Abstract
The malaria elimination programme in Kayin State (Myanmar) uses malaria posts for rapid detection and treatment, together with mass drug administration in high-transmission villages, which has reduced transmission by 97%. Here we examine the impact of control on parasite genomic parameters to inform future control efforts. Using 2,270 genome-sequenced Plasmodium falciparum infections from 283 malaria posts, sampled over 58 months (2015–2020), we find that parasite effective population size decreased over the study period, but there was minimal change in artemisinin resistance allele frequency until 2020, when just one predominant genotype (carrying kelch13-R561H) remained. We observed sustained localized transmission of unique parasite genotypes revealing transmission chains and positive correlations in parasite relatedness for ≤20 km. Mass drug administration resulted in parasite founder effects, providing genomic evidence for the efficacy of this control tool. These results reveal changes in population structure driven by control and rapid shifts in allele frequency in a parasite population close to elimination.
Similar content being viewed by others
Main
Regions of low malaria transmission intensity predominate in Southeast (SE) Asia and South America and are becoming increasingly common in Africa1. A central challenge for malaria control is to develop efficient approaches to eliminate malaria from such regions. Rapid selection of drug-resistant parasites is a central concern for intensive malaria control programmes. For example, kelch13 mutations conferring artemisinin (ART) resistance increased from a 0% to 90% frequency in parasites collected from patients visiting Shoklo Malaria Research Unit (SMRU) clinics on the Thailand–Myanmar border between 2003 and 2014 (ref. 2). The use of mass drug administration (MDA) is controversial for malaria treatment owing to concerns about resistance. Previous use of chloroquine-treated salt is thought to have accelerated the selection of chloroquine resistance in the last century3. However, MDA is effective for treating submicroscopic malaria infections that constitute the majority of infections in many SE Asian locations4 and are missed by passive malaria surveillance. Submicroscopic infections may be cured effectively by MDA because there are few parasites per patient and so treatment is more likely to be completely successful than in high-parasitaemia infections. It has therefore been argued that MDA is not likely to promote resistance spread in low-prevalence regions such as SE Asia5. A previous paper examined the epidemiology of kelch13 haplotypes in Kayin State and showed limited changes of kelch13 ART resistance alleles between 2013 and 2019 (ref. 6). The spread of resistance was much less rapid than that which occurred in the SMRU clinics (2000–2014); this is consistent with near-exhaustive coverage of communities with malaria posts (MPs, which refer to community-based malaria clinics) and MDA imposing limited selection for ART resistance.
Here, we examine the genomic epidemiology of Plasmodium falciparum samples collected between November 2015 and August 2020 during malaria elimination efforts in Kayin State. Between 2014 and 2020, the Malaria Elimination Task Force (METF) targeted four townships (Myawaddy, Kawkareik, Hlaingbwe and Hpapun) in Kayin State, eastern Myanmar, for malaria elimination7,8. This was done using a combination of interventions: (1) 1,475 village MPs were opened, providing rapid diagnosis and malaria treatment (artemether–lumefantrine plus single low-dose primaquine); (2) MDA (dihydroartemisinin (DHA)–piperaquine (PPQ) plus single-dose primaquine once per month for three consecutive months) was used for 69 ‘hotspot’ villages where malaria remained prevalent (>40% malaria and >20% P. falciparum)8. The combination of these two approaches reduced P. falciparum cases by 97% from an incidence of 39 cases per 1,000 person-years (May 2014 to April 2015)8 to 1 case per 1,000 person-years (May 2019 to April 2020)9.
Our central goal was to use genome sequence data to understand parasite transmission, population genomics and resistance evolution and to use these data to inform future control efforts in elimination settings of SE Asia. Our key questions were to: (1) determine the distribution and stability of malaria populations within the Kayin State target region; (2) evaluate evidence for long-distance gene flow between Kayin State, Myanmar, and other eastern SE Asian countries (Cambodia, Vietnam and Laos); (3) document the origins of kelch13 resistance alleles; (4) determine the impact of MDA on parasite population structure; and (5) evaluate appropriate genomic metrics for assessing transmission intensity.
Results
Extreme clonal expansion and inbreeding in a region under massive drug selection
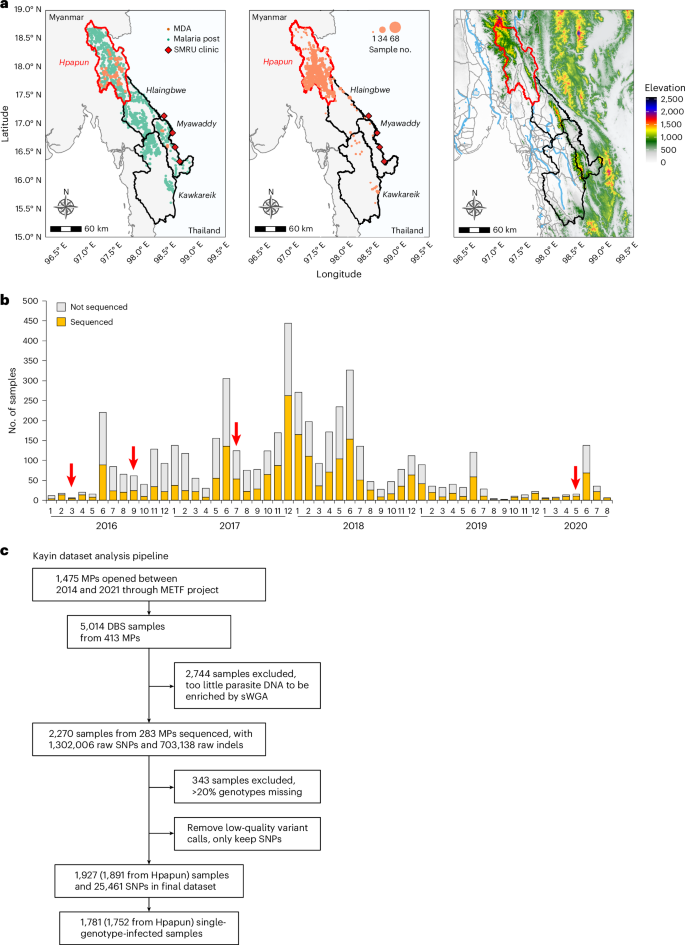
A total of 5,014 dried blood spot (DBS) samples with geographic references from 413 MPs were collected between November 2015 and August 2020 as part of the METF malaria elimination programme in eastern Myanmar, led by the SMRU (Fig. 1 and Extended Data Fig. 1). We processed and analysed genome-wide sequencing data of 2,270 DBS samples collected from 283 MPs (Supplementary Table 1). After filtration of low-coverage samples and low-quality genotypes, the final Kayin State dataset contains 1,927 P. falciparum samples with a set of 25,461 high-quality single-nucleotide polymorphisms (SNPs). Collectively, 92.4% (1,781/1,927) of the samples were from single-genotype infections (Fws ≥ 0.95; Extended Data Fig. 2 and Supplementary Table 2).
a, Physical geography of the METF intervention region. The METF project was performed at four townships (Myawaddy, Kawkareik, Hlaingbwe and Hpapun) of Kayin State, Myanmar. Left: the distribution of MPs (green dots). Vermillion dots indicate locations where MDAs were applied. Middle: location of sequenced samples. Over 96% of the sequenced sample were from Hpapun township, the northern part of Kayin State. Right: elevation map. Elevation data were downloaded from the US Geological Survey (https://earthexplorer.usgs.gov/). The locations of roads (grey) and rivers (light blue) were obtained from Myanmar Information Management Unit (https://themimu.info/). b, Temporal distribution of samples collected through the METF project. Red arrows indicate the time when MDAs were applied. c, Analysis pipeline for samples collected from Kayin State.
