Abstract
As freshwater lakes undergo rapid anthropogenic change, long-term studies reveal key microbial dynamics, evolutionary shifts and biogeochemical interactions, yet the vital role of viruses remains overlooked. Here, leveraging a 20 year time series from Lake Mendota, WI, USA, we characterized 1.3 million viral genomes across time, seasonality and environmental factors. Double-stranded DNA phages from the class Caudoviricetes dominated the community. We identified 574 auxiliary metabolic gene families representing over 140,000 auxiliary metabolic genes, including important genes such as psbA (photosynthesis), pmoC (methane oxidation) and katG (hydrogen peroxide decomposition), which were consistently present and active across decades and seasons. Positive associations and niche differentiation between virus–host pairs, including keystone Cyanobacteria, methanotrophs and Nanopelagicales, emerged during seasonal changes. Inorganic carbon and ammonium influenced viral abundances, underscoring viral roles in both ‘top-down’ and ‘bottom-up’ interactions. Evolutionary processes favoured fitness genes, reduced genomic heterogeneity and dominant sub-populations. This study transforms understanding of viral ecology and evolution in Earth’s microbiomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The metagenomic datasets (including assemblies and raw reads) are all available under JGI Proposal ID 504350 at the platform of the Integrated Microbial Genomes & Microbiomes system (https://img.jgi.doe.gov/m/). The retrieved viral genomes were deposited in NCBI Bioproject PRJNA1130067. At the same time, the TYMEFLIES viral genomes and related properties, including annotations for viral proteins, taxonomic classification, host prediction and virus clustering results, are available via Figshare at https://figshare.com/articles/dataset/TYMEFLIES_vMAGs_and_related_properties/24915750 (ref. 84). The raw environmental parameter spreadsheets are available in the Environmental Data Initiative (https://edirepository.org/) database.
Code availability
Codes used in this project are available via GitHub at https://github.com/AnantharamanLab/TYMEFLIES_Viral.
References
Tran, P. Q. & Anantharaman, K. Biogeochemistry goes viral: towards a multifaceted approach to study viruses and biogeochemical cycling. mSystems 6, e01138–21 (2021).
Rosenwasser, S., Ziv, C., van Creveld, S. G. & Vardi, A. Virocell metabolism: metabolic innovations during host–virus interactions in the ocean. Trends Microbiol. 24, 821–832 (2016).
Camargo, A. P. et al. IMG/VR v4: an expanded database of uncultivated virus genomes within a framework of extensive functional, taxonomic, and ecological metadata. Nucleic Acids Res. 51, D733–D743 (2023).
McMahon, K. D. & Newton, R. J. Pelagic bacteria, archaea, and viruses. in Wetzel’s Limnology 4th edn (eds Jones, I. D. & Smol, J. P.) Ch. 23, 705–757 (Academic, 2024).
Karl, D. M. & Church, M. J. Microbial oceanography and the Hawaii Ocean Time-series programme. Nat. Rev. Microbiol. 12, 699–713 (2014).
Steinberg, D. K. et al. Overview of the US JGOFS Bermuda Atlantic Time-series Study (BATS): a decade-scale look at ocean biology and biogeochemistry. Deep Sea Res. II 48, 1405–1447 (2001).
Shade, A. et al. Interannual dynamics and phenology of bacterial communities in a eutrophic lake. Limnol. Oceanogr. 52, 487–494 (2007).
Bendall, M. L. et al. Genome-wide selective sweeps and gene-specific sweeps in natural bacterial populations. ISME J. 10, 1589–1601 (2016).
Roux, S. et al. Ecogenomics of virophages and their giant virus hosts assessed through time series metagenomics. Nat. Commun. 8, 858 (2017).
Bragg, J. G. & Chisholm, S. W. Modeling the fitness consequences of a cyanophage-encoded photosynthesis gene. PLoS ONE 3, e3550 (2008).
Mann, N. H., Cook, A., Millard, A., Bailey, S. & Clokie, M. Bacterial photosynthesis genes in a virus. Nature 424, 741 (2003).
Kieft, K. et al. Ecology of inorganic sulfur auxiliary metabolism in widespread bacteriophages. Nat. Commun. 12, 1–16 (2021).
Puxty, R. J., Evans, D. J., Millard, A. D. & Scanlan, D. J. Energy limitation of cyanophage development: implications for marine carbon cycling. ISME J. 12, 1273–1286 (2018).
