Abstract
Severe fever with thrombocytopenia syndrome (SFTS) is an emerging tick-borne disease caused by a phlebovirus in the Bunyaviridae family. Infection can result in systemic inflammatory response syndrome with a high fatality rate, and there are currently no treatments or vaccines available. The microbiota has been implicated in host susceptibility to systemic viral infection and disease outcomes, but whether the gut microbiota is implicated in severe fever with thrombocytopenia syndrome virus (SFTSV) infection is unknown. Here, we analysed faecal and serum samples from patients with SFTS using 16S ribosomal RNA-sequencing and untargeted metabolomics, respectively. We found that the gut commensal Akkermansia muciniphila increased in relative abundance over the course of infection and was reduced in samples from deceased patients. Using germ-free or oral antibiotic-treated mice, we found that A. muciniphila produces the β-carboline alkaloid harmaline, which protects against SFTSV infection by suppressing NF-κB-mediated systemic inflammation. Harmaline indirectly modulated the virus-induced inflammatory response by specifically enhancing bile acid-CoA: amino acid N-acyltransferase expression in hepatic cells to increase conjugated primary bile acids, glycochenodeoxycholic acid and taurochenodeoxycholic acid. These bile acids induced transmembrane G-protein coupled receptor-5-dependent anti-inflammatory responses. These results indicate the probiotic potential of A. muciniphila in mitigating SFTSV infection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
16S rRNA sequence data are available in the Sequence Read Archive (SRA) under BioProject accession no. PRJNA890424 and PRJNA888451. RNA-seq data are available in the SRA under BioProject accession no. PRJNA889171. Source data can be accessed on FigShare: https://doi.org/10.6084/m9.figshare.21334422. Source data are provided with this paper.
Code availability
No custom code was used.
References
Skelly, A. N., Sato, Y., Kearney, S. & Honda, K. Mining the microbiota for microbial and metabolite-based immunotherapies. Nat. Rev. Immunol. 19, 305–323 (2019).
Rooks, M. G. & Garrett, W. S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 16, 341–352 (2016).
Alwin, A. & Karst, S. M. The influence of microbiota-derived metabolites on viral infections. Curr. Opin. Virol. 49, 151–156 (2021).
Woods Acevedo, M. A. & Pfeiffer, J. K. Microbiota–immune system interactions and enteric virus infection. Curr. Opin. Virol. 46, 15–19 (2020).
Grau, K. R. et al. The intestinal regionalization of acute norovirus infection is regulated by the microbiota via bile acid-mediated priming of type III interferon. Nat. Microbiol. 5, 84–92 (2019).
Steed, A. L. et al. The microbial metabolite desaminotyrosine protects from influenza through type I interferon. Science 357, 498–502 (2017).
Stefan, K. L., Kim, M. V., Iwasaki, A. & Kasper, D. L. Commensal microbiota modulation of natural resistance to virus infection. Cell 183, 1312–1324 e1310 (2020).
Winkler, E. S. et al. The intestinal microbiome restricts alphavirus infection and dissemination through a bile acid–type I IFN signaling axis. Cell 182, 901–918 e918 (2020).
Yu, X. J. et al. Fever with thrombocytopenia associated with a novel bunyavirus in China. N. Engl. J. Med. 364, 1523–1532 (2011).
Zhuang, L. et al. Transmission of severe fever with thrombocytopenia syndrome virus by Haemaphysalis longicornis ticks, China. Emerg. Infect. Dis. 24, 868–871 (2018).
Che, T. L. et al. The role of selenium in severe fever with thrombocytopenia syndrome: an integrative analysis of surveillance data and clinical data. Int. J. Infect. Dis. 122, 38–45 (2022).
Zhao, G. P. et al. Mapping ticks and tick-borne pathogens in China. Nat. Commun. 12, 1075 (2021).
Li, H. et al. Epidemiological and clinical features of laboratory-diagnosed severe fever with thrombocytopenia syndrome in China, 2011–17: a prospective observational study. Lancet Infect. Dis. 18, 1127–1137 (2018).
Kim, Y. R. et al. Severe fever with thrombocytopenia syndrome virus infection, South Korea, 2010. Emerg. Infect. Dis. 24, 2103–2105 (2018).