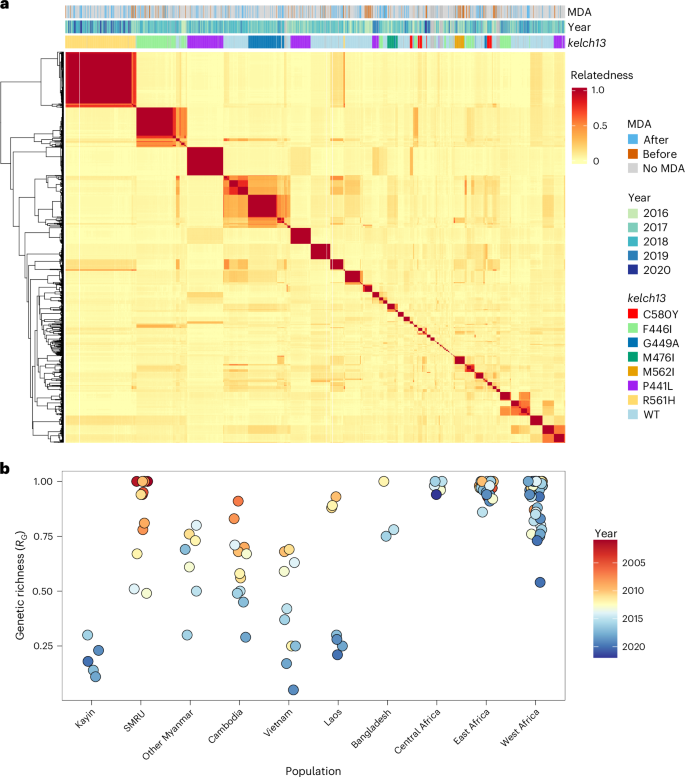
Pairwise genetic relatedness (r, proportion of genome that was identity by descent (IBD)) was used to identify highly related parasites, and samples sharing ≥90% genome IBD (r ≥ 0.90) were grouped into clonal clusters (Fig. 2a and Extended Data Fig. 3). From the 1,781 single-genotype Kayin samples, we identified 93 IBD clusters (2–229 samples per cluster) and 73 singletons, giving 166 unique genomes. We then calculated genetic richness (RG, number of unique genomes divided by the total number of parasites) to quantify the proportion of genetically distinct parasites within each population. The Kayin population exhibited very low RG values (0.10–0.30 across 2015–2020; 166 unique genomes among 1,781 single-genotype infections), reflecting extensive clonal expansion, including 14 large clonal clusters (>30 samples each; Fig. 2a). RG values have declined over time in SE Asian parasite populations (Fig. 2b and Supplementary Tables 3 and 4). The Kayin population showed comparable RG to population samples collected during the same time period (2015–2020) from Vietnam and Laos but were lower than those observed in nearby SMRU clinics (2001–2014) or in other Myanmar regions (excluding Kayin, 2007–2018). For comparison, African P. falciparum populations showed consistently high RG values (0.54–1.00) (Fig. 2b).
a, Heat map showing relatedness among Kayin samples. Pairwise parasite relatedness (r) was measured as the proportion of the genome with shared IBD between pairs of samples. Samples with r ≥ 0.9 are considered as IBD and share the same unique genome. Colour bars at the top of the heat map indicate information for each sample: MDA, if the sample came from an MP with (orange) or without (blue) MDA; year, the year of sampling; and kelch13, the genotype of kelch13—only alleles with frequency >2% in at least one sampling year were coloured. b, Level of clonal transmission. Genetic richness (RG) was defined as the number of unique parasite genotypes divided by the total number of parasite genotypes.
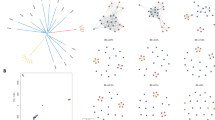
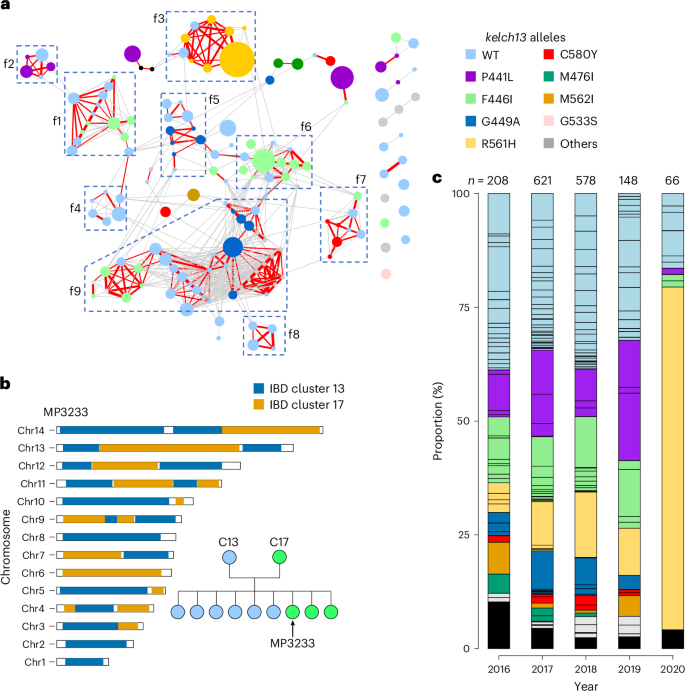
The ten largest IBD clusters account for 51.6% of the population, indicating low parasite diversity in Kayin State. In total, 151 out of 166 (91.0%) unique genomes were genetically related (r ≥ 0.25) to at least one other genome, indicating high levels of inbreeding (Fig. 3a). Nine closely related families account for 110/166 unique genomes and 66.9% (1,192/1,781) of single-genotype infections, with members sharing r ≥ 0.45. Analysis of chromosomal IBD segments revealed genealogical relationships within families (Fig. 3b and Extended Data Fig. 4). For two families (family 1 and 7), both parents and F1 progeny were identified, indicating an extremely small parasite population size in Kayin State.
a, IBD network of unique genomes from Kayin population. Nodes: each circle indicates one unique genome and is colour-coded on the basis of its kelch13 alleles; circle size indicates sample size (ranged from 1 to 229). Edges: connections with relatedness (r) ≥0.25; thicker lines indicate higher relatedness; red lines are connections with r ≥ 0.45. Parasites from closely related families (f1 to f9) are labelled using boxes. In total, 152 of 166 unique genomes were included in the network, representing 98.6% of single-genotype-infected samples. b, Pedigree tree of parasites from f1 and chromosome plot for an estimated progeny (MP3233). See Extended Data Fig. 7 for chromosome plots for all progeny. We infer that the parents of f1 are C13 (IBD cluster 13, kelch13-WT) and C17 (IBD cluster 17, kelch13-F446I). c, Proportion of unique genomes across time. Each segment within a bar represents one unique genome, which is coloured on the basis of its kelch13 allele. Black blocks indicate number of unique genomes that were recovered only once (‘singletons’). A clonal expansion of IBD cluster 1 (kelch13-R561H) parasites was detected in 2020.
Localized transmission and regional stability of haplotypes
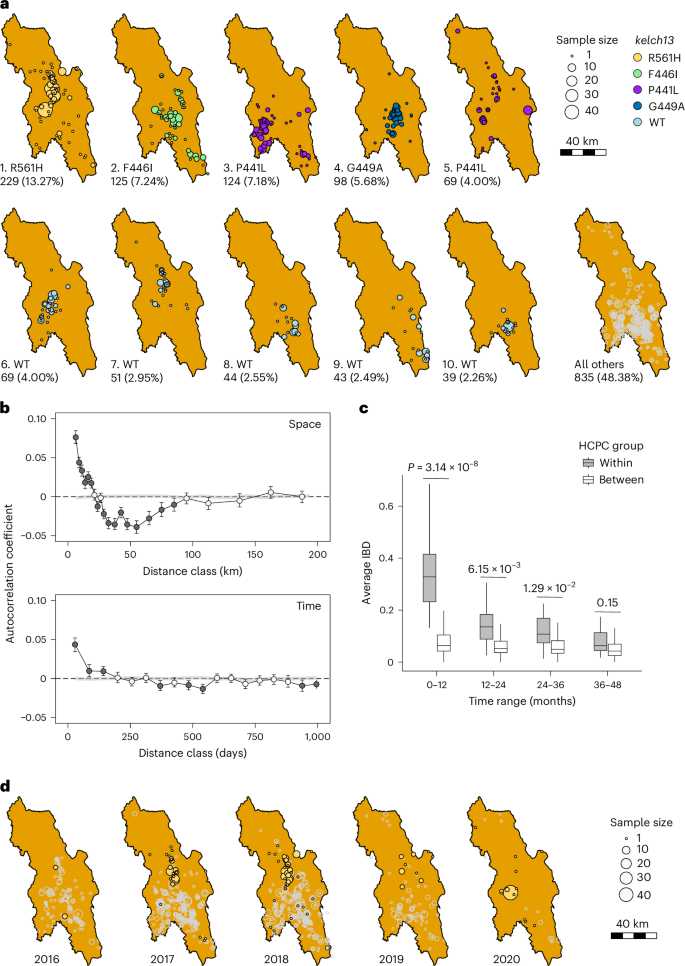
Patterns of genetic relatedness across space reveal localized transmission dynamics of malaria parasites. Spatial groups of IBD clusters reflect direct or indirect transmission chains and show clear geographic structure (Fig. 4a and Supplementary Fig. 1). For example, 86% of samples from the largest IBD cluster (carrying kelch13 R561H) were collected to the north of Hpapun Township (where >96% sequenced samples were collected; Fig. 1a). The second-largest IBD cluster (carrying kelch13 F446I) was found mainly in the centre of Hpapun Township, while the third-largest IBD cluster (carrying kelch13 P441L) was found in the west of the same township. Similarly, different IBD clusters carrying kelch13 wild type (WT) alleles show localized distribution. Spatial correlograms confirm that parasite relatedness is positively correlated at distances ≤20 km (Fig. 4b), consistent with local transmission of IBD clusters. We also observed significant negative correlations in relatedness between 27.5 km and 90 km, reflecting boundaries between distinct genetic clusters or transmission foci. The negative correlations further indicate the spatial pattern of parasite relatedness. Not only are parasites that are geographically close more likely to be related—those that are geographically distant (27.5–90 km) are less likely to be related.
a, Spatial distribution of different IBD clusters. Each circle represents samples collected from an individual MP. IBD cluster IDs and corresponding kelch13 alleles are indicated at the bottom left of each panel; for example, 1_R561H indicates IBD cluster number 1 carrying the kelch13 R561H allele. b, Correlogram analysis of pairwise parasite relatedness across space and time. A total of 1,718 samples with available location and collection time were included in the Mantel correlogram analysis. Bars indicate 95% confidence intervals from 500 bootstrap replicates. c, Comparison of relatedness between within-group and between-group HCPC region pairs. The x axis indicates time intervals in months. Relatedness was measured as average IBD among samples within each HCPC group or between pairs of the 29 groups (more than 10 samples per group). Boxes show interquartile ranges, centre lines indicate medians, whiskers extend to 1.5× the interquartile range, and outliers are not shown. P values were calculated using a two-sided Welch’s two-sample t-test; no adjustment for multiple comparisons was applied. d, Temporal dynamics and spatial distribution of the IBD cluster 1 (kelch13 R561H) through the sampling year.
