Abstract
Horizontal transfer of small non-conjugative plasmids is primarily attributed to transformation, transduction or comobilization with conjugative elements; however, transfer through intercellular membranous nanotube conduits can also occur. Here we show that nanotube-dependent plasmid exchange (NPex) operates bidirectionally between bacteria, enabling plasmid donation and, to a lesser extent, plasmid acquisition. We identified a Bacillus subtilis isolate, BSB1, deficient in NPex and show that a prophage-encoded factor, YokF, blocks plasmid transmission. YokF is an endonuclease that localizes to the membrane of donor bacteria, where it interacts with the nanotube component, FlhA, to impede plasmid transfer through DNA degradation. We further show that YokF provides an advantage to donor bacteria by restricting the sharing of beneficial plasmids with competing neighbouring cells. Bioinformatics and functional analyses revealed that YokF homologues are widespread across Gram-positive bacteria, representing a conserved family of gatekeepers that restrict plasmid flow via NPex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All relevant data supporting the findings of this study are available within the article and its Supplementary Information. The sequencing data generated in this study have been deposited in the DDBJ/EMBL/GenBank database under BioProject accession number PRJNA1098594, with SRA accession numbers SRX24206248 (PY79) and SRX24206250 (BSB1). Additional source data can be found via figshare at https://figshare.com/s/8d959561f0c8861f8081 (ref. 90). Correspondence and requests for materials should be addressed to the corresponding authors. Source data are provided with this paper.
Code availability
The MATLAB script generated for this study is available via Zenodo at https://doi.org/10.5281/zenodo.17564647 (ref. 78).
References
Partridge, S. R., Kwong, S. M., Firth, N. & Jensen, S. O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. 31, e00088-17 (2018).
Ochman, H., Lawrence, J. G. & Groisman, E. A. Lateral gene transfer and the nature of bacterial innovation. Nature 405, 299–304 (2000).
Grohmann, E., Muth, G. & Espinosa, M. Conjugative plasmid transfer in Gram-positive bacteria. Microbiol. Mol. Biol. Rev. 67, 277–301 (2003).
Bottery, M. J. Ecological dynamics of plasmid transfer and persistence in microbial communities. Curr. Opin. Microbiol. 68, 102152 (2022).
Rodriguez-Beltran, J., DelaFuente, J., Leon-Sampedro, R., MacLean, R. C. & San Millan, A. Beyond horizontal gene transfer: the role of plasmids in bacterial evolution. Nat. Rev. Microbiol. 19, 347–359 (2021).
Stecher, B. et al. Gut inflammation can boost horizontal gene transfer between pathogenic and commensal Enterobacteriaceae. Proc. Natl Acad. Sci. USA 109, 1269–1274 (2012).
Ghigo, J. M. Natural conjugative plasmids induce bacterial biofilm development. Nature 412, 442–445 (2001).
Waksman, G. From conjugation to T4S systems in Gram-negative bacteria: a mechanistic biology perspective. EMBO Rep. 20, e47012 (2019).
Goessweiner-Mohr, N., Arends, K., Keller, W. & Grohmann, E. Conjugation in Gram-positive bacteria. Microbiol. Spectr. 2, PLAS-0004-2013 (2014).
Virolle, C., Goldlust, K., Djermoun, S., Bigot, S. & Lesterlin, C. Plasmid transfer by conjugation in Gram-negative bacteria: from the cellular to the community level. Genes 11, 1239 (2020).
Redondo-Salvo, S. et al. Pathways for horizontal gene transfer in bacteria revealed by a global map of their plasmids. Nat. Commun. 11, 3602 (2020).
Acman, M., van Dorp, L., Santini, J. M. & Balloux, F. Large-scale network analysis captures biological features of bacterial plasmids. Nat. Commun. 11, 2452 (2020).
Guglielmetti, S., Mora, D. & Parini, C. Small rolling circle plasmids in Bacillus subtilis and related species: organization, distribution, and their possible role in host physiology. Plasmid 57, 245–264 (2007).
Zhuang, M. et al. Horizontal plasmid transfer promotes antibiotic resistance in selected bacteria in Chinese frog farms. Environ. Int. 190, 108905 (2024).
Ramsay, J. P. & Firth, N. Diverse mobilization strategies facilitate transfer of non-conjugative mobile genetic elements. Curr. Opin. Microbiol. 38, 1–9 (2017).
von Wintersdorff, C. J. H. et al. Dissemination of antimicrobial resistance in microbial ecosystems through horizontal gene transfer. Front. Microbiol. 7, 173 (2016).
Coluzzi, C., Garcillán-Barcia, M. P., de la Cruz, F. & Rocha, E. P. C. Evolution of plasmid mobility: origin and fate of conjugative and nonconjugative plasmids. Mol. Biol. Evol. https://doi.org/10.1093/molbev/msac115 (2022).
Marinacci, B., Krzyzek, P., Pellegrini, B., Turacchio, G. & Grande, R. Latest update on outer membrane vesicles and their role in horizontal gene transfer: a mini-review. Membranes https://doi.org/10.3390/membranes13110860 (2023).
Schwechheimer, C. & Kuehn, M. J. Outer-membrane vesicles from Gram-negative bacteria: biogenesis and functions. Nat. Rev. Microbiol. 13, 605–619 (2015).
Kahn, M. E., Barany, F. & Smith, H. O. Transformasomes: specialized membranous structures that protect DNA during Haemophilus transformation. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.80.22.6927 (1983).
Dubey, G. P. & Ben-Yehuda, S. Intercellular nanotubes mediate bacterial communication. Cell 144, 590–600 (2011).
Bhattacharya, S. et al. A ubiquitous platform for bacterial nanotube biogenesis. Cell Rep. 27, 334–342 e310 (2019).
Morawska, L. P. & Kuipers, O. P. Cell-to-cell non-conjugative plasmid transfer between Bacillus subtilis and lactic acid bacteria. Microb. Biotechnol. 16, 784–798 (2023).