Takahashi, T. et al. The first identification and retrospective study of severe fever with thrombocytopenia syndrome in Japan. J. Infect. Dis. 209, 816–827 (2014).
Tran, X. C. et al. Endemic severe fever with thrombocytopenia syndrome, Vietnam. Emerg. Infect. Dis. 25, 1029–1031 (2019).
Win, A. M. et al. Genotypic heterogeneity of Orientia tsutsugamushi in scrub typhus patients and thrombocytopenia syndrome co-infection, Myanmar. Emerg. Infect. Dis. 26, 1878–1881 (2020).
Luo, L. M. et al. Haemaphysalis longicornis ticks as reservoir and vector of severe fever with thrombocytopenia syndrome virus in China. Emerg. Infect. Dis. 21, 1770–1776 (2015).
Liu, Q., He, B., Huang, S. Y., Wei, F. & Zhu, X. Q. Severe fever with thrombocytopenia syndrome, an emerging tick-borne zoonosis. Lancet Infect. Dis. 14, 763–772 (2014).
Yang, X. L. et al. The intestinal microbiome primes host innate immunity against enteric virus systemic infection through type I interferon. mBio. 12, e00366–21 (2021).
Li, S. et al. SFTSV infection induces BAK/BAX-dependent mitochondrial DNA release to trigger NLRP3 inflammasome activation. Cell Rep. 30, 4370–4385 e4377 (2020).
Chen, M. L., Takeda, K. & Sundrud, M. S. Emerging roles of bile acids in mucosal immunity and inflammation. Mucosal Immunol. 12, 851–861 (2019).
Thomas, C., Pellicciari, R., Pruzanski, M., Auwerx, J. & Schoonjans, K. Targeting bile-acid signalling for metabolic diseases. Nat. Rev. Drug Discov. 7, 678–693 (2008).
Ridlon, J. M., Kang, D. J. & Hylemon, P. B. Bile salt biotransformations by human intestinal bacteria. J. Lipid Res. 47, 241–259 (2006).
Yoon, H. S. et al. Akkermansia muciniphila secretes a glucagon-like peptide-1-inducing protein that improves glucose homeostasis and ameliorates metabolic disease in mice. Nat. Microbiol. 6, 563–573 (2021).
Li, H. et al. Single-cell landscape of peripheral immune responses to fatal SFTS. Cell Rep. 37, 110039 (2021).
Qu, B. et al. Suppression of the interferon and NF-kappaB responses by severe fever with thrombocytopenia syndrome virus. J. Virol. 86, 8388–8401 (2012).
Jia, W., Xie, G. & Jia, W. Bile acid–microbiota crosstalk in gastrointestinal inflammation and carcinogenesis. Nat. Rev. Gastroenterol. Hepatol. 15, 111–128 (2018).
Kawamata, Y. et al. A G protein-coupled receptor responsive to bile acids. J. Biol. Chem. 278, 9435–9440 (2003).
Vaquero, J., Monte, M. J., Dominguez, M., Muntané, J. & Marin, J. J. Differential activation of the human farnesoid X receptor depends on the pattern of expressed isoforms and the bile acid pool composition. Biochem. Pharm. 86, 926–939 (2013).
Deriu, E. et al. Influenza virus affects intestinal microbiota and secondary Salmonella infection in the gut through type I interferons. PLoS Pathog. 12, e1005572 (2016).
Hu, X. et al. Akkermansia muciniphila improves host defense against influenza virus infection. Front. Microbiol. 11, 586476 (2020).
Derrien, M., Vaughan, E. E., Plugge, C. M. & de Vos, W. M. Akkermansia muciniphila gen. nov., sp. nov., a human intestinal mucin-degrading bacterium. Int. J. Syst. Evol. Microbiol. 54, 1469–1476 (2004).
Zhang, T., Li, Q., Cheng, L., Buch, H. & Zhang, F. Akkermansia muciniphila is a promising probiotic. Microb. Biotechnol. 12, 1109–1125 (2019).
Pierre, J. F. et al. Activation of bile acid signaling improves metabolic phenotypes in high-fat diet-induced obese mice. Am. J. Physiol. Gastrointest. Liver Physiol. 311, G286–G304 (2016).
