Abstract
For cellular functions such as division and polarization, protein pattern formation driven by NTPase cycles is a central spatial control strategy. Operating far from equilibrium, no general theory links microscopic reaction networks and parameters to the pattern type and dynamics in these protein systems. Here we discover a generic mechanism giving rise to an effective interfacial tension organizing the macroscopic structure of non-equilibrium steady-state patterns. Namely, maintaining protein-density interfaces by cyclic protein attachment and detachment produces curvature-dependent protein redistribution, which straightens the interface. We develop a non-equilibrium Neumann angle law and Plateau vertex conditions for interface junctions and mesh patterns, thus introducing the concepts of ‘Turing mixtures’ and ‘Turing foams’. In contrast to liquid foams and mixtures, these non-equilibrium patterns can select an intrinsic wavelength by interrupting an equilibrium-like coarsening process. Data from in vitro experiments with the Escherichia coli Min protein system verify the vertex conditions and support the wavelength dynamics. Our study shows how interface laws with correspondence to thermodynamic relations can arise from distinct physical processes in active systems. It allows the design of specific pattern morphologies with potential applications as spatial control strategies in synthetic cells.
Similar content being viewed by others
Main
Heterogeneous systems are governed by interfaces that separate spatially differentiated regions such as grain boundaries in crystals1, membranes2, liquid–liquid boundaries3 and foams4,5,6. Interfaces are also central to biological processes, such as tissue segregation7, morphogenesis8,9 and selection in microbial colonies10,11. Moreover, macroscopic phenomena in key models of active matter are governed by interface tension12,13. Can an analogous concept emerge far from equilibrium without mechanical interactions? How may resulting laws, such as the Gibbs–Thomson and Young–Laplace relations, differ and induce new behaviour?
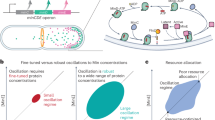
Reaction–diffusion systems describe ensembles of particles that diffuse independently and undergo chemical reactions without mechanical interactions. In biological cells, nucleoside-triphosphatase (NTPase)-driven reaction–diffusion mechanisms form heterogeneous non-equilibrium protein patterns14,15,16,17,18 (Fig. 1a). The diversity of macroscopic, highly nonlinear patterns in these systems—nicely exemplified by the paradigmatic Escherichia coli Min protein system19,20,21,22,23,24—remains largely unexplained.
a, Coupling membrane attachment and detachment driven by NTP hydrolysis with fast cytosolic and slow membrane diffusion creates intracellular reaction–diffusion patterns. b, Patterns of diverse McRD systems are described by the movement of the interface (black line) between domains of high- and low-density or domains dominated by proteins A and B. The simulated membrane density of species A is shown in blue in the symmetric PAR system (f) on the surface of an ellipsoid [(a, b) = (1, 0.6)]. c, The straight pattern interface is sustained by cyclic fluxes of attachment and detachment fa,d(0) in an attachment and detachment zone, respectively, which induce a shallow cytosolic gradient counteracting membrane diffusion. d, In phase-separating liquids, the (exchange) chemical potential at the interface shows a gradient from outwards- to inwards-curved interfaces (orange). e, In McRD systems, an area difference between the attachment and detachment zones arises at curved interfaces. The blue shading symbolizes the membrane-density gradient (left). The (local) reactive turnover balance of the total attachment–detachment fluxes Ja(κ) = Jd(κ) can only be fulfilled if the local attachment–detachment fluxes fa,d(κ) change compared with their values fa,d(0) at a straight interface. The resulting curvature-induced cytosolic density gradients (equations (4) and (5)) induce mass transport from outwards- to inwards-bending regions (orange). f, In the symmetric PAR model, anterior (A, blue) and posterior (B, red) PAR proteins detach each other mutually (see ‘PAR system’ in Methods). g,h, The non-equilibrium Gibbs–Thomson relation equation (5) is verified in numerical simulations of a circular interface (inset) of the symmetric PAR system. The scaling for varying interface curvature \((\delta {\eta }_{{\rm{A}}}^{{\rm{stat}}}-\delta {\eta }_{{\rm{B}}}^{{\rm{stat}}}) \approx \sigma \kappa\) and the approximate scaling \(\sigma \approx \sqrt{{D}_{{\mathrm{m}}}^{{\rm{A}}}}+\sqrt{{D}_{{\mathrm{m}}}^{{\rm{B}}}}\) with the membrane diffusion coefficients are shown (κ ≈ 0.14; Supplementary Sections 3.6.4 and 3.6.5). The model equations and simulation parameters for panels b, g and h are given in ‘PAR system’ in Methods.
The partitioning-defective (PAR) protein system in Caenorhabditis elegans suggests a mechanism generating interface tension in intracellular protein patterns, as it aligns the anterior–posterior PAR interface along the shortest cell circumference25. Although the actomyosin pseudocleavage furrow aids the alignment in wild type, it may be dispensable26. A curvature preference of PAR protein binding has been suggested27, but simulations show that interface-length minimization can occur solely via a reaction–diffusion mechanism28 (Fig. 1b).
Moreover, mathematical models of mass-conserving reaction–diffusion (McRD) systems, which generically describe intracellular protein pattern formation15,16,17,29,30,31, and exhibit sharp interfaces emerging through mechanisms such as wave-pinning15,30, suggest the presence of an effective interfacial tension. For instance, elementary McRD systems minimize the interface length by pattern reorientation32,33 and coarsening29,34,35. Interface-length minimization has been shown mathematically in single-species McRD models36 (T. A. Roth, H. Weyer and E. Frey, manuscript in preparation). However, introducing a third component or coupling to a reservoir can interrupt coarsening35,37,38 and induce interface instabilities39,40,41,42. This raises the central question: Can the self-organization of multi-species and multi-component reaction–diffusion systems be comprehensively understood through the concept of an effective interfacial tension?
Here, by exploring multi-species intracellular protein systems, we discover a generic mechanism for effective interfacial tension in non-equilibrium patterns: cyclic protein attachment and detachment at interfaces—driven by an NTPase cycle. This effective interfacial tension organizes multi-species systems via a non-equilibrium Gibbs–Thomson relation, Neumann law and Plateau vertex conditions, differing systematically from their thermodynamic counterparts. We term the resulting polyhedral patterns ‘Turing mixtures’ and ‘Turing foams’, as they resemble phase-separated liquid mixtures and foams but arise entirely from a reaction–diffusion mechanism. Unlike classical Turing patterns, typically associated with small amplitudes near a supercritical onset, these are highly nonlinear and characterized by sharp interfaces43, which can be understood through a wave-pinning mechanism30 in two-component McRD systems. We demonstrate the Turing foam and its dynamics in experimental data of the in vitro Min protein system20 by comparison with a molecular model based on known biochemistry of the Min system.
Protein reaction–diffusion patterns
Intracellular protein patterns form by the regional accumulation and depletion of proteins on the cell membrane due to their attachment and detachment from and to the cytosol17,44,45 (Fig. 1a,b). We therefore model the dynamics of multiple protein species α = 1, …, Nspecies with one membrane density mα(x, t) and one cytosolic density cα(x, t) each:
Here, x parameterizes the membrane and we neglect the extent of the cytosol perpendicular to the membrane, which can yield intriguing additional effects28,44,46,47. The diffusion coefficients of the membrane-bound and cytosolic components are denoted by \(D_{m,c}^{\alpha}\). The term fα, describing attachment and detachment, typically takes the form fα = aα(mβ)cα − dα(mβ)mα, modelling density-dependent recruitment and enzymatic detachment with rates aα(mβ) and dα(mβ). Importantly, total protein densities ρα = mα + cα are conserved and follow continuity equations
This introduces the ‘mass-redistribution potentials’ \({\eta }_{\alpha }={c}_{\alpha }+{D}_{{\mathrm{m}}}^{\alpha }/{D}_{{\mathrm{c}}}^{\alpha }{m}_{\alpha }\) whose gradients drive the redistribution of the protein mass analogously to chemical-potential gradients in liquid mixtures29,45.
