Abstract
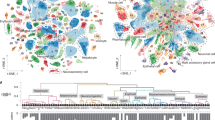
Animal cell types are defined by differential access to genomic information—a process orchestrated by the combinatorial activity of transcription factors that bind to cis-regulatory elements (CREs) to control gene expression. Changes in these gene regulatory networks (GRNs) underlie the origin and diversification of cell types, yet the regulatory logic and specific GRNs that define cell identities remain poorly resolved across the animal tree of life. Cnidarians, as early-branching metazoans, provide a critical window into the early evolution of cell type-specific genome regulation. Here we profiled chromatin accessibility in 60,000 cells from whole adults and gastrula-stage embryos of the sea anemone Nematostella vectensis. We identified 112,728 putative CREs and quantified their activity across cell types, revealing pervasive combinatorial enhancer usage and distinct promoter architectures. To decode the underlying regulatory grammar, we trained sequence-based models predicting CRE accessibility and used these models to infer cell type similarities that reflect known ontogenetic relationships. By integrating sequence motifs, transcription factor expression and CRE accessibility, we reconstructed the GRNs that define cnidarian cell types. Our results show the regulatory complexity underlying cell differentiation in a morphologically simple animal and highlight conserved principles in animal gene regulation. This work provides a foundation for comparative regulatory genomics to understand the evolutionary emergence of animal cell type diversity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw and processed files will be available in GEO repository under accession number GEO: GSE294388. In addition, the atlas can be explored in an interactive database: https://sebelab.crg.eu/nematostella-cis-regulatory-atlas/ and also in an interactive genome browser: https://sebelab.crg.eu/nematostella-cis-reg-jb2/.
Code availability
Scripts to reproduce the data processing and downstream analysis are available via Zenodo at https://doi.org/10.5281/zenodo.17425383 (ref. 93). Unless otherwise specified, scripts are based on R v.4.2.2 and Python v.3.8.10, and the language-specific libraries specified in Methods.
References
Musser, J. M. et al. Profiling cellular diversity in sponges informs animal cell type and nervous system evolution. Science 374, 717–723 (2021).
Sebé-Pedrós, A. et al. Cnidarian cell type diversity and regulation revealed by whole-organism single-cell RNA-seq. Cell 173, 1520–1534 (2018).
Sebé-Pedrós, A. et al. Early metazoan cell type diversity and the evolution of multicellular gene regulation. Nat. Ecol. Evol. 2, 1176–1188 (2018).
Fincher, C. T., Wurtzel, O., de Hoog, T., Kravarik, K. M. & Reddien, P. W. Cell type transcriptome atlas for the planarian Schmidtea mediterranea. Science 360, eaaq1736 (2018).
Plass, M. et al. Cell type atlas and lineage tree of a whole complex animal by single-cell transcriptomics. Science 1723, eaaq1723 (2018).
Cao, J. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Levy, S. et al. A stony coral cell atlas illuminates the molecular and cellular basis of coral symbiosis, calcification, and immunity. Cell 184, 2973–2987.e18 (2021).
Najle, S. R. et al. Stepwise emergence of the neuronal gene expression program in early animal evolution. Cell 186, 4676–4693 (2023).
Tanay, A. & Sebé-Pedrós, A. Evolutionary cell type mapping with single-cell genomics. Trends Genet. 37, 919–932 (2021).
Janssens, J. et al. Decoding gene regulation in the fly brain. Nature 601, 630–636 (2022).
Cusanovich, D. A. et al. Multiplex single-cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Calderon, D. et al. The continuum of Drosophila embryonic development at single-cell resolution. Science 377, eabn5800 (2022).
Domcke, S. et al. A human cell atlas of fetal chromatin accessibility. Science 370, eaba7612 (2020).
Sarropoulos, I. et al. Developmental and evolutionary dynamics of cis-regulatory elements in mouse cerebellar cells. Science 373, eabg4696 (2021).
Li, Y. E. et al. A comparative atlas of single-cell chromatin accessibility in the human brain. Science 382, eadf7044 (2023).
Zhang, K. et al. A single-cell atlas of chromatin accessibility in the human genome. Cell 184, 5985–6001 (2021).