Sayin, S. I. et al. Gut microbiota regulates bile acid metabolism by reducing the levels of tauro-beta-muricholic acid, a naturally occurring FXR antagonist. Cell Metab. 17, 225–235 (2013).
Khan, F. A., Maalik, A., Iqbal, Z. & Malik, I. Recent pharmacological developments in beta-carboline alkaloid ‘harmaline’. Eur. J. Pharm. 721, 391–394 (2013).
Nakagawa, Y., Suzuki, T., Ishii, H., Ogata, A. & Nakae, D. Mitochondrial dysfunction and biotransformation of beta-carboline alkaloids, harmine and harmaline, on isolated rat hepatocytes. Chem. Biol. Interact. 188, 393–403 (2010).
Zhao, T. et al. Metabolic pathways of the psychotropic-carboline alkaloids, harmaline and harmine, by liquid chromatography/mass spectrometry and NMR spectroscopy. Food Chem. 134, 1096–1105 (2012).
Chen, Q., Zhang, S. & Xie, Y. Characterization of a new microbial Pictet-Spenglerase NscbB affording the β-carboline skeletons from Nocardiopsis synnemataformans DSM 44143. J. Biotechnol. 281, 137–143 (2018).
Pols, T. W. et al. TGR5 activation inhibits atherosclerosis by reducing macrophage inflammation and lipid loading. Cell Metab. 14, 747–757 (2011).
Wen, A. Y., Sakamoto, K. M. & Miller, L. S. The role of the transcription factor CREB in immune function. J. Immunol. 185, 6413–6419 (2010).
Li, H. et al. Calcium channel blockers reduce severe fever with thrombocytopenia syndrome virus (SFTSV) related fatality. Cell Res. 29, 739–753 (2019).
Zhande, R. et al. Dephosphorylation by default, a potential mechanism for regulation of insulin receptor substrate-1/2, Akt, and ERK1/2. J. Biol. Chem. 281, 39071–39080 (2006).
Liu, W. et al. Case–fatality ratio and effectiveness of ribavirin therapy among hospitalized patients in China who had severe fever with thrombocytopenia syndrome. Clin. Infect. Dis. 57, 1292–1299 (2013).
Zhang, Q. et al. Influenza infection elicits an expansion of gut population of endogenous Bifidobacterium animalis which protects mice against infection. Genome Biol. 21, 99 (2020).
Magoc, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Acknowledgements
We thank W. Han (Department of Medical Oncology, Sir Run Run Shaw Hospital, College of Medicine) for providing a valuable TGR5 knockout mouse strain. This work was supported by grants from the National Key Research and Development Plan of China (2021YFC230200–02) to W.L., the National Natural Science Foundation of China (No. 32172864 and No. 82172270) to S.J.Z. and H.L., and the National Natural Science Fund for Distinguished Young Scholars (No. 81825019) to W.L.
Author information
Authors and Affiliations
Contributions
S.J.Z. and W.L. designed the experiments. J.X., H.L., X.Z., T.Y., M.Y., Y.Z. and S.C. performed the experiments. J.X., H.L., X.Z. and T.Y. conducted the bioinformatics analysis. N.C. and C.Y. collected samples and data. J.L. commented on and revised the drafts of the manuscript. S.J.Z., W.L., J.X., H.L. and T.Y. wrote the paper. S.J.Z. and W.L. supervised the research, coordination and strategy.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The intestinal microbiota of surviving SFTSV-infected individuals.
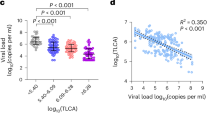
(a) Relative abundance of phyla among the four groups. (b) Relative abundance of Lactobacillus, Parabacteroides and Enterococcus among the four groups (HC n = 19, Non-SF n = 176, SF-S n = 233, SF-D n = 27). (c) Serum concentration of TNF-α among the four groups and the association with the relative abundance of Akkermansia (HC n = 10, non-SF n = 25, SF-S n = 107, SF-D n = 21). HC, healthy controls; Non-SF, febrile patients without SFTSV infection; SF-S, surviving patients with SFTSV infection; SF-D, deceased patients with SFTSV infection. (d) Schematic representation of treatment with Abx, FMT, bacteria, HAL, acetate, propionate or TCDCA. The two-sided P values were examined by Student’s t test and data were presented as mean values ± SD (b, and c). R2 and exact two-sided P values calculated by Pearson’s test are shown (c).