Temporal patterns of genetic relatedness provide insight into the frequency of outbreeding within malaria parasite populations. We detected clonal IBD clusters (r ≥ 0.90, contain two or more samples) that were sampled across the 56-month study period, as well as new genome haplotypes generated through recombination (Extended Data Fig. 5). Of 93 IBD clusters, 9 lasted ≥36 months, while 34 lasted ≤6 months; the mean cluster duration in the Kayin population was 13.8 (s.e.m, 1.4) months. The mean duration of closely related parasite family members (r ≥ 0.45) was 48.7 (s.e.m, 3.0) months, consistent with limited outbreeding. In two families where both parents and progeny were identified, there were only 12 recombination events among the 109 samples (14 unique genomes, 41 months of family duration) in family (f)1 and 6 among 31 samples (8 unique genomes, 48 months of family duration) in f7 (Extended Data Fig. 4), further supporting with low parasite outbreeding frequency.
We used a temporal correlogram to examine the number of days over which correlations in relatedness were observed (Fig. 4b). This revealed positive correlations in relatedness between parasites sampled ≤170 days apart. To evaluate the relative impact of space and time on parasite relatedness, we divided the control region into 50 regions using hierarchical clustering on principal components (HCPC; Extended Data Fig. 6); 29 of these HCPC regions contained 10–161 parasites. We examined relatedness in parasites sampled within and between HCPC regions for parasites sampled 1–12, 13–24, 25–36 and 37–48 months apart (Fig. 4c). We observed significantly greater relatedness among parasites sampled from the same HCPC unit, relative to those sampled from different HCPC units. This remained significant for parasites sampled up to ≤36 months apart, demonstrating spatial stability of parasite populations within Hpapun Township.
Long-distance connectivity of parasite populations in SE Asia
We used parasite relatedness to measure connectivity within west SE Asian populations and between west and east SE Asia. Genetic similarity and population structure revealed two subpopulations: west SE Asia (Kayin State, SMRU clinics and other Myanmar regions) and east SE Asia (Cambodia, Vietnam and Laos) (Extended Data Fig. 7 and Supplementary Fig. 2). Evidence of gene flow was detected between SMRU clinics and Kayin State: 38.3% of Kayin samples had ≥25% IBD (r ≥ 0.25) with at least one sample from SMRU clinics and 18.1% had ≥35% genome IBD (r ≥ 0.35).
We found no evidence for clonal transmission or recent recombination (r ≥ 0.15) between west and east SE Asia. The low connectivity is further supported by the distribution of pfcrt mutations conferring PPQ resistance. Collectively, 51.21% of east SE Asian (Cambodia, Vietnam and Laos) parasites carried PPQ-resistant pfcrt alleles between 2015 and 2020 (MalariaGEN Pf8 (ref. 10)). These included T93S (22.91%), I218F (13.97%)11,12, H97Y (3.38%), F145I (5.91%), G353V (1.33%)13 and G367C (3.70%)14. By contrast, these pfcrt mutations were absent from the Kayin dataset (Supplementary Table 5) and from other west SE Asian regions (other Myanmar regions and west Thailand).
Genomic measures of parasite population size
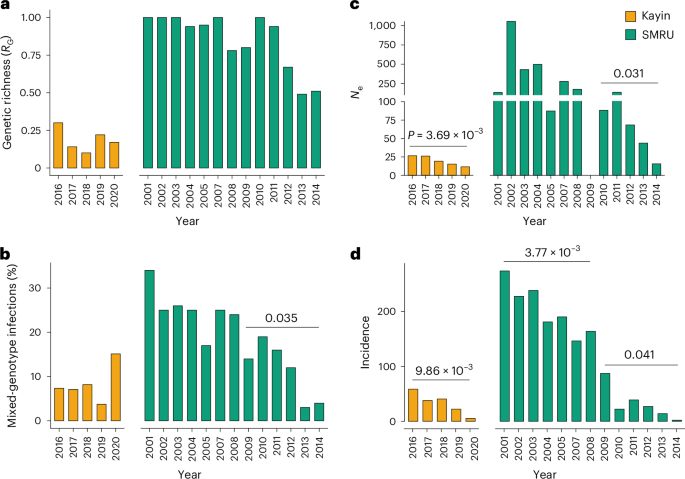
We evaluated three genetic metrics—proportion of multiple-genotype infections, RG and effective population size (Ne) in both Kayin State and SMRU parasite populations, for assessing how control efforts impact parasite population size (Fig. 5). Malaria incidence declined sharply in both regions: from 273.9 to 2.2 cases per 1,000 person-years at SMRU clinics (2001–2014) and from 58.8 to 5.3 cases per 1,000 person-years in Hpapun Township, northern Kayin State (2016–2020), where >96% of sequenced samples originated (Supplementary Tables 4 and 6). To facilitate direct comparison with Kayin, we stratified the SMRU dataset into high-transmission (2001–2008) and low-transmission (2009–2014) periods using an incidence threshold of 100 cases per 1,000 population per year15.
a–c, Population size was estimated using genetic richness (RG) (a), the proportion of samples from mixed-genotype infections (b) and effective population size (Ne) (c). d, Malaria incidence. For Kayin State, data were from Landier et al.8 and Legendre et al.9; the incidence data for SMRU clinics (regions near Mae Sot) was from Nkhoma et al.16. Temporal trends were assessed using linear regression against year; P values are shown only for statistically significant trends; no adjustment for multiple comparisons was applied.
The proportion of multiple genotype infections (7.6%, 146/1927) in Kayin samples was low compared with other P. falciparum populations and remained low throughout the year (range, 3.7–15.1%; Supplementary Table 6). By contrast, SMRU clinics showed a marked decline in multiple genotype infections, from 34.3% in 2001 to 4.3% in 2014, consistent with previous findings16. This decline was not significant during the high-transmission period before 2008 (linear regression, P = 0.23, R2 = 0.27) but was significant during the low-transmission period (P = 0.04, R2 = 0.71). No significant temporal change was observed in Kayin (P = 0.61, R2 = 0.10).
The RG ratio is expected to be negatively correlated with the level of clonal expansion and positively correlated with transmission intensity16. RG ratios were consistently lower in Kayin (0.10–0.30; 2016–2020) than in the SMRU clinics (0.50–1.00; 2001–2014). In SMRU, RG was higher in 2001–2008 (high transmission) than in 2002–2014 (low transmission) (two-sided Welch’s two-sample t-test, P = 0.03). However, RG was not correlated with sampling year in either low- or high-transmission time periods. In Kayin, RG showed little temporal change (P = 0.61, R2 = 0.10).
We calculated single-sample estimates of Ne using unique genomes from each population for each sampling year. Ne estimates were lower in Kayin (11.5–26.6) compared with SMRU clinics (15.5 to infinite). We detected significant reductions in Ne in both Kayin (P = 3.69 × 10−3, R2 = 0.96) and SMRU clinics (P = 0.03, R2 = 0.94) when transmission was low (<100 cases per 1,000 population per year).
The evolution of pfkelch13 alleles
Patterns of drug resistance provide insight into the impact of malaria elimination efforts in Kayin State. Overall, 61.32% of samples from Kayin State carried non-synonymous SNP mutations in kelch13, with P441L (15.19%), F446I (15.01%), R561H (14.02%) and G449A (7.87%) being the most common alleles; only 2.28% carried C580Y (Extended Data Fig. 8). By comparison, at adjacent SMRU clinics, C580Y was the dominant allele, reaching 71.05% frequency in 2014 (Supplementary Table 3). Few samples were collected from SMRU after 2014 owing to the extremely low malaria incidence, limiting temporally matched comparisons. There were 47 IBD clusters (r ≥ 0.90) carrying mutant kelch13 alleles and 46 IBD clusters carrying WT kelch13 in the Kayin samples. Cluster sizes did not differ significantly between mutant and WT kelch13 IBD groups (Extended Data Fig. 9). These results suggest that ART selection was not the main driver for clonal expansion in Kayin State.