With suitable feedback in membrane accumulation and depletion, these reaction–diffusion systems form patterns with sharp interfaces on the membrane between high- and low-density protein regions, while the amplitude of cytosolic gradients remains shallower due to faster diffusion17,29,30,31,45,48. For a single species, patterns form high- and low-density regions on the membrane while in multi-species systems a different species may be enriched in the low-density region of the first (Fig. 1b). Specifically, the continuity equations, equation (2), ensure uniform mass-redistribution potentials (\({\eta }_{\alpha }={\eta }_{{\rm{stat}}}^{\alpha }\)) in stationary patterns under no-flux conditions, balancing diffusive fluxes between membrane and cytosol (Fig. 1c). The biologically relevant limit \({D}_{{\mathrm{c}}}^{\alpha }\gg {D}_{{\mathrm{m}}}^{\alpha }\) then implies weakly varying cytosolic densities \({c}_{{\rm{stat}}}^{\alpha }={\eta }_{{\rm{stat}}}^{\alpha }-{D}_{{\mathrm{m}}}^{\alpha }/{D}_{{\mathrm{c}}}^{\alpha }\,{m}_{{\rm{stat}}}\)\(\approx {\eta }_{{\rm{stat}}}^{\alpha }\). Thereby, the profile of a weakly curved interface, with a curvature radius κ−1 that is large compared with the interface width ℓint, follows from equation (1a) for \({\partial }_{t}{m}_{\alpha }^{{\rm{stat}}}=0\) as
with d ≡ Dm/Dc and r the coordinate perpendicular to the interface. For simplicity, we consider a single species, dropping the index 1. The balance of membrane diffusion with attachment and detachment fluxes (equation (3)) implies dominant attachment in high-membrane-density regions and dominant detachment in low-density regions17,45 (zones in Fig. 1c). We define the position of the interface, analogous to the Gibbs dividing surface in liquid mixtures49, as the boundary between attachment and detachment zones (Supplementary Section 2). The width of the interface is estimated as \({\ell }_{{\rm{int}}} \approx \sqrt{{D}_{{\rm{m}}}{\tau }_{{\rm{r}}}}\), given a timescale τr of reactive attachment and detachment45. Thus, a fixed pattern interface is sustained as a non-equilibrium steady state through an attachment–diffusion–detachment cycle (Fig. 1c) driven by NTP hydrolysis.
Effective interfacial tension
In phase-separating liquid mixtures, surface tension σ governs interface dynamics: outwardly curved regions (κ > 0) have higher chemical potential δμ ≈ σκ than inwardly curved ones, inducing fluxes that straighten the interface (Gibbs–Thomson relation; Fig. 1d). However, this energetic argument does not apply to reaction–diffusion systems, where particles interact via chemical reactions rather than mechanical potentials.
Instead, for the interface to be stationary, total attachment must balance total detachment (Fig. 1c). For a straight interface, multiplying equation (3) by ∂rmstat and integrating over the interface region, one obtains the total turnover balance \(0=\mathop{\int}\nolimits_{{m}_{-}}^{{m}_{+}}{\rm{d}}m\,f(m,{\eta }_{{\rm{stat}}}(0)-d\,m)\) fixing ηstat(0) at vanishing curvature in terms of the plateau densities m± (ref. 45). For κ ≠ 0, the first term in equation (3) gives a correction to this balance and, thereby, to the mass-redistribution potential. An analogous integration then yields the curvature-induced shifts35 (Supplementary Section 3.4).
where Δρ = ρ+ − ρ− is the density jump across the interface, and \({\bar{f}}_{\eta }\) describes the average change of the reaction term in the interface region with η.
Why does the curvature alter ηstat(κ)? Curving the interface changes the relative area Aa/Ad of the attachment and detachment zones. Outwards (inwards) curvature enlarges (reduces) the detachment zone, enlarging (decreasing) the detachment flux Jd(κ) (Fig. 1e), and thus requires a balancing increase (decrease) in local attachment flux fa(κ) (Fig. 1e). Because attachment increases with cytosolic density (\({\bar{f}}_{\eta } > 0\) required for stability of the interface; Supplementary Section 3.3.2 and ref. 35), the stationary value ηstat(κ) ≈ cstat increases (decreases). The ratio \({\tau }_{{\rm{r}}}{\bar{f}}_{\eta }\) determines how much the cytosolic density (ηstat(κ)) must change to induce the required change of the total turnover.
Finally, the shifts δη(κ) result in protein gradients that, due to the continuity equation, equation (2), induce protein fluxes from outwards- to inwards-curved interface regions, straightening the interface (within a quasi-steady-state approximation). The analogy of equation (4) to the Gibbs–Thomson relation suggests that cyclic attachment and detachment of proteins at the pattern interface induces large-scale dynamics governed by an effective interfacial-tension scaling with the interface width \(\sigma \sim {\ell }_{\rm{int}} \sim \sqrt{D_{\rm{m}}}\). In McRD systems with one species, the interface dynamics has been found in closed form and analysed numerically34,35,36.
Importantly, the geometric area change of the attachment and detachment zones at curved interfaces suggests that the effective interfacial tension extends beyond single-species systems to biologically relevant multi-species systems. Indeed, for the McRD systems, equation (1a,b), with symmetric, second-order mutual detachment (Fig. 1f), which describe, for instance, the PAR polarity system in C. elegans48, we find that the curvature-induced shifts δηstat(κ) of the mass-redistribution potentials fulfil
The matrix Fint of (scaled) attachment rates is diagonal with positive entries for Dc ≫ Dm. The derivation and the relation for general systems equation (1a,b) are given in Supplementary Section 3. For the PAR polarity model with symmetric rates48 and Dc ≫ Dm, the relation equation (5) simplifies to \(\delta {\eta }_{{\rm{A}}}^{{\rm{stat}}}-\delta {\eta }_{{\rm{B}}}^{{\rm{stat}}}\approx -\sigma \kappa /(\Delta {\rho }_{{\rm{A}}}{k}_{{\rm{c}}}^{{\rm{A}}})\), where \({k}_{{\rm{c}}}^{{\rm{A}}}\) denotes the attachment rate, and agrees with numerical simulations (Fig. 1f–h and ‘PAR system’ in Methods). Thus, except for the rescaling of the shifts δηstat by the rate matrix Fint, the non-equilibrium relation equation (5) resembles the thermodynamic Gibbs–Thomson relation Δρ ⋅ δμ = −σκ for multi-component liquid mixtures50. Here, Δρα and δμα represent the (total) density jump across the interface and the chemical potential shift of species α. In contrast to the balance of attachment and detachment in the McRD systems, the thermodynamic relation follows in liquid mixtures from the balance between the Laplace pressure and the sum of the osmotic pressures of all species (Supplementary Section 3.1 and ref. 50).
Thus, in a quasi-steady-state limit, the interface dynamics of multi-species, two-component reaction–diffusion systems closely resembles multi-species Cahn–Hilliard systems50, a mapping that will be exciting to derive mathematically.