Extended Data Fig. 2 Microbiota reconstitution with A. muciniphila prevents SFTSV early infection and inhibits resulting systemic inflammatory responses.
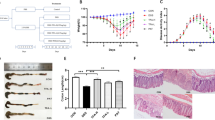
(a) qPCR of 16 s rRNA genomic copies in faeces from PBS-treated, Abx-treated or FMT-transferred mice at 2 days after transfer (n = 6). (b) qPCR of IL-1β and IL-6 mRNA (right) and SFTSV RNA (left) in spleens from Abx-treated mice with or without FMT (faeces from recovered SFTSV mice) and infected with SFTSV at 3 days post-infection (n = 6). (c) H. E. staining of lung, liver or spleen cross sections from Abx-treated mice with or without FMT (faeces from recovered SFTSV mice) and infected with SFTSV at 3 dpi. Boxed areas are magnified immediately in the top right corner. (d) qPCR of A. muciniphila genomic copies in faeces from SFTSV infected mice at 0- and 3-days post infection (n = 4). (e) qPCR of A. muciniphila, L. reuteri or E. faecalis genomic copies in faeces from Abx-treated or bacteria-colonized mice at 2 days post colonization (A. muciniphila n = 4, L. reuteri and E. faecalis n = 5). (f) qPCR of SFTSV RNA in the spleen, liver or lung from PBS-treated, Abx-treated or A. muciniphila-colonized mice infected with SFTSV at 3 and 5 dpi (n = 8). (g) IFA of spleen sections from Abx-treated mice colonized with or without A. muciniphila and infected with SFTSV at 3 dpi. SFTSV protein NP and IL-6 protein were double stained with the respective antibodies. (h) H. E. staining of lung, liver or spleen cross sections from Abx-treated mice colonized with or without A. muciniphila and infected with SFTSV at 5 dpi. Boxed areas are magnified immediately in the top right corner. The two-sided P values were examined by Student’s t test and data were presented as mean values ± SD (a, b, d, e and f).
Extended Data Fig. 3 A. muciniphila-associated conjugated primary bile acids GCDCA and TCDCA suppress SFTSV-induced inflammatory responses in human PBMCs in vitro.
(a) Relative mRNA levels of IL-1β and IL-6 in SFTS patients (n = 3) and (b) healthy donor PBMCs treated with GCDCA and then infected with SFTSV (MOI 1) at 24 hpi (n = 6). Data were presented as mean values ± SD (a and b).
Extended Data Fig. 4 A. muciniphila colonization in antibiotic-pretreated mice upregulates TCDCA secretion.
(a) PCA of serum from four groups: PBS-treated, Abx-treated and A. muciniphila-colonized mice infected with SFTSV at 3 dpi and nontreated mice. (b) KEGG analysis of differentially regulated metabolites between the PBS group and the Abx group. The two-sided P values were examined by Student’s t test.
Extended Data Fig. 5 TCDCA protects Abx mice by prohibiting SFTSV-induced systemic inflammation.
(a) qPCR of IL-1β or TNF-α mRNA and (b) SFTSV RNA in mouse PMBCs pretreated with TCDCA, GCDCA, TCA, TDCA or T-α-MCA and then infected with SFTSV (MOI 1) at 24 hpi (IL-1β n = 4, TNF-α and SFTSV RNA n = 6). (C) Relative mRNA transcripts of proinflammatory cytokines in SFTSV-infected mouse PBMCs treated with different doses of TCDCA and (d) respective viral loads at 24 hpi (n = 4). (e) Serum concentration of TCDCA in Abx mice treated with TCDCA (Abx n = 4, TCDCA n = 2). (f) qPCR of SFTSV RNA in the spleen and liver from PBS-treated and Abx-treated mice treated with or without TCDCA and infected with SFTSV at 3 and 5 dpi (Abx 3dpi n = 7, Abx 5dpi n = 5, TCDCA 3dpi n = 8, TCDCA 3dpi n = 7). The two-sided P values were examined by Student’s t test (a, and c). Data were presented as mean values ± SD (a to f).
Extended Data Fig. 6 A. muciniphila-associated metabolites enhance the expression of BAAT both in vivo and in vitro.