Benomar, S. et al. Nutritional stress induces exchange of cell material and energetic coupling between bacterial species. Nat. Commun. 6, 6283 (2015).
Pande, S. et al. Metabolic cross-feeding via intercellular nanotubes among bacteria. Nat. Commun. 6, 6238 (2015).
Stempler, O. et al. Interspecies nutrient extraction and toxin delivery between bacteria. Nat. Commun. 8, 315 (2017).
Baidya, A. K., Rosenshine, I. & Ben-Yehuda, S. Donor-delivered cell wall hydrolases facilitate nanotube penetration into recipient bacteria. Nat. Commun. 11, 1938 (2020).
Takahashi, C. et al. Electron microscopy of Staphylococcus epidermidis fibril and biofilm formation using image-enhancing ionic liquid. Anal. Bioanal. Chem. 407, 1607–1613 (2015).
Wei, X., Vassallo, C. N., Pathak, D. T. & Wall, D. Myxobacteria produce outer membrane-enclosed tubes in unstructured environments. J. Bacteriol. 196, 1807–1814 (2014).
Chakravortty, D., Rohde, M., Jager, L., Deiwick, J. & Hensel, M. Formation of a novel surface structure encoded by Salmonella pathogenicity island 2. EMBO J. 24, 2043–2052 (2005).
McCaig, W. D., Koller, A. & Thanassi, D. G. Production of outer membrane vesicles and outer membrane tubes by Francisella novicida. J. Bacteriol. 195, 1120–1132 (2013).
Patel, N., Yamada, Y. & Azam, F. Bacterial nanotubes as intercellular linkages in marine assemblages. Front. Mar. Sci. https://doi.org/10.3389/fmars.2021.768814 (2021).
Boopathi, S. et al. Unveiling nanotubes-mediated communication: Enterococcus faecalis countering Salmonella ser. Typhi—in vitro and in vivo insights. Micro. Pathog. 184, 106387 (2023).
Kaplan, M. et al. In situ imaging of bacterial outer membrane projections and associated protein complexes using electron cryo-tomography. Elife https://doi.org/10.7554/eLife.73099 (2021).
Gerdes, H. H. & Carvalho, R. N. Intercellular transfer mediated by tunneling nanotubes. Curr. Opin. Cell Biol. https://doi.org/10.1016/j.ceb.2008.03.005 (2008).
Rosenshine, I., Tchelet, R. & Mevarech, M. The mechanism of DNA transfer in the mating system of an archaebacterium. Science https://doi.org/10.1126/science.2818746 (1989).
Angulo-Canovas, E. et al. Direct interaction between marine cyanobacteria mediated by nanotubes. Sci. Adv. 10, eadj1539 (2024).
Pal, R. R. et al. Pathogenic E. coli extracts nutrients from infected host cells utilizing injectisome components. Cell 177, 683–696.e618 (2019).
Minamino, T. Protein export through the bacterial flagellar type III export pathway. Biochim. Biophys. Acta 1843, 1642–1648 (2014).
Pospisil, J. et al. Bacterial nanotubes as a manifestation of cell death. Nat. Commun. 11, 4963 (2020).
Meijer, W. J. et al. Rolling-circle plasmids from Bacillus subtilis: complete nucleotide sequences and analyses of genes of pTA1015, pTA1040, pTA1050, and pTA1060, and comparisons with related plasmids from Gram-positive bacteria. FEMS Microbiol. Rev. 21, 337–368 (1998).
Tanaka, T. & Koshikawa, T. Isolation and characterization of four types of plasmids from Bacillus subtilis (natto). J. Bacteriol. https://doi.org/10.1128/jb.131.2.699-701.1977 (1977).
Meijer, W. J. et al. The endogenous Bacillus subtilis (natto) plasmids pTA1015 and pTA1040 contain signal peptidase-encoding genes: identification of a new structural module on cryptic plasmids. Mol. Microbiol. 17, 621–631 (1995).
Uozumi, T., Ozaki, A., Beppu, T. & Arima, K. New cryptic plasmid of Bacillus subtilis and restriction analysis of other plasmids found by general screening. J. Bacteriol. https://doi.org/10.1128/jb.142.1.315-318.1980 (1980).
Dubey, G. P. et al. Architecture and characteristics of bacterial nanotubes. Dev. Cell 36, 453–461 (2016).
Sørensen, S. J. et al. Studying plasmid horizontal transfer in situ: a critical review. Nat. Rev. Microbiol. 3, 700–710 (2005).
Thorne, C. B. Transduction in Bacillus subtilis. J. Bacteriol. 83, 106–111 (1962).
Smith, D. R., Kearns, D. B. & Burton, B. M. ComI inhibits transformation in Bacillus subtilis by selectively killing competent cells. J. Bacteriol. 206, e0041323 (2024).
Holscher, T. et al. Impaired competence in flagellar mutants of Bacillus subtilis is connected to the regulatory network governed by DegU. Environ. Microbiol. Rep. 10, 23–32 (2018).
Nicolas, P. et al. Condition-dependent transcriptome reveals high-level regulatory architecture in Bacillus subtilis. Science https://doi.org/10.1126/science.1206848 (2012).
Auchtung, M. et al. Regulation of a Bacillus subtilis mobile genetic element by intercellular signaling and the global DNA damage response. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.0505835102 (2005).
Kohm, K. & Hertel, R. The life cycle of SPβ and related phages. Arch. Virol. 166, 2119–2130 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Söding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. https://doi.org/10.1093/nar/gki408 (2005).
Coudert, E. et al. Annotation of biologically relevant ligands in UniProtKB using ChEBI. Bioinformatics https://doi.org/10.1093/bioinformatics/btac793 (2023).
Kiedrowski, M. R. et al. Staphylococcus aureus Nuc2 is a functional, surface-attached extracellular nuclease. PLoS ONE 9, e95574 (2014).