Non-equilibrium Neumann law
In multi-species McRD systems, equation (1a,b), multiple pattern domains can emerge (Fig. 2a,b), for instance, due to mutually antagonistic interactions, such as mutual detachment or inhibited attachment at the membrane (Fig. 2e,f). We find a pattern morphology with interfaces and their junctions that resembles multi-component liquid mixtures (Fig. 2b), and we refer to these patterns as Turing mixtures.
a, McRD systems with three or more species can be constructed by introducing pairwise antagonistic interactions resembling the PAR system (d) or inhibited attachment (e). b, Junction angles θA,B,C are constrained by the non-equilibrium Neumann law (equation (6)) for the effective interfacial tensions (times the interface normal vectors) \({\tilde{{\bf{\upsigma }}}}_{{\rm{AB}},{\rm{AC}},{\rm{BC}}}\). c, In steady state, protein attachment (dark shadings) and detachment (in the attachment regions of the opposing species) must balance. In the junction core, all three domains meet. d, The core turnover measures the differences \({\int}_{\varGamma (y)}{\rm{d}}{\bf{\uprho }}\cdot \tilde{{\bf{f}}}={\sum }_{\alpha = 1,2}{\int}_{\tilde{\varGamma }(y)}{\rm{d}}{x}_{\alpha }({\partial }_{{x}_{\alpha }}{\bf{\uprho }})\cdot \tilde{{\bf{f}}}\) in the reactive turnover within the junction (black) core (solid red path between red dots) and integrated around the junction core (dashed red path cutting single interfaces perpendicularly), summing over all parallel paths through the core (parameterized by coordinate y; light red). Here, we set \({\tilde{f}}_{\alpha }=(1-{D}_{{\mathrm{m}}}^{\alpha }/{D}_{{\mathrm{c}}}^{\alpha }){f}_{\alpha }\). The path Γ(y) is the path in the phase space of total concentrations ρA,B,C corresponding to the real-space path \({\tilde{\varGamma}}(y)\). The core turnover vector Tcore (blue) points in the direction in which the integrated differences are largest (Supplementary Section 4). e, The junction angle θA increases as a function of the membrane diffusion coefficient \({D}_{{\rm{m}}}^{{\rm{A}}}\) (keeping \({D}_{{\rm{m}}}^{{\rm{B}}}={D}_{{\rm{m}}}^{{\rm{C}}}\) constant) for a symmetric three-species system with mutual detachment (inset) where all rates are equal; based on the solution of equations (9) and (12) with parameters given in Extended Data Table 1 (circles; ‘Pattern analysis’ in Methods). The non-equilibrium Neumann law including the core turnover Tcore (black line) and for setting Tcore = 0 (dashed line) are shown. The circular inset shows the membrane densities of the steady-state pattern. f, The junction angle θA decreases as a function of the attachment inhibition kBC between species B and C (circles; ‘Pattern analysis’ in Methods) until the BC interface vanishes (red-shaded area) in a three-species system with inhibited attachment (blunt arrows in the inset). The model equations and parameters are given in ‘Inhibited attachment’ in Methods. g,h, Two four-species systems based on attachment inhibition are designed (interaction network shown in f; ‘Inhibited attachment’ in Methods) in which the non-equilibrium analogue of full wetting only allows interfaces to form between the species connected in the contact graphs. The membrane-density patterns are shown at time t = 10,000. The simulation parameters are given in Extended Data Table 1 and we choose kαβ = kβα with k12 = k14 = k23 = k34 = 3, k13 = k24 = 600 in g and k12 = k23 = k24 = k34 = 0.6, k13 = k14 = 2,400, μαβ = 4 in h. The system in g forms stable four-fold junctions as in phase-separating liquids55. The splitting into two three-fold junctions is suppressed because it would create AC or BD interfaces, which were tuned to violate the wetting condition equation (7).
In liquid mixtures, the meeting angles at stationary interface junctions are determined analogously to a force balance condition: the sum of the three surface-tension vectors (surface tensions multiplied by the interface tangent vectors) must vanish, known as the Neumann law5. In contrast, in reaction–diffusion systems, these interface junctions are stationary if the overall attachment–detachment turnover of each protein species is balanced (Fig. 2c). For the McRD systems equation (1a,b), we derive the non-equilibrium Neumann law (Supplementary Section 4.1):
Here, all four terms are vectors with the three interfacial-tension vectors parallel to the interfaces and the core turnover Tcore (Fig. 2b). The mathematical expressions for the effective interfacial tensions \({\widetilde{\sigma }}_{ij}\) between domains i, j = A, B, C differ quantitatively from those in the non-equilibrium Gibbs–Thomson relation but also scale with the interface widths \({\ell }_{{\rm{int}}}^{ij}\) (and typical reaction rates \({({\tau }_{{\rm{r}}}^{ij})}^{-1}\); Supplementary Sections 3.6.5 and 4). Several different non-equilibrium interfacial tensions were also found in scalar active matter13,51,52.
The turnover (attachment–detachment) balance leading to equation (6) replaces the osmotic-pressure balance that must be fulfilled in liquid mixtures. In equilibrium systems, the osmotic-pressure construction aligns with the effective surface-tension force balance50 (Supplementary Section 4.2.1), but this does not hold for non-equilibrium reaction–diffusion systems. The reactive turnover in the junction core—where all three domains meet—is qualitatively different from the reactive flows at the interfaces, at each of which only two domains meet. Thus, the balance of attachment and detachment for all protein species at the junction not only depends on the turnover balance at the individual interfaces, represented by the effective interfacial tensions, but also the additional contribution from the core. The source term Tcore describes this additional contribution of the junction core to the reactive turnover (projected onto the junction’s translation mode) compared with the integrated turnover on a contour around the junction crossing only individual interfaces (Fig. 2d and Supplementary Section 4.1). The core turnover vector Tcore points in the direction in which the additional contributions integrated over the junction are largest, and its amplitude is set by their integrated strength. Thus, the core turnover depends on the density profiles in the junction core, and thereby on the diffusion coefficients and reaction rates. A description of the density profiles is required to explicitly determine this dependence, as, for instance, the interface profiles are required to determine the surface tensions even in equilibrium systems. The non-equilibrium Neumann law equation (6) ensures that including this additional turnover contribution Tcore from the core total attachment and detachment is balanced, resulting in a stationary junction. Similarly, non-equilibrium currents modify the Young–Dupré equation in motility-induced phase separation and active–passive interfaces in microtubule–motor mixtures53,54.
Figure 2e,f shows how the non-equilibrium Neumann law prescribes the junction angles in two different three-species systems. The junction angle θA of species A increases as the effective interfacial tensions \({\widetilde{\sigma }}_{{\rm{AB,AC}}}\) rise with \({D}_{{\mathrm{m}}}^{{\rm{A}}}\), as expected from their dependence on ℓint. In contrast, increasing the rate of mutual antagonism between species B and C (‘Inhibited attachment’ in Methods) enlarges \({\widetilde{\sigma }}_{{\rm{BC}}}\), as expected from its dependence on \({({\tau }_{{\rm{r}}}^{{\rm{BC}}})}^{-1}\), reducing the angle θA until the BC interface vanishes when αA = 0. This represents a non-equilibrium version of the transition to full wetting in phase separation. Full wetting occurs, akin to thermal-equilibrium systems5,55, if
The core turnover Tcore does not contribute because, at the transition, the junction core vanishes and thus also its contribution to the non-equilibrium Neumann law (Fig. 2f and Supplementary Section 4.2.3). Choosing the effective interfacial tensions relative to each other to fulfil or not the bound \({\tilde{\sigma }}_{{\rm{BC}}} < {\tilde{\sigma }}_{{\rm{AB}}}+{\tilde{\sigma }}_{{\rm{AC}}}\) determines which species contact each other55. Thus, a central finding of our theory is that it enables the design of specific patterns by linking the microscopic parameters, reaction rates and diffusion coefficients, to the macroscopic pattern morphology (Fig. 2g,h). Note that the quantitative parameter dependence of the effective interfacial tensions beyond the scaling \({\widetilde{\sigma }}_{ij} \sim {\ell }_{{\rm{int}}}^{ij}/{\tau }_{{\rm{r}}}^{ij}\) is difficult to obtain because they depend on the interface profile. Also in liquid mixtures, exact relations only exist in special cases.
