Abstract
Class A penicillin-binding proteins (aPBPs) are involved in the biosynthesis and remodelling of peptidoglycan (PG). The human bacterial pathogen Streptococcus pneumoniae produces three aPBPs, which are regulated to maintain the bacterium’s ovoid shape. Evidence suggests that PBP1a and PBP2a activities are closely coordinated; however, their precise functions remain unclear. Here we characterized the pneumococcal S protein, which contains a LysM-PG-binding domain and a GpsB-interacting domain. Using S protein fusion constructs or mutant bacterial strains, we show that S protein localizes to the division ring and is required to prevent premature cell lysis and minicell formation due to aberrant division site placement. S protein interacts with PBP1a and activates its PG synthesis activity. Co-immunoprecipitation experiments combined with biochemical, genetic, structural prediction and microscopy analyses suggest that S protein is part of a larger multiprotein complex containing aPBPs and PG-modifying enzymes, and coordinated by the scaffolding protein GpsB. Together, these findings suggest that a GpsB-associated complex orchestrates PG biosynthesis and remodelling in S. pneumoniae.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author on request. The mass spectrometry proteomics data have been deposited to the Center for Computational Mass Spectrometry repository (University of California, San Diego) via the MassIVE tool with the dataset identifier MassIVE MSV000098614. Strains constructed here are available from the corresponding author on request. Raw western blots and SDS–PAGE for all figures are provided as Source data with this paper. Source data are provided with this paper.
References
Egan, A. J. F., Errington, J. & Vollmer, W. Regulation of peptidoglycan synthesis and remodelling. Nat. Rev. Microbiol. 18, 446–460 (2020).
Kumar, S., Mollo, A., Kahne, D. & Ruiz, N. The bacterial cell wall: from lipid II flipping to polymerization. Chem. Rev. 122, 8884–8910 (2022).
Meeske, A. J. et al. SEDS proteins are a widespread family of bacterial cell wall polymerases. Nature 537, 634–638 (2016).
Sjodt, M. et al. Structural coordination of polymerization and crosslinking by a SEDS–bPBP peptidoglycan synthase complex. Nat. Microbiol. 5, 813–820 (2020).
Straume, D., Piechowiak, K. W., Kjos, M. & Håvarstein, L. S. Class A PBPs: it is time to rethink traditional paradigms. Mol. Microbiol. 116, 41–52 (2021).
Vigouroux, A. et al. Class-A penicillin binding proteins do not contribute to cell shape but repair cell-wall defects. eLife 9, e51998 (2020).
Typas, A. et al. Regulation of peptidoglycan synthesis by outer membrane proteins. Cell 143, 1097–1109 (2010).
Navarro, P. P. et al. The aPBP-type cell wall synthase PBP1b plays a specialized role in fortifying the Escherichia coli division site against osmotic rupture. Preprint at bioRxiv https://doi.org/10.1101/2025.04.02.646830 (2025).
Paradis-Bleau, C. et al. Lipoprotein cofactors located in the outer membrane activate bacterial cell wall polymerases. Cell 143, 1110–1120 (2010).
Fenton, A. K., El Mortaji, L., Lau, D. T. C., Rudner, D. Z. & Bernhardt, T. G. CozE is a member of the MreCD complex that directs cell elongation in Streptococcus pneumoniae. Nat. Microbiol. 2, 16237 (2016).
Lenoir, C. et al. The morphogenic protein CopD controls the spatio-temporal dynamics of PBP1a and PBP2b in Streptococcus pneumoniae. mBio 14, e01411-23 (2023).
Costa, S. F. et al. The role of GpsB in Staphylococcus aureus cell morphogenesis. mBio 15, e03235-23 (2024).
Cleverley, R. M. et al. The cell cycle regulator GpsB functions as cytosolic adaptor for multiple cell wall enzymes. Nat. Commun. 10, 261 (2019).
Paik, J., Kern, I., Lurz, R. & Hakenbeck, R. Mutational analysis of the Streptococcus pneumoniae bimodular class A penicillin-binding proteins. J. Bacteriol. 181, 3852–3856 (1999).
