Abstract
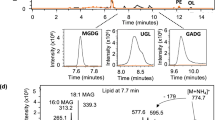
Per- and polyfluoroalkyl substances (PFASs), also known as forever chemicals, are global contaminants, but understanding of microbiota–PFAS interactions is limited. Here we show that bacteria covalently incorporate n:3 fluorotelomer carboxylates (FTCAs) into phosphatidylethanolamine and phosphatidylglycerol, two prominent components of bacterial lipid bilayers. Lipidomics of the soil bacterium Pseudomonas sp. strain 273 grown in the presence of 7:3 FTCA or 8:3 FTCA estimated that 7–12% of the bacterium’s glycerophospholipid pool contains the respective polyfluoroacyl chains. This covalent incorporation was observed in five other axenic bacterial cultures tested, including other Pseudomonas species, Escherichia coli and Enterococcus faecalis, albeit with lower incorporation percentages. Incorporation occurred over a broad concentration range, and n:3 FTCAs with varying chain length were covalently incorporated into membranes. Biotransformation of polyfluoroalkyl substances (also known as precursors) results in n:3 FTCA intermediates, which bacteria can covalently incorporate into their glycerophospholipid pools. We conclude that bacteria can form fluoromembranes when exposed to precursors and are a potential PFAS sink.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All MS proteomics raw data are available at the ProteomeXchange Consortium via the MassIVE database with the unique dataset identifier MSV000098309 (https://doi.org/10.25345/C5SB3XB59). The ProteomeXchange project identifier is PXD065413. The genome of strain 273 is available in GenBank (CP116775) and the JGI IMG system (ID no. 2814122901). The mass spectrometry lipidomic raw data and sequence files are available in the MassIVE database with the unique dataset identifier MSV000100274 (https://doi.org/10.25345/C53T9DM05). The raw MS data for LC–QToF–MS suspect screening analysis are available in the Texas Data Repository (https://doi.org/10.18738/T8/BDF7TU). All other data are available in this Article and its Supplementary Information. Source data are provided with this paper.
References
Evich, M. G. et al. Per- and polyfluoroalkyl substances in the environment. Science 375, eabg9065 (2022).
Glüge, J. et al. An overview of the uses of per-and polyfluoroalkyl substances (PFAS). Environ. Sci. Process. Impacts 22, 2345–2373 (2020).
Schymanski, E. L. et al. Per-and polyfluoroalkyl substances (PFAS) in PubChem: 7 million and growing. Environ. Sci. Technol. 57, 16918–16928 (2023).
Hartz, W. F. et al. Levels and distribution profiles of per-and polyfluoroalkyl substances (PFAS) in a high Arctic Svalbard ice core. Sci. Total Environ. 871, 161830 (2023).
Nielsen, F., Fischer, F. C., Leth, P. M. & Grandjean, P. Occurrence of major perfluorinated alkylate substances in human blood and target organs. Environ. Sci. Technol. 58, 143–149 (2023).
Sunderland, E. M. et al. A review of the pathways of human exposure to poly- and perfluoroalkyl substances (PFASs) and present understanding of health effects. J. Expo. Sci. Environ. Epidemiol. 29, 131–147 (2019).
Dasu, K., Xia, X., Siriwardena, D., Klupinski, T. P. & Seay, B. Concentration profiles of per- and polyfluoroalkyl substances in major sources to the environment. J. Environ. Manage. 301, 113879 (2022).
LaFond, J. A., Hatzinger, P. B., Guelfo, J. L., Millerick, K. & Jackson, W. A. Bacterial transformation of per- and poly-fluoroalkyl substances: a review for the field of bioremediation. Environ. Sci. Adv. 2, 1019–1041 (2023).
Fitzgerald, N. J. et al. Partitioning and accumulation of perfluoroalkyl substances in model lipid bilayers and bacteria. Environ. Sci. Technol. 52, 10433–10440 (2018).