Turing foam in the in vitro Min protein system
Do these interface laws apply to other biochemical systems beyond the PAR system? The E. coli Min protein system is well studied in vitro19,20,21,23. Patterns emerge from the interaction of two proteins: MinD, an ATPase that attaches to the lipid membrane in its ATP-bound state; and MinE, the ATPase-activating protein recruited by MinD onto the membrane17,56,57. Using MinE proteins with a C-terminal His tag or wild-type MinE, the Min system forms (quasi-)stationary mesh (Fig. 3a) and labyrinthine patterns for most protein concentrations20. Currently, we lack both a model that robustly reproduces these network patterns and a comprehensive theoretical framework that elucidates their macroscopic dynamics and morphology. While the proposed in vitro Min model (Fig. 3g) is more complex than equation (1a,b), we expect that the geometric mechanism inducing the effective interfacial tension (Fig. 1e) generalizes beyond the concrete mathematical model equation (1a,b) (Supplementary Section 2). It remains an open problem to derive mathematically under which assumptions a reaction–diffusion system is governed by positive effective interfacial tensions. Our interface theory then predicts a distinct foam-like morphology for the mesh patterns, which we refer to as Turing foams.
a, In vitro, the E. coli Min system can form mesh patterns made up of thin MinE branches (cyan) separating MinD domains (magenta). Total concentrations: [MinD] = 1.5 μM and [MinE-His] = 3 μM (compare Extended Data Fig. 1; experimental data obtained and described in ref. 20). Branches of triple vertices are marked in white and four-fold vertices are labelled with black circles (‘Pattern analysis’ in Methods). b,c, The instability of four-fold and higher-order vertices (b) and branch rearrangement at triple vertices towards 120° junction angles (c) can be explained based on the curvature dependence of net attachment (blue) and detachment (red). d–f, Histograms of the angles at the triple vertices (d), the number of edges of each MinD domain (e) and the MinD domain areas (f) are determined from the experimental data in a (Extended Data Fig. 1a and see ‘Pattern analysis’ in Methods). For comparison, the vertex-angle histogram of vertices with random angles is shown (black line; Supplementary Section 5). The largest MinD domains have an internal MinE domain disconnected from the mesh (green). g, Reaction network of a minimal model of MinD (magenta) and MinE (cyan) membrane attachment and detachment supplemented by a conformational MinE switch and persistent membrane binding of MinE. h, This model reproduces stationary mesh patterns of MinD (magenta) and MinE (cyan) in two-dimensional simulations (model equations and parameters given in ‘Min system’ in Methods). i,j, Histograms of the vertex angles (i) and the edge number of the MinD domains (j) are determined from six independent simulations (‘Pattern analysis’ in Methods).
The mesh pattern is formed by thin MinE branches separating extended MinD domains and meeting at vertices (Fig. 3a; further data analysed in Extended Data Fig. 1). Each branch has two MinD–MinE interfaces. Assuming branch stability, our theory predicts that only triple vertices with three branches are stable (Fig. 3b,c): deviations from the symmetric configuration of equal vertex angles change the curvatures of the interfaces bounding the MinD domains meeting at the vertex. According to the Gibbs–Thomson relation and its heuristic explanation (‘Effective interfacial tension’ section), increased curvature at sharper angles leads to enhanced MinD detachment, while decreased curvature at shallower angles favours attachment. This imbalance causes the interface segments at shallow angles to move inwards until the stable branch width is restored, while those at sharp angles retract. As a result, higher-order vertices become unstable and break up into multiple triple vertices (Fig. 3b) while asymmetric triple vertices evolve towards symmetric 120° vertex angles (Fig. 3c). These vertex conditions, which are referred to as Plateau vertex laws in two-dimensional liquid foams4, are our predictions for the structure of the Min mesh pattern. It is an exciting future challenge to derive the microscopic mechanism—the ‘surfactant’—that stabilizes the MinE branches in the Min system despite the effective interfacial tension, allowing the formation of the Turing foam instead of droplet patterns. This understanding will guide a mechanistic classification of the conditions under which Turing foams can form in reaction–diffusion systems.
Reanalysing the experimental data in ref. 20, we find that the Min mesh pattern shows about 94% triple junctions (Fig. 3a) whose vertex angles are distributed with a small standard deviation of about 11.4° around 120° (Fig. 3d; further data analysed in Extended Data Fig. 1). The mean number of domain edges is 5.7 ± 1.1 (Fig. 3e). In contrast to equilibrium foams, the largest MinD domains show disconnected, internal MinE branches (Fig. 3f), which we discuss in the next section.
Using an extended model based on biochemically suggested interactions of the Min proteins (Fig. 3g and ‘Min system’ in Methods), we are able to reproduce the statistics of Turing foams in numerical simulations (Fig. 3h–j). Moreover, during the development of the pattern, four-fold vertices decay into pairs of triple vertices (Supplementary Video 1) as predicted.
Von Neumann’s law and domain splitting
Two-dimensional liquid foams follow von Neumann’s law: bubbles with fewer than six edges shrink and collapse, while those with more than six edges grow4,58. In contrast, the Min patterns do not coarsen continuously but larger MinD domains contain internal MinE branches (Fig. 3a,f and Extended Data Fig. 1). This implies a mechanism that limits the maximum size of stationary domains beyond which nucleation of internal MinE branches leads to the division of these MinD domains.
In simulations, the mesh pattern undergoes coarsening that follows von Neumann’s law at domain sizes that are small compared with the final stationary pattern (Fig. 4a). This behaviour is explained by branch dynamics driven by the effective interfacial tension (Fig. 4b,e,f and Supplementary Section 6). However, this process is interrupted when the original domain size is reached (Fig. 4c,d), and reversed at larger scales by the splitting of domains due to the growth of new branches (Fig. 4g–i). Both numerically predicted dynamic behaviours of domain collapse and splitting are observed in the time course of the in vitro Min mesh pattern (Extended Data Fig. 2 and Supplementary Video 4).
a, The dynamics of a stationary mesh pattern is shown that is reduced in size by rescaling the simulation-domain length L to \(L/\sqrt{2}\) (simulation parameters given in Extended Data Table 2; MinE membrane density shown in cyan and MinD in magenta; Supplementary Video 2). White pentagons label pattern domains with five edges (considering the reflective boundary conditions to label boundary domains). b, The trajectories of the areas A of one-half of the domains with n = 5, 6, 7 (blue, black, red) in 6 independent simulations are shown. c,d,f, The statistics from these six independent simulations are aggregated to calculate the distribution of initial (c) and final (d) domain areas, as well as the area change A(t) − A(0) compared with the initial domain area A(0) independently averaged over the domains with n = 5, 6, 7 edges (f, shaded regions depict the standard deviation around the mean). e, The curvature-induced branch movement with a normal velocity vκ is caused by the curvature-induced transport of MinD (red shaded) through the MinE branch (blue shaded) from the inside to the outside (black double arrow; compare Fig. 1e). Sections of the MinD attachment and detachment zones are sketched as in Fig. 1e. g, The dynamics of a stationary mesh pattern on an enlarged domain \(L\to \sqrt{2}L\) is shown (Supplementary Video 3). Domains split by the growth of an internal MinE branch (domain i) or by the growth of a branch out of an existing branch (domain iii). Small domains do not split (domain ii). The splitting domains i and iii are enlarged below. h, One-half of the domain-area trajectories from six independent simulations are shown. The trajectories of the domains i, ii and iii are labelled. i, For domains with a given maximal domain area over time, the fraction that splits is shown (black circles). It is calculated from the histogram of the maximal areas of all domains and those that eventually split in the six simulations. The threshold of domain splitting is approximately given by the area at which a single circular domain forms an internal MinE domain (dotted line in h and i; ‘Numerical simulation’ in Methods).