Rued, B. E. et al. Suppression and synthetic-lethal genetic relationships of ΔgpsB mutations indicate that GpsB mediates protein phosphorylation and penicillin-binding protein interactions in Streptococcus pneumoniae D39. Mol. Microbiol. 103, 931–957 (2017).
Midonet, C. et al. MacP bypass variants of Streptococcus pneumoniae PBP2a suggest a conserved mechanism for the activation of bifunctional cell wall synthases. mBio 14, e02390-23 (2023).
Wierzbicki, I. H. et al. Group A streptococcal S protein utilizes red blood cells as immune camouflage and is a critical determinant for immune evasion. Cell Rep. 29, 2979–2989.e15 (2019).
Buist, G., Steen, A., Kok, J. & Kuipers, O. P. LysM, a widely distributed protein motif for binding to (peptido)glycans. Mol. Microbiol. 68, 838–847 (2008).
Hoskins, J. et al. Genome of the bacterium Streptococcus pneumoniae strain R6. J. Bacteriol. 183, 5709–5717 (2001).
Piovesan, D. et al. MOBIDB in 2025: integrating ensemble properties and function annotations for intrinsically disordered proteins. Nucleic Acids Res. 53, D495–D503 (2025).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Alcorlo, M., Martínez-Caballero, S., Molina, R. & Hermoso, J. A. Carbohydrate recognition and lysis by bacterial peptidoglycan hydrolases. Curr. Opin. Struct. Biol. 44, 87–100 (2017).
Fleurie, A. et al. MapZ marks the division sites and positions FtsZ rings in Streptococcus pneumoniae. Nature 516, 259–262 (2014).
Holečková, N. et al. LocZ is a new cell division protein involved in proper septum placement in Streptococcus pneumoniae. mBio https://doi.org/10.1128/mbio.01700-14 (2014).
Manuse, S. et al. Structure–function analysis of the extracellular domain of the pneumococcal cell division site positioning protein MapZ. Nat. Commun. 7, 12071 (2016).
Kuru, E. et al. Mechanisms of incorporation for d-amino acid probes that target peptidoglycan biosynthesis. ACS Chem. Biol. 14, 2745–2756 (2019).
Berg, K. H., Stamsås, G. A., Straume, D. & Håvarstein, L. S. Effects of low PBP2b levels on cell morphology and peptidoglycan composition in Streptococcus pneumoniae R6. J. Bacteriol. 195, 4342–4354 (2013).
Tsui, H.-C. T. et al. Pbp2x localizes separately from Pbp2b and other peptidoglycan synthesis proteins during later stages of cell division of Streptococcus pneumoniae D39. Mol. Microbiol. 94, 21–40 (2014).
Tsui, H.-C. T. et al. Suppression of a deletion mutation in the gene encoding essential PBP2b reveals a new lytic transglycosylase involved in peripheral peptidoglycan synthesis in Streptococcus pneumoniae D39. Mol. Microbiol. 100, 1039–1065 (2016).
Schultz, B. J., Snow, E. D. & Walker, S. Mechanism of d-alanine transfer to teichoic acids shows how bacteria acylate cell envelope polymers. Nat. Microbiol. 8, 1318–1329 (2023).
Land, A. D. & Winkler, M. E. The requirement for pneumococcal MreC and MreD is relieved by inactivation of the gene encoding PBP1a. J. Bacteriol. 193, 4166–4179 (2011).
Álvarez-Salmoral, D. et al. AlphaBridge: tools for the analysis of predicted biomolecular complexes. Preprint at bioRxiv https://doi.org/10.1101/2024.10.23.619601 (2025).
Vollmer, W. & Tomasz, A. The pgdA gene encodes for a peptidoglycan N-acetylglucosamine deacetylase in Streptococcus pneumoniae. J. Biol. Chem. 275, 20496–20501 (2000).
Vollmer, W. & Tomasz, A. Peptidoglycan N-acetylglucosamine deacetylase, a putative virulence factor in Streptococcus pneumoniae. Infect. Immun. 70, 7176–7178 (2002).
Egan, A. J. F. et al. Outer-membrane lipoprotein LpoB spans the periplasm to stimulate the peptidoglycan synthase PBP1B. Proc. Natl Acad. Sci. USA 111, 8197–8202 (2014).