Lindell, D., Jaffe, J. D., Johnson, Z. I., Church, G. M. & Chisholm, S. W. Photosynthesis genes in marine viruses yield proteins during host infection. Nature 438, 86–89 (2005).
Hellweger, F. L. Carrying photosynthesis genes increases ecological fitness of cyanophage in silico. Environ. Microbiol. 11, 1386–1394 (2009).
Chen, L.-X. et al. Large freshwater phages with the potential to augment aerobic methane oxidation. Nat. Microbiol. 5, 1504–1515 (2020).
Lindell, D. et al. Transfer of photosynthesis genes to and from Prochlorococcus viruses. Proc. Natl Acad. Sci. USA 101, 11013–11018 (2004).
Ruiz-Perez, C. A., Tsementzi, D., Hatt, J. K., Sullivan, M. B. & Konstantinidis, K. T. Prevalence of viral photosynthesis genes along a freshwater to saltwater transect in Southeast USA. Environ. Microbiol. Rep. 11, 672–689 (2019).
Sullivan, M. B. et al. Prevalence and evolution of core photosystem II genes in marine cyanobacterial viruses and their hosts. PLoS Biol. 4, e234 (2006).
Roux, S. et al. Ecology and evolution of viruses infecting uncultivated SUP05 bacteria as revealed by single-cell- and meta-genomics. eLife 3, e03125 (2014).
Anantharaman, K. et al. Sulfur oxidation genes in diverse deep-sea viruses. Science 344, 757–760 (2014).
Ahlgren, N. A., Fuchsman, C. A., Rocap, G. & Fuhrman, J. A. Discovery of several novel, widespread, and ecologically distinct marine Thaumarchaeota viruses that encode amoC nitrification genes. ISME J. 13, 618–631 (2019).
Cassman, N. et al. Oxygen minimum zones harbour novel viral communities with low diversity. Environ. Microbiol. 14, 3043–3065 (2012).
Rohwer, R. R., Hale, R. J., Vander Zanden, M. J., Miller, T. R. & McMahon, K. D. Species invasions shift microbial phenology in a two-decade freshwater time series. Proc. Natl Acad. Sci. USA 120, e2211796120 (2023).
Rohwer, R. R. et al. Two decades of bacterial ecology and evolution in a freshwater lake. Nat. Microbiol. https://doi.org/10.1038/s41564-024-01888-3 (2025).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585 (2021).
Sharon, I. et al. Comparative metagenomics of microbial traits within oceanic viral communities. ISME J. 5, 1178–1190 (2011).
Heyerhoff, B., Engelen, B. & Bunse, C. Auxiliary metabolic gene functions in pelagic and benthic viruses of the Baltic Sea. Front. Microbiol. 13, 863620 (2022).
Kieft, K. et al. Virus-associated organosulfur metabolism in human and environmental systems. Cell Rep. 36, 109471 (2021).
Nguyen, A. A. et al. CpeT is the phycoerythrobilin lyase for Cys-165 on β-phycoerythrin from Fremyella diplosiphon and the chaperone-like protein CpeZ greatly improves its activity. Biochim. Biophys. Acta 1861, 148284 (2020).
Puxty, R. J., Millard, A. D., Evans, D. J. & Scanlan, D. J. Viruses inhibit CO2 fixation in the most abundant phototrophs on earth. Curr. Biol. 26, 1585–1589 (2016).
Thompson, L. R. et al. Phage auxiliary metabolic genes and the redirection of cyanobacterial host carbon metabolism. Proc. Natl Acad. Sci. USA 108, E757–E764 (2011).
Sachindra, N. M. et al. Radical scavenging and singlet oxygen quenching activity of marine carotenoid fucoxanthin and its metabolites. J. Agric. Food Chem. 55, 8516–8522 (2007).
Parada, V., Herndl, G. J. & Weinbauer, M. G. Viral burst size of heterotrophic prokaryotes in aquatic systems. J. Mar. Biol. Assoc. UK 86, 613–621 (2006).
Middelboe, M. Bacterial growth rate and marine virus–host dynamics. Microb. Ecol. 40, 114–124 (2000).
Burson, A., Stomp, M., Greenwell, E., Grosse, J. & Huisman, J. Competition for nutrients and light: testing advances in resource competition with a natural phytoplankton community. Ecology 99, 1108–1118 (2018).