Clonal expansion of parasite carrying kelch13-R561H was observed in 2020 (Fig. 3c). Despite strong drug selection, frequencies of mutant kelch13 alleles remained stable between 2016 and 2019 (ref. 6) (Extended Data Fig. 8). However, in 2020, one of the kelch13 alleles—R561H—reached 74.2%. This clonal expansion results from near elimination of parasites from most areas in Kayin, except for northern Hpapun Township where parasites carrying kelch13-R561H predominate (Fig. 4d). In total, 54.8% (40/73) of samples collected between January and August 2020 before the coronavirus disease 2019 pandemic lockdowns were from one single MP (LH-0266B) (Supplementary Table 1).
The changes in kelch13 allele frequencies were reflected by changes in diversity in this gene and its flanking regions. Expected heterozygosity (He) in kelch13 remained high between 2016 and 2019 (He = 0.78) but declined to 0.41 in 2020, with parallel reductions in flanking-region diversity from 0.48 to 0.19 (Extended Data Fig. 8). The sharp decline probably reflects the combined impact of MDA and intensive case management, which together drove a major reduction in malaria transmission. By 2020, infections were detected only in a few remaining villages, indicating a dramatic reduction in parasite population size. This severe bottleneck, and the corresponding drop in Ne, probably led to the marked loss of kelch13 allelic diversity.
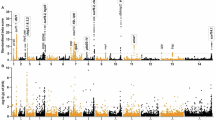
Haplotype reconstruction around kelch13 (±100 kb) revealed diverse genetic backgrounds underlying resistance alleles (Extended Data Fig. 10 and Supplementary Fig. 3). Two P441L, one F446I, one G449A and one R561H haplotypes had shared ancestry between Kayin State and SMRU clinics or other Myanmar regions. However, none of these alleles had a high frequency in SMRU clinics or other Myanmar regions. Two F446I and one C580Y haplotypes were uniquely observed in Kayin State, consistent with local origin. For two Kelch13 resistance alleles (G449A and R561H), the WT kelch13 haplotypes on which these resistance mutations arose were sampled in the early 2000s in SMRU clinics. Of the three C580Y haplotypes identified in Kayin, two were also found in SMRU clinics, while one was unique to Kayin State. The two C580Y haplotypes shared with SMRU clinics were only found to the south of Hpapun Township and 60–120 km north of the SMRU clinics (Supplementary Fig. 4). Malaria incidence was extremely low in the SMRU catchment (12 samples sequenced) between 2015 and 2017; hence, direct comparison of contemporaneous parasite genomes from SMRU and Kayin was not possible. None of these haplotypes shared IBD with east SE Asian C580Y haplotypes (Supplementary Fig. 5).
Impact of mass drug use on parasite population structure
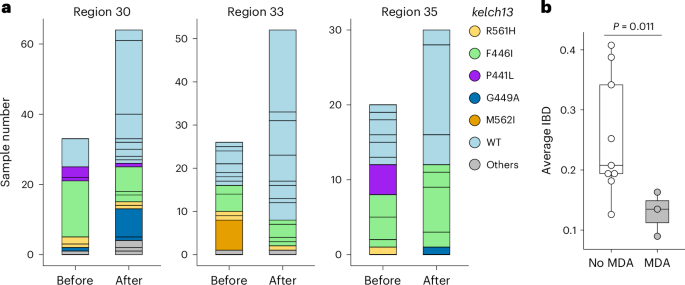
We predicted that MDA would reduce relatedness between pre- and post-MDA parasite populations, by clearing local infections and enabling repopulation by new parasite genotypes, leading to founder effects. There were three HCPC units in which sufficient parasites (n ≥ 20) were sequenced both before and after MDA (Fig. 6a). The relatedness between pre- and post-MDA parasites from these three HCPC units was significantly lower than observed between malaria parasites collected during the sampling time period from HCPC regions where MDA was not used (Fig. 6b). Hence, MDA impacted parasite relatedness, consistent with efficient control of pre-MDA parasite genotypes and post-MDA recolonization with unrelated genotypes.
a, Population relatedness before and after MDA. MDAs were implemented between 24 June 2017 and 12 October 2017 in HCPC regions 30, 33 and 35, using DHA–PPQ plus a single dose of primaquine administered monthly for three consecutive months. Bar segments represent unique genomes identified within the 6 months before (12 April 2017 to 12 October 2017) or after the intervention (13 October 2017 to 12 April 2018), colour-coded by kelch13 allele type. b, Comparison of parasite genetic relatedness between HCPC regions with (n = 3) and without (n = 9) MDA intervention. Parasites collected from the same time windows from HCPC regions where MDA was not used provided ‘no MDA’ controls. P values were calculated using a two-sided Welch’s two-sample t-test; no adjustment for multiple comparisons was applied.
We further assessed drug-resistance markers before and after MDA (Supplementary Table 5). No pfcrt SNPs associated with PPQ resistance were detected. Copy number variation (CNV) analysis of plasmepsin II/III (pm2/3) identified three samples with fold-change >1.5 among 281 with sufficient depth; however, manual inspection in Integrative Genomics Viewer showed no clear CNV boundaries, suggesting that these signals were artefacts of selective whole-genome amplification (sWGA). Thus, we found no evidence of pm2/3 amplification. Analysis of mutations in aat1, mdr1, dhfr, dhps and gch1, as well as CNVs in mdr1, revealed no signatures of MDA-associated selection. Genome-wide IBD-based selection scans similarly showed no strong signals of recent selection (Supplementary Fig. 6).
Discussion
The METF elimination efforts, combining community MPs and MDA, significantly decreased malaria case numbers in the target area (near the Myanmar–Thailand border) between 2014 and 2020 (refs. 8,9). We described the key results from genomic surveillance during these elimination efforts, including parasite transmission patterns, population diversity and genomics, evolution of drug resistance and genomic impacts of MDA using over 2,000 whole genome-sequenced P. falciparum samples collected between November 2015 and August 2020.
Parasite sequences from the METF study region reveal extremely high levels of inbreeding and low levels of genetic variation. We identified 166 genotype clusters (≥90% genome-wide IBD) among 1,781 single-clone samples, indicating that only ~10% of parasites were genetically unique (RG = 0.10). In regions without MDA, 1,408 single-clone samples were divided among 153 clusters, yielding a similar RG of 0.11. The RG in Kayin is comparable to values from Vietnam and Laos during the same period (2015–2020) but lower than those reported from earlier SMRU clinic collections (2001–2014) and from other Myanmar regions outside Kayin (2007–2018). By contrast, genetic richness remained high in African populations (RG = 0.54–1.00)10. Of the 166 unique genomes, 110 formed nine closely related families (r ≥ 0.45). Parasites within these families typically carry one or two kelch13 alleles, which most likely inherited from the parents. This is clearly the case in the two families with both parents identified—f1 (F446I and WT kelch13) and f7 (C580Y and WT kelch13). Hence, recombination is rare in these populations and most infections are clonally related.
We observed strong spatial structure in the parasite population. This is particularly clear from the distribution of unique parasite genotypes. Such parasites result from self-fertilization of male and female gametes of the same genotype and allow spatial tracing of transmission networks. Visually, it is clear that these transmission networks are clustered in space (Fig. 4a), and statistically, it is evident from autocorrelation analyses (Fig. 4b), which reveal positive correlations in relatedness between genotypes for up to 20 km. The local distribution of unique genotypes indicates (1) local transmission and very few long-distance transmission events and (2) reintroduction of circulating genotypes from asymptomatic carriers.
Unique parasite genotypes were long lived in this low-transmission setting, with some IBD clusters observed over the complete 58-month study period. This is clear from (1) the temporal autocorrelation, which reveals positive correlation in relatedness between parasites collected 170 days apart, and (2) the elevated relatedness observed in parasites collected from the same HCPC regions but up 3 years apart. The strong spatial and temporal sub-structure of parasite populations in Kayin State is comparable to that observed in Cambodia17 and Guyana18.
As elimination approaches, genetic drift is expected to play an increasingly important role and expansions of parasite lineages may occur19. We observed a sudden increase in the frequency of IBD cluster 1 parasites carrying kelch13-R561H in 2020. In this case, the increase in kelch13-R561H frequency resulted from elimination of malaria from all regions of Kayin other than northern Hpapun Township (Fig. 4a), where IBD cluster 1 carrying kelch13-R561H is at high frequency. Despite predominating in northern Hpapun Township since 2017, the kelch13-R561H did not spread into other areas of Kayin State. The rapid-frequency increase in IBD cluster 1 in 2020 is consistent with bottlenecks and genetic drift in a P. falciparum population nearing elimination. Further sampling will determine whether this parasite genotype spreads further in Kayin State and elsewhere in Myanmar and Thailand. By contrast, Wasakul et al.19 describe a classic outbreak driven by a selective sweep in Laos, where a lineage carrying kelch13-R539H (named LAA1) rose from a low frequency to replace the previously dominant KEL1/PLA1 (C580Y) population.