Lupoli, T. J. et al. Lipoprotein activators stimulate Escherichia coli penicillin-binding proteins by different mechanisms. J. Am. Chem. Soc. 136, 52–55 (2014).
Sauvage, E. & Terrak, M. Glycosyltransferases and transpeptidases/penicillin-binding proteins: valuable targets for new antibacterials. Antibiotics 5, 12 (2016).
Sauvage, E., Kerff, F., Terrak, M., Ayala, J. A. & Charlier, P. The penicillin-binding proteins: structure and role in peptidoglycan biosynthesis. FEMS Microbiol. Rev. 32, 234–258 (2008).
Di Guilmi, A. M. et al. Identification, purification, and characterization of transpeptidase and glycosyltransferase domains of Streptococcus pneumoniae penicillin-binding protein 1a. J. Bacteriol. 180, 5652–5659 (1998).
Lanie, J. A. et al. Genome sequence of Avery’s virulent serotype 2 strain D39 of Streptococcus pneumoniae and comparison with that of unencapsulated laboratory strain R6. J. Bacteriol. 189, 38–51 (2007).
Trouve, J. et al. Nanoscale dynamics of peptidoglycan assembly during the cell cycle of Streptococcus pneumoniae. Curr. Biol. 31, 2844–2856.e6 (2021).
Burnier, J. et al. Pneumococcal S protein coordinates cell wall modification and repair to resist host antimicrobials. Nat. Microbiol. https://doi.org/10.1038/s41564-025-02184-4 (2026).
Claessen, D. et al. Control of the cell elongation–division cycle by shuttling of PBP1 protein in Bacillus subtilis. Mol. Microbiol. 68, 1029–1046 (2008).
Fenton, A. K. et al. Phosphorylation-dependent activation of the cell wall synthase PBP2a in Streptococcus pneumoniae by MacP. Proc. Natl Acad. Sci. USA 115, 2812–2817 (2018).
Berg, K. H., Biørnstad, T. J., Straume, D. & Håvarstein, L. S. Peptide-regulated gene depletion system developed for use in Streptococcus pneumoniae. J. Bacteriol. 193, 5207–5215 (2011).
Sung, C. K., Li, H., Claverys, J. P. & Morrison, D. A. An rpsL cassette, Janus, for gene replacement through negative selection in Streptococcus pneumoniae. Appl. Environ. Microbiol. 67, 5190–5196 (2001).
Zucchini, L. et al. PASTA repeats of the protein kinase StkP interconnect cell constriction and separation of Streptococcus pneumoniae. Nat. Microbiol. 3, 197–209 (2018).
Ducret, A., Quardokus, E. M. & Brun, Y. V. MicrobeJ, a tool for high throughput bacterial cell detection and quantitative analysis. Nat. Microbiol. 1, 16077 (2016).
Morlot, C., Zapun, A., Dideberg, O. & Vernet, T. Growth and division of Streptococcus pneumoniae: localization of the high molecular weight penicillin-binding proteins during the cell cycle. Mol. Microbiol. 50, 845–855 (2003).
Fleurie, A. et al. Mutational dissection of the S/T-kinase StkP reveals crucial roles in cell division of Streptococcus pneumoniae. Mol. Microbiol. 83, 746–758 (2012).
Cortay, J. C. et al. In vitro asymmetric binding of the pleiotropic regulatory protein, FruR, to the ace operator controlling glyoxylate shunt enzyme synthesis. J. Biol. Chem. 269, 14885–14891 (1994).
Elfmann, C. & Stülke, J. PAE viewer: a webserver for the interactive visualization of the predicted aligned error for multimer structure predictions and crosslinks. Nucleic Acids Res. 51, W404–W410 (2023).
Sehnal, D. et al. Mol* Viewer: modern web app for 3D visualization and analysis of large biomolecular structures. Nucleic Acids Res. 49, W431–W437 (2021).