Lee, S. et al. Methane-derived carbon flows into host–virus networks at different trophic levels in soil. Proc. Natl Acad. Sci. USA 118, e2105124118 (2021).
Yoon, H., Kim, H.-C. & Kim, S. Long-term seasonal and temporal changes of hydrogen peroxide from cyanobacterial blooms in fresh waters. J. Environ. Manage. 298, 113515 (2021).
Kim, S., Kang, I., Seo, J.-H. & Cho, J.-C. Culturing the ubiquitous freshwater actinobacterial acI lineage by supplying a biochemical ‘helper’ catalase. ISME J. 13, 2252–2263 (2019).
Zheng, Q., Jiao, N., Zhang, R., Chen, F. & Suttle, C. A. Prevalence of psbA-containing cyanobacterial podoviruses in the ocean. Sci. Rep. 3, 3207 (2013).
Warwick-Dugdale, J., Buchholz, H. H., Allen, M. J. & Temperton, B. Host-hijacking and planktonic piracy: how phages command the microbial high seas. Virol. J. 16, 15 (2019).
Nei, M., Suzuki, Y. & Nozawa, M. The neutral theory of molecular evolution in the genomic era. Annu. Rev. Genomics Hum. Genet. 11, 265–289 (2010).
Howard-Varona, C. et al. Phage-specific metabolic reprogramming of virocells. ISME J. https://doi.org/10.1038/s41396-019-0580-z (2020).
Agarkova, I. V., Dunigan, D. D. & Van Etten, J. L. Virion-associated restriction endonucleases of chloroviruses. J. Virol. 80, 8114–8123 (2006).
Jeudy, S. et al. The DNA methylation landscape of giant viruses. Nat. Commun. 11, 2657 (2020).
Cohan, F. M. Bacterial species and speciation. Syst. Biol. 50, 513–524 (2001).
Cohan, F. M. & Perry, E. B. A systematics for discovering the fundamental units of bacterial diversity. Curr. Biol. 17, R373–R386 (2007).
Shapiro, B. J. & Polz, M. F. Microbial speciation. Cold Spring Harb. Perspect. Biol. 7, a018143 (2015).
Hatfull, G. F. Bacteriophage genomics. Curr. Opin. Microbiol. 11, 447–453 (2008).
Mavrich, T. N. & Hatfull, G. F. Bacteriophage evolution differs by host, lifestyle and genome. Nat. Microbiol. 2, 17112 (2017).
Schulz, E. C. et al. Structural basis for the recognition and cleavage of polysialic acid by the bacteriophage K1F tailspike protein EndoNF. J. Mol. Biol. 397, 341–351 (2010).
Bork, P. & Doolittle, R. F. Proposed acquisition of an animal protein domain by bacteria. Proc. Natl Acad. Sci. USA 89, 8990–8994 (1992).
Parisien, A., Allain, B., Zhang, J., Mandeville, R. & Lan, C. Q. Novel alternatives to antibiotics: bacteriophages, bacterial cell wall hydrolases, and antimicrobial peptides. J. Appl. Microbiol. 104, 1–13 (2008).
Ahern, K. S., Ahern, C. R. & Udy, J. W. In situ field experiment shows Lyngbya majuscula (cyanobacterium) growth stimulated by added iron, phosphorus and nitrogen. Harmful Algae 7, 389–404 (2008).
Kang, R. et al. Interactions between organic and inorganic carbon sources during mixotrophic cultivation of Synechococcus sp. Biotechnol. Lett. 26, 1429–1432 (2004).
Muñoz-Marín, M. D. C., López-Lozano, A., Moreno-Cabezuelo, J. Á., Díez, J. & García-Fernández, J. M. Mixotrophy in cyanobacteria. Curr. Opin. Microbiol. 78, 102432 (2024).
Flores, E. & Herrero, A. Nitrogen assimilation and nitrogen control in cyanobacteria. Biochem. Soc. Trans. 33, 164–167 (2005).
Rohwer, R. R. & McMahon, K. D. Lake iTag measurements over nineteen years, introducing the limony dataset. Preprint at bioRxiv https://doi.org/10.1101/2022.08.04.502869 (2022).
Breitbart, M., Bonnain, C., Malki, K. & Sawaya, N. A. Phage puppet masters of the marine microbial realm. Nat. Microbiol. 3, 754–766 (2018).