Sobolewski, T. N., Trousdale, R. C., Gauvin, C. L., Lawrence, C. M. & Walker, R. A. Nanomolar PFOA concentrations affect lipid membrane structure: consequences for bioconcentration mechanisms. Environ. Sci. Technol. 59, 709–718 (2025).
Lindell, A. E. et al. Human gut bacteria bioaccumulate per- and polyfluoroalkyl substances. Nat. Microbiol. 10, 1630–1647 (2025).
Xie, Y. et al. Pseudomonas sp. strain 273 degrades fluorinated alkanes. Environ. Sci. Technol. 54, 14994–15003 (2020).
Xie, Y. et al. Genome-wide expression analysis unravels fluoroalkane metabolism in Pseudomonas sp. strain 273. Environ. Sci. Technol. 57, 15925–15935 (2023).
Xie, Y. et al. Pseudomonas sp. strain 273 incorporates organofluorine into the lipid bilayer during growth with fluorinated alkanes. Environ. Sci. Technol. 56, 8155–8166 (2022).
Neidhardt, F. C. Escherichia coli and Salmonella: Cellular and Molecular Biology (ASM Press, 1996).
Yao, J. & Rock, C. O. How bacterial pathogens eat host lipids: implications for the development of fatty acid synthesis therapeutics. J. Biol. Chem. 290, 5940–5946 (2015).
Soupene, E. & Kuypers, F. A. Mammalian long-chain acyl-CoA synthetases. Exp. Biol. Med. 233, 507–521 (2008).
Beld, J., Finzel, K. & Burkart, M. D. Versatility of acyl-acyl carrier protein synthetases. Chem. Biol. 21, 1293–1299 (2014).
Parsons, J. B., Frank, M. W., Jackson, P., Subramanian, C. & Rock, C. O. Incorporation of extracellular fatty acids by a fatty acid kinase-dependent pathway in Staphylococcus aureus. Mol. Microbiol. 92, 234–245 (2014).
Xie, Y., Chen, G., Ramirez, D., Yan, J. & Löffler, F. E. Complete genome sequence of Pseudomonas sp. strain 273, a haloalkane-degrading bacterium. Microbiol. Resour. Announc. 12, e00176–00123 (2023).
Mothersole, R. G., Mothersole, M. K., Goddard, H. G., Liu, J. & Van Hamme, J. D. Enzyme catalyzed formation of CoA adducts of fluorinated hexanoic acid analogues using a long-chain acyl-CoA synthetase from Gordonia sp. strain NB4-1Y. Biochemistry 63, 2153–2165 (2024).
Liu, J., Lee, L. S., Nies, L. F., Nakatsu, C. H. & Turco, R. F. Biotransformation of 8:2 fluorotelomer alcohol in soil and by soil bacteria isolates. Environ. Sci. Technol. 41, 8024–8030 (2007).
Rankin, K., Lee, H., Tseng, P. J. & Mabury, S. A. Investigating the biodegradability of a fluorotelomer-based acrylate polymer in a soil–plant microcosm by indirect and direct analysis. Environ. Sci. Technol. 48, 12783–12790 (2014).
Lang, J. R., Allred, B. M., Field, J. A., Levis, J. W. & Barlaz, M. A. National estimate of per-and polyfluoroalkyl substance (PFAS) release to US municipal landfill leachate. Environ. Sci. Technol. 51, 2197–2205 (2017).
Wu, C., Wang, Q., Chen, H. & Li, M. Rapid quantitative analysis and suspect screening of per-and polyfluorinated alkyl substances (PFASs) in aqueous film-forming foams (AFFFs) and municipal wastewater samples by Nano-ESI-HRMS. Water Res. 219, 118542 (2022).
Munoz, G. et al. Zwitterionic, cationic, and anionic PFAS in freshwater sediments from AFFF-impacted and non-impacted sites of Eastern Canada. J. Hazard. Mater. 484, 136634 (2025).