This numerical study demonstrates that the Turing foam dynamics closely resembles the wavelength dynamics in two-component reaction–diffusion systems previously explained by us35,38. Motivated by these results, an empirical model balancing interfacial-tension-driven coarsening with domain-area-dependent terms may allow for the description of the statistics of interrupted coarsening and domain splitting. Nonetheless, identifying the mechanisms that counteract interfacial-tension-driven coarsening in the Min system—resulting in its arrest and domain splitting—will be required to predict the selected wavelength, which thus remains a key challenge.
Discussion
Both the instability of four-fold vertices and 120° vertex angles have been observed in experiments and simulations based on (non-mass-conserving) pH-feedback reactions59,60 related to the classical ferrocyanide–iodate–sulfite (FIS) reaction61,62. The FIS reaction also shows the Turing-foam dynamics in simulations (Supplementary Video 5 and ‘FIS system’ in Methods). Notably, these systems are not mass-conserving, suggesting a broader relevance of Turing foams beyond strict mass conservation. In addition, one-component reaction–diffusion models describing bistable media are mathematically equivalent to Model A (Allen–Cahn) dynamics, allowing the definition of an interfacial tension, for example, for competing microbial populations11,63,64.
Further experimental investigations should probe patterns on complex geometries, for example, on structured membranes or vesicles. While liquid foams are always unstable with respect to coarsening on surfaces with positive Gaussian curvature65, simulations suggest that polyhedral domain arrangements on spheres are stabilized by interrupted coarsening in Turing foams (Extended Data Fig. 3).
Foam-like structures are observed as well in scalar13, polar66 and nematic active matter67, and in experimental active matter systems68. In scalar active matter, several distinct interfacial tensions arise even in single-species systems due to distinct Gibbs–Thomson relations for the high- and low-density sides of interfaces13,51. In contrast, the non-equilibrium Gibbs–Thomson relation is unique for each interface in McRD systems. However, higher-order relations such as the non-equilibrium Neumann law introduce quantitatively different interfacial tensions in multi-species systems. By revealing how far-from-equilibrium microscopic processes can yield complex equilibrium-like interface dynamics, our findings offer an approach to mechanistically understand the pattern dynamics and macroscopic morphologies in non-equilibrium systems more broadly.
Methods
Reaction–diffusion dynamics
In reaction–diffusion systems, n different components ui, i = 1, ..., n, spread diffusively with diffusion constants Di (assuming the absence of density dependence and cross diffusion) and undergo (nonlinear) reactions fi(u). The components’ evolution with time t is described by coupled, nonlinear partial differential equations
Here, the diffusion matrix is D = diag(D1, …, Dn), and we define \({\bf{u}}={({u}_{i})}_{i = 1,\ldots ,n}\) and \({\bf{f}}={({f}_{i})}_{i = 1,\ldots ,n}\).
In McRD systems, the components constitute different (conformational) states or complexes of N different types of molecule (species) whose total number is conserved. Thus, the reactions solely describe the conversion of the molecules between the different components, for example, membrane attachment and detachment (examples in the following subsections). We refer to N as the number of species while n is the number of components of the system. The mass-conservation constraint is formalized by introducing the stoichiometric vectors sα for the molecular species α = 1, …, N (or A, B, …), which count how many of those molecules are contained in the different components69. Consequently, the total density ρα(x, t) of the molecular species α reads
For example, a system with components A, AB and B2 would have the stoichiometric vectors sA = (1, 1, 0) and sB = (0, 1, 2). Conservation of the total molecule numbers (‘total mass’) implies that the evolution of the total densities ρα(x, t) follow the continuity equations29,45,70
Note, that the Greek index is not summed over. The constraint
describes that molecules leaving one component are added in a different component such that the individual terms sαi fi cancel. As a result, the average total densities
are set by the initial conditions on domains \(\varOmega \subset {{\mathbb{R}}}^{d}\) (volume ∣Ω∣) with reflective or periodic boundary conditions.
Moreover, equation (8) defines the mass-redistribution potentials ηα(x, t) (α = 1, …, N) as a sum of the different molecular states weighted by their diffusion coefficients29,44,45. The gradients of the mass-redistribution potential(s) determine the redistribution of the total densities analogously to the chemical potential(s) in Model B dynamics although they are not given as derivatives of a free energy71,72. The mass-redistribution potentials are normalized by one diffusion coefficient \({D}_{{\mathrm{c}}}^{\alpha }\), for example, the largest one whose component contributes most strongly to the redistribution of the total density. For intracellular protein patterns, this is the cytosolic diffusion coefficient (assuming different cytosolic states of the same molecule diffuse similarly quickly).
Multi-species McRD systems with two components per species
Consider first, as introduced in equation (1a,b), McRD systems where each molecular species α has only two states mα and cα (for example, cytosolic and membrane-bound states) with diffusion coefficients \({D}_{{\mathrm{m}}}^{\alpha }\) and \({D}_{c}^{\alpha } > {D}_{m}^{\alpha }\), respectively. With reaction terms fα(m, c) for each species, the reaction–diffusion equations for species α read
The total densities ρα = mα + cα follow the continuity equations
with the mass-redistribution potential
where \({d}_{\alpha }={D}_{{\mathrm{m}}}^{\alpha }/{D}_{{\mathrm{c}}}^{\alpha } < 1\) defines the ratio of the diffusion constants. In the intracellular context, one has dα ≪ 1 because diffusion on the membrane is much slower than diffusion in the cytosol.
The mass-redistribution potential satisfies
with
PAR system
The C. elegans PAR polarity system is modelled by McRD systems describing a membrane-bound and a cytosolic component (mα, cα) for the anterior PAR (aPAR) and posterior PAR (pPAR) protein species labelled by α = A, B, respectively48,73. The reaction term includes linear attachment and detachment at the cell membrane as well as mutual detachment:
Commonly, the parameters are chosen symmetrically, that is, \({k}_{{\rm{c}}}^{{\rm{A}}}={k}_{{\rm{c}}}^{{\rm{B}}}\), \({k}_{{\rm{m}}}^{{\rm{A}}}={k}_{{\rm{m}}}^{{\rm{B}}}\) and kAB = kBA, with the exponents for the mutual detachment term taken as μ = ν = 2 (ref. 48). In what we call the ‘symmetric PAR model’, we maintain this symmetry by choosing identical diffusion coefficients on the membrane (\({D}_{{\rm{m}}}^{{\rm{A}}}={D}_{{\rm{m}}}^{{\rm{B}}}\)) and in the cytosol (\({D}_{{\rm{c}}}^{{\rm{A}}}={D}_{{\rm{c}}}^{{\rm{B}}}\)). The time and length scales are fixed by setting the attachment rate and cytosolic diffusion coefficient to 1. The standard parameter values are given in Extended Data Table 1.
We extend the PAR system to the interaction of N > 2 different molecule species by introducing the mutual detachment between any pair of species. In this case, the reaction terms read
In the simulations we chose the exponents μαβ to be the same for all pairs of species.
Inhibited attachment
Another scenario of mutual antagonism arises when molecular species hinder each other’s attachment to the membrane. We consider the reaction terms
which describe the attachment inhibition by a Hill term. The strength of inhibition is set by the parameters kαβ and the Hill exponents are μαβ. In the simulations, we employed the parameter values given in Extended Data Table 1 for all pairs of species. Parameters modified in individual simulations are specified in the respective figure captions.
Min system
Pattern formation in the Min system has been studied based on reaction–diffusion models using various biochemical networks and rate constants23,74,75,76,77. For our analysis, we started with a core reaction network comprising biochemically validated or plausible reactions75,76, complemented by a molecular MinE switch78, which has been shown to be important in both in vivo22 and in vitro77 settings. However, our numerical analysis of this core reaction network revealed that achieving stationary Turing foams requires, in addition, persistent membrane binding of MinE, as previously hypothesized19,77,79,80 (see full reaction network in Fig. 3g). A parameter study and comparison of the experimental and simulated phase diagrams will be published elsewhere. For simplicity, we simulated the patterns on a two-dimensional membrane coupled to a two-dimensional cytosolic layer. Thus, the model disregards any effects due to a (non-uniform) bulk height18,28,44.