Two control measures were used in Kayin State by METF: MPs (early diagnosis and community case management) and regional MDA in malaria hot spots8. The combination of these approaches significantly decreased malaria incidence8,9. Encouragingly, there was minimal evidence of selection for drug-resistance parasites through these elimination efforts, which agrees with observations by Imwong et al.20, McLean et al.21 and Thu et al.6. At the genomic level, these combined control measures reduced parasite effective population size (Fig. 5). This study provided an opportunity to examine the impact of one component of this elimination strategy—MDA—on parasite population structure. MDA is expected to generate bottlenecks between pre- and post-MDA malaria populations because reservoirs of asymptomatic malaria are removed. We therefore expect to see founder effects resulting from newly colonizing parasites and large divergence between pre- and post-MDA populations, when compared with populations with no MDA. Our power to detect an impact was limited because most parasites sequenced were collected after MDA: only three HCPC units had sufficient numbers of infection sampled both before and after MDA to examine this hypothesis. Nevertheless, as predicted, we saw lower relatedness between pre- and post-treatment parasites in these three HCPC regions than in control regions where MDA was not implemented. These results provide genomic evidence for the effectiveness of MDA in removing local parasite populations through effective clearance of both asymptomatic and symptomatic parasite infections.
Genetic metrics, such as proportion of multiple clone infections, can provide useful metrics for assessing control efforts15,16,22. Such metrics are particularly useful in low-transmission regions, where a high prevalence of low-density asymptomatic infections complicates assessment of transmission using standard epidemiological methods. However, genetic metrics perform poorly when transmission levels are extremely low. In Senegal15,22, complexity of infection (number of genetically distinct parasite strains co-infecting an individual) provided an unreliable metric for evaluating transmission when transmission is incidence <100 cases per 1,000 person-years. Consistent with this, we observed that both RG and proportion of multiple infections worked poorly in Kayin where incidence ranged from 1 to 39 cases per 1,000 person-years. However, we found that another metric, effective population size (Ne), calculated from linkage disequilibrium between unlinked markers, showed a significant decline in both the Kayin State parasite population and from SMRU clinics. We suggest that Ne may be a useful genomic indicator of transmission dynamics, particularly in parasite populations in which transmission has been reduced to near elimination levels. Ne is typically used in conservation biology to assess viability in endangered populations of animals and plants. Our results suggest that this metric may also have utility for assessing whether parasite populations are approaching local extinction.
The current frontline treatments for P. falciparum malaria parasites have been failing in east SE Asia20,23,24, owing to the spread of the multidrug-resistant parasites carrying kelch13-C580Y mutation and plasmepsin 2 amplifications, named KEL1/PLA1. The KEL1/PLA1 lineage was first detected in Cambodia, as DHA–PPQ was heavily used. When Cambodia withdrew DHA–PPQ and adopted artesunate–mefloquine, KEL1/PLA1 subgroups with acquired pfcrt mutations conferring PPQ resistance rapidly spread to other east SE Asian countries, such as Laos and Vietnam23. There is a concern that this parasite lineage will further spread to west SE Asia, which has the majority of malaria cases in SE Asia21 and where DHA–PPQ is the frontline treatment for P. falciparum. Two lines of evidence suggest minimal gene flow between east and west SE Asia. First, we did not detect pfcrt mutations associated with PPQ resistance on the Thailand–Myanmar border or in the Kayin State sampling sites. Second, examination of genome-wide IBD sharing among 9,115 infections (2,302 unique genomes) revealed no recent recombination or clonal transmission between east and west SE Asia (Extended Data Fig. 7), consistent with previous studies by Shetty et al.25 and Zhu et al.26.
Kelch13 mutations conferring ART resistance are established in both east and west SE Asia. C580Y is the major mutation in regions other than northern Myanmar, where F446I predominates20. By contrast, the dominant kelch13 mutations in the Kayin State include P441L, F446I, R561H and G449A, depending on the location (Fig. 4). While the majority of infections from nearby SMRU clinics carry C580Y (71.05% in 2014), the C580Y frequency in Kayin State was only 2.28%. The most widespread F446I haplotypes in Kayin State originated independently from the dominant F446I haplotype in northern Myanmar.
What factors led to the patterns of ART resistance evolution seen in Kayin? Longitudinal studies in both Cambodia and from SMRU clinics have revealed that multiple independent kelch13 mutations emerged and spread initially (soft selective sweeps). Single kelch13 genotypes (typically kelch13-C580Y) eventually outcompeted other lineages, leading to hard selective sweeps2,17,20,23. By contrast, we found limited evidence that strong drug selection drove drug resistance evolution in Kayin State: (1) We found no significant increase in kelch13 mutant allele frequencies before 2020 (ref. 6) (Fig. 3c and Extended Data Fig. 8) and (2) the size of clonal clusters was not significantly different when comparing kelch13 WT and mutant parasites (Extended Data Fig. 9). The small effective population size of malaria parasite populations may contribute to the patterns observed because selection is inefficient when population sizes are small and genetic drift is enhanced27. The initial effective population size of malaria parasites in the Kayin State dataset was much smaller (Ne = 11.5–26.6) compared with SMRU clinics (15.5 to infinite) (Fig. 5). Other factors that may also limit the impact of drug selection. Human population movements were more limited in Kayin compared with nearby SMRU clinics, especially in Northern Hpapun Township where human movement is limited by difficult terrain, the heavily militarized landscape and a lack of year-round roads7,28, which can hinder transmission of resistance alleles. Similarly, low levels of recombination in Kayin State limits the rate of formation of new multilocus parasite genotypes. The small parasite population size, limited population movement and minimal recombination enhance the role of genetic drift rather than selection in determining drug resistance evolution in the Kayin State region. Our results, and those from other studies20, illustrate how genetic drift can result in rapid stochastic changes in parasite population genomics and drug resistance status in small parasite populations close to elimination.
This study had several limitations. First, we analysed malaria genomes collected from 2015 onwards. However, control efforts began earlier than this in 2014. Hence, we were unable to examine malaria population structure and diversity before initiation of control efforts. Second, the use of finger-prick blood samples and whole-genome amplification resulted in bias towards sequencing high-parasitaemia infections. However, the sustained sampling of a high proportion of blood spots collected over a 5-year period provides a unique dataset for examining the impact of malaria control efforts on parasite population structure and resistance evolution.
Methods
Ethics statement
The METF program was approved by the Department of Medical Research (Lower Myanmar) 73/Ethics 2014 and OxTREC (reference no. 1017-13).
Study area and sample origins
The samples for this analysis were collected during routine diagnosis and treatment efforts in Kayin State as part of the METF malaria elimination effort led by the SMRU (based on the Thailand–Myanmar border) (Fig. 1 and Extended Data Fig. 1). This METF project was established in 2014 and used two primary P. falciparum-focused interventions: the establishment of a large network of community-based malaria diagnosis and treatment posts (MPs) and targeted MDA in communities determined to have a high prevalence of asymptomatic P. falciparum infections. The MPs were stocked with filter papers (Whatman 3 mm blotting paper) and were asked to collect DBSs from finger-prick blood samples for patients with rapid diagnostic test-confirmed P. falciparum infection. Each DBS sample is linked to the MP from which it originated, and all MPs have geographic references (latitude and longitude). A total of 5,014 DBS samples were collected between November 2015 and August 2020 (Supplementary Table 1). The DBS samples were then transported to the SMRU and subsequently shipped to the Texas Biomedical Research Institute (in the USA) for molecular analyses.
Sequencing library preparation
We extracted DNA from the DBSs and enriched parasite genomes using sWGA following Li et al.29 and Oyola et al.30. In short, we extracted and purified genomic DNA using QIAamp 96 DNA Blood Kit or QIAamp DNA Mini Kit, following the instruction manual for DNA isolation from DBSs. The DNA was then eluted in 100 µl of 10 mM Tris–HCl (pH 8.0–8.5) buffer. We used real-time quantitative PCR (qPCR) to estimate the numbers of parasite genomes in each DNA sample29.
For samples with more than 200 copies of parasite genome per µl, we used sWGA to enrich parasite DNA. sWGA reactions were performed following Oyola et al.30. In short, each 25-μl reaction contained at least 1,000 copies of Plasmodium DNA, 1× BSA (New England Biolabs), 1 mM deoxynucleotide triphosphates (dNTPs; New England Biolabs), 3.5 μM of each amplification primer, 1× Phi29 reaction buffer (New England Biolabs) and 15 units of Phi29 polymerase (New England Biolabs). We used a PCR machine (SimpliAmp, Applied Biosystems) programmed to run a ‘stepdown’ protocol: 35 °C for 10 min, 34 °C for 10 min, 33 °C for 10 min, 32 °C for 10 min, 31 °C for 10 min, 30 °C for 6 h and then heating at 65 °C for 10 min to inactivate the enzymes before cooling to 4 °C. Sample were cleaned with AMPure XP Beads (Beckman Coulter), at a 1:1 ratio.