Ackerman Grunfeld, D. et al. Underestimated burden of per- and polyfluoroalkyl substances in global surface waters and groundwaters. Nat. Geosci. 17, 340–346 (2024).
Oviedo-Vargas, D., Anton, J., Coleman-Kammula, S. & Qin, X. Quantification of PFAS in soils treated with biosolids in ten northeastern US farms. Sci. Rep. 15, 5582 (2025).
Schüßler, M., Capitain, C., Bugsel, B., Zweigle, J. & Zwiener, C. Non-target screening reveals 124 PFAS at an AFFF-impacted field site in Germany specified by novel systematic terminology. Anal. Bioanal. Chem. 417, 6049–6064 (2024).
Mejia-Avendaño, S. et al. Novel fluoroalkylated surfactants in soils following firefighting foam deployment during the Lac-Mégantic railway accident. Environ. Sci. Technol. 51, 8313–8323 (2017).
Dunn, M. et al. Unregulated active and closed textile mills represent a significant vector of PFAS contamination into coastal rivers. ACS ES&T Water 4, 114–124 (2024).
Yang, S.-H., Shi, Y., Strynar, M. & Chu, K.-H. Desulfonation and defluorination of 6:2 fluorotelomer sulfonic acid (6:2 FTSA) by Rhodococcus jostii RHA1: carbon and sulfur sources, enzymes, and pathways. J. Hazard. Mater. 423, 127052 (2022).
Chi, F. et al. Unveiling behaviors of 8:2 fluorotelomer sulfonic acid (8:2 FTSA) in Arabidopsis thaliana: bioaccumulation, biotransformation and molecular mechanisms of phytotoxicity. Sci. Total Environ. 927, 172165 (2024).
Shaw, D. M. et al. Degradation and defluorination of 6:2 fluorotelomer sulfonamidoalkyl betaine and 6:2 fluorotelomer sulfonate by Gordonia sp. strain NB4-1Y under sulfur-limiting conditions. Sci. Total Environ. 647, 690–698 (2019).
Yan, P.-F. et al. Aerobic biotransformation of 6:2 fluorotelomer sulfonate in soils from two aqueous film-forming foam (AFFF)-impacted sites. Water Res. 249, 120941 (2024).
Griebler, C. & Lueders, T. Microbial biodiversity in groundwater ecosystems. Freshw. Biol. 54, 649–677 (2009).
Nouhi, S. et al. Interactions of perfluoroalkyl substances with a phospholipid bilayer studied by neutron reflectometry. J. Colloid Interface Sci. 511, 474–481 (2018).
Zhao, L. et al. Insight into the binding model of per- and polyfluoroalkyl substances to proteins and membranes. Environ. Int. 175, 107951 (2023).
Naumann, A., Alesio, J., Poonia, M. & Bothun, G. D. PFAS fluidize synthetic and bacterial lipid monolayers based on hydrophobicity and lipid charge. J. Environ. Chem. Eng. 10, 107351 (2022).
Forsthuber, M. et al. Albumin is the major carrier protein for PFOS, PFOA, PFHxS, PFNA and PFDA in human plasma. Environ. Int. 137, 105324 (2020).
Ng, C. A. & Hungerbühler, K. Bioaccumulation of perfluorinated alkyl acids: observations and models. Environ. Sci. Technol. 48, 4637–4648 (2014).
Sundström, M. et al. A temporal trend study (1972–2008) of perfluorooctanesulfonate, perfluorohexanesulfonate, and perfluorooctanoate in pooled human milk samples from Stockholm, Sweden. Environ. Int. 37, 178–183 (2011).
Wang, Q. et al. Target, nontarget, and suspect screening and temporal trends of per-and polyfluoroalkyl substances in marine mammals from the South China Sea. Environ. Sci. Technol. 55, 1045–1056 (2021).
De Silva, A. O. et al. PFAS exposure pathways for humans and wildlife: a synthesis of current knowledge and key gaps in understanding. Environ. Toxicol. Chem. 40, 631–657 (2021).