Arber, W. & Linn, S. DNA modification and restriction. Annu. Rev. Biochem. https://doi.org/10.1146/annurev.bi.38.070169.002343 (1969).
Shaw, L. P., Rocha, E. P. C. & MacLean, R. C. Restriction-modification systems have shaped the evolution and distribution of plasmids across bacteria. Nucleic Acids Res. https://doi.org/10.1093/nar/gkad452 (2023).
Kohm, K. et al. The Bacillus phage SPβ and its relatives: a temperate phage model system reveals new strains, species, prophage integration loci, conserved proteins and lysogeny management components. Environ. Microbiol. https://doi.org/10.1111/1462-2920.15964 (2022).
Sakamoto, J. J., Sasaki, M. & Tsuchido, T. Purification and characterization of a Bacillus subtilis 168 nuclease, YokF, involved in chromosomal DNA degradation and cell death caused by thermal shock treatments. J. Biol. Chem. 276, 47046–47051 (2001).
Konkol, M. A., Blair, K. M. & Kearns, D. B. Plasmid-encoded ComI inhibits competence in the ancestral 3610 strain of Bacillus subtilis. J. Bacteriol. 195, 4085–4093 (2013).
Durieux, I. et al. Diverse conjugative elements silence natural transformation in Legionella species. Proc. Natl Acad. Sci. USA 116, 18613–18618 (2019).
Jaskolska, M., Adams, D. W. & Blokesch, M. Two defence systems eliminate plasmids from seventh pandemic Vibrio cholerae. Nature 604, 323–329 (2022).
Loeff, L. et al. Molecular mechanism of plasmid elimination by the DdmDE defense system. Science 385, 188–194 (2024).
Yang, X. Y., Shen, Z., Wang, C., Nakanishi, K. & Fu, T. M. DdmDE eliminates plasmid invasion by DNA-guided DNA targeting. Cell 187, 5253–5266.e5216 (2024).
Bravo, J. P. K., Ramos, D. A., Fregoso Ocampo, R., Ingram, C. & Taylor, D. W. Plasmid targeting and destruction by the DdmDE bacterial defence system. Nature 630, 961–967 (2024).
Hausner, M. & Wuertz, S. High rates of conjugation in bacterial biofilms as determined by quantitative in situ analysis. Appl. Environ. Microbiol. 65, 3710–3713 (1999).
Li, B. et al. Real-time study of rapid spread of antibiotic resistance plasmid in biofilm using microfluidics. Environ. Sci. Technol. 52, 11132–11141 (2018).
Madsen, J. S., Burmolle, M., Hansen, L. H. & Sorensen, S. J. The interconnection between biofilm formation and horizontal gene transfer. FEMS Immunol. Med. Microbiol. 65, 183–195 (2012).
Savage, V. J., Chopra, I. & O’Neill, A. J. Staphylococcus aureus biofilms promote horizontal transfer of antibiotic resistance. Antimicrob. Agents Chemother. 57, 1968–1970 (2013).
Bhattacharya, S. et al. Flagellar rotation facilitates the transfer of a bacterial conjugative plasmid. EMBO J. 44, 587–611 (2025).
Sambrook, J., Fritsch, E. R. & Maniatis, T. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 1989).
Harwood, C. R. & Cutting, S. M. Molecular Biological Methods for Bacillus (Wiley, 1990).
Branda, S. S., González-Pastor, J. E., Ben-Yehuda, S., Losick, R. & Kolter, R. Fruiting body formation by Bacillus subtilis. Proc. Natl Acad. Sci. USA 98, 11621–11626 (2001).
Kearns, D. B. & Losick, R. Swarming motility in undomesticated Bacillus subtilis. Mol. Microbiol. https://doi.org/10.1046/j.1365-2958.2003.03584.x (2003).
Bejerano-Sagie, M. et al. A checkpoint protein that scans the chromosome for damage at the start of sporulation in Bacillus subtilis. Cell https://doi.org/10.1016/j.cell.2006.03.039 (2006).
Dieterich, D. C. et al. Labeling, detection and identification of newly synthesized proteomes with bioorthogonal non-canonical amino-acid tagging. Nat. Protoc. 2, 532–540 (2007).
Yakovian, O. Bacteria simulation. Zenodo https://doi.org/10.5281/zenodo.17564647 (2025).
Stülke, J., Grüppen, A., Bramkamp, M. & Pelzer, S. Bacillus subtilis, a swiss army knife in science and biotechnology. J. Bacteriol. https://doi.org/10.1128/jb.00102-23 (2023).
Teufel, F. et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. 40, 1023–1025 (2022).
Blum, M. et al. The InterPro protein families and domains database: 20 years on. Nucleic Acids Res. https://doi.org/10.1093/nar/gkaa977 (2021).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Terzian, P. et al. PHROG: families of prokaryotic virus proteins clustered using remote homology. NAR Genom. Bioinform. https://doi.org/10.1093/nargab/lqab067 (2021).
Youngman, P., Perkins, J. B. & Losick, R. Construction of a cloning site near one end of Tn917 into which foreign DNA may be inserted without affecting transposition in Bacillus subtilis or expression of the transposon-borne erm gene. Plasmid https://doi.org/10.1016/0147-619x(84)90061-1 (1984).
Itaya, M., Sakaya, N., Matsunaga, S., Fujita, K. & Kaneko, S. Conjugational transfer kinetics of pLS20 between Bacillus subtilis in liquid medium. Biosci. Biotechnol. Biochem. https://doi.org/10.1271/bbb.70.740 (2006).
Zhou, B. et al. Dormant bacterial spores encrypt a long-lasting transcriptional program to be executed during revival. Mol. Cell https://doi.org/10.1016/j.molcel.2023.10.010 (2023).
Ando, R. et al. StayGold variants for molecular fusion and membrane-targeting applications. Nat. Methods 21, 648–656 (2024).
Bron, S. et al. Protein secretion and possible roles for multiple signal peptidases for precursor processing in bacilli. J. Biotechnol. 64, 3–13 (1998).