We used the Quant-iT PicoGreen Assay (Invitrogen) to determine the total amount of sWGA product and quantified the proportion of Plasmodium DNA by qPCR. Only sWGA products with more than 50% DNA from Plasmodium were used for further library preparation and Illumina sequencing. We constructed PCR-free next-generation sequencing libraries using 300 ng sWGA product following the KAPA HyperPlus Kit protocol. All libraries were sequenced to an average coverage of 60× using Illumina Hiseq X or Novaseq sequencers.
Whole genome-sequencing data generation
We individually mapped whole-genome sequencing reads for each library against the P. falciparum 3D7 reference genome (PlasmoDB, release 46) and human GRCh38 reference genome, using the alignment algorithm BWA mem (http://bio-bwa.sourceforge.net/) under the default parameters. The resulting alignments were then converted to SAM format, sorted to BAM format and deduplicated using picard tools v2.0.1 (ref. 31). Reads that mapped to the human genome were discarded before genotyping.
We used Genome Analysis Toolkit GATK v3.7 (ref. 32) to recalibrate the base quality score on the basis of a set of verified known variants33. We called variants for each sample using HaplotypeCaller, and calls from every 100 samples were merged using CombineGVCFs with default parameters. Variants were further called at all sample-level using GenotypeGVCFs, with the following parameters: --max_alternate_alleles 6 --variant_index_type LINEAR --variant_index_parameter 128000 --sample_ploidy 2 -nt 20.
The recalibrated variant quality scores were calculated by comparing the raw variant distribution with the known and verified Plasmodium-variant dataset. SNPs and indels with a variant quality score lower than 1 or located outside of the core genome regions (21 Mb, defined in ref. 33) were removed from further analysis. Samples with less than 50% of the core genome callable were also excluded from further analysis. Only biallelic SNPs that pass all the quality filter were used, unless otherwise specified. The final variants in VCF format were annotated at functional effect to genes and proteins using snpEff v4.3 (ref. 34) with 3D7 (PlasmoDB, release 46) as reference.
We initially identified 1,302,006 SNPs and 703,138 indels (Fig. 1c). We removed 343 samples with >20% genotypes missing. We then filtered the SNP calls following a ‘stringent’ filtering method35, to generate a final list of 447,435 high-quality, biallelic, core genome-located (defined in ref. 33) SNPs. For further population structure analysis, we removed SNPs that were genotyped in less than 50% of samples or with a minor allele frequency <0.05.
Within-infection genetic diversity
We measured multiplicity of P. falciparum infections using the within-infection inbreeding coefficient (Fws)10,36. Only coding biallelic SNPs and loci with coverage more than one read for either the reference or alternative allele (more than six total reads) were included in the Fws calculation. Loci with within-sample allele frequencies <0.05 or >0.95 were rounded to 0 or 1, respectively. Samples with Fws ≥ 0.95 were assumed to come from single-genotype infections for samples from Kayin State. Allele frequencies across the genome were plotted and manually inspected to detect further possible complex infections.
IBD analysis
We used relatedness—r, defined as the fraction of the genome that is IBD between a pair of individuals37—to estimate parasite relationships, performed using hmmIBD v2.0 (ref. 38). On the basis of the distribution of relatedness among F1 progeny from malaria parasite genetic crosses (Supplementary Fig. 3), we assume that parasites are genetically related if ≥25% of their genome is identical (r ≥ 0.25); parasites are closely related (such as siblings or parent and progeny) if their relatedness is greater than 45% (r ≥ 0.45). We considered samples to be clonal if their relatedness is over 90% (r ≥ 0.90). We visualized relatedness among samples using the R package pheatmap v1.0.13 and the Cytoscape v3.10.3 software. We also examined the recombination patterns between closely related parasites and plotted shared IBD regions between estimated parents and progeny using karyoploteR v1.28.0.
To detect potential post-MDA selective sweeps, we performed genome-wide scans of enhanced relatedness to identify potential signals of selection following MDA using isoRelate v0.9.7 (ref. 39). We compared IBD relatedness patterns by sampling year and between pre- and post-MDA periods.
Genotyping of drug resistance loci
We evaluated drug-resistance markers in P. falciparum isolates from Kayin State by examining SNPs in pfcrt and additional loci, as well as CNV in pm2/3, gch1 and mdr1. pfcrt polymorphisms were called using Illumina short-read data, and variants were compared with previously reported alleles associated with chloroquine or PPQ resistance. CNV analysis of pm2/3 followed established amplification boundaries defined in MalariaGEN Pf8 (ref. 10) and by Kane et al.14. Read coverage within the CNV region was compared with 10-kb upstream and downstream flanks, and coverage fold-change was calculated as the ratio of mean coverage inside versus outside the CNV region. Samples with a fold-change ≥1.5 and ≥20-fold coverage across all regions were flagged as potential amplifications; these were further evaluated by manual inspection in Integrative Genomics Viewer to confirm the presence or absence of step-like CNV boundaries. The same CNV detection approach was applied to gch1 and mdr1 using boundaries defined in MalariaGEN Pf8 (ref. 10). To identify hrp2 and hrp3 deletions, we visually inspected read coverage across each locus, classifying samples with <3× coverage across the gene and ≥20× coverage in the β-tubulin control region as putative deletions (Supplementary Table 5).
Surveillance of kelch13 haplotypes
We extracted SNPs distributed within 100 kb upstream and 100 kb downstream of the kelch13 gene. We measured expected heterozygosity (He) at the kelch13 locus by treating kelch13 as a single locus with multiple alleles. We also measured He over the 200-kb kelch13 haplotype region. To compare the relationships between different kelch13 haplotypes, we measured pairwise IBD sharing among all kelch13 haplotypes. We assume that haplotypes with IBD sharing ≥0.90 originated from the same mutational event; that when 0.35≤ IBD <0.90, there was a one least recombination event to break the original haplotype and when IBD <0.35, these haplotypes have emerged independently.
Comparisons of malaria parasite populations
We compared the Kayin State parasite population with other worldwide malaria parasite populations (Fig. 1 and Supplementary Table 3). The SMRU clinics are located around Mae Sot, in Tak Province, along the international Thailand–Myanmar border. We used ‘other Myanmar’ to represent sampling sites in Myanmar but not from Kayin State. West SE Asia population includes samples from Kayin State, SMRU clinics and other Myanmar regions, while east SE Asia population includes Cambodia, Vietnam and Laos.
We merged raw SNP genotypes from the Kayin dataset with those from the MalariaGEN P. falciparum Community Project (Pf8)10. We performed ‘stringent’ filtration as described above and selected loci with minor allele frequency ≥0.05. We calculated genetic richness (RG = (G – 1)/(S – 1))40,41 to quantify the richness of clonal parasites in each population, where G is the number of unique genomes and S is the total number of single-genotype-infected samples. For samples with relatedness ≥0.9, only one representative sample per population per year with the highest genotype rate was selected and used for further analysis (Supplementary Table 3). We conducted principal component analyses using R package SNPRelate42. We estimated effective population size (Ne) on the basis of patterns of linkage disequilibrium at unlinked loci, using methods implemented in NeEstimator v2.0 (ref. 43).
Statistical analysis
All statistical analyses were performed using R version 4.1.3. For pairwise comparisons between groups, we used Welch’s two-sample t-test. We measured correlations (autocorrelation coefficients) between parasite genetic relatedness and geographic distance or time with the Mantel statistic using the mantel function in the vegan v2.6-6 package. The autocorrelation coefficient ranges from +1 to −1, where positive values indicate that individuals or populations at a given distance are more genetically similar than expected by chance, zero indicates similarity expected at random and negative values indicate they are more genetically different than expected. P < 0.05 was considered statistically significant. We used HCPC following scripts in FactoMineR v2.11 (ref. 44) to divide the 283 MPs with samples sequenced into 50 HCPC regions on the basis of latitude and longitude. We then compared parasite relatedness within and between HCPC regions for parasites collected in the same year, between 1–2, 2–3 and 3–4 years apart. We compared relatedness between parasites collected from HCPC regions 6 months before and 6 months after MDA. As controls, we examined relatedness of parasites collected from HCPC regions where MDA was not used in the same time windows.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw sequencing data for the 2,270 sequenced samples collected by the Malaria Elimination Task Force project from Myanmar used in the present analysis have been submitted to the NABI Sequence Read Archive (SRA; https://www.ncbi.nlm.nih.gov/sra) under project no. PRJNA864839. The data matrices (genetic distances, geographic distances and temporal distances) are available via GitHub at https://github.com/emilyli0325/Malaria-genomics-in-Eastern-Myanmar. This publication also uses data from the MalariaGEN P. falciparum Community Project (www.malariagen.net/data_package/open-dataset-plasmodium-falciparum-v80/). Source data are provided with this paper.