Cusanovich, D. A. et al. A single-cell atlas of in vivo mammalian chromatin accessibility. Cell 174, 1309–1324 (2018).
Minnoye, L. et al. Chromatin accessibility profiling methods. Nat. Rev. Methods Prim. 1, 10 (2021).
Hecker, N. et al. Enhancer-driven cell type comparison reveals similarities between the mammalian and bird pallium. Science 387, eadp3957 (2025).
Parker, J. & Pennell, M. The cellular substrate of evolutionary novelty. Curr. Biol. 35, R626–R637 (2025).
Schwaiger, M. et al. Evolutionary conservation of the eumetazoan gene regulatory landscape. Genome Res. 24, 639–650 (2014).
Kim, I. V. et al. Chromatin loops are an ancestral hallmark of the animal regulatory genome. Nature 642, 1097–1105 (2025).
Chari, T., et al. Whole-animal multiplexed single-cell RNA-seq reveals transcriptional shifts across Clytia medusa cell types. Sci. Adv. 7, eabh1683 (2021).
Siebert, S. et al. Stem cell differentiation trajectories in Hydra resolved at single-cell resolution. Science 365, eaav9314 (2019).
Li, Y. et al. Single-cell transcriptomic analyses reveal the cellular and genetic basis of aquatic locomotion in scyphozoan jellyfish. Preprint at bioRxiv https://doi.org/10.1101/2023.02.06.527379 (2023).
Hu, M., Zheng, X., Fan, C.-M. & Zheng, Y. Lineage dynamics of the endosymbiotic cell type in the soft coral Xenia. Nature 582, 534–538 (2020).
Steger, J. et al. Single-cell transcriptomics identifies conserved regulators of neuroglandular lineages. Cell Rep. 40, 111370 (2022).
Steinmetz, P. R. H. et al. Independent evolution of striated muscles in cnidarians and bilaterians. Nature 487, 231–234 (2012).
Cole, A. G. et al. Muscle cell-type diversification is driven by bHLH transcription factor expansion and extensive effector gene duplications. Nat. Commun. 14, 1747 (2023).
Denner, A. et al. Nanos2 marks precursors of somatic lineages and is required for germline formation in the sea anemone Nematostella vectensis. Sci. Adv. 10, eado0424 (2024).
Miramón-Puértolas, P., Pascual-Carreras, E. & Steinmetz, P. R. H. A population of Vasa2 and Piwi1 expressing cells generates germ cells and neurons in a sea anemone. Nat. Commun. 15, 8765 (2024).
Babonis, L. S. & Martindale, M. Q. Old cell, new trick? Cnidocytes as a model for the evolution of novelty. Integr. Comp. Biol. 54, 714–722 (2014).
Babonis, L. S., Enjolras, C., Ryan, J. F. & Martindale, M. Q. A novel regulatory gene promotes novel cell fate by suppressing ancestral fate in the sea anemone Nematostella vectensis. Proc. Natl Acad. Sci. USA 119, e2113701119 (2022).
Babonis, L. S. et al. Single-cell atavism reveals an ancient mechanism of cell type diversification in a sea anemone. Nat. Commun. 14, 885 (2023).
Persad, S., et al. SEACells infers transcriptional and epigenomic cellular states from single-cell genomics data. Nat. Biotechnol. 41, 1746–1757 (2023).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 935 (2021).
Cole, A. G. et al. Updated single cell reference atlas for the starlet anemone Nematostella vectensis. Front Zool. 21, 8 (2024).
Richards, G. S. & Rentzsch, F. Regulation of Nematostella neural progenitors by SoxB, Notch and bHLH genes. Development 142, 3332–3342 (2015).
Lemaître, Q. I. B. et al. NvPrdm14d-expressing neural progenitor cells contribute to non-ectodermal neurogenesis in Nematostella vectensis. Nat. Commun. 14, 4854 (2023).
Steinmetz, P. R. H., Aman, A., Kraus, J. E. M. & Technau, U. Gut-like ectodermal tissue in a sea anemone challenges germ layer homology. Nat. Ecol. Evol. 1, 1535–1542 (2017).
Rentzsch, F., Fritzenwanker, J. H., Scholz, C. B. & Technau, U. FGF signalling controls formation of the apical sensory organ in the cnidarian Nematostella vectensis. Development 135, 1761–1769 (2008).