Garcia, P. S. et al. An early origin of iron–sulfur cluster biosynthesis machineries before Earth oxygenation. Nat. Ecol. Evol. 6, 1564–1572 (2022).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Hallgren, J. et al. DeepTMHMM predicts alpha and beta transmembrane proteins using deep neural network. Preprint at bioRxiv https://doi.org/10.1101/2022.04.08.487609 (2022).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Acknowledgements
We thank N. Campo for providing us with the silica micropillar mould for the preparation of microhole agarose pads. Support for this work comes from the CNRS, the Université Lyon I, the foundation Bettencourt-Schueller to C.G. and the Agence Nationale de la Recherche (ANR-24-CE11-4058 to C.G., ANR-23-CE11-0029 and ANR-19-CE15-0011 to C.G. and C.M., and ANR-20-CE07-0012 to A.Z.). Research in the laboratory of D.I.R. is supported by MRC grants MR/Z504245/1 and BBSRC grant BB/Y003187/1. We acknowledge the contribution of the microscopy facility (PLATIM) and we thank A. Page and F. Delolme from the Protein Science Facility (PSF) at the SFR Biosciences (University Claude Bernard Lyon1, CNRS UAR3444, Inserm US8, ENS de Lyon) for the mass spectrometry analyses. We gratefully acknowledge support from the CNRS/IN2P3 Computing Center (Lyon, France) for providing computing and data-processing resources needed for this work. IBS acknowledges integration into the Interdisciplinary Research Institute of Grenoble (IRIG, CEA).
Author information
Authors and Affiliations
Contributions
H.M., N.S.B., D.I.R., A.D. and C.G. designed the research. H.M., C.L., C.F., P.S.G., S.M., N.S.B., J.S., A.D., C.F., C.C. and A.Z. performed experiments. H.M., A.D., P.S.G., N.S.B., A.Z., C.M., D.I.R. and C.G. analysed data. H.M., A.D. and C.G. wrote the paper. All authors revised the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 AlphaFold3 prediction of S Protein structure.
The five best model predictions of S protein by AlphaFold3. The residues are colored according to the pLDDT score and the corresponding predicted alignment error (PAE) matrices are shown for each model.
Extended Data Fig. 2 S Protein is a membrane associated and forms a ring at midcell.
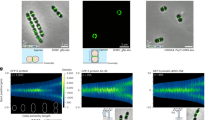
(a) Western immunoblot of whole-cell lysates of WT and mgfp-ess strains with specific anti-GFP antibody and anti-enolase antibody as a loading control. The expected mGFP-S Protein band is indicated by the black triangle. (b) Growth of WT and mgfp-ess strains in C + Y media at 37 °C. (c) Representative phase contrast microscopy images of WT and mgfp-ess strains. Scale bars, 2 µm. (d) Violin SuperPlots showing the distribution of the maximum cell width from three independent experiments, (n = 2000 cells for each strain). ns, not significant, p > 0.05 (e) Western immunoblot of whole-cell lysates of mgfp-ess, ∆ess and PcomX-mgfp-ess ∆ess cells, grown in the presence (3 µM) or absence (0 µM) of inducer (ComS), revealed with anti-GFP antibody and anti-enolase antibody as a loading control. The expected band for mGFP-S Protein is indicated with the black triangle. (f) Merge between phase contrast and GFP fluorescent signal of mgfp-ess, ∆ess and PcomX-mgfp-ess ∆ess cells, grown in the presence (3 µM) or absence (0 µM) of ComS inducer. Minicell-like compartments are highlighted by a white triangle. Scale bars, 2 µm. (g) Violin SuperPlots showing the distribution of the maximum cell width from three independent experiments, (n = 2000 cells for each strain). ***p < 0.001 and ns, not significant, p > 0.05. (h) Growth of mgfp-ess, ∆ess and PcomX-mgfp-ess ∆ess in C + Y medium at 37 °C. (i, j) Representative montage of images showing the dynamics of FtsZ-GFP (i) or mGFP-S Protein (j) in WT cells observed by conventional fluorescence microscopy with cells positioned vertically in agarose microholes for 60 s (9 s intervals). A summation of all the images and a kymograph are shown on the right and below the montage respectively. The kymographs (1 frame/s) were generated from a circular line around the circumference of the cell, highlighted in orange on the summation images. Scale bars, 0.5 µm. (k) Western immunoblot of whole-cell lysates of mgfp-ess fusion (Full length) and derivatives with specific anti-GFP antibody and anti-enolase antibody as a loading control.
