Abstract
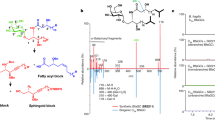
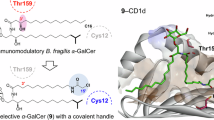
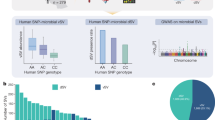
Gut symbiont Bacteroides fragilis can produce α-galactosylceramides (BfaGCs), sphingolipids with immunomodulatory functions that regulate colonic natural killer T (NKT) cells. However, their synthesis pathway and whether other human gut bacteria can produce them are unclear. Here, using genetic and metabolomic approaches, we mapped the sphingolipid biosynthesis pathway of B. fragilis and determined that α-galactosyltransferase (agcT) is essential and sufficient for colonic NKT cell regulation in mice. The distribution of agcT is restricted to only a few species among Bacteroidales. However, structural homologues of AgcT, such as BgsB, are widely distributed in gut microbiota and produce α-glycosyldiacylglycerols (aGDGs), particularly in Enterococcus. Analysis of infant gut metagenomes revealed that B. fragilis predominantly accounts for agcT abundance regardless of the cohort, but bgsB-encoding bacteria were taxonomically diverse and showed dynamic changes with host age. In addition, aGDGs from bgsB-encoding species act as antagonistic ligands for BfaGC-mediated NKT cell activation in vitro and in vivo. Our findings highlight the distinct natures of immunoactive glycolipid-producing symbionts and their relevance in the human gut microbiome, particularly in early life.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Lipidomics datasets supporting this study are available via Harvard Dataverse (https://doi.org/10.7910/DVN/DYJJQJ). TEDDY microbiome data are available from dbGaP (phs001443.v1.p1) under dbGaP-controlled access. Curated human microbiome data are accessible via curatedMetagenomicData (https://doi.org/10.18129/B9.bioc.curatedMetagenomicData). The RNA-sequencing data are available in the NCBI database under BioProject accession number PRJNA1309292. Source data are provided with this paper.
References
Skelly, A. N., Sato, Y., Kearney, S. & Honda, K. Mining the microbiota for microbial and metabolite-based immunotherapies. Nat. Rev. Immunol. 19, 305–323 (2019).
Brutkiewicz, R. R. CD1d ligands: the good, the bad, and the ugly. J. Immunol. 177, 769–775 (2006).
Dorrestein, P. C., Mazmanian, S. K. & Knight, R. Finding the missing links among metabolites, microbes, and the host. Immunity 40, 824–832 (2014).
An, D. et al. Sphingolipids from a symbiotic microbe regulate homeostasis of host intestinal natural killer T cells. Cell 156, 123–133 (2014).
Morita, M. et al. Structure–activity relationship of alpha-galactosylceramides against B16-bearing mice. J. Med. Chem. 38, 2176–2187 (1995).
Oh, S. F. et al. Host immunomodulatory lipids created by symbionts from dietary amino acids. Nature 600, 302–307 (2021).
Goodman, A. L. et al. Identifying genetic determinants needed to establish a human gut symbiont in its habitat. Cell Host Microbe 6, 279–289 (2009).
Comstock, L. E. et al. Analysis of a capsular polysaccharide biosynthesis locus of Bacteroides fragilis. Infect. Immun. 67, 3525–3532 (1999).
Coyne, M. J., Chatzidaki-Livanis, M., Paoletti, L. C. & Comstock, L. E. Role of glycan synthesis in colonization of the mammalian gut by the bacterial symbiont Bacteroides fragilis. Proc. Natl Acad. Sci. USA 105, 13099–13104 (2008).
Moremen, K. W. & Haltiwanger, R. S. Emerging structural insights into glycosyltransferase-mediated synthesis of glycans. Nat. Chem. Biol. 15, 853–864 (2019).
Okino, N. et al. Two bacterial glycosphingolipid synthases responsible for the synthesis of glucuronosylceramide and α-galactosylceramide. J. Biol. Chem. 295, 10709–10725 (2020).
Stankeviciute, G. et al. Convergent evolution of bacterial ceramide synthesis. Nat. Chem. Biol. 18, 305–312 (2021).
Naidenko, O. V. et al. Binding and antigen presentation of ceramide-containing glycolipids by soluble mouse and human CD1d molecules. J. Exp. Med. 190, 1069–1080 (1999).
Brennan, P. J. et al. Invariant natural killer T cells recognize lipid self antigen induced by microbial danger signals. Nat. Immunol. 12, 1202–1211 (2011).
Zeissig, S., Olszak, T., Melum, E. & Blumberg, R. S. Analyzing antigen recognition by natural killer T cells. Methods Mol. Biol. 960, 439–453 (2013).
Olszak, T. et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336, 489–493 (2012).