In the cytosol, the model includes ADP-bound MinD cDD and ATP-bound MinD cDT states, as well as the reactive and latent MinE states cE,r, cE,l. On the membrane, the model includes membrane-bound MinD md, MinD–MinE complexes mde, and membrane-bound MinE me. The stoichiometric vectors sD,E for MinD and MinE and the components u = (cDD, cDT, cE,r, cE,l, md, mde, me) are
The average total densities \({\bar{\rho }}_{{\rm{D}}},{\bar{\rho }}_{{\rm{E}}}\) of MinD and MinE added to the system are control parameters in the simulation and experiment. The reaction terms read
Extended Data Table 2 explains and gives the chosen values of the rates, diffusion coefficients and average densities.
FIS system
We show in simulations that Turing foams also arise in the classical FIS reaction61,62. In the simulation, the four-species Gáspár–Showalter model with the parameters employed in ref. 62, Fig. 17, was used. This model includes the four species x, y, z and a with diffusion coefficients Dx > Dy = Dz = Da. The parameters are given in Extended Data Table 3. This model is not mass-conserving because the reactants are supplied with a rate k0, modelling the exchange between the gel matrix and a reservoir. The reaction terms are62
Numerical simulation
We performed simulations of the different models on a two-dimensional domain with no-flux boundary conditions using COMSOL Multiphysics 6.1. For Extended Data Fig. 3, the simulation was performed on a spherical, closed surface. These simulations employ a finite-element discretization on a triangular mesh with linear Lagrange elements.
To estimate the threshold size of splitting in the simulated Min protein Turing foam (dashed line in Fig. 4h,i), a single, circular MinD domain surrounded by a MinE branch at the boundary of the simulation domain with a radius of R0 = 5 μm was simulated for 105 s. Instead of adiabatically increasing the radius of the simulation domain (and thereby the radius of the MinD domain), and observing the threshold size at which an internal MinE domain grows, all diffusion coefficients were adiabatically decreased with time t by the common factor f = 1 + t/105 s. Thereby, the length unit was rescaled, resulting in an increasing radius R(t) = R0 f−1/2. The pattern was recorded every 500 s, and the size was determined when an internal MinE domain of comparable density to the surrounding MinE branch had formed.
The parameter dependence of the non-equilibrium Gibbs–Thomson relation of a radially symmetric interface in the symmetric PAR system in Fig. 1g,h was determined using numerical pseudo-arclength continuation implemented in Mathematica previously35,45: Assuming radial symmetry, the stationary reaction–diffusion equations were converted into an algebraic system of equations by a finite-differences discretization in the radial direction. The algebraic steady-state equations are solved by Newton iteration.
The configuration files for the COMSOL simulations and the Mathematica notebooks are available at https://github.com/henrikweyer/Turing-foams (ref. 81).
Pattern analysis
The statistical analysis of the simulation and experimental data was performed using Mathematica 13.1 and is explained in detail in the Supplementary Sections 6 and 7. To measure the junctions and vertices in simulation and experimental images, the skeleton network of the interfaces or branches was determined using Mathematica’s morphological algorithms. Its branch points give rough junction or vertex positions. In a second step, a junction or vertex model was fitted to the density fields and their gradients to determine precise junction and vertex angles. The domain areas were determined from the skeleton network.
To compare these measurements of the junction angles with the non-equilibrium Neumann relation, the effective interfacial tensions are measured at the interfaces far from the junction (in the same simulation). In addition, the core turnover is determined as integral over the junction core (Supplementary Section 7.2). Importantly, one has to correct for the interface curvature, which shifts the turnover balance and results in a non-zero contribution compared with straight interfaces. To this end, the curvature-induced contribution is determined by integrating the core turnover expression at the interfaces far away from the junction. This contribution is subtracted from the integral over the junction core, assuming that the interfaces are circular arcs up to their meeting point.
The Mathematica notebooks are available at https://github.com/henrikweyer/Turing-foams (ref. 81).
Effective interfacial tension and non-equilibrium Neumann law
The derivation of the non-equilibrium Gibbs–Thomson relation equation (4) for the single-species system is elaborated in Supplementary Section 3.4. For multi-component systems, in Supplementary Section 2, we show that a shift δηstat ≈ ℓintκ arises under conditions on the monotonicity of the component densities at the interface if the slow-diffusing components induce the pattern-forming feedback, that is, they increase their production (attachment) at high density and lead to net detachment at low densities while the fast-diffusing components only enter linearly in their attachment terms. In Supplementary Section 3, we then derive explicit expressions for the effective interfacial tension for multi-species systems with two components per protein species. In Supplementary Section 4, the non-equilibrium Neumann law is derived using methods developed for multi-component Cahn–Hilliard systems50,72,82,83,84,85.
Data availability
Data generated in this study are available from the corresponding author upon reasonable request. The experimental data analysed in this study were obtained from the authors of ref. 20.
Code availability
The analysis code and exemplary set-up codes to perform the numeric simulations of this study are available at https://github.com/henrikweyer/Turing-foams (ref. 81).
References
Godrèche, C. Solids Far from Equilibrium Aléa-Saclay No. 1 (Cambridge Univ. Press, 1992).
Safran, S. A. Statistical Thermodynamics of Surfaces, Interfaces, and Membranes 1st edn (CRC Press, 2018).
Mao, S., Kuldinow, D., Haataja, M. P. & Košmrlj, A. Phase behavior and morphology of multicomponent liquid mixtures. Soft Matter 15, 1297–1311 (2019).
Weaire, D. & Hutzler, S. The Physics of Foams (Oxford Univ. Press, 2001).
De Gennes, P.-G., Brochard-Wyart, F. & Quéré, D. Capillarity and Wetting Phenomena (Springer, 2004).
Cantat, I. et al. (eds) Foams: Structure and Dynamics (Oxford Univ. Press, 2013).
Foty, R. A. & Steinberg, M. S. The differential adhesion hypothesis: a direct evaluation. Dev. Biol. 278, 255–263 (2005).
Noll, N., Mani, M., Heemskerk, I., Streichan, S. J. & Shraiman, B. I. Active tension network model suggests an exotic mechanical state realized in epithelial tissues. Nat. Phys. 13, 1221–1228 (2017).
Fernández, P. A. et al. Surface-tension-induced budding drives alveologenesis in human mammary gland organoids. Nat. Phys. 17, 1130–1136 (2021).
Kayser, J., Schreck, C. F., Gralka, M., Fusco, D. & Hallatschek, O. Collective motion conceals fitness differences in crowded cellular populations. Nat. Ecol. Evol. 3, 125–134 (2018).
Giometto, A., Nelson, D. R. & Murray, A. W. Antagonism between killer yeast strains as an experimental model for biological nucleation dynamics. Elife 10, e62932 (2021).
Cates, M. E. & Tailleur, J. Motility-induced phase separation. Annu. Rev. Condens. Matter Phys. 6, 219–244 (2015).
Fausti, G., Tjhung, E., Cates, M. E. & Nardini, C. Capillary interfacial tension in active phase separation. Phys. Rev. Lett. 127, 068001 (2021).
Loose, M., Kruse, K. & Schwille, P. Protein self-organization: lessons from the Min system. Annu. Rev. Biophys. 40, 315–336 (2011).
Edelstein-Keshet, L., Holmes, W. R., Zajac, M. & Dutot, M. From simple to detailed models for cell polarization. Phil. Trans. R. Soc. B 368, 20130003 (2013).