Rohwer, R. R. & McMahon, K. D. Lake Mendota Microbial Observatory temperature, dissolved oxygen, pH, and conductivity data, 2006-present. Environmental Data Initiative https://doi.org/10.6073/PASTA/7E533C197ED8EBD27777A89A2C8D7DFE (2022).
Magnuson, J. J., Carpenter, S. R. & Stanley, E. H. North temperate lakes LTER: physical limnology of primary study lakes 1981 - current. Environmental Data Initiative https://doi.org/10.6073/PASTA/316203040EA1B8ECE89673985AB431B7 (2021).
Magnuson, J., Carpenter, S. & Stanley, E. North temperate lakes LTER: high frequency water temperature data - Lake Mendota Buoy 2006 - current. Environmental Data Initiative https://doi.org/10.6073/PASTA/8CEFF296AD68FA8DA6787076E0A5D992 (2020).
Robertson, D. Lake Mendota water temperature secchi depth snow depth ice thickness and meterological conditions 1894 - 2007. Environmental Data Initiative https://doi.org/10.6073/PASTA/F20F9A644BD12E4B80CB288F1812C935 (2016).
Magnuson, J. J., Carpenter, S. R. & Stanley, E. H. Lake Mendota multiparameter sonde profiles: 2017 - current. Environmental Data Initiative https://doi.org/10.6073/PASTA/5F15BF453851987FC030B2F07A110B21 (2021).
Rohwer, R. R. & McMahon, K. D. Lake Mendota microbial observatory secchi disk measurements 2012-present. Environmental Data Initiative https://doi.org/10.6073/PASTA/3B650E19D28CBC7B9ED631F0A7878033 (2022).
Magnuson, J. J., Carpenter, S. R. & Stanley, E. H. North temperate lakes LTER: secchi disk depth; other auxiliary base crew sample data 1981 - current. Environmental Data Initiative https://doi.org/10.6073/PASTA/26FA98B39F9758FDA2109021F5B88076 (2021).
Magnuson, J. J., Carpenter, S. R. & Stanley, E. H. North temperate lakes LTER: chemical limnology of primary study lakes: major ions 1981 - current. Environmental Data Initiative https://doi.org/10.6073/pasta/bb563f16c7338fdb3ddf82057ef43cc6 (2023).
Magnuson, J. J., Carpenter, S. R. & Stanley, E. H. North temperate lakes LTER: chemical limnology of primary study lakes: nutrients, pH and carbon 1981 - current. Environmental Data Initiative https://doi.org/10.6073/PASTA/325232E6E4CD1CE04025FA5674F7B782 (2023).
Magnuson, J., Carpenter, S. & Stanley, E. North temperate lakes LTER: chlorophyll - Madison Lakes area 1995 - current. Environmental Data Initiative https://doi.org/10.6073/PASTA/F9C2E1059BCF92F138E140950A3632F2 (2022).
Magnuson, J. J., Carpenter, S. R. & Stanley, E. H. North temperate lakes LTER: phytoplankton - Madison Lakes area 1995 - current. Environmental Data Initiative https://doi.org/10.6073/PASTA/43D3D401AF88CC05C6595962BDB1AB5C (2022).
Magnuson, J., Carpenter, S. & Stanley, E. North temperate lakes LTER: zooplankton - Madison Lakes area 1997 - current. Environmental Data Initiative https://doi.org/10.6073/PASTA/D5ABE9009D7F6AA87D1FCF49C8C7F8C8 (2022).
Nurk, S., Meleshko, D., Korobeynikov, A. & Pevzner, P. A. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017).
Chen, I.-M. A. et al. IMG/M v.5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 47, D666–D677 (2019).
Kieft, K., Zhou, Z. & Anantharaman, K. VIBRANT: automated recovery, annotation and curation of microbial viruses, and evaluation of viral community function from genomic sequences. Microbiome 8, 90 (2020).
Kieft, K., Adams, A., Salamzade, R., Kalan, L. & Anantharaman, K. vRhyme enables binning of viral genomes from metagenomes. Nucleic Acids Res. 50, e83 (2022).
Nayfach, S. et al. Metagenomic compendium of 189,680 DNA viruses from the human gut microbiome. Nat. Microbiol. 6, 960–970 (2021).
van Dongen, S. & Abreu-Goodger, C. Using MCL to extract clusters from networks. Methods Mol. Biol. 804, 281–295 (2012).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
O’Leary, N. A. et al. Reference Sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Finn, R. D. et al. HMMER web server: 2015 update. Nucleic Acids Res. 43, W30–W38 (2015).