(a) Total hepatic cholesterol level of PBS-treated, Abx-treated or A. muciniphila-colonized mice at 5 days post-colonization (n = 5). (b) qPCR of Asbt mRNA in the ileum of PBS-treated, Abx-treated or A. muciniphila-colonized mice at 5 days post-colonization (n = 5). (c) qPCR of BAAT mRNA in liver from PBS-treated, Abx-treated, A. muciniphila-colonized or L. reuteri-colonized mice at 5 days post-colonization (n = 5). (d) Quantification of the proteins in Fig. 5a. The protein levels of enzymes in the bile acid biosynthesis pathways were normalized to those of ACTIN. The amount of each protein in the fasted state was defined as 1 (n = 3). (e) qPCR of BAAT mRNA in liver from GF mice colonized with or without A. muciniphila at 5 days post-colonization (n = 6). (f) Quantification of the proteins in Fig. 5b. The protein levels of enzymes in the bile acid biosynthesis pathways were normalized to those of TUBULIN. The amount of each protein in the fasted state was defined as 1 (n = 3). (g) Serum bile acid concentration of Abx-treated, A. muciniphila-colonized or L. reuteri-colonized mice at 5 days post colonization (n = 4). (h) qPCR of BAAT mRNA in Huh-7 cells treated with different fractions of A. muciniphila culture supernatants at 24 h (n = 6). (i) qPCR of BAAT mRNA in Huh-7 cells treated with different doses of 10 kDa or less filtrate with or without proteinase K digestion and the correlations with the content of 10 kDa or less filtrate (n = 6). (j) The content of acetate or propionate in the supernatant of A. muciniphila (n = 4). The two-sided P values were examined by Student’s t test and data were presented as mean values ± SD (a–j).
Extended Data Fig. 7 The A. muciniphila metabolite harmaline upregulates the expression of BAAT.
(a) Relative abundance changes of positive or negative metabolites in 10 kDa or less filtrate of A. muciniphila cultured supernatants. (b) The correlations between the content of HAL and A. muciniphila cell numbers. (c, d) qPCR of BAAT mRNA in (c) Huh-7 or (d) MPH cells treated with different doses of HAL or butyrate at 24 h (n = 5). (e) qPCR of BAAT mRNA in MPH cells transfected with siNC or siBAAT at 24 h (n = 6). (f) qPCR of BAAT mRNA in liver from PBS-treated, Abx-treated, HAL-treated or acetate-treated mice at 5 days post-treatment (n = 6). (g) Quantification of the proteins in Fig. 5g. The proteins of the bile acid biosynthesis pathway were normalized to that of ACTIN. The amount of each protein in the fasted state was defined as 1 (n = 3). The two-sided P values were examined by Student’s t test and data were presented as mean values ± SD (c–g).
Extended Data Fig. 8 The A. muciniphila metabolite harmaline prohibits systemic inflammatory responses in Abx-treated and GF mice.
(a) Serum bile acid concentration of Abx- and HAL-treated mice and (b) GF mice treated with or without A. muciniphila or HAL at 5 days post-colonization (n = 8). (c) Serum HAL concentration of PBS-treated, Abx-treated, A. muciniphila-colonized, L. reuteri-colonized, HAL-treated mice at 2 days post colonization (n = 3). (d) Serum HAL concentrations of recovered patients with SFTSV infection and recovered febrile patients without SFTSV infection. Serum samples were collected approximately two weeks after symptom onset (n = 13). (e) IFA images of spleen sections from GF mice treated with or without HAL at 3 dpi. SFTSV protein NP and IL-6 protein were double stained with the respective antibodies. (f) H. E staining of lung, liver or spleen cross sections from GF mice treated with or without HAL and infected with SFTSV at 3 dpi. Boxed areas are magnified immediately in the top right corner. (g) Relative mRNA levels of IL-1β, IL-6, TNF-α (left) and SFTSV RNA (right) in mouse PBMCs that were pretreated with 10 μM HAL and then infected with SFTSV (MOI 1) at 24 hpi or HAL was added to the PBMCs simultaneously with SFTSV infection (n = 6). (h) Expression of Baat and actin in the livers of Abx-treated wild-type B6 mice transiently transfected with siNC or siBAAT (N/P ratio is 6 to 8). The two-sided P values were examined by Student’s t test (a, b and d). Data were presented as mean values ± SD (a–d and g).