Haillot, E. et al. Segregation of endoderm and mesoderm germ layer identities in the diploblast Nematostella vectensis. Nat. Commun. 16, 7979 (2025).
Lebedeva, T. et al. Cnidarian-bilaterian comparison reveals the ancestral regulatory logic of the β-catenin dependent axial patterning. Nat. Commun. 12, 4032 (2021).
Reddington, J. P. et al. Lineage-resolved enhancer and promoter usage during a time course of embryogenesis. Dev. Cell 55, 648–664 (2020).
Zhu, F. et al. The interaction landscape between transcription factors and the nucleosome. Nature 562, 76–81 (2018).
Bulyk, M. L., Drouin, J., Harrison, M. M., Taipale, J. & Zaret, K. S. Pioneer factors—key regulators of chromatin and gene expression. Nat. Rev. Genet. https://doi.org/10.1038/s41576-023-00648-z (2023).
Yanai, I. et al. Genome-wide midrange transcription profiles reveal expression level relationships in human tissue specification. Bioinformatics 21, 650–659 (2005).
Marlétaz, F. et al. Amphioxus functional genomics and the origins of vertebrate gene regulation. Nature 564, 64–70 (2018).
Weintraub, A. S. et al. YY1 is a structural regulator of enhancer-promoter loops. Cell 171, 1573–1588 (2017).
Lenhard, B., Sandelin, A. & Carninci, P. Metazoan promoters: emerging characteristics and insights into transcriptional regulation. Nat. Rev. Genet. 13, 233–245 (2012).
Haberle, V. & Lenhard, B. Promoter architectures and developmental gene regulation. Semin. Cell Dev. Biol. 57, 11–23 (2016).
Ghandi, M., Lee, D., Mohammad-Noori, M. & Beer, M. A. Enhanced regulatory sequence prediction using gapped k-mer features. PLoS Comput. Biol. 10, e1003711 (2014).
Pampari, A. et al. ChromBPNet: bias factorized, base-resolution deep learning models of chromatin accessibility reveal cis-regulatory sequence syntax, transcription factor footprints and regulatory variants. Preprint at bioRxiv https://doi.org/10.1101/2024.12.25.630221 (2025).
De Winter, S., Konstantakos, V. & Aerts, S. Modelling and design of transcriptional enhancers. Nat. Rev. Bioeng. https://doi.org/10.1038/s44222-025-00280-y (2025).
Shrikumar, A., Prakash, E. & Kundaje, A. GkmExplain: fast and accurate interpretation of nonlinear gapped k-mer SVMs. Bioinformatics 35, i173–i182 (2019).
Shrikumar, A. et al. Technical note on transcription factor motif discovery from importance scores (TF-MoDISco) version 0.5.6.5. Preprint at https://arxiv.org/abs/1811.00416v5 (2018).
Vierstra, J. et al. Global reference mapping of human transcription factor footprints. Nature 583, 729–736 (2020).
Spitz, F. & Furlong, E. E. M. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Lambert, S. A. et al. Similarity regression predicts evolution of transcription factor sequence specificity. Nat. Genet. 51, 981–989 (2019).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Argelaguet, R. et al. Decoding gene regulation in the mouse embryo using single-cell multi-omics. Preprint at bioRxiv https://doi.org/10.1101/2022.06.15.496239 (2022).
Tournière, O. et al. NvPOU4/Brain3 functions as a terminal selector gene in the nervous system of the cnidarian Nematostella vectensis. Cell Rep. 30, 4473–4489 (2020).
Babonis, L. S. & Martindale, M. Q. PaxA, but not PaxC, is required for cnidocyte development in the sea anemone Nematostella vectensis. Evodevo 8, 14 (2017).
Danladi, B. et al. Conserved and lineage-restricted gene regulatory programs modulate developmental cnidocyte specification in Nematostella vectensis. Preprint at bioRxiv https://doi.org/10.1101/2025.05.08.652877 (2025)
Jahnel, S. M., Walzl, M. & Technau, U. Development and epithelial organisation of muscle cells in the sea anemone Nematostella vectensis. Front. Zool. 11, 44 (2014).