Camargo, A. P. et al. Identification of mobile genetic elements with geNomad. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01953-y (2023).
Roux, S. et al. iPHoP: an integrated machine learning framework to maximize host prediction for metagenome-derived viruses of archaea and bacteria. PLoS Biol. 21, e3002083 (2023).
Gregory, A. C. et al. MetaPop: a pipeline for macro- and microdiversity analyses and visualization of microbial and viral metagenome-derived populations. Microbiome 10, 49 (2022).
Zhou, Z. et al. TYMEFLIES vMAGs and related properties. Figshare https://figshare.com/articles/dataset/TYMEFLIES_vMAGs_and_related_properties/24915750 (2023).
Acknowledgements
We thank the local support for fieldwork conducted in Lake Mendota, WI, as a site of a long-term lake ecological study for North Temperate Lakes Long Term Ecological Research, including the following people as sampling leads: A. Kent, T. Yannarell, A. Shade, S. Jones, R. Newton, G. Wolfe, E. K. Read, L. Beversdorf and J. Mutschler; and initial Microbial Observatory lead, E. W. Triplett. This research was supported by the National Science Foundation grant number DBI2047598 (K.A.), US Department of Agriculture National Institute of Food and Agriculture under Hatch project 1025641 (K.A.), Simons Foundation Investigator in Aquatic Microbial Ecology Award LI-SIAME-00002001 (B.J.B.) and research funds from Synthetic Biology Research Center of Shenzhen University (Z.Z.). C.M. was supported by a National Science Foundation Graduate Research Fellowship. R.R.R. was supported by a National Science Foundation Postdoctoral Research Fellowship in Biology (DBI-2011002). Sequencing and initial sequence datasets processing were carried out at the US DOE JGI (CSP 504350). The work (proposal: CSP 504350) conducted by the US DOE JGI (https://ror.org/04xm1d337), a Department of Energy Office of Science User Facility, is supported by the Office of Science of the US Department of Energy operated under contract number DE-AC02-05CH11231.
Author information
Authors and Affiliations
Contributions
Z.Z., K.D.M. and K.A. conceived the project. R.R.R. performed DNA extraction and sequencing. Z.Z., C.M. and P.Q.T. conducted bioinformatic analyses, statistical analyses, visualization of results and content organization. Z.Z. and K.A. wrote the manuscript draft. All authors (Z.Z., K.D.M., K.A., R.R.R., B.J.B., C.M., P.Q.T.) reviewed the results and edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Timothy Ghaly, Andrew Millard and David Pearce for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Summary of viral scaffolds and genomes.
a Binned/unbinned scaffold percentage after binning by vRhyme and bin member number frequency for all bins (vMAGs). Only bin member numbers with frequencies > 1% are shown in the bar plot. Numbers of scaffolds and numbers of bins are labeled accordingly. b Length and completeness change after binning, and CheckV quality to viral genome length distribution. Viral scaffold or/and vMAG (viral genome) numbers are labeled accordingly. “Viral scaffolds”: total viral scaffolds before binning; “vMAGs+unbinned scaffolds”: vMAGs and unbinned scaffolds after binning; “vMAGs”: vMAGs after binning; “Binned scaffolds (within vMAGs)”: binned scaffolds (the scaffolds that are in the vMAGs) after binning; “Unbinned scaffolds”: unbinned scaffolds after binning. Statistical significance was assessed using two-sided t-tests for the indicated comparisons, with p-values indicating significance between comparisons. c The number of viral scaffolds, vMAGs, species, and genera. d The rarefaction curve of species-level vOTU numbers. Ten replicates with a random starting sample were made to generate error bars.