Code availability
The analysis codes are available via GitHub at https://github.com/emilyli0325/Malaria-genomics-in-Eastern-Myanmar.
References
Venkatesan, P. The 2023 WHO world malaria report. Lancet Microbe 5, e214 (2024).
Anderson, T. J. et al. Population parameters underlying an ongoing soft sweep in Southeast Asian malaria parasites. Mol. Biol. Evol. 34, 131–144 (2017).
Wellems, T. E. & Plowe, C. V. Chloroquine-resistant malaria. J. Infect. Dis. 184, 770–776 (2001).
Imwong, M. et al. Numerical distributions of parasite densities during asymptomatic malaria. J. Infect. Dis. 213, 1322–1329 (2016).
White, N. J. Does antimalarial mass drug administration increase or decrease the risk of resistance?. Lancet Infect. Dis. 17, e15–e20 (2017).
Thu, A. M. et al. Molecular markers of artemisinin resistance during falciparum malaria elimination in Eastern Myanmar. Malar. J. 23, 138 (2024).
Parker, D. M. et al. Scale up of a Plasmodium falciparum elimination program and surveillance system in Kayin State, Myanmar. Wellcome Open Res. 2, 98 (2017).
Landier, J. et al. Effect of generalised access to early diagnosis and treatment and targeted mass drug administration on Plasmodium falciparum malaria in Eastern Myanmar: an observational study of a regional elimination programme. Lancet 391, 1916–1926 (2018).
Legendre, E. et al. ‘Forest malaria’ in Myanmar? Tracking transmission landscapes in a diversity of environments. Parasit. Vectors 16, 324 (2023).
Malaria Genomic Epidemiology, N. et al. Pf8: an open dataset of Plasmodium falciparum genome variation in 33,325 worldwide samples. Wellcome Open Res. 10, 325 (2025).
Boonyalai, N. et al. Piperaquine resistant Cambodian Plasmodium falciparum clinical isolates: in vitro genotypic and phenotypic characterization. Malar. J. 19, 269 (2020).
Small-Saunders, J. L. et al. Evidence for the early emergence of piperaquine-resistant Plasmodium falciparum malaria and modeling strategies to mitigate resistance. PLoS Pathog. 18, e1010278 (2022).
Ross, L. S. et al. Emerging Southeast Asian PfCRT mutations confer Plasmodium falciparum resistance to the first-line antimalarial piperaquine. Nat. Commun. 9, 3314 (2018).
Kane, J. et al. A Plasmodium falciparum genetic cross reveals the contributions of pfcrt and plasmepsin II/III to piperaquine drug resistance. mBio 15, e0080524 (2024).
Wong, W. et al. Evaluating the performance of Plasmodium falciparum genetic metrics for inferring National Malaria Control Programme reported incidence in Senegal. Malar. J. 23, 68 (2024).
Nkhoma, S. C. et al. Population genetic correlates of declining transmission in a human pathogen. Mol. Ecol. 22, 273–285 (2013).
Miotto, O. et al. Multiple populations of artemisinin-resistant Plasmodium falciparum in Cambodia. Nat. Genet. 45, 648–655 (2013).
Vanhove, M. et al. Temporal and spatial dynamics of Plasmodium falciparum clonal lineages in Guyana. PLoS Pathog. 20, e1012013 (2024).
Wasakul, V. et al. Malaria outbreak in Laos driven by a selective sweep for Plasmodium falciparum kelch13 R539T mutants: a genetic epidemiology analysis. Lancet Infect. Dis. 23, 568–577 (2023).
Imwong, M. et al. Molecular epidemiology of resistance to antimalarial drugs in the Greater Mekong subregion: an observational study. Lancet Infect. Dis. 20, 1470–1480 (2020).
McLean, A. R. D. et al. Mass drug administration for the acceleration of malaria elimination in a region of Myanmar with artemisinin-resistant falciparum malaria: a cluster-randomised trial. Lancet Infect. Dis. 21, 1579–1589 (2021).
Schaffner, S. F. et al. Malaria surveillance reveals parasite relatedness, signatures of selection, and correlates of transmission across Senegal. Nat. Commun. 14, 7268 (2023).
Hamilton, W. L. et al. Evolution and expansion of multidrug-resistant malaria in southeast Asia: a genomic epidemiology study. Lancet Infect. Dis. 19, 943–951 (2019).
Amaratunga, C. et al. Dihydroartemisinin-piperaquine resistance in Plasmodium falciparum malaria in Cambodia: a multisite prospective cohort study. Lancet Infect. Dis. 16, 357–365 (2016).
Shetty, A. C. et al. Genomic structure and diversity of Plasmodium falciparum in Southeast Asia reveal recent parasite migration patterns. Nat. Commun. 10, 2665 (2019).
Zhu, L. et al. The origins of malaria artemisinin resistance defined by a genetic and transcriptomic background. Nat. Commun. 9, 5158 (2018).
Barton, N. H. Natural selection and random genetic drift as causes of evolution on islands. Philos. Trans. R. Soc. Lond. B 351, 785–794 (1996).
Parker, D. M., Carrara, V. I., Pukrittayakamee, S., McGready, R. & Nosten, F. H. Malaria ecology along the Thailand–Myanmar border. Malar. J. 14, 388 (2015).
Li, X. et al. Genetic mapping of fitness determinants across the malaria parasite Plasmodium falciparum life cycle. PLoS Genet. 15, e1008453 (2019).
Oyola, S. O. et al. Whole genome sequencing of Plasmodium falciparum from dried blood spots using selective whole genome amplification. Malar. J. 15, 597 (2016).
Picard. Broad Institute http://broadinstitute.github.io/picard/ (2025).
Genome Analysis Toolkit. Broad Institute https://software.broadinstitute.org/gatk/ (2026).
Miles, A. et al. Indels, structural variation, and recombination drive genomic diversity in Plasmodium falciparum. Genome Res. 26, 1288–1299 (2016).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
McDew-White, M. et al. Mode and tempo of microsatellite length change in a malaria parasite mutation accumulation experiment. Genome Biol. Evol. 11, 1971–1985 (2019).
Manske, M. et al. Analysis of Plasmodium falciparum diversity in natural infections by deep sequencing. Nature 487, 375–379 (2012).
Brown, T. S., Arogbokun, O., Buckee, C. O. & Chang, H. H. Distinguishing gene flow between malaria parasite populations. PLoS Genet. 17, e1009335 (2021).
Schaffner, S. F., Taylor, A. R., Wong, W., Wirth, D. F. & Neafsey, D. E. hmmIBD: software to infer pairwise identity by descent between haploid genotypes. Malar. J. 17, 196 (2018).
Henden, L., Lee, S., Mueller, I., Barry, A. & Bahlo, M. Identity-by-descent analyses for measuring population dynamics and selection in recombining pathogens. PLoS Genet. 14, e1007279 (2018).
Eckert, C. G., Dorken, M. E. & Mitchell, S. A. Loss of sex in clonal populations of a flowering plant, Decodon verticillatus (Lythraceae). Evolution 53, 1079–1092 (1999).
Echeverry, D. F. et al. Long term persistence of clonal malaria parasite Plasmodium falciparum lineages in the Colombian Pacific region. BMC Genet. 14, 2 (2013).
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012).
Do, C. et al. NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol. Ecol. Resour. 14, 209–214 (2014).
Lê, S., Josse, J. & Husson, F. FactoMineR: an R package for multivariate analysis. J. Stat. Softw. 25, 1–18 (2008).
Button-Simons, K. A. et al. The power and promise of genetic mapping from Plasmodium falciparum crosses utilizing human liver-chimeric mice. Commun. Biol. 4, 734 (2021).
Kumar, S. et al. A malaria parasite cross reveals genetic determinants of Plasmodium falciparum growth in different culture media. Front. Cell. Infect. Microbiol. 12, 878496 (2022).
Amambua-Ngwa, A. et al. Chloroquine resistance evolution in Plasmodium falciparum is mediated by the putative amino acid transporter AAT1. Nat. Microbiol. 8, 1213–1226 (2023).
Acknowledgements
This work was supported by National Institutes of Health (NIH) grant nos. R01 AI048071 (to T.J.C.A.) and P01 AI127338 (to T.J.C.A.: Core B PI). Work at Texas Biomedical Research Institute was conducted in facilities constructed with support from Research Facilities Improvement Program grant no. C06 RR013556 from the National Center for Research Resources. The Shoklo Malaria Research Unit (SMRU) is part of the Mahidol Oxford University Research Unit supported by the Wellcome Trust of Great Britain. The malaria elimination programme (Malaria Elimination Task Force (METF)) in Kayin State, Myanmar, is supported by the Regional Artemisinin Initiative (Global Fund to Fight AIDS, Tuberculosis and Malaria) and the Bill and Melinda Gates Foundation (award no. OPP1117507, to F.N.). We thank the contributions of members of the METF and SMRU, collaborators and colleagues who have supported the elimination programme. We also thank N. White and R. Pearson for providing code essential to the Fws calculations.