Yeung, L. W. et al. Perfluorinated compounds and total and extractable organic fluorine in human blood samples from China. Environ. Sci. Technol. 42, 8140–8145 (2008).
Spaan, K. M. et al. Fluorine mass balance and suspect screening in marine mammals from the northern hemisphere. Environ. Sci. Technol. 54, 4046–4058 (2020).
Yeung, L. W. et al. Perfluoroalkyl substances and extractable organic fluorine in surface sediments and cores from Lake Ontario. Environ. Int. 59, 389–397 (2013).
Yamashita, A., Sugiura, T. & Waku, K. Acyltransferases and transacylases involved in fatty acid remodeling of phospholipids and metabolism of bioactive lipids in mammalian cells. J. Biochem. 122, 1–16 (1997).
Shobnam, N., Sun, Y., Mahmood, M., Löffler, F. E. & Im, J. Biologically mediated abiotic degradation (BMAD) of bisphenol A by manganese-oxidizing bacteria. J. Hazard. Mater. 417, 125987 (2021).
Danne-Rasche, N., Coman, C. & Ahrends, R. Nano-LC/NSI MS refines lipidomics by enhancing lipid coverage, measurement sensitivity, and linear dynamic range. Anal. Chem. 90, 8093–8101 (2018).
Clasquin, M. F., Melamud, E. & Rabinowitz, J. D. LC–MS data processing with MAVEN: a metabolomic analysis and visualization engine. Curr. Protoc. Bioinformatics 37, 1–23 (2012).
Wischnak, C., Löffler, F. E., Li, J., Urbance, J. W. & Müller, R. Pseudomonas sp. strain 273, an aerobic α, ω-dichloroalkane-degrading bacterium. Appl. Environ. Microbiol. 64, 3507–3511 (1998).
LaFond, J. A. et al. Biotransformation of PFAA precursors by oxygenase-expressing bacteria in AFFF-impacted groundwater and in pure-compound studies with 6:2 FTS and EtFOSE. Environ. Sci. Technol. 58, 13820–13832 (2024).
Charbonnet, J. A. et al. Communicating confidence of per-and polyfluoroalkyl substance identification via high-resolution mass spectrometry. Environ. Sci. Technol. Lett. 9, 473–481 (2022).
Werner, A. Z. et al. A distinct subpopulation of membrane vesicles in Pseudomonas putida is enriched in enzymes for lignin catabolism. Appl. Environ. Microbiol. 91, e01617–e01625 (2025).
Kassambara, A. rstatix: Pipe-Friendly Framework for Basic Statistical Tests. R Package Version 0.7.2. (CRAN, 2023).
Bustin, S. A. et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622 (2009).
Lee, H., D’eon, J. & Mabury, S. A. Biodegradation of polyfluoroalkyl phosphates as a source of perfluorinated acids to the environment. Environ. Sci. Technol. 44, 3305–3310 (2010).
D’Agostino, L. A. & Mabury, S. A. Aerobic biodegradation of 2 fluorotelomer sulfonamide-based aqueous film-forming foam components produces perfluoroalkyl carboxylates. Environ. Toxicol. Chem. 36, 2012–2021 (2017).
Harding-Marjanovic, K. C. et al. Aerobic biotransformation of fluorotelomer thioether amido sulfonate (Lodyne) in AFFF-amended microcosms. Environ. Sci. Technol. 49, 7666–7674 (2015).
Royer, L. A., Lee, L. S., Russell, M. H., Nies, L. F. & Turco, R. F. Microbial transformation of 8:2 fluorotelomer acrylate and methacrylate in aerobic soils. Chemosphere 129, 54–61 (2015).
Dasu, K., Lee, L. S., Turco, R. F. & Nies, L. F. Aerobic biodegradation of 8:2 fluorotelomer stearate monoester and 8:2 fluorotelomer citrate triester in forest soil. Chemosphere 91, 399–405 (2013).