Extended Data Fig. 2 AMG metabolisms and functions.
a AMGs involved in photosynthesis, b AMGs involved in methane and related metabolism, c AMGs involved in nitrogen metabolism, d AMGs involved in CO2 fixation, e AMGs involved in sulfur and related metabolism, f AMGs involved in nucleotide metabolism, g AMGs involved in amino sugar and nucleotide sugar metabolism, h AMGs involved in pyruvate metabolism, i AMGs involved in porphyrin, haem, and cobalamin metabolism, j AMGs involved in folate biosynthesis, k AMGs involved in nicotinate and nicotinamide metabolism, l AMGs involved in riboflavin metabolism, m AMGs assigned to AMG clusters with other important functions (distributed in >300 metagenomes), n AMGs assigned to clusters with other important functions. Gene symbols, the corresponding enzyme name, and CAZy ID for genes in n were depicted in brown together with the occurrence and abundance values (labeled as “occurrence|abundance”; occurrence, the number of metagenomes in which an AMG cluster can be found; abundance, the mean normalized abundance of AMG carrying viruses in the metagenomes in which this AMG can be found). Dotted arrows indicate steps that are not encoded by AMGs. Detailed information on each AMG cluster can be found in Supplementary Table S6.
Extended Data Fig. 3 AMG cluster variation in species and high occurrence AMG cluster distribution across different seasons.
a AMG cluster variation in viral species. The left bar plot represents the AMG cluster presence ratio pattern among all AMG cluster and species combinations. The x-axis indicates the size category of species and the number of AMG cluster and species combinations. The y-axis indicates the fractions of four quartiles of AMG cluster presence ratios. The right scatter plot represents the AMG cluster count fraction (the percentage of one AMG cluster being encountered among all AMG clusters within a species) to the mean AMG cluster presence ratio (the percentage that one AMG cluster appears among all members within a species) across all species. This scatter plot used the AMG cluster and species combinations of the 1st quartile (75-100%) of AMG cluster presence ratio category (the highest presence ratio) with the species size in the 4th quartile (the largest species size), which was shown as the connection by dash lines. High occurrence AMG clusters (distributed > 400 metagenomes) were colored red, and other AMG clusters were colored green. b Seasonal distribution of high occurrence AMG clusters (distributed > 400 metagenomes)) across metagenomes. The percentage indicates the AMG cluster containing metagenome number over the total metagenome number in each season.
Extended Data Fig. 4 The taxonomic distribution (classified to the family level) of predicted hosts for eight AMG cluster-containing viruses with low Simpson indices.
Unclassified hosts were not depicted and low abundance families (with abundance < 5% in all eight AMG clusters) were integrated into a group named “Others”.
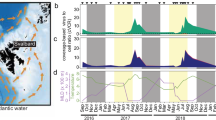
Extended Data Fig. 5 Species and AMG abundance across the time-series.
The seasonal abundance distribution, species and AMG abundance percentage, and total species and AMG abundance across 20 years for psbA- (a), pmoC- (b), katG- (c), ahbD-containing (d) viruses are summarized. In each subpanel, high occurrence species were picked according to the occurrence across 20 years, high abundance AMGs were picked according to the non-zero mean relative abundance across 20 years, and the abundance for each year was represented by the season with the highest/second to the highest species abundance in each year (Late Summer for psbA, Fall for pmoC, Late Summer for katG, and Early Summer for ahbD). Species and AMGs were colored in blue and orange, respectively. Star-labeled AMGs indicate the overlap of the high occurrence species and high abundance AMG in subpanels a, c, d. The abundance values (for both species and AMGs) were normalized by 100 M reads/metagenome. For psbA- and ahbD-containing viruses, only species with ≥ 20 occurrences out of 471 metagenomes were included in the analysis; for pmoC- and katG-containing viruses, only species with ≥ 5 occurrences out of 471 metagenomes were included in the analysis. The species and AMG abundance percentage calculation was based on the total occurrence-filtered viral species.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1, Methods, Results and references.
Supplementary Tables (download XLSX )
Supplementary Tables 1–14.
Supplementary Dataset (download ZIP )
Supplementary Dataset 1.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, Z., Tran, P.Q., Martin, C. et al. Unravelling viral ecology and evolution over 20 years in a freshwater lake. Nat Microbiol 10, 231–245 (2025). https://doi.org/10.1038/s41564-024-01876-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-024-01876-7
This article is cited by
-
Diversity and ecological roles of hidden viral players in groundwater microbiomes
Nature Communications (2026)
-
A genomic view of Earth’s biomes
Nature Reviews Genetics (2026)
-
Viromics approaches for the study of viral diversity and ecology in microbiomes
Nature Reviews Genetics (2026)
-
Metagenomic analysis reveals how multiple stressors disrupt virus–host interactions in multi-trophic freshwater mesocosms
Nature Communications (2025)
-
Crucial stepping stones in freshwater microbiology
Nature Microbiology (2025)