Author information
Authors and Affiliations
Contributions
X.L., F.N. and T.J.C.A. conceived of and designed the study. J.L., A.M.T., G.D., D.M.P., K.M.L., K.S. and F.N. coordinated the sample and data collection. X.L., G.A.A. and A.R. processed the samples and generated genomic data. X.L. analysed and interpreted the sequencing data, with input from T.J.C.A., D.M.P., J.L. and F.N. K.M.L., K.S., J.L., D.M.P. and F.N. were involved in the management and coordination of the genetic surveillance project. X.L. and T.J.C.A. wrote the initial paper. X.L., D.M.P., J.L., F.N. and T.J.C.A. revised the paper. X.L., D.M.P., F.N. and T.J.C.A. accessed and verified all the data. All authors provided critical revision of the paper. All authors had full access to all the data in the study and had final responsibility for the decision to submit for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Locations of the Malaria Elimination Task Force (METF) project.
a, The METF project was performed at four townships (Myawaddy, Kawkareik, Hlaingbwe, and Hpapun) of Kayin state (orange shaded region), Myanmar. The control activities were started in 2014. The malaria clinics run by the Shoklo Malaria Research Unit (SMRU) were located along the western border of Thailand (red squares), opened since 2001. The majority of patients of SMRU clinics came from adjacent Myanmar. Blue cycles indicate other Myanmar sampling sites that were not located in Kayin state. b, Location of malaria posts and mass drug administration. Red dots, locations where mass drug administration (MDA) was applied; green dots, location of malaria posts.
Extended Data Fig. 2 Within-infection genetic diversity measured by inbreeding coefficient (FWS).
a, Cumulative distributions of FWS values across geographic regions. The curves plateau at approximately 0.95 for all populations; therefore, we classified infections with FWS ≥ 0.95 as single-genotype infections. b, Distribution of FWS values across populations. c, Temporal changes in FWS among Kayin samples, with the proportion of single-genotype infections indicated above each panel.
Extended Data Fig. 3 IBD distribution in malaria parasite genetic crosses and in Kayin samples.
a-c, Distribution of pairwise genome-wide IBD proportions between any two progeny or between progeny and parents. Panels A and C show strong peaks at IBD = 1, reflecting the high frequency of self-fertilization in the MKK2835×NHP1337 and Mal31×KH004 crosses. In contrast, no peak at IBD = 1 is observed in panel B, consistent with the NF54×NHP4026 cross yielding very few selfed progeny. We used these experimental crosses to establish thresholds for investigating relatedness in field collected samples. Filtered genotypes for cloned progeny from the three crosses (MKK2835xNHP1337, NF54xNHP4026, and Mal31xKH004) are available at https://github.com/emilyli0325/P01-cloned.progeny. Raw sequencing data has been published at: MKK2835xNHP1337 (Li et al.29 and Button-Simons et al.45), NF54xNHP4026 (Button-Simons et al.45, Kumar et al.46. and Amambua-Ngwa et al.47), Mal31xKH004 (Kane et al.14). d, Distribution of pairwise genome-wide IBD proportions in Kayin parasites. We used a conservative cutoff of 0.9 (red dotted line) to identify clonal genotypes.
Extended Data Fig. 4 Chromosome plots for estimated progeny from family 1.
We infer that the parents of family 1 are C13 (parent A, IBD cluster 13, kelch13-wildtype) and C17 (parent B, IBD cluster 17, kelch13-F446I).
Extended Data Fig. 5 Closed related parasite families and their duration.
a, Duration of each parasite family. One dot indicates one sample, and each line indicates one unique genome. Families are defined as a group of parasites that share over 45% of their genome (r ≥ 0.45). b, Density distribution of unique genome durations.
Extended Data Fig. 6 Grouping of malaria posts (MPs) based on hierarchical clustering on principal components (HCPC) of geographic location.
MPs geographically close to each other, determined by HCPC analysis, were grouped together (circles) for further population relatedness analyses. This map shows the 29 HCPC-defined regions, each containing more than 10 samples collected across the entire sampling period. Dots represent locations of malaria posts from which sequenced samples were obtained.
Extended Data Fig. 7 Parasite connection network based on pair-wise genetic relatedness.
Networks were illustrated using Cytoscape and were colored according to the geographical origin (a) or kelch13 alleles (b). Lines indicate connections with relatedness (r) ≥ 0.35. There was no connection detected between east and west of Southeast Asia with r ≥ 0.15. Closely related parasites (r ≥ 0.35) were found between Kayin state and SMRU clinics on Thailand-Myanmar border. To assess the directionality of long-distance connectivity between Kayin State and the SMRU clinics, we used these closely related parasite pairs (r ≥ 0.35) sampled from both locations. Two Kayin parasites were closely related to common parasite genotypes from SMRU, consistent with geneflow from SMRU to Kayin. Similarly, five SMRU parasites were related to common genotypes from Kayin, consistent with geneflow from SMRU to Kayin. These findings indicate bidirectional connectivity, although interpretation is limited by lack of contemporary sampling at SMRU.
Extended Data Fig. 8 Prevalence of kelch13 mutations.
a, Percentages of kelch13 alleles in samples from Kayin State. b, Temporal changes of kelch13 allele frequency. The number of samples in each year is labeled on top of the plot. There is a slow decrease in wild-type kelch13 alleles. An outbreak of R561H allele was detected in 2020. Colored lines, alleles with at least one year > 2%, grey lines, other alleles. c, Expected heterozygosity of kelch13 gene across time. d, Expected heterozygosity of kelch13 spanning region. The analysis was based on single nucleotide polymorphisms (SNPs) located 100 kb up and down stream of the kelch13 gene. Only SNPs with minor allele frequency (MAF) > 0.3 in 2016 were used.
Extended Data Fig. 9 Temporal changes in unique genome proportion.
a, Sample size of parasites with different unique genomes across time. The barplot was separated by IBD genome groups (unique genomes, with ≥ 90% of genome identical), colored based on kelch13 alleles; See Fig. 3a for color legend of kelch13 alleles; black blocks indicate number of unique genomes. b, Comparison of the size of IBD groups carrying kelch13 wild-type and mutant alleles. No significant difference was detected (Welch Two Sample t-test, p-value = 0.34). c, Proportion distribution of IBD groups with wild-type and mutant alleles kelch13 alleles across time. No significant difference was detected (p-value = 0.54, 0.17, 0.37, 0.20 and 0.41 for 2016-2020, separately).
Extended Data Fig. 10 Analyses of kelch13 haplotypes in Kayin region.
Left panel, haplotype diagram showing 100 kb upstream and down-stream of the kelch13 gene. Each column represents a SNP with minor allele frequency (MAF) ≥ 0.05 in Kayin; each row represents a unique genome; color represents genotype: reference (green), alternative (orange), and white (ungenotyped). Middle panel, kelch13 haplotypes: orange and blue blocks represent haplotypes that have high allele frequency in Kayin; green blocks indicate major haplotype from other Myanmar regions; and red blocks are C580Y haplotypes dominating SMRU clinics. Right panel, frequency of unique genomes from Kayin.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–6.
Supplementary Tables 1–6 (download XLSX )
Supplementary Table 1: Information of samples from Kayin State. Supplementary Table 2: Information of samples from other SE Asian populations. Supplementary Table 3: Comparison of estimated parasite population size. Supplementary Table 4: Genotyping and summary of drug resistance loci in Kayin samples. Supplementary Table 5: Worldwide malaria parasite datasets used in this study. Supplementary Table 6: Comparison of genetic metrics between Kayin State and Thailand–Myanmar border (SMRU clinics) populations.
Source data
Source Data Fig. 1 (download XLSX )
Source data for Fig. 1a,b.
Source Data Fig. 2 (download XLSX )
Source data for Fig. 2.
Source Data Fig. 3 (download XLSX )
Source data for Fig. 3c.
Source Data Fig. 4 (download XLSX )
Source data for Fig. 4.
Source Data Fig. 5 (download XLSX )
Source data for Fig. 5.
Source Data Fig. 6 (download XLSX )
Source data for Fig. 6.
Source Data Extended Data Fig. 1 (download XLSX )
Source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2 (download XLSX )
Source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3 (download XLSX )
Source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4 (download XLSX )
Source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5 (download XLSX )
Source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6 (download XLSX )
Source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7 (download XLSX )
Source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8 (download XLSX )
Source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 9 (download XLSX )
Source data for Extended Data Fig. 9.
Source Data Extended Data Fig. 10 (download XLSX )
Source data for Extended Data Fig. 10.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, X., Arya, G.A., Thu, A.M. et al. Impact of intensive control on malaria population genomics under elimination settings in Southeast Asia. Nat Microbiol (2026). https://doi.org/10.1038/s41564-026-02327-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41564-026-02327-1