Almeida, A. et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat. Biotechnol. 39, 105–114 (2021).
Kawahara, K. et al. Chemical structure of glycosphingolipids isolated from Sphingomonas paucimobilis. FEBS Lett. 292, 107–110 (1991).
Naka, T. et al. A novel sphingoglycolipid containing galacturonic acid and 2-hydroxy fatty acid in cellular lipids of Sphingomonas yanoikuyae. J. Bacteriol. 182, 2660–2663 (2000).
Kinjo, Y. et al. Recognition of bacterial glycosphingolipids by natural killer T cells. Nature 434, 520–525 (2005).
Mayassi, T., Barreiro, L. B., Rossjohn, J. & Jabri, B. A multilayered immune system through the lens of unconventional T cells. Nature 595, 501 (2021).
Oh, S. F., Jung, D.-J. & Choi, E. Gut microbiota-derived unconventional T cell ligands: contribution to host immune modulation. Immunohorizons 6, 476–487 (2022).
Pasolli, E. et al. Accessible, curated metagenomic data through ExperimentHub. Nat. Methods 14, 1023–1024 (2017).
Hölzl, G. & Dörmann, P. Structure and function of glycoglycerolipids in plants and bacteria. Prog. Lipid Res. 46, 225–243 (2007).
Theilacker, C. et al. Deletion of the glycosyltransferase bgsB of Enterococcus faecalis leads to a complete loss of glycolipids from the cell membrane and to impaired biofilm formation. BMC Microbiol. 11, 67 (2011).
Stewart, C. J. et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562, 583–588 (2018).
Vatanen, T. et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 165, 842–853 (2016).
Vatanen, T. et al. The human gut microbiome in early-onset type 1 diabetes from the TEDDY study. Nature 562, 589–594 (2018).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015).
Brooks, B. et al. Strain-resolved analysis of hospital rooms and infants reveals overlap between the human and room microbiome. Nat. Commun. 8, 1814 (2017).
Shao, Y. et al. Stunted microbiota and opportunistic pathogen colonization in caesarean-section birth. Nature 574, 117–121 (2019).
Yassour, M. et al. Strain-level analysis of mother-to-child bacterial transmission during the first few months of life. Cell Host Microbe 24, 146–154 (2018).
Turnbaugh, P. J. et al. The Human Microbiome Project. Nature 449, 804–810 (2007).
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Kinjo, Y. et al. Natural killer T cells recognize diacylglycerol antigens from pathogenic bacteria. Nat. Immunol. 7, 978–986 (2006).
Yoo, J. S. & Oh, S. F. Unconventional immune cells in the gut mucosal barrier: regulation by symbiotic microbiota. Exp. Mol. Med. 55, 1905–1912 (2023).
Huang, S. et al. CD1 lipidomes reveal lipid-binding motifs and size-based antigen-display mechanisms. Cell 186, 4583–4596.e13 (2023).
Baldridge, M. T. et al. Commensal microbes and interferon-λ determine persistence of enteric murine norovirus infection. Science 347, 266–269 (2015).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Lu, S. et al. CDD/SPARCLE: the conserved domain database in 2020. Nucleic Acids Res. 48, D265–D268 (2020).
Katoh, K. & Standley, D. M. MAFFT Multiple Sequence Alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T. ggtree: an r package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Blanco-Míguez, A. et al. Extending and improving metagenomic taxonomic profiling with uncharacterized species using MetaPhlAn 4. Nat. Biotechnol. 41, 1633–1644 (2023).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Zhang, Y. & Skolnick, J. TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res. 33, 2302–2309 (2005).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 42, 243–246 (2024).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
García-Bayona, L. & Comstock, L. E. Streamlined genetic manipulation of diverse Bacteroides and Parabacteroides isolates from the human gut microbiota. mBio 10, 4 (2019).
Lim, B., Zimmermann, M., Barry, N. A. & Goodman, A. L. Engineered regulatory systems modulate gene expression of human commensals in the gut. Cell 169, 547–558.e15 (2017).
Thurlow, L. R., Thomas, V. C. & Hancock, L. E. Capsular polysaccharide production in Enterococcus faecalis and contribution of CpsF to capsule serospecificity. J. Bacteriol. 191, 6203–6210 (2009).
Behar, S. M., Podrebarac, T. A., Roy, C. J., Wang, C. R. & Brenner, M. B. Diverse TCRs recognize murine CD1. J. Immunol. 162, 161–167 (1999).
Buchfink, B., Reuter, K. & Drost, H. G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Acknowledgements
This study was supported by the National Institutes of Health (K01-DK102771, R01-AT010268, R01-AI165987: S.F.O.) and the National Research Foundation of Korea (2021R1A6A3A14044113: J-S.Y.; RS-2024-00411992: B.G.; 2021R1A6A3A14039202: D-J.J.; RS-2024-00348702: K.H.; RS-2023–00217123: J.I.S.; 2014R1A3A2030423 and 2012M3A9C4048780: S.B.P.). CD1d tetramers were provided by the NIH Tetramer Core Facility (contract number 75N93020D00005). We thank T. Yanostang for technical support and N. Surana (Duke University) for the helpful discussion and comments.