Krásný, L., Vacík, T., Fučík, V. & Jonák, J. Cloning and characterization of the str operon and elongation factor Tu expression in Bacillus stearothermophilus. J. Bacteriol. https://doi.org/10.1128/jb.182.21.6114-6122.2000 (2000).
Gopu, V. et al. A family of endonucleases that block nanotube-mediated plasmid dissemination. figshare https://figshare.com/s/8d959561f0c8861f8081 (2026).
Acknowledgements
We thank the members of the S.B.-Y. and I.R. laboratories for their insightful discussions. We are also grateful to D. Rudner (Harvard University), O. P. Kuipers (University of Groningen) and L. Krásný (Academy of Sciences of the Czech Republic) for providing bacterial strains and plasmids. We thank M. Kumar (Hebrew University) and A. Nasereddin (Core Research Facilities, Hebrew University) for technical support. V.G. was supported by the Planning and Budgeting Committee of Israel postdoctoral fellowship, and S.B. by the Golda Meir postdoctoral fellowship. This work was funded by the European Research Council Synergy grant (810186) awarded to S.B.-Y. and I.R., and by the German Research Foundation (DFG) Priority Program SPP2389 awarded to S.B.-Y. and B.M.
Author information
Authors and Affiliations
Contributions
Conceptualization: V.G., S.B., I.R. and S.B.-Y. Methodology: V.G., S.B., M.B.-S., M.Z., Y.N., O.Y., B.S., M.R., M.K.G., T.K., B.M., I.R. and S.B.-Y. Investigation: V.G., S.B., M.B.-S., O.Y., M.Z., M.R., I.R. and S.B.-Y. Visualization: V.G., S.B., M.B.-S., M.K.G., I.R. and S.B.-Y. Funding acquisition: B.M., S.B.-Y. and I.R. Project administration: S.B.-Y., M.R. and I.R. Supervision: S.B.-Y. and I.R. Writing – original draft: V.G., S.B., M.B.-S., M.R., I.R. and S.B.-Y.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks James Boedicker, Ines Mandic Mulec, Galain Williams and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 NPex is inhibited by SPβ prophage.
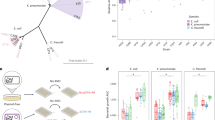
(A) NPex assay controls: PY79-derived donor strains: WT (PY79), or isogenic ΔCORE (ΔfliO-flhA::tet, Pfla/che-flhF) carrying p6.6 kb (pHB201/cat, erm) (GV478 and GV491, respectively) were mixed in 1:1 ratio with kanamycin resistant WT (SB513: amyE::PIPTG-gfp-kan) or isogenic ΔCORE (SH13: ∆fliO-flhA::tet, Pfla/che-flhF, amyE::PIPTG-gfp-kan) recipients. The used donor (black) and recipient (blue) are indicated. The mixtures were incubated for 4 h without selection. Equal volumes of cells were then spread over LB plates without antibiotics (10−5 dilution, left panel) or over plates containing chloramphenicol, erythromycin, and kanamycin to select for trans-recipients (middle panels). As controls, donor and recipient strains were grown separately under similar conditions and plated in parallel (right panels). Plates were imaged using a fluorescent imaging system, in colorimetric and fluorescence modes, as indicated. All colonies grown on the triple-selective plates were GFP-positive, indicating that they only derived from recipient cells, harbouring GFP, that acquired the plasmid. Shown are representative images of selective plates, based on 3 independent experiments. (B) Confirmation of plasmid transfer to recipients by NPex: NPex assay was conducted as in (A) using WT PY79-derived donor (GV478) carrying p6.6 kb (pHB201/cat, erm) and kanamycin resistance recipient (SB513: amyE::PIPTG-gfp-kan). DNA was extracted from 10 randomly isolated trans-recipients, and from donor (D) and recipient (R) parental strains, and subjected to PCR using p6.6 kb specific primers. Shown is an agarose gel electrophoresis analysis of a PCR product obtained from p6.6 kb. M: Molecular weight marker [bp]. (C) NPex is competence and motility independent: PY79-derived donor strains: WT (PY79) (GV478), and Δhag (GB01: ∆hag::erm), carrying p6.6 kb (pHB201/cat, erm), were mixed in a 1:1 ratio with kanamycin resistance recipient strains: WT (SB513: amyE::PIPTG-gfp-kan), ΔcomK (GB252: ΔcomK::tet, amyE::PIPTG-gfp-kan), and Δhag (GD268: ∆hag::erm, amyE::PIPTG-gfp-kan). The used donor (black) and recipient (blue) pairs are indicated. The mixtures were incubated for 4 h without selection, and equal volumes of cells were then spread over plates containing chloramphenicol, erythromycin, and kanamycin to select for trans-recipients. NPex efficiency was determined as the ratio of trans-recipients (CFU)/total recipients (CFU). Shown is NPex efficiency (%) relative to WT → WT pair. Data is presented as mean values and SEM, based on 4 independent experiments. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to the WT → WT samples. P values: WT→ΔcomK (ns) = 1.55×10−1, Δhag→Δhag (ns) = 1.89×10−1. (D) Natural transformation is negligible under NPex assay conditions: Wild-type PY79-derived donor (GV478) and kanamycin resistance recipient (SB513: amyE::PIPTG-gfp-kan) were mixed in 1:1 ratio and incubated for 4 h without selection, in the absence (Control) or presence of DNaseI. In parallel, recipient cells were incubated for 4 h in LB medium supplemented with 100 ng plasmid DNA p6.6 kb (pHB201/cat, erm), without selection. Equal volumes of cells were then spread over plates containing chloramphenicol, erythromycin, and kanamycin to select for trans-recipients. NPex efficiency was calculated as in (C). Data is presented as mean values and SEM, based on 3 independent experiments. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to the control samples. P values: DNaseI (ns) = 3.68×10−1, p6.6KB (***) = 1.93×10−5. (E) NPex is transduction- and membrane vesicle-independent: Wild-type PY79-derived donor (GV478) and kanamycin resistance recipient (SB513: amyE::PIPTG-gfp-kan) were mixed in 1:1 ratio and incubated for 4 h without selection. Equal volumes of cells were then spread over plates containing chloramphenicol, erythromycin, and kanamycin to select for trans-recipients (Control). In parallel, donor and recipient cells were cultured separately for 3 h without selection. The spent medium of the recipient strain was then replaced with filtered supernatant from the donor strain culture, followed by incubation for 1 h. After total incubation of 4 h, equal volumes of cells were then spread over plates containing chloramphenicol, erythromycin, and kanamycin to select for trans-recipients (Test). Plasmid transfer efficiency was determined as the ratio of trans-recipients (CFU)/total recipients (CFU). Data is presented as mean values and SEM, based on 4 independent experiments. (F) NPex is limited by plasmid size: PY79-derived donor strains: WT (PY79) carrying pLS20ΔvirB4/cm (SH346) or pLS20-mini (MR34) were mixed in 1:1 ratio with kanamycin resistance WT recipient strain (SB513: amyE::PIPTG-gfp-kan). The mixtures were incubated for 4 h without selection, and equal volumes of cells were then spread over plates containing chloramphenicol, and kanamycin to select for recipients that acquired the plasmids. Plasmid transfer efficiencies were determined as in (C). Data are presented as mean values and SEM, based on 4 independent experiments. ND, not detected. (G) NPex efficiency is elevated under biofilm inducing conditions: PY79-derived donor strain (GV478) and kanamycin resistance recipient (SB513: amyE::PIPTG-gfp-kan) were mixed in 1:1 ratio and incubated for 4 h or 20 h (biofilm inducing condition) without selection. In parallel, a 10:1 ratio of donor:recipient was incubated for 20 h without selection. Equal volumes of cells were then spread over plates containing chloramphenicol, and kanamycin to select for recipients that acquired the plasmids. Plasmid transfer efficiencies were determined as in (C). Data are presented as mean values and SEM, based on 3 independent experiments. Statistical significance was calculated using Student’s t-test (two-sided) for each sample. P values: Compared to the control sample (1:1 at 4 h) 1:1 20 h- (***) = 1.53×10−4, 10:1 20 h- (**) = 5.06×10−3, Compared to the 1:1 at 20 h- (*) = 3.29×10−2. (H) Mapping NPex inhibition genomic locus: Donor strains: WT (PY79) (GV478), and PY79-derived strains harboring V1BSB1(GV669), or V2BSB1 (GV671), and WT (BSB1) (GV345), and BSB1-derived strains harboring V1PY79 (GV377), or V2PY79 (GV384), all carrying p6.6 kb (pHB201/cat, erm), were mixed in a 1:1 ratio with kanamycin resistance recipient strains (SB513, GV670, GV672, GV372, GV375, and GV382), harboring equivalent genotypes, but lacking a plasmid. The mixtures were incubated for 4 h without selection, and equal volumes of cells were then spread over plates selective for trans-recipients. A detailed description of the used donor and recipient pairs and antibiotic selections employed are listed in Supplementary Table 1. NPex efficiency was calculated as in (C). Shown is NPex efficiency (%) relative to WT → WT (PY79) pair. Data are presented as mean values and SEM, based on 4 independent experiments. ND, not detected. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to the WT (PY79) samples. P values: V1BSB1 (ns) = 9.98×10−1, V2BSB1 (ns) = 7.50×10−1. (I) SPβ prophage inhibits NPex: Donor strains: WT (PY79) (GV478), WT (BSB1) (GV345) and BSB1-derived strains, ΔL1 (GV674: ΔlrpC-mntH::tet), ΔL2 (SH624: ΔydcL-yddM::tet), and ΔL3 (SH625: ΔsprB-sprA::tet), all carrying p6.6 kb (pHB201/cat, erm), were mixed in a 1:1 ratio with kanamycin resistance recipient strains (amyE::PIPTG-gfp-kan) harboring equivalent genotypes (SB513, GV372, GV675, SH620, and SH621), but lacking a plasmid. The mixtures were incubated for 4 h without selection, and equal volumes of cells were then spread over plates containing chloramphenicol, erythromycin, and kanamycin to select for trans-recipients. NPex efficiency was calculated as described in (C). Data are presented as mean values and SEM, based on 4 independent experiments. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to the WT (PY79) samples. P values: ΔSPβ (ns) = 1.47×10−1.
Extended Data Fig. 2 NPex is inhibited by YokF.