Nakanishi, N., Renfer, E., Technau, U. & Rentzsch, F. Nervous systems of the sea anemone Nematostella vectensis are generated by ectoderm and endoderm and shaped by distinct mechanisms. Development 139, 347–357 (2012).
Weirauch, M. T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Jolma, A. et al. Multiplexed massively parallel SELEX for characterization of human transcription factor binding specificities. Genome Res. 20, 861–873 (2010).
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013).
Jolma, A. et al. Perspectives on Codebook: sequence specificity of uncharacterized human transcription factors. Preprint at bioRxiv https://doi.org/10.1101/2024.11.11.622097 (2024).
Jolma, A. et al. GHT-SELEX demonstrates unexpectedly high intrinsic sequence specificity and complex DNA binding of many human transcription factors. Preprint at bioRxiv https://doi.org/10.1101/2024.11.11.618478 (2024).
Wang, M. et al. Distinct gene regulatory dynamics drive skeletogenic cell fate convergence during vertebrate embryogenesis. Nat. Commun. 16, 2187 (2025).
Hand, C. & Uhlinger, K. R. The culture, sexual and asexual reproduction, and growth of the sea anemone Nematostella vectensis. Biol. Bull. 182, 169–176 (1992).
Fritzenwanker, J. H. & Technau, U. Induction of gametogenesis in the basal cnidarian Nematostella vectensis (Anthozoa). Dev. Genes Evol. 212, 99–103 (2002).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959 (2017).
De Rop, F. V., et al. Hydrop enables droplet-based single-cell ATAC-seq and single-cell RNA-seq using dissolvable hydrogel beads. eLife 11, e73971 (2022).
Iglesias, M. Gastrula_Nvectensis_scATAC-seq, v.1, https://doi.org/10.17504/protocols.io.81wgbwrpogpk/v1 (2025).
Drokhlyansky, E., et al. The human and mouse enteric nervous system at single-cell resolution. Cell 182, 1606–1622 (2020).
Iglesias, M. Adult_Nvectensis_scATAC-seq, v.1, https://doi.org/10.17504/protocols.io.261gek5mwg47/v1 (2025).
Torres-Méndez, A. et al. A novel protein domain in an ancestral splicing factor drove the evolution of neural microexons. Nat. Ecol. Evol. 3, 691–701 (2019).
García-Castro, H. et al. ACME dissociation: a versatile cell fixation-dissociation method for single-cell transcriptomics. Genome Biol. 22, 89 (2021).
Yu, W., Uzun, Y., Zhu, Q., Chen, C. & Tan, K. scATAC-pro: a comprehensive workbench for single-cell chromatin accessibility sequencing data. Genome Biol. 21, 94 (2020).
Fletcher, C. & Pereira da Conceicoa, L. The genome sequence of the starlet sea anemone, Nematostella vectensis (Stephenson, 1935). Wellcome Open Res. 8, 79 (2023).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Lun, A. T. L. et al. EmptyDrops: distinguishing cells from empty droplets in droplet-based single-cell RNA sequencing data. Genome Biol. 20, 63 (2019).
van den Oord, J. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Zhang, Y. et al. Model-based analysis of ChIP–Seq (MACS). Genome Biol. 9, R137 (2008).
Huber, B. R. & Bulyk, M. L. Meta-analysis discovery of tissue-specific DNA sequence motifs from mammalian gene expression data. BMC Bioinform. 7, 229 (2006).
Machlab, D., et al. monaLisa: an R/Bioconductor package for identifying regulatory motifs. Bioinformatics 38, 2624–2625 (2022).
Lee, D. LS-GKM: a new gkm-SVM for large-scale datasets. Bioinformatics 32, 2196–2198 (2016).
Shrikumar, A., Greenside, P. & Kundaje, A. Learning important features through propagating activation differences. Preprint at https://arxiv.org/abs/1704.02685v2 (2017).
Renfer, E. & Technau, U. Meganuclease-assisted generation of stable transgenics in the sea anemone Nematostella vectensis. Nat. Protoc. 12, 1844–1854 (2017).
Elek, A. sebepedroslab/nvec-scatac: Nematostella_scATAC_atlas. Zenodo https://doi.org/10.5281/zenodo.17425383 (2025).