Dasu, K., Liu, J. & Lee, L. S. Aerobic soil biodegradation of 8:2 fluorotelomer stearate monoester. Environ. Sci. Technol. 46, 3831–3836 (2012).
Ruan, T. et al. Aerobic soil biotransformation of 6:2 fluorotelomer iodide. Environ. Sci. Technol. 47, 11504–11511 (2013).
Martin, J. W., Mabury, S. A. & O’Brien, P. J. Metabolic products and pathways of fluorotelomer alcohols in isolated rat hepatocytes. Chem. Biol. Interact. 155, 165–180 (2005).
Wang, N. et al. 6:2 Fluorotelomer sulfonate aerobic biotransformation in activated sludge of waste water treatment plants. Chemosphere 82, 853–858 (2011).
Yaqoob, S., Wu, C., Li, C., Antunes, J. M. D. & Li, M. Anaerobic biotransformation and biodefluorination of 6:2 fluorotelomer carboxylic acid by biosolids under the nitrate-reducing condition. ACS ES&T Water 5, 2412–2420 (2025).
Wu, C., Goodrow, S., Chen, H. & Li, M. Distinctive biotransformation and biodefluorination of 6:2 versus 5:3 fluorotelomer carboxylic acids by municipal activated sludge. Water Res. 254, 121431 (2024).
Acknowledgements
This work was supported by the US Department of Defense, Strategic Environmental Research and Development Program (project no. ER23-3601 to G.C., R.L.H. and F.E.L.) and, in part, by ExxonMobil Environmental and Property Solutions Company (to F.E.L.) and the US National Science Foundation Graduate Research Fellowship Program (grant no. DGE 2140745 to J.A.L.).
Author information
Authors and Affiliations
Contributions
Y.X. and F.E.L. conceived the study. Y.X., G.C., D.R., C.M.S. and B.M.W. performed the cultivation experiments, enumerated bacteria with qPCR, extracted lipids for lipidomic analysis and prepared samples for proteomic measurements and untargeted LC–MS analysis. M.J.K. and A.J.S. performed mass spectrometry analysis of lipidomic samples. D.L.C. and P.E.A. conducted proteomic analysis. J.A.L. and J.L.G. measured 8:2 FTSA transformation products. Y.X., F.E.L., A.L.M. and S.R.C. performed the initial lipidomic measurements. All authors contributed methodology and supported data analysis and interpretation. Y.X., G.C., R.L.H. and F.E.L. synthesized information. Y.X. and F.E.L. wrote the manuscript. All authors reviewed manuscript drafts and provided input.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Schematic illustrating the covalent incorporation of aqueous n:3 FTCAs into the bacterial lipid bilayer and the formation of fluoromembranes.
The fluoromembrane structure displays a fluorinated glycerophospholipid with an n:3 FTCA (n = 5, 7, 8, 9) acyl tail covalently attached to the glycerol backbone at the sn1 position. Free n:3 FTCA and n:3 FTCA that partitions into membranes due to hydrophobic interactions are shown in blue color, while covalently incorporated n:3 FTCA is depicted in red. Note that the covalent incorporation and the non-covalent partitioning of n:3 FTCA into lipid membrane are mechanistically distinct and likely occur simultaneously.
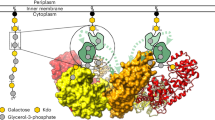
Extended Data Fig. 2 Proposed pathway for covalent incorporation of n:3 FTCAs into membrane glycerophospholipids during de novo lipid biosynthesis.