(A) Sequential deletion analysis of SPβ genome: Schematic illustration of the sequential deletion analysis of SPβ prophage genome in BSB1 for identifying the NPex inhibitory region. SPβ was divided into 3 regions (R1: sprB-yopS, R2: yopP-youB, and R3: yomK-sprA). YokFE operon is located in R3. (B) Sequential deletion analysis: BSB1-derived donor strains: WT (BSB1) (GV345), ΔSPβ (SH625), ΔR1 (GV119), ΔR2 (GV239), ΔR3 (SH627), ΔyokFE (GV428), ΔyokF (GV417), and ΔyokE (GV419), all carrying p6.6 kb (pHB201/cat, erm), were mixed in a 1:1 ratio with kanamycin resistance recipient strains: (GV372, SH621, GV118, GV238, SH623, GV427, GV416, and GV418), harboring equivalent genotypes but lacking a plasmid. The mixtures were incubated for 4 h without selection, and equal volumes of cells were then spread over plates selective for trans-recipients. R1-R3 regions correspond to (A). A detailed description of the used donor and recipient pairs and antibiotic selections employed are listed in Supplementary Table 1. NPex efficiency was determined as the ratio of trans-recipients (CFU)/total recipients (CFU). Data are presented as mean values and SEM, based on 4 independent experiments. ND, not detected. (C) NPex is unaffected by conditioned medium derived from YokF expressing strain: PY79-derived donor (GV478) and kanamycin resistance recipient (SB513: amyE::PIPTG-gfp-kan) were mixed in a 1:1 ratio, and resuspended in 0.22 µm filter-sterilized culture supernatants (sup) derived from either PY79 (lacking yokF) or BSB1 (harboring yokF) cultures. The mixtures were incubated for 4 h without selection, and equal volumes of cells were then spread over plates containing chloramphenicol, erythromycin, and kanamycin to select for trans-recipients. NPex efficiency was calculated as in (B). Data is presented as mean values and SEM, based on 3 independent experiments. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to the PY79 sup samples. P-values: BSB1 sup (ns) = 3.73×10−1. (D) Natural transformation is unaffected by YokF: WT (PY79) or PY79 expressing YokF, WT::yokF (GV462: sacA::PyokF-yokFBSB1-spec) cells were grown under competence-inducing conditions with either genomic DNA (100 ng) encoding a tetracycline resistance gene isolated from (GV600: ΔhelD::tet), or p6.6 kb (pHB201/cat, erm) (100 ng). An equal volume of cells was spread over plates containing tetracycline or chloramphenicol to select for transformants, and on antibiotic-free media for the total population. Transformation efficiency was determined as the ratio of transformants (CFU)/total population (CFU). Data are presented as mean values and SEM, based on 4 independent experiments. Statistical significance was calculated using Student’s t-test (two-sided) for each set compared to their respective WT samples. P-values: yokF p6.6 (*) = 2.22×10−2, yokF gDNA (ns) = 1.27×10−1. (E) YokF does not affect phage sensitivity: WT (PY79) or PY79 expressing YokF, WT::yokF (GV462: sacA::PyokF-yokFBSB1-spec) cells were grown to mid-logarithmic phase and infected with SPO1 or SPP1 phages. After phage attachment, infected cells were plated onto MB agar plates. Infection efficiency was determined as the ratio of plaque forming units (PFU) between WT::yokF and WT. Shown is relative infection efficiency (%). Data are presented as mean values and SEM, based on 3 independent experiments. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to the WT samples. P-values: yokF (SPO1) (ns) = 4.28×10−1, yokF (SPP1) (*) = 4.28×10−2. (F) YokF does not affect conjugation: PY79-derived donor strains: WT (PY79) or PY79 expressing YokF, WT::yokF (sacA::PyokF-yokFEBSB1-spec) carrying pLS20/cm (SH337 and GV454, respectively) were mixed in 1:1 ratio with kanamycin resistance WT recipient strain (SB513: amyE::PIPTG-gfp-kan). The mixtures were incubated for 20 min without selection, and equal volumes of cells were then spread over plates containing chloramphenicol and kanamycin to select for recipients that acquired the plasmids. Conjugation efficiency was determined as the ratio of transconjugants (CFU)/total recipients (CFU). Data is presented as mean values and SEM, based on 4 independent experiments. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to the WT samples. P-values: yokF (ns) = 5.80×10−1. (G) Plasmid stability is unaffected by YokF: WT (PY79) (GV478), PY79 expressing YokF PY79::yokF (GV469: sacA:: PyokF-yokFBSB1-spec), PY79::PIPTG-yokF (GV480: amyE::PIPTG-yokFBSB1-spec), WT (BSB1) (GV345), and isogenic ΔyokF (GV417: ΔyokF::tet) strains, all carrying p6.6 kb (pHB201/cat, erm), were grown with antibiotics (generation 0). PY79::PIPTG-yokF was grown in the presence of IPTG. Cells were refreshed every 3 h ( ~ 10 generations) without antibiotics until 30 generations. Every 10 generations, equal volumes of cells were spread over plates containing chloramphenicol and antibiotic-free media to estimate the numbers of plasmid-containing cells and the total population, respectively. The graph illustrates plasmid stability as the ratio of plasmid-containing cells (CFU)/total population (CFU). Data is presented as mean values and SEM, based on 4 independent experiments. (H) YokF does not affect plasmid copy number: WT (GV478) or PY79 expressing YokF, WT::yokF (GV469: sacA::PyokF-yokFBSB1-spec) strains, carrying p6.6 kb (pHB201/cat, erm) were grown to OD600 1.0 and plasmid levels were determined by qPCR. Shown are relative plasmid abundance normalized to chromosomal DNA level (amyE locus). Data is presented as average values and SEM of 3 independent biological repeats. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to the WT samples. P-values: yokF (ns) = 5.82×10−1.
Extended Data Fig. 3 YokF does not inhibit nanotube formation or protein exchange.
(A) Nanotube formation appears unaffected by YokF: WT (PY79), PY79 expressing YokF PY79::yokF (GV462: sacA:: PyokF-yokFBSB1-spec), WT (BSB1) (GV345), and isogenic ΔyokF (GV411: ΔyokF::tet) were grown to the mid-logarithmic phase, spotted onto EM grids, incubated on LB agar plates for 4 h at 37 °C, and visualized by HR-SEM. Arrows indicate intercellular nanotubes connecting neighbouring cells. This experiment was repeated 3 times with similar results. Scale bar 0.5 µm. (B) Protein exchange is unaffected by YokF: WT (PY79) (SB463: amyE::PIPTG-cat-spec), isogenic ΔCORE (SH17: ΔfliO-flhA::tet, Pfla/che-flhF, amyE::PIPTG-cat-spec), WT::yokF (GV274: sacA::yokFBsB1-spec, amyE::PIPTG-cat-spec), WT (BSB1) (GV373: amyE::PIPTG-cat-spec), isogenic ΔCORE (GV494: ΔfliO-flhA::tet, Pfla/che-flhF, amyE::PIPTG-cat-spec), and isogenic ΔyokF (GV456: ΔyokF::tet, amyE::PIPTG-cat-spec) were mixed in a 1:1 ratio with kanamycin resistance strains (amyE::PIPTG-gfp-kan) harboring equivalent genotypes (SB513, SH13, GV468, GV372, GV107, and GV416). The mixtures were incubated for 4 h without selection, and equal volumes of cells were spotted onto LB agar (control) and LB agar with chloramphenicol and kanamycin to evaluate protein exchange. Plates were photographed after 18 h of incubation. Due to surface tension, bacterial cells are deposited in the circumference of the spots at a higher density, thereby leading to growth of bacterial cells in a ring-like pattern. (C) Motility is unaffected by YokF: For motility assay, PY79 (WT), isogenic ΔCORE (SH9: ΔfliO-flhA::tet, Pfla/che-flhF), WT::yokF (GV462: sacA::yokFBsB1-spec), BSB1(WT), isogenic ΔCORE (GV582: ΔfliO-flhA::tet, Pfla/che-flhF), and isogenic ΔyokF (GV411: ΔyokF::tet) strains were grown to the mid-logarithmic phase and spotted onto 0.3% agar LB plates. Plates were incubated for 7 h at 37 °C before being photographed.