Gross, P. et al. Guiding self-organized pattern formation in cell polarity establishment. Nat. Phys. 15, 293–300 (2019).
Halatek, J., Brauns, F. & Frey, E. Self-organization principles of intracellular pattern formation. Phil. Trans. R. Soc. B 373, 20170107 (2018).
Burkart, T., Wigbers, M. C., Würthner, L. & Frey, E. Control of protein-based pattern formation via guiding cues. Nat. Rev. Phys. 4, 511–527 (2022).
Vecchiarelli, A. G. et al. Membrane-bound MinDE complex acts as a toggle switch that drives Min oscillation coupled to cytoplasmic depletion of MinD. Proc. Natl Acad. Sci. USA 113, E1479 (2016).
Glock, P. et al. Stationary patterns in a two-protein reaction-diffusion system. ACS Synth. Biol. https://doi.org/10.1021/acssynbio.8b00415 (2019).
Brauns, F. et al. Bulk-surface coupling identifies the mechanistic connection between Min-protein patterns in vivo and in vitro. Nat. Commun. 12, 3312 (2021).
Ren, Z. et al. Robust and resource-optimal dynamic pattern formation of Min proteins in vivo. Nat. Phys. https://doi.org/10.1038/s41567-025-02878-w (2025).
Loose, M., Fischer-Friedrich, E., Ries, J., Kruse, K. & Schwille, P. Spatial regulators for bacterial cell division self-organize into surface waves in vitro. Science 320, 789–792 (2008).
Wu, F. et al. Multistability and dynamic transitions of intracellular Min protein patterns. Mol. Syst. Biol. https://doi.org/10.15252/msb.20156724 (2016).
Mittasch, M. et al. Non-invasive perturbations of intracellular flow reveal physical principles of cell organization. Nat. Cell Biol. 20, 344–351 (2018).
Bhatnagar, A. et al. Axis convergence in C. elegans embryos. Curr. Biol. https://doi.org/10.1016/j.cub.2023.10.050 (2023).
Klinkert, K. et al. Aurora A depletion reveals centrosome-independent polarization mechanism in Caenorhabditis elegans. Elife 8, e44552 (2019).
Geßele, R., Halatek, J., Würthner, L. & Frey, E. Geometric cues stabilise long-axis polarisation of PAR protein patterns in C. elegans. Nat. Commun. 11, 539 (2020).
Otsuji, M. et al. A mass conserved reaction–diffusion system captures properties of cell polarity. PLoS Comput. Biol. 3, e108 (2007).
Mori, Y., Jilkine, A. & Edelstein-Keshet, L. Wave-pinning and cell polarity from a bistable reaction–diffusion system. Biophys. J. 94, 3684–3697 (2008).
Frey, E. & Brauns, F. in Active Matter and Nonequilibrium Statistical Physics 1st edn (eds Tailleur, J. et al.) 347–445 (Oxford Univ. Press, 2022).
Singh, A. R., Leadbetter, T. & Camley, B. A. Sensing the shape of a cell with reaction diffusion and energy minimization. Proc. Natl Acad. Sci. USA 119, e2121302119 (2022).
Marée, A. F. M., Grieneisen, V. A. & Edelstein-Keshet, L. How cells integrate complex stimuli: the effect of feedback from phosphoinositides and cell shape on cell polarization and motility. PLoS Comput. Biol. 8, e1002402 (2012).
Tateno, M. & Ishihara, S. Interfacial-curvature-driven coarsening in mass-conserved reaction–diffusion systems. Phys. Rev. Res. 3, 023198 (2021).
Brauns, F., Weyer, H., Halatek, J., Yoon, J. & Frey, E. Wavelength selection by interrupted coarsening in reaction–diffusion systems. Phys. Rev. Lett. 126, 104101 (2021).
Miller, P. W., Fortunato, D., Novaga, M., Shvartsman, S. Y. & Muratov, C. B. Generation and motion of interfaces in a mass-conserving reaction–diffusion system. SIAM J. Appl. Dyn. Syst. 22, 2408–2431 (2023).
Chiou, J.-G., Moran, K. D. & Lew, D. J. How cells determine the number of polarity sites. Elife 10, e58768 (2021).
Weyer, H., Brauns, F. & Frey, E. Coarsening and wavelength selection far from equilibrium: a unifying framework based on singular perturbation theory. Phys. Rev. E 108, 064202 (2023).
Ohta, T., Mimura, M. & Kobayashi, R. Higher-dimensional localized patterns in excitable media. Physica D 34, 115–144 (1989).
Petrich, D. M. & Goldstein, R. E. Nonlocal contour dynamics model for chemical front motion. Phys. Rev. Lett. 72, 1120–1123 (1994).
Muratov, C. B. & Osipov, V. V. Scenarios of domain pattern formation in a reaction–diffusion system. Phys. Rev. E 54, 4860–4879 (1996).
Carter, P., Doelman, A., Lilly, K., Obermayer, E. & Rao, S. Criteria for the (in)stability of planar interfaces in singularly perturbed 2-component reaction–diffusion equations. Physica D 444, 133596 (2023).
Veerman, F., Mercker, M. & Marciniak-Czochra, A. Beyond Turing: far-from-equilibrium patterns and mechano-chemical feedback. Phil. Trans. R. Soc. A 379, 20200278 (2021).
Halatek, J. & Frey, E. Rethinking pattern formation in reaction–diffusion systems. Nat. Phys. 14, 507–514 (2018).
Brauns, F., Halatek, J. & Frey, E. Phase-space geometry of mass-conserving reaction–diffusion dynamics. Phys. Rev. X 10, 041036 (2020).
Thalmeier, D., Halatek, J. & Frey, E. Geometry-induced protein pattern formation. Proc. Natl Acad. Sci. USA 113, 548–553 (2016).
Villar-Sepúlveda, E., Champneys, A. R., Cusseddu, D. & Madzvamuse, A. Pattern formation of bulk-surface reaction–diffusion systems in a ball. Preprint at https://arxiv.org/abs/2409.06826 (2024).
Trong, P. K., Nicola, E. M., Goehring, N. W., Kumar, K. V. & Grill, S. W. Parameter-space topology of models for cell polarity. N. J. Phys. 16, 065009 (2014).
Rowlinson, J. S. & Widom, B. Molecular Theory of Capillarity (Dover Publications, 2002).
Bronsard, L., Garcke, H. & Stoth, B. A multi-phase Mullins–Sekerka system: matched asymptotic expansions and an implicit time discretisation for the geometric evolution problem. Proc. R. Soc. Edinb. A 128, 481–506 (1998).
Langford, L. & Omar, A. K. Theory of capillary tension and interfacial dynamics of motility-induced phases. Phys. Rev. E 110, 054604 (2024).
Langford, L. & Omar, A. K. The mechanics of nucleation and growth and the surface tensions of active matter. J. Chem. Phys. https://doi.org/10.1063/5.0263060 (2025).
Adkins, R. et al. Dynamics of active liquid interfaces. Science 377, 768–772 (2022).
Zhao, Y. et al. Active Young–Dupré equation: how self-organized currents stabilize partial wetting. Preprint at https://arxiv.org/abs/2405.20651 (2024).
Mao, S., Chakraverti-Wuerthwein, M. S., Gaudio, H. & Košmrlj, A. Designing the morphology of separated phases in multicomponent liquid mixtures. Phys. Rev. Lett. 125, 218003 (2020).
Lutkenhaus, J. Assembly dynamics of the bacterial MinCDE system and spatial regulation of the Z ring. Annu. Rev. Biochem. 76, 539–562 (2007).
Ramm, B., Heermann, T. & Schwille, P. The E. coli MinCDE system in the regulation of protein patterns and gradients. Cell. Mol. Life Sci. 76, 4245–4273 (2019).
von Neumann, J. in Metal Interfaces 108–110 (American Society for Metals, 1952).