Extended Data Fig. 3 Expression, cell growth and cell morphology of FLAG-ess and PcomX-FLAG-ess ∆ess cells.
(a) Western immunoblot of whole-cell lysates of FLAG-ess, ∆ess and PcomX-FLAG-ess ∆ess cells, grown in the presence (3 µM) or absence (0 µM) of inducer (ComS), revealed with anti-FLAG antibody and anti-enolase antibody as a loading control. The expected band for FLAG-S Protein is indicated by the black triangle. (b) Representative phase contrast microscopy images of FLAG-ess, ∆ess and PcomX-FLAG-ess ∆ess cells, grown in the presence (3 µM) or absence (0 µM) of ComS inducer. Minicell-like compartments are highlighted by a white triangle. Scale bars, 2 µm. (c) Violin SuperPlots showing the distribution of the maximum cell width from three independent experiments, (n = 2000 cells for each strain). ***p < 0.001 and ns, not significant, p > 0.05. (d) Growth of FLAG-ess, ∆ess and PcomX-FLAG-ess ∆ess in C + Y medium at 37 °C. (e) Representative montage of images acquired by time-lapse microscopy, showing the formation of a minicell-like compartments in a ∆ess cell. The position of the ectopic division site is indicated by a white triangle. Time is given in min. Scale bar, 0.5 µm.
Extended Data Fig. 4 Minicell-like compartment formation results from aberrant ectopic division septa and the absence of S Protein leads to reduced TDL incorporation.
(a) Representative montage of images of WT and ∆ess cells after a TDL long-pulse labeling. The phase contrast (PC), the TDL channels and the merged image (Merge) are shown. Scale bars, 2 µm. (b) Violin SuperPlots showing the distribution of the TDL fluorescence intensity for WT and ∆ess cells after a long period of TDL labeling from three independent experiments, (n = 2000 cells for each strain). ***p < 0.001.
Extended Data Fig. 5 S Protein interacts with and promotes the activity of PBP1a.
(a) Representative images of WT, ∆ess, ∆pbp1a, ∆pbp2a and ∆pbp1b cells after a TDL long-pulse labeling. The phase contrast, the TDL channels and the merged images are shown. Scale bars, 2 µm. (b) Violin SuperPlots showing the distribution of the FDAA fluorescence intensity for WT, ∆ess, ∆pbp1a, ∆pbp2a and ∆pbp1b cells after a long period of TDL labeling, from three independent experiments, (n = 2000 cells for each strain). ***p < 0.001 and ns, not significant, p > 0.05. (c) Western immunoblot of whole-cell lysates of WT, mgfp-ess and ∆ess strains revealed with specific anti-aPBPs antibodies and anti-enolase antibody as a loading control. (d) Bacterial two-hybrid analyses. Plasmids, expressing the T25 fragment of the adenylate cyclase protein fused to the N-terminus of PBP1a, DltX or PBP1a-TMDltX or the T18 fused to the N-terminus of S Protein, were co-transformed in E. coli BTH101. The blue coloration indicates positive interactions. The colonies were pictured every 5 h after 24 h of incubation at 25 °C. The data shown are representative of three independent experiments. (e) Coomassie-stained SDS-PAGE of the recombinant proteins used this study (S Protein, PBP1aD39, PBP1aR6 and PBP1aR6(A124T). Proteins are indicated by black triangles. (f) Gel electrophoresis analysis of PBP1a activity upon addition of moenomycin. Synthetic Lipid II (Lys-Dansyl, isoGln) was incubated in presence of either PBP1aD39 or PBP1aR6 or PBP1a(A124T) in absence (- S Protein) or presence of S Protein or S Protein∆PαH (+) with addition of moenomycin 5 µM. The gel was imaged under UV-transillumination. Lipid II (LII) migrates at the front, uncross-linked glycan chains (GC) migrate as bands within the gel. Incubation of S Protein or S Protein∆PαH alone is shown on the right of the gel.
Extended Data Fig. 6 The PαH of S Protein interacts specifically with PBP1a.
(a, b) Predicted complexes of S Protein with PBP1a (a) and either PBP1b, PBP2a, PBP2b, or PBP2x (b) by AlphaFold3. The residues are colored according to the pLDDT score and the corresponding predicted alignment error (PAE) matrices are shown. The AlphaBridge diagram on the right shows the predicted surface interaction between S Protein and the corresponding aPBP, if present.