Extended Data Fig. 4 YokF provides growth advantage by limiting plasmid donation.
(A) Effect of YokF on strain competition at varying donor-to-recipient ratios: Computational simulation of a competition between donor bacteria (kin) harboring (+) or lacking (-) yokF, both carrying a beneficial (antibiotic resistance) non-conjugative plasmid, and recipient bacteria (non-kin) lacking plasmid. Simulations were performed either under non-restricting conditions (-Antibiotics) or under conditions mimicking sub-lethal antibiotic pressure ( + Antibiotics). Plasmid transfer rate was set to 0% ( + yokF) or 5% (-yokF) every 10 min. The simulations started with different ratios of donor and recipient cells as indicated, donor: recipient 100:500 (upper panels), or donor: recipient 500:100 (lower panels). The simulations continued until the total bacterial count exceeded 106 cells. Division times were fixed at 40 min for donor cells (with or without antibiotic), 38 min (-Antibiotic) or 60 min ( + Antibiotic) for plasmid-free recipient cells, and 38 min for trans-recipient (non-kin+plasmid) cells. (B) Impact of trans-recipient generation time on strain competition following NPex: Simulations started with 100 cells for each donor or recipient bacteria and continued until the total bacterial count exceeded 106 cells. Plasmid transfer rate was set to 5% every 10 min in the presence of antibiotic. Division times were fixed at 40 min for donor cells (blue), 60 min for plasmid-free recipient cells, and varied from 30 to 50 min for trans-recipient (non-kin+plasmid) cells (red). (C) Impact of NPex rate on strain competition: Simulations started with 100 cells for each donor or recipient bacteria and continued until the total bacterial count exceeded 106 cells. Plasmid transfer rate varied from 0% to 15% every 10 min (0% represents donor containing yokF) in the presence of antibiotic. Division times were fixed at 40 min for donor cells (blue), 60 min for plasmid-free recipient cells, and 30 min for trans-recipient (non-kin+plasmid) cells (red).
Extended Data Fig. 5 YokF nuclease activity is modulated and essential for NPex inhibition.
(A) YokF TNase domain structure: AlphaFold-predicted structure of YokF TNase domain, emphasizing the potential active and binding site residues D79, R93, D98, T99, E101, and R144 clustered within the TNase domain. High pLDDT (predicted local difference test) value is shown in a blue ribbon, indicating a high level of confidence in the structural prediction. (B) YokF TNase mutants are NPex deficient: Upper panel: BSB1-derived donor strains: ΔyokF (GV417), yokFD79H (GV602), yokFR93D (GV576), yokFD98H-T99P (GV604), yokFE101H (GV578), and yokFR144D (GV580), carrying p6.6 kb (pHB201/cat, erm), were mixed in a 1:1 ratio with kanamycin resistance recipient strains (amyE::PIPTG-gfp-kan) (GV416, GV601, GV575, GV603, GV577, and GV579), harboring equivalent genotypes, but lacking a plasmid. The mixtures were incubated for 4 h without selection, and equal volumes of cells were then spread over plates selective for trans-recipients. A detailed description of the used donor and recipient pairs and antibiotic selections employed are listed in Supplementary Table 1. NPex efficiency was determined as the ratio of trans-recipients (CFU)/total recipients (CFU). Shown is NPex efficiency (%) relative to ΔyokF pair. Data is presented as mean values and SEM, based on 4 independent experiments. Lower panels: BSB1-derived strains expressing the indicated YokF mutants (GV411, GV598, GV566, GV599, GV567, and GV568) were grown to mid-logarithmic phase and the total lysates were extracted. Shown is a Western blot analysis using anti-YokF antibodies, with a stain-free image as a loading control. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to Δ samples. P-values: D79H (***) = 1.17×10−4, R93D (***) = 1.69×10−5, D98H T99P (ns) = 7.75×10−1, E101H (***) = 1.20×10−4, R144D (*) = 3.35×10−2. (C) Schematic illustration of YokF truncations and their associated toxicity when expressed from the native chromosomal locus in BSB1. Toxicity (+) was manifested by the failure to obtain colonies following transformation, whereas lack of toxicity (-) showed no impact on transformation efficiency. (D) YokF exhibits a self-inhibiting TNM domain: BSB1-derived cells expressing YokF (WT), PIPTG-TNM (GV573: amyE::PIPTG-yokF216-296aa-spec), and ΔTNM, PIPTG-TNM (GV619: yokF1-215aa-tet, amyE::PIPTG-yokF216-296aa-spec), were streaked on LB agar plates with (+) or without (-) IPTG as indicated, and incubated overnight at 37 °C. Shown are representative images of at least 3 independent experiments. The residual growth for ΔTNM, PIPTG-TNM without IPTG was found to emanate from suppressor mutations in the yokF region as determined by sequence analysis. (E) Uninhibited TNase domain causes chromosome degradation: BSB1 cells expressing YokF (WT), PIPTG-TNM (GV573: amyE::PIPTG-yokF216-296aa-spec), and ΔTNM, PIPTG-TNM (GV619: yokF1-215aa-tet, amyE::PIPTG-yokF216-296aa-spec), were grown for 2 h with (+) or without (-) IPTG, as indicated. Shown are FM4-64 (red), DAPI (blue) fluorescence images, their overlay (overlay 1), and the enlargement of the inset (overlay 2). Arrows highlight cells lacking a signal from DAPI staining. This experiment was repeated 3 times with similar results. Scale bars 1 µm.