Acknowledgements
We thank I. Kim, A. de Mendoza, S. Montgomery, M. Irimia and N. Maeso for critical comments on the paper, as well as all members of the Sebe-Pedros group for discussion and suggestions. We thank F. Rentzsch for access to Nematostella Elav1::mOrange transgenic line. We are grateful to D. Cañas-Armenteros for taking care of Nematostella cultures and to the CRG Flow Cytometry, Genomics and ALMU facilities for technical support. Research in A.S-P. group has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (grant agreement number 851647) and the Spanish Ministry of Science, Innovation and Universities (PID2021-124757NB-I00 funded by MICIU /AEI /10.13039/501100011033 / FEDER, UE). We acknowledge support of the Spanish Ministry of Science and Innovation through the Centro de Excelencia Severo Ochoa (CEX2020-001049-S, MCIN/AEI /10.13039/501100011033), the Generalitat de Catalunya through the CERCA program and to the EMBL partnership. A.E. was supported by FPI PhD fellowship from the Spanish Ministry of Science and Innovation (PRE2019-087793SO funded by MCIN/AEI/10.13039/501100011033 and FSE+). M.I. has received funding from the European Union’s Horizon 2020 research and innovation program under the Marie Sklodowska-Curie grant agreement number 75442. X.G-B. was supported by the European Union’s H2020 research and innovation program under Marie Skłodowska-Curie grant agreement 101031767.
Author information
Authors and Affiliations
Contributions
A.S.-P. conceived and supervized the study. M.I. performed single-cell experiments and generated transgenic reporter lines. A.E. analysed scATAC-seq data, performed motif analyses and trained sequence models with the support of L.M. and S.A. G.Z. and X.G.-B. performed phylogenetic and comparative genomics analyses. A.E. created visualizations. A.E., M.I. and A.S.-P. interpreted the data and wrote the paper with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Maria Ina Arnone, Ferdinand Marlétaz and Juan Tena for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 scATAC-seq dataset QC, clustering and annotation.
a, Number of cells (top) and unique fragments per cell (bottom), b, scATAC-seq fragment size distribution for each sample. c, TSS enrichment signal for each sample. d, UMAP projection of single cells and metacells for gastrula dataset. e, UMAP projection of single cells and metacells for adult dataset. f, NJ clustering of metacells for adult and gastrula together, only for adult (g) and only for gastrula (h). i, Heatmap showing peak accessibility per cell type. j, Annotation transfer heatmap for adult scATAC-seq clusters. k, Annotation transfer heatmap for gastrula scATAC-seq clusters. l, Comparison of ATAC and RNA correlations for multiome (left) and separately profiled scATAC-seq and scRNA-seq data (right).
Extended Data Fig. 2 Cell type markers.
Comparison between accessibility scores and expression for selected marker genes.
Extended Data Fig. 3 Cis-regulatory element classification.
a, Decision tree used to classify CRE into different promoter types. b, Fraction of gastrula CREs classified as promoters and enhancers. Promoters are further classified as constitutive promoters (CP), specific promoters (SP) and alternative promoters (AP). Enhancers are classified based on their overlap with different genomic regions. c, Distance to the nearest TSS distributions for different types of CREs. d, Summary peak statistics. Number of peaks per gene (top-left) and number of genes that each peak gets assigned to (bottom-left), correlation across metacells between peak accessibility and expression of the genes they are assigned to (top-right), and co-accessibility across cell clusters of all pairs of peaks (bottom-right). e, V-plots showing tagmentation fragment size distributions (y-axis) at different distances (x-axis) around CP, SP and distal CREs in adult and gastrula stages.
Extended Data Fig. 4 Motif enrichment analysis.
Dotmap of representative motifs enriched in different cell types. Significantly enriched motifs (fold change > 1, padj < 0.001) are grouped by motif-motif similarity and top enriched representative motifs for each cluster are indicated (hypergeometric test, FDR adjustment for multiple testing).
Extended Data Fig. 5 Sequence models.
a, Five-fold cross-validation (CV) area under the curve (AUC, top) and test set AUC (bottom) for gkm-SVM classifiers trained on adult cnidocytes. b, Area under receiver operator curve (AUC ROC, top) and area under precision recall curve (AUC PR, bottom) for all cell type gkm-SVM classifiers. c, Spearman correlation for ChromBPNet predicted accessibility counts in test set peaks. d, Spearman correlation for crested predicted accessibility counts in test set peaks. e, Nucleotide importance scores for top scored CREs in different cell types.