Goldstein, R. E., Muraki, D. J. & Petrich, D. M. Interface proliferation and the growth of labyrinths in a reaction–diffusion system. Phys. Rev. E 53, 3933–3957 (1996).
Szalai, I., Cuiñas, D., Takács, N., Horváth, J. & De Kepper, P. Chemical morphogenesis: recent experimental advances in reaction–diffusion system design and control. Interface Focus 2, 417–432 (2012).
Lee, K. J., McCormick, W. D., Ouyang, Q. & Swinney, H. L. Pattern formation by interacting chemical fronts. Science 261, 192–194 (1993).
Lee, K. J. & Swinney, H. L. Lamellar structures and self-replicating spots in a reaction–diffusion system. Phys. Rev. E 51, 1899–1915 (1995).
Schlögl, F. Chemical reaction models for non-equilibrium phase transitions. Z. Phys. 253, 147–161 (1972).
Lavrentovich, M. O. & Nelson, D. R. Nucleation of antagonistic organisms and cellular competitions on curved, inflating substrates. Phys. Rev. E 100, 042406 (2019).
Avron, J. E. & Levine, D. Geometry and foams: 2D dynamics and 3D statics. Phys. Rev. Lett. 69, 208–211 (1992).
De Luca, F., Maryshev, I. & Frey, E. Supramolecular assemblies in active motor-filament systems: micelles, bilayers, and foams. Phys. Rev. X 14, 031031 (2024).
Maryshev, I., Morozov, A., Goryachev, A. B. & Marenduzzo, D. Pattern formation in active model C with anchoring: bands, aster networks, and foams. Soft Matter 16, 8775–8781 (2020).
Lemma, B., Mitchell, N. P., Subramanian, R., Needleman, D. J. & Dogic, Z. Active microphase separation in mixtures of microtubules and tip-accumulating molecular motors. Phys. Rev. X 12, 031006 (2022).
Brauns, F., Halatek, J. & Frey, E. Diffusive coupling of two well-mixed compartments elucidates elementary principles of protein-based pattern formation. Phys. Rev. Res. 3, 013258 (2021).
Ishihara, S., Otsuji, M. & Mochizuki, A. Transient and steady state of mass-conserved reaction-diffusion systems. Phys. Rev. E 75, 015203 (2007).
Bray, A. J. Theory of phase-ordering kinetics. Adv. Phys. 51, 481–587 (2002).
Elliott, C. M. & Luckhaus, S. A generalised diffusion equation for phase separation of a multi-component mixture with interfacial free energy. Preprint at University Digital Conservancy https://hdl.handle.net/11299/1733 (1991).
Goehring, N. W. et al. Polarization of PAR proteins by advective triggering of a pattern-forming system. Science 334, 1137–1141 (2011).
Howard, M., Rutenberg, A. D. & de Vet, S. Dynamic compartmentalization of bacteria: accurate division in E. coli. Phys. Rev. Lett. 87, 278102 (2001).
Huang, K. C., Meir, Y. & Wingreen, N. S. Dynamic structures in Escherichia coli: spontaneous formation of MinE rings and MinD polar zones. Proc. Natl Acad. Sci. USA 100, 12724–12728 (2003).
Halatek, J. & Frey, E. Highly canalized MinD transfer and MinE sequestration explain the origin of robust MinCDE-protein dynamics. Cell Rep. 1, 741–752 (2012).
Denk, J. et al. MinE conformational switching confers robustness on self-organized Min protein patterns. Proc. Natl Acad. Sci. USA 115, 4553–4558 (2018).
Park, K.-T. et al. The Min oscillator uses MinD-dependent conformational changes in MinE to spatially regulate cytokinesis. Cell 146, 396–407 (2011).
Loose, M., Fischer-Friedrich, E., Herold, C., Kruse, K. & Schwille, P. Min protein patterns emerge from rapid rebinding and membrane interaction of MinE. Nat. Struct. Mol. Biol. 18, 577–583 (2011).
Fu, M. et al. Mechanochemical feedback loop drives persistent motion of liposomes. Nat. Phys. https://doi.org/10.1038/s41567-023-02058-8 (2023).
Weyer, H., Roth, T. & Frey, E. Henrikweyer/Turing-foams: code associated with publication ‘Deciphering the interface laws of turing mixtures and foams’. Zenodo https://doi.org/10.5281/zenodo.17336403 (2025).
Cahn, J. W. & Hilliard, J. E. Free energy of a nonuniform system. I. Interfacial free energy. J. Chem. Phys. 28, 258–267 (1958).
Eyre, D. J. Systems of Cahn–Hilliard equations. SIAM J. Appl. Math. 53, 1686–1712 (1993).
Bronsard, L. & Reitich, F. On three-phase boundary motion and the singular limit of a vector-valued Ginzburg–Landau equation. Arch. Ration. Mech. Anal. 124, 355–379 (1993).
Garcke, H. & Novick-Cohen, A. A singular limit for a system of degenerate Cahn–Hilliard equations. Adv. Differ. Equ. 5, 401–434 (2000).
Acknowledgements
We thank P. Glock and P. Schwille for providing us with previously unpublished data on the mesh patterns obtained in the experiments for ref. 20; and F. Brauns and M. Predel for inspiring discussions. We also thank J. Kayser, B. Nettuno, S. Rulands and D. Toffenetti for valuable feedback on the paper. This work was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) through the Excellence Cluster ORIGINS under Germany’s Excellence Strategy - EXC-2094 - 390783311, the European Union (ERC, CellGeom, project number 101097810), and the Chan-Zuckerberg Initiative (CZI) (E.F.). This research was supported in part by grant NSF PHY-2309135 to the Kavli Institute for Theoretical Physics (KITP) (E.F.).
Funding
Open access funding provided by Ludwig-Maximilians-Universität München.
Author information
Authors and Affiliations
Contributions
H.W., T.A.R. and E.F. conceived the study. H.W. developed the theory and performed the analysis. T.A.R. assisted in the analysis. H.W., T.A.R. and E.F. discussed and interpreted the results and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Physics thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Vertex angles, edge numbers, and domain areas for different experimental conditions.
The statistics of 3-fold vertices are analyzed as in Fig. 3 for several snapshots under different total concentrations of MinD and MinE indicated in the figure. The data was recorded in the experiments for ref. 20. The snapshots for each concentration combination stem from different fields of view of the same sample, which were prepared separately for the different concentration combinations. The fitted triple vertices are overlayed over the pattern (MinE fluorescence intensity in blue, MinD fluorescence intensity in red) in white. The 4-fold vertices are marked by black circles. a The aggregated statistics for the snapshot analyzed in Fig. 3 (first panel in this figure) and a second one are shown. b The three histograms are obtained from three snapshots, two of which are shown. At these total concentrations of MinD and MinE, the pattern transitions to a labyrinthine structure (second panel) which is seen in the histograms by the large domains with many edges. c, d Two further concentration combinations (two analyzed images each) show the same statistics of Turing foams as the condition a.
Extended Data Fig. 2 Domain collapse and splitting in the in vitro Min system.
a The time series of single domains in the in vitro Min protein Turing foam at the total concentrations of MinD [MinD] = 2 μM and MinE-His [MinE-His] = 8 μM are shown. The fluorescence intensity of labeled MinD (red) and MinE (blue) from the experiments for ref. 20 are shown for a domain with five (top row) and four (lower row) edges. Movie 4 shows the full pattern evolution recorded every 10 seconds. b Two domains are shown that split by the growth of new MinE branches (cf. Movie 4). c The domain-size trajectories for domains with less (blue) or more (red) than six edges are given (see Methods Sec. C). d The histograms of the final domain sizes of domains with less (blue), equal to (black), and more (red) than six edges are shown in d (see Methods Sec. C).
Extended