Extended Data Fig. 6 YokF colocalizes with FlhA.
(A) flhA-sg is motile: BSB1 (WT), and flhA-sg (GV618: flhA-lin4x-sg-Kan-Pfla/che-flhF) strains were grown to the mid-logarithmic phase, spotted onto 0.3% agar LB plates, and photographed after 12 h of incubation at 37 °C. (B) YokF and FlhA colocalization analysis: BSB1-derived strain expressing FlhA-SG (GV617: flhA-lin4x-sgd-kan- Pfla/che-flhF) was treated with anti-YokF primary antibodies followed by Alexa 647-conjugated secondary antibodies and subjected to fluorescence microscopy. Shown are fluorescence images of YokF (magenta), FlhA-SG (green), their overlay (Overlay 1), and their overlay with phase contrast (grey) (Overlay 2). Arrows highlight YokF and FlhA colocalization sites. This experiment was repeated 3 time with similar results. Scale bar 1 µm.
Extended Data Fig. 7 Comparing YokF with its putative paralogue YncB.
(A) YncB-YokF sequence comparison: Comparative sequence alignment of YokF and YncB from B. subtilis. Protein sequences were aligned using T-COFFEE and annotated with ESPript. Similar and identical amino acid residues are shown in red, highlighted within blue frames. Domain boundaries are delineated by coloured bars: SP (green), TNase (dark blue), and TNM (light blue). (B) YncB does not impact nanotube-mediated protein exchange: PY79-derived strains WT (SB463: amyE::PIPTG-cat-spec), and ΔyncB (GV473: ΔyncB::tet, amyE::PIPTG-gfp-kan), were mixed in a 1:1 ratio with kanamycin resistance recipient strains (amyE::PIPTG-gfp-kan) harboring equivalent genotypes (SB513, and GV474). The mixtures were incubated for 4 h without selection, and equal volumes of cells were spotted onto LB agar (control) and LB agar with chloramphenicol and kanamycin to evaluate protein exchange. Plates were photographed after 18 h of incubation. (C) YncB does not affect NPex: PY79-derived donor strains: WT (GV478), and ΔyncB (GV472: ΔyncB::tet), carrying p6.6 kb (pHB201/cat, erm), were mixed in a 1:1 ratio with kanamycin resistance recipient strains (amyE::PIPTG-gfp-kan), harboring equivalent genotypes (SB513, and GV474). The mixtures were incubated for 4 h without selection, and equal volumes of cells were then spread over plates containing chloramphenicol, erythromycin, and kanamycin to select for trans-recipients. Shown are representative images of NPex selective plates. (D) Quantitative analysis of NPex efficiencies between the donor and recipient pairs shown in (C). NPex efficiency=Trans-recipients CFU/Total recipients CFU. Shown is NPex efficiency (%) relative to the WT pair. Data are presented as mean values and SEM, based on 4 independent experiments. Statistical significance was calculated using Student’s t-test (two-sided) for each sample compared to the WT samples. P-values: ΔyncB (ns) = 4.37×10−1.
Extended Data Fig. 8 Characterization of YokF homologs.
(A) Multiple sequence alignment of YokF homologs: Comparative sequence alignment of YokF from various bacterial species obtained from GenBank (Supplementary Table 3). YokF homologs were aligned using T-COFFEE, and annotated with ESPript. Similar and identical amino acid residues are shown in red, highlighted within blue frames, with the percentage of identity and similarity between the YokF sequence from B. subtilis and its homologs indicated. Secondary structural elements derived from the B. subtilis YokF predicted structure are depicted above the sequences. (B) Expression of YokF homologs has no growth impact: PY79-derived cells expressing YokF homologs from B. subtilis (BSB1) (Bs) (GV480: amyE::PIPTG-yokFBs-spec), B. atrophaeus (Ba) (GV653: amyE::PIPTG-yokFBa-spec), B. vallismortis (Bv) (GV655: amyE::PIPTG-yokFBv-spec), B. halotolerans (Bh) (GV657: amyE::PIPTG-yokFBh-spec), and S. aureus (Sa) (GV659: amyE::PIPTG-yokFSa-spec) were grown with (+) or without (-) IPTG. Cell growth was followed by measuring OD600nm at 15-minute intervals. Data is presented as mean values and SEM, based on 3 independent repeats.
Extended Data Fig. 9 A model for YokF anti-NPex activity.
Schematic illustrating the mechanism by which YokF inhibits NPex. Following membrane localization, YokF undergoes a proteolytic cleavage, leading to TNM secretion and TNase activation. Activated YokF interacts with the CORE component FlhA, degrading plasmid DNA in the nanotube vicinity, thereby impeding NPex. An enlarged view shows YokF action site. Figure created in BioRender; Ben-Yehuda, S. https://BioRender.com/gum4ra9 (2026).
Supplementary information
Supplementary Data 1 (download XLSX )
Source file for MS results (Supplementary Table 2).
Source data
Source Data Fig. 3 (download PDF )
Unprocessed western blots for Fig. 3b,c,g,h.
Source Data Fig. 4 (download PDF )
Unprocessed western blots and gels for Fig. 4d,e.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed gel for Extended Data Fig. 1b.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blot and gel for Extended Data Fig. 5b.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gopu, V., Bhattacharya, S., Bejerano-Sagie, M. et al. A family of endonucleases blocks nanotube-mediated plasmid exchange. Nat Microbiol 11, 960–975 (2026). https://doi.org/10.1038/s41564-026-02293-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-026-02293-8