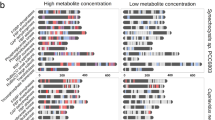
n:3 FTCAs are activated in a fatty acyl-CoA synthetase (FadD) reaction, and the resulting CoA thioesters are then incorporated into the sn-1 and/or the sn-2 positions of the glycerophospholipid backbone, that is, glycerol-3-phosphate (G3P). These esterification reactions are catalyzed by G3P acyltransferase (PlsB) and/or 1-acyl-sn-G3P acyltransferase(s) (PlsC), forming phosphatidic acids with one or two n:3 FTCA acyl tails. Subsequent headgroup assembly generates complete PFASylated glycerophospholipids, which become part of the lipidome during growth of strain 273 in the presence of a n:3 FTCAs. The heatmaps show enzyme expression changes in Pseudomonas sp. strain 273 cells grown with versus without 8:3 FTCA. Notably, PlsC-1 (2814126759) and PlsC-2 (2814131665) were only detected during growth in the presence of 8:3 FTCA, suggesting higher abundance, particularly of PlsC-2, in the presence of the n:3 FTCA. The asterisks denote significant up-regulation, defined as a Log2 (fold change) > 1 and a p value < 0.05, in the presence of 8:3 FTCA.
Extended Data Fig. 3 Illustration of precursor subgroups as sources of n:3 FTCAs.
Biotransformation of (from left to right) PAPs58, FTSAms59, FTTAoSs60, FTACs61, FTABs34, FTSs62,63, TBCs62, miscellaneous precursors23, FTIs64, FTSAs32,33,53, FTOHs22, FTALs65,66, and n:2 FTCAs67,68 to n:3 FTCAs has been demonstrated. n:3 FTCAs are subject to incorporation into glycerophospholipids (solid red arrow) or biotransformation to PFCAs68 (solid black arrow). Cell lysis and lipid hydrolysis would release (fluoro)fatty acids, including n:3 FTCAs, the latter available for recycling (that is, renewed covalent incorporation into glycerophospholipids, dashed red arrow) or biotransformation to PFCAs (dashed black arrow).
Extended Data Fig. 4 Molecular structure of PE(7:3FTCA_16:0) detected in the Pseudomonas sp. strain 273 lipidome following growth with 8:2 FTSA as a sulfur source.
The molecular structure (A), tandem mass spectra (B) and associated ion fragments (C) of the target molecule PE(7:3 FTCA_16:0) are shown.The fragmentation pattern of PE(7:3FTCA_16:0) corroborated structure elucidation as seen by [sn1]− at 441 m/z, [sn2]− at 255 m/z, [M-sn1]− at 452 m/z, and a [M-sn2]− at 620 m/z. The fragments with m/z at 357, 337, 317 and 267, correspond to rearranged fragments from the FTCA tail. The mass spectra are normalized to the intensity of the ion fragment with the highest intensity in the selected m/z range.
Supplementary information
Supplementary Information (download PDF )
List of Supplementary Tables 1–4, Notes 1 and 2 and Fig. 1.
Supplementary Tables (download XLSX )
Supplementary Table 1. Phosphatidylethanolamine (PE) abundances in different bacteria grown in the presence and in the absence of 8:3 FTCA. Entries 1–12 correspond to different bacterial species grown with and without 8:3 FTCA, whereas entry 13 and entry 14 represent controls where the same concentrations of 8:3 FTCA were added immediately before lipid extraction. Supplementary Table 2. Phosphatidylglycerol (PG) abundances in different bacteria grown in the presence and in the absence of 8:3 FTCA. Entries 1–12 correspond to different bacterial species grown with and without 8:3 FTCA, whereas entry 13 and entry 14 represent controls where the same concentrations of 8:3 FTCA were added immediately before lipid extraction. Supplementary Table 3. Details of the calibrant–internal standard pairs used for the quantification or semi-quantification of 8:2 FTSA transformation products. Supplementary Table 4. Internal standard recoveries for 8:2 FTSA transformation products reported as a relative percentage to the average peak area of the internal standards in the calibration curve.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xie, Y., Chen, G., Keller, M.J. et al. Bacteria covalently incorporate polyfluoroalkyl carboxylates into membrane lipids. Nat Microbiol (2026). https://doi.org/10.1038/s41564-026-02301-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41564-026-02301-x