Extended Data Fig. 6 Sequence motif discovery.
a, Heatmap showing pairwise motif similarities used to generate motif archetypes from enriched motifs. b, Examples of motif clusters. c, Examples of motif archetypes. d, Same as (a) and (b) for patterns discovered with sequence models. e, Number of motifs per archetype (left) and number of archetypes composed of motifs from different sources (right). f, Number of patterns per archetype (left) and number of archetypes composed of patterns from different sequence models (right). g, Comparison of motif enrichment fold change (left) and adjusted p-value (right) for archetypes versus best scoring motif in each archetype cluster (hypergeometric test, FDR adjustment for multiple testing). h, For all pattern archetypes and their most similar motif archetype, Jensen-Shannon divergence (JSD) calculated across the best pairwise alignment of archetypes (x-axis, JSDcomplete), and calculated across the best alignment spanning the length of shorter archetype (y-axis, JSDmin). Based on these two metrics, pattern archetypes are classified as being novel motifs, having novel context or resembling known motifs from motif enrichment analysis. i, Fraction of pattern archetype classes defined in h, Euler diagrams summarizing the source of pattern archetypes (that is sequence models) for each of these three categories are shown on top. j, Fraction of enriched motifs found with each of the sequence models, k, Co-occurrence network of pattern archetypes with contribution in cnidocytes. Size of the node reflects its cell type contribution, and width of the connection scales with the number of CREs in which two motifs co-occur. l, Correlation-based approach for assigning motifs to TFs. For each TF, we rank motifs based on correlation of motif activity to TF accessibility and expression, and assign it top ranking motif of the same structural class. m, Euler diagram showing TF coverage using different motif-to-TF assignment methods for expressed Nematostella TFs. n, Final motif assignment sources for expressed Nematostella TFs.
Extended Data Fig. 7
Examples of TF expression and TF motif activity correlations for selected marker genes.
Extended Data Fig. 8 Cell type gene regulatory networks.
a, Number of TFs in GRNs inferred for each broad cell type (top), number of genes targeted by each TF (middle), and fraction of overlapping target genes for each pair of TFs (bottom). b, Overlap of target genes for the same TF across cell types, plotted for groups of TFs active in different number of cell types. Selected TFs are highlighted on the plot and overlap of their target genes is shown as Euler diagrams below. c, Number of CREs per target gene (x-axis) compared to number of CREs of the same gene with any single TF motif (y-axis). Most TFs have binding motif in a single CREs of their target genes. d-g, Additional inferred GRN and TF connectivity measurements for neuro-secretory cell types: GATA/Islet neurons (d-e), Pou4/FoxL2 neurons (f-g) and gland cells (h-i). Asterisks highlight TFs known to be involved in neurosecretory development.
Supplementary information
Supplementary Tables 1–3 (download XLSX )
Table 1. scATAC-seq library information. Table 2. Annotation of putative CREs assigned to Nematostella genes in broad cell types. Peaks are classified as CP, SP, AP or NO (not a promoter, putative enhancer). Table 3. N. vectensis TF motif assignment.
Supplementary Data 1 (download XLSX )
Active TFs per cell type. Thresholds used for TF filtering: gene expression FC > 0.4 quantile of cell type expression values (exp_thrs column), ChromVAR deviation Z-score > 4.
Supplementary Data 2 (download XLSX )
Cell type-specific GRNs. Thresholds used for filtering cell type-specific networks: gene expression FC > 0.4 quantile of cell type expression values (exp_thrs column), gene accessibility > 0.4 quantile of cell type accessibility values (acc_thrs column), ChromVAR deviation Z-score > 4. Note that in Fig. 3 and Extended Data Fig. 8, for simplicity, cell type GRNs are further filtered for genes with expression FC > 1.2.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Elek, A., Iglesias, M., Mahieu, L. et al. Decoding cnidarian cell type gene regulation. Nat Ecol Evol 10, 140–153 (2026). https://doi.org/10.1038/s41559-025-02906-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41559-025-02906-1