Extended Data Fig. 9 GCDCA suppresses SFTSV-induced inflammatory cytokine expression in THP-1 cells in a dose-dependent manner.
(a) Relative mRNA levels of IL-1β and IL-6 in THP-1PMA cells that were pretreated with 50, 100, or 150 μM GCDCA and then infected with SFTSV (MOI 1) at 24 hpi (n = 3). (b) Relative protein levels of IL-1β and IL-6 in THP-1PMA cells pretreated with 100 μM GCDCA and then infected with SFTSV (MOI 1) at 24 hpi (n = 3). (c) Relative mRNA levels of IL-1β, IL-6 (n = 3) and (d) SFTSV RNA in THP-1PMA cells (n = 6) pretreated with 10, 50, or 100 μM CDCA and then infected with SFTSV (MOI 1) at 24 hpi. The two-sided P values were examined by Student’s t test (a and c). Data were presented as mean values ± SD (a to d).
Extended Data Fig. 10 GCDCA does not hinder SFTSV-induced inflammatory responses via the TLR8-MyD88 or NLRP3-inflammasome signalling pathways.
THP-1PMA cells were pretreated with 100 μM GCDCA and then infected with SFTSV (MOI 1) for 24 hours. (a) Relative mRNA levels of IL1B and IL6 in TLR8- or MyD88-knockdown THP-1PMA cells (n = 6). (b) Western blot of pro-IL-1β, TLR8, MyD88 and tubulin in TLR8- or MyD88-knockdown THP-1PMA cells. (c) Relative protein levels of P50, P65 and phos-P65 in THP-1PMA cells (n = 3). (d) Western blot of MyD88, IKKβ, NLRP3, ASC, CASP1 and tubulin in THP-1PMA cells. (e) SFTSV RNA level in TGR5 knockdown THP-1PMA cells (n = 6). (f) Relative protein levels of pro-IL-1β, TGR5, p50, p65 and phos-p65 in TGR5 knockdown THP-1PMA cells (n = 3). (g and h) THP-1PMA cells were pretreated with 100 μM GCDCA and then infected with SFTSV (MOI 1) for 24 hours. (g) Relative mRNA levels of IL1B and IL6 in FXR knockdown THP-1PMA cells (n = 6). (h) Western blot of pro-IL-1β, FXR and tubulin in FXR knockdown THP-1PMA cells. (i) Serum TCDCA concentration of Abx-treated TGR5 CKO mice with or without TCDCA treatment at 5 dpi. (j) qPCR of A. muciniphila genomic copies in faeces from Abx-treated or bacteria-colonized mice at 2 days post colonization (n = 3). (k) Schematic illustration of the Akkermansia muciniphila metabolite harmaline protecting against severe fever with thrombocytopenia syndrome through the bile acid-TGR5-NF-κB axis. The two-sided P values were examined by using Student’s t test (a, g and j). Data were presented as mean values ± SD (a, c, e–g and j).
Supplementary information
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Unprocessed western blots.
Source Data Fig. 6 (download PDF )
Statistical source data.
Source Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Unprocessed western blots.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download PDF )
Statistical source data.
Source Data Extended Data Fig. 10 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xie, J., Li, H., Zhang, X. et al. Akkermansia muciniphila protects mice against an emerging tick-borne viral pathogen. Nat Microbiol 8, 91–106 (2023). https://doi.org/10.1038/s41564-022-01279-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-022-01279-6
This article is cited by
-
Navigating the duality of Akkermansia muciniphila
Nature Microbiology (2026)
-
Intermittent fasting and metabolic dysfunction-associated steatotic liver disease: the potential role of the gut-liver axis
Cell & Bioscience (2025)
-
Short-term and long-term high-fat diet promote metabolic disorder through reprogramming mRNA m6A in white adipose tissue by gut microbiota
Microbiome (2025)
-
Isoliquiritigenin ameliorates Parkinson’s disease via gut microbiota remodeling: potential mediating role of Parabacteroides distasonis
Journal of Translational Medicine (2025)
-
Gut microbiota-derived isovaleric acid ameliorates influenza virus infection via gut-lung axis
npj Biofilms and Microbiomes (2025)