertson Foundation and an Allen Distinguished Investigator Award from Paul G. Allen Family Foundation and Kellen Women’s Entrepreneurship Fund to L.Z.
Author information
Authors and Affiliations
Contributions
J.P., L.Z. and N.S. conceived the project. J.P. designed and performed large-scale scRNA-seq and regulon analysis. N.S. designed the genetic cross scheme. B.-J.W. and N.S. carried out the genetic experiments and RNA-seq analysis. J.P., L.Z., N.S. and B.-J.W. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Anna Grandchamp, Mohammad Siddiq and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Random permutation analysis of tissue- and cell-type- specificity.
(a). Random permutation with simulated annealing can be used to randomly sample genes with similar expression distribution (blue line) to that of de novo genes (dashed line). Randomly sampled genes before random permutation with simulated annealing show very different expression distributions (red solid line). (b) Differences in distributions before (red solid line) and after random permutation with simulated annealing (blue solid line). Tissue (c) and cell-type- (d) specificity of de novo genes (bootstrapped n = 100) were significantly larger than randomly sampled genes (bootstrapped n = 100) with similar expression distribution, with one-sided t-test p = 3e-162 and 4e-163, respectively. Boxplots show the median (center line), the 25th-75th percentiles (box), and whiskers extending to 1.5× IQR.
Extended Data Fig. 2 Clustering analysis of de novo gene expression across 250 different cell types.
The expression shows clusters in spermatogenesis, gland cells, sensory neurons, hemocytes, ovary cells, midguts, etc.
Extended Data Fig. 3 Regulon profiles of current and additional SCENIC predictions.
(a) and (bb) Current SCENIC predictions. (a) The ratio of de novo genes and testis-specific genes in each regulon. (b) Number of de novo genes and de novo and non-de novo genes in each regulon. (c)–(e) Additional SCENIC analysis with a lower TF expression cutoff of 10 (Methods). (c) Top regulons from this additional SCENIC analysis were similar to the current SCENIC analysis (Fig. 2a). (d) The ratio of de novo genes and testis-specific genes in each regulon. (e) Number of de novo genes and de novo and non-de novo genes in each regulon. (g)–(h) Additional SCENIC analysis with two additional gene sets except for de novo genes and TF genes, which are non-testis tissue specific genes and non-tissue specific genes, with the same cutoff as the original SCENIC analysis. (f) Top regulons from this additional SCENIC analysis were also similar to the current SCENIC analysis (Fig. 2a). (g) The ratio of de novo genes and non-testis tissue-specific genes in each regulon (n = 83). (h) The ratio of de novo genes and non-tissue specific genes in each regulon (n = 83). In (g) and (h), p-values were calculated by Kendall’s Tau and Spearman rank correlations, and shaded areas indicate 95% confident intervals.
Extended Data Fig. 4 Expression pattern of achi, Jra, and de novo genes predicted to be regulated by Achi and Jra.
(a) De novo genes that were predicted to be regulated by Achi were mostly expressed in the testis. (b) 30 out of 45 of the de novo genes predicted to be regulated by Achi have significant correlations with achi in the testis (p < 0.05), with 25 of them being positively correlated and 5 of them negatively correlated. (c) De novo genes that were predicted to be regulated by Jra were mostly expressed in the testis, but also show some expression in the fat body, heart, etc. (d) 49 out of 66 of the de novo genes predicted to be regulated by Jra have significant correlations in testis (p < 0.05), with 48 being positively correlated and only 1 negatively correlated. In (b) and (d), for each predicted target, the p-value was determined as the smallest obtained from a two-sided t-test, Pearson Correlation Coefficient, and Kendall Tau rank test, with the sample size (n) of 7, corresponding to the 7 different spermatogenesis stages.
Extended Data Fig. 5 Expression correlation between vis and Vis targets.
The slope and the coefficient of determination between vis and Vis targets, including (a) testis-specific genes, (b) de novo genes, and (c) TFs.
Extended Data Fig. 6 Prevalence of Vis/Achi binding motifs across the genome.
(a) (a) The number matches found for the binding motifs of TFs in the whole genome. (b) The number of ATAC-seq peaks harboring matches found for different TFs. (c) The number of genes that have matches for different TFs in their promoter regions. (d) The number of de novo gene candidates that have matches for different TFs in their promoter regions.
Extended Data Fig. 7 Gene tree of vis/achi inferred from protein sequence alignments in melanogaster subgroup species.
The protein sequences were obtained from blastp search against non-redundant protein sequence database. Protein sequences from suppressed genome assemblies and annotations were shown in gray.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–8 and Table 1.
Supplementary Data 1 (download CSV )
Tissue specificity of all annotated genes (including coding and non-coding) in Fly Cell Atlas dataset.
Supplementary Data 2 (download CSV )
Predicted de novo gene regulons.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Peng, J., Wang, BJ., Svetec, N. et al. Gene regulatory networks and essential transcription factors for de novo-originated genes. Nat Ecol Evol 9, 1487–1498 (2025). https://doi.org/10.1038/s41559-025-02747-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41559-025-02747-y