Author information
Authors and Affiliations
Contributions
S.F.O., D.L.K., S.B.P. and J-S.Y. conceived the idea and designed the outline of the research. J.-S.Y., K.H., J.I.S. and B.G. carried out the lipidomic profiling of gut symbionts. W.Z., C.C.L., J-S.Y., M.W. and S.F.O. carried out loss-of-function screening. K.H. and B.G. carried out gain-of-function screening. J.-S.Y. and N.G.-Z. carried out targeted bacterial knockout generation. D.-J.J., J.-S.Y. and S.F.O. carried out in vivo NKT analysis. J.-S.Y. carried out in silico structural analysis, in vitro NKT activation assays and metagenomic analysis. J.-S.Y. and S.F.O. wrote the paper, and all authors contributed to the relevant discussion.
Corresponding author
Ethics declarations
Competing interests
S.F.O. and D.L.K. filed a patent on the functions of BfaGCs and related structures (US patent 10,329,315). S.F.O., S.B.P. and D.L.K. filed a patent on the functions of BfaGCs and related structures (US patent application 17/427,756). The other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Peter Turnbaugh and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of screened genes and generation of knockout and transformant strains.
a) Graphical scheme of transposon insertion (loss-of-function) and heterologous expression (gain-of-function) library generation, followed by targeted metabolomic analysis to characterize α-galactosyltransferase, was presented. b) Loss-of-function screening identified a transposon insertion mutant (Tn8A7) with depleted BfaGC production. The relative abundance was quantified by UHPLC-MS/MS based on peak area. c) BfaGC production was abrogated in the B. fragilis Δ3069 mutant (n = 2) compared to B. fragilis WT (n = 3). The abundance of BfaGC (C34:0) was quantified by LC-MS/MS based on peak area. d) RNA-seq volcano plot comparing B. fragilis Δ3069 to wild type revealed multiple genes downregulated in Δ3069 mutant. e) The expression of agcT is reduced in B. fragilis Δ3069 (n = 3) compared to B. fragilis WT (n = 3). Transcript levels of agcT, normalized to leuB, were measured by qRT-PCR. Significance was determined by an unpaired two-tailed t-test. f) Gain-of-function screening identified P. vulgatus transformant producing aGCs. The relative abundance of aGC/Cer was quantified by UHPLC-MS/MS based on peak area. g) agcT-transformed P. vulgatus produced molecules identical to BfaGC. XICs of aGCs from B. fragilis, P. vulgatus wild type and P. vulgatus expressing agcT were displayed. Chromosome-integrated expressing system pNBU2-tetR was used to expressing agcT in P. vulgatus. h) MS/MS spectra mirror plot demonstrated a match between aGCs from P. vulgatus expressing agcT and those from B. fragilis. i) B. thetaiotaomicron and P. vulgatus heterologously expressing agcT produced aGCs. Chromosome-integrated expressing system pNBU2-tetR was used to expressing agcT in B. thetaiotaomicron and P. vulgatus (n = 2 per group). j) Relative abundance of KDS, ketoCer, Cer, and BfaGC in B. fragilis wild type (n = 4), Δspt (n = 4), ΔcerS (n = 4), ΔcerR (n = 4), and ΔagcT (n = 3) were shown. The abundance was quantified by UHPLC-MS/MS based on peak area.
Extended Data Fig. 2 Tandem mass spectra analysis determined sphingolipid intermediate structures.
To assign the structures unambiguously, we chose chain length variants of major sphingolipid species, whose synthetic version was available. a) MS/MS spectra of B. fragilis KDS (C18:0) matched those of the synthetic standard. b) MS/MS spectra of synthetic C35:1 ceramide (m/z 568.53) was displayed. c) The parent ion with an m/z value of 570.53 from B. fragilis was structurally assigned as C35:0 dihydroceramide. d) The parent ion of m/z 568.53 from B. fragilis ΔcerR was assigned as C35:0 ketoCer. Comparison with isobaric C35:1 ceramide (Panel b) exhibits a distinct fragmentation pattern, as shown by the presence and absence of m/z 264 (300-2H2O). e-f) MS/MS spectra of C34:0 ketoCer and dhCer from B. fragilis were shown.