Extended Data Fig. 7 The PαH of S Protein is required for PBP1a activation.
(a, b) Different representations of the interface between the PαH of S Protein and the helix of GT domain of PBP1a (light green) bearing the catalytic residue (E91) highlighted in red. (a) Transparent surface and ribbon representations for both molecules. (b) Hydrophobicity surface representation. (c) Hydrophobicity surface and ribbon diagram representation for S Protein and PBP1a, respectively. (d) Ribbon diagram and hydrophobicity surface representations for S Protein and PBP1a, respectively. (e) Growth of WT, ess-∆PαH, and all PαH alanine mutant strains in C + Y medium at 37 °C. (f) Western immunoblot of whole-cell lysates of WT, ess-∆PαH, L68A and F71A strains with specific anti-S Protein antibody and anti-enolase antibody as a loading control.
Extended Data Fig. 8 Western immunoblots revealing the interactions between GpsB, PBP1a, PBP2a, S Protein, MacP, MpgA and PgdA.
All Samples were analyzed by immunoblotting using anti-GFP for the detection of bait proteins and either anti-PBP1a, anti-PBP2a, anti-S Protein, anti-MacP, anti-MpgA, anti-PgdA or anti-GpsB antibodies to determine the presence of the corresponding protein. The data shown are representative of three independent experiments. (a) Co-immunoprecipitation of PBP1a, PBP2a, S Protein, MacP, MpgA and PgdA with GFP-GpsB and GFP (control) using anti-GFP antibodies. The green triangle indicates the GFP-GpsB bait. (b) Co-immunoprecipitation of PBP1a, PBP2a, S Protein, MacP, MpgA and PgdA with either GFP-MacP or GFP-S Protein in WT (left panel) or ∆gpsB (right panel) cells. The blue and grey triangle indicate the GFP-MacP and mGFP-S Protein baits, respectively. Native S Protein and MacP are indicated by black triangle whereas their respective GFP-fusions are indicated by green triangle. (c) Co-immunoprecipitation of PBP1a, PBP2a, S Protein, MacP, MpgA and PgdA with either GFP-PgdA or GFP-MpgA in WT (left side) or ∆gpsB (right side) cells. The blue and grey triangle indicate the GFP-PgdA and mGFP-MpgA baits, respectively. Native PgdA and MpgA are indicated by black triangle whereas their respective GFP-fusions are indicated by green triangle. (d) Interaction diagram of GpsB corresponding to (a). Weak interactions are shown as red dashed lines, and strong interactions as red solid lines (e) Interaction diagram of MacP and S Protein corresponding to (b). Weak interactions are shown as black dashed lines, and strong interactions as black solid lines. Interactions strongly affected in absence of GpsB are represented by blue dashed lines. (f) Interaction diagram of PgdA and MpgA corresponding to (c). Weak interactions are shown as black dashed lines, and strong interactions as black solid lines. Interactions strongly affected in absence of GpsB are represented by blue dashed line. (g) Interaction diagram combining data from (d–f).
Extended Data Fig. 9 Structural homology between S Protein (S. pneumoniae) and YpbE (B. subtilis).
(a) AlphaFold3 models of S Protein and YpbE. The predicted transmembrane segment and the LysM domain of both proteins are shown in blue and green, respectively. Intrinsically disordered regions are shown in gray. The predicted α-helix of S Protein is shown in orange. (b) The best model prediction of YpbE by AlphaFold3. The residues are colored according to the pLDDT score and the corresponding predicted alignment error (PAE) matrices are shown for each model.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–3.
Supplementary Tables (download XLSX )
Supplementary Tables 1–6.
Source data
Source Data Fig. 4 (download PDF )
Unprocessed western blots and gels.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots and gels.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 8 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Millat, H., Falcou, C., Lenoir, C. et al. Streptococcus pneumoniae S protein activates PBP1a to regulate peptidoglycan remodelling and cell division. Nat Microbiol 11, 301–316 (2026). https://doi.org/10.1038/s41564-025-02210-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02210-5