Extended Data Fig. 3 In vitro and in vivo assays for NKT cell modulation by BfaGCs.
a) Schematic process illustrates the in vitro antigen presentation assay. Bacterial lipid extracts were loaded onto biotinylated CD1d molecules, which were subsequently immobilized on streptavidin-coated plates. After washing, NKT cell hybridomas (24.7) were added and incubated overnight. IL-2 secretion was measured by ELISA to assess NKT cell activation. b) Representative gating strategies for colonic NKT cell analysis by flow cytometry are shown. NKT cell refers to the CD1d tetramer⁺ cells within the CD3⁺CD45⁺TCRβ⁺ T cell population. c) Representative flow cytometry plots of unloaded CD1d tetramer controls are shown.
Extended Data Fig. 4 B. fragilis and B. salyersiae produce aGCs and induce NKT cell activation.
a) A MS/MS mirror plot of B. fragilis and B. salyersiae C34:0 aGC species showed an exact match. b) All B. fragilis and B. salyersiae strains, including type strains and clinical isolates, synthesize aGC, in contrast, no tested B. nordi, P. gordornii, and P. copri strains produce aGCs. c) Only aGC-producing bacterial lipid extract can elicit NKT cell activation. NKT cell hybridomas (24.7) were incubated with CD1d-lipid complex and IL-2 secretion was measured by ELISA. Each dot indicates a biologically independent replicate (n = 4 per group). Bars and error bars depict the mean ± s.e.m. Data are representative of three independent experiments showing consistent trends. Statistical analysis was performed using one-way ANOVA. d) Symbiotic Bacteroidales species did not produce aGlcACers, whereas Sphingomonas species served as positive controls and produced aGlcACers.
Extended Data Fig. 5 Homologs of AgcT share conserved structures.
a) The histogram displays the distribution of e-values from the search for proteins containing cd03817 domain. The red dashed vertical line indicates the cutoff threshold used for identifying cd03817 family hits by HMMER search. b) An AgcT homolog from B. salyersiae and a cd03817 family protein from E. faecalis belong to the same protein family and have comparable structures.
Extended Data Fig. 6 aGDG production by cd03817-encoding bacteria.
a) Mirror plot of MS/MS spectra of E. faecalis aGDGs and a commercially available standard (Avanti #840522p) showed a match. b) The retention time of aGDGs of E. faecalis matches that of standard molecule (Avanti #840522p). c) aGDGs from cd03817-domain-containing gut bacterial species exhibit chain-length variation. XICs of aGDGs from Streptococcus mitis, Lacticaseibacillus rhamnosus, and E. faecalis were shown. d) Various Enterococcus strains produce aGDGs. XICs of C34:1 aGDGs were displayed. e) E. faecalis WT and bgsB mutant exhibit comparable colonization abilities in 4-week-old male C57BL/6 mice (n = 3 per group). f) aGDG levels in stool samples from 5-week-old male/female C57BL/6 SPF (n = 6), conventionalized (n = 6), and GF (n = 7) mice were shown. “Conventionalized” indicates GF mice conventionalized via cohousing with SPF mice.
Extended Data Fig. 7 Metagenomic profilies of agcT and bgsB in multiple cohorts.
a) Skimming process to search species with homologs of AgcT or BgsB was shown. b) Histograms show sequence identity distributions from the search for homologs of AgcT or BgsB, with vertical dashed lines indicating identity cutoffs. c) In the DIABIMMUNE cohort, B. fragilis consistently dominates agcT abundance, while bgsB abundance shows dynamic changes and is contributed by a taxonomically diverse set of species. Pie charts show the proportion of species abundance in the dataset. d-e) Across multiple infant and adult cohorts, agcT abundance is consistently driven by B. fragilis, whereas bgsB is distributed among a broader array of species.
Extended Data Fig. 8 aGDGs species inhibit the effects of BfaGCs on NKT cells.
a) Bacterial lipid extracts from various Lactobacillales bgsB-encoding species exhibit dose-dependent antagonism (n = 4 per group) to SB2217-induced (n = 2) NKT cell activation. b) aGDGs competed with BfaGCs for CD1d binding in a dose-dependent manner. After loading the lipids onto CD1d, CD1d-lipids complexes were purified, and the bound lipids were extracted and analyzed by LC-MS (n = 4 per group). c) Representative flow cytometry plots show the gating strategy used for analysis of splenic dendritic cells.
Extended Data Fig. 9 Overview of screening and exploration of related metabolites.
Unannotated metabolites were linked to their biosynthetic gene agcT through forward genetics-based metabolomic screening. Protein structural homologs were identified in gut symbionts, revealing structurally related glycolipids with contrasting immunological activities. Species-level contributions of agcT and its homologs bgsB were profiled in human gut metagenomes. Figure created with BioRender.com.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yoo, JS., Jung, DJ., Goh, B. et al. Human gut bacteria produce structurally related monoglycolipids with contrasting immune functions. Nat Microbiol 10, 2797–2807 (2025). https://doi.org/10.1038/s41564-025-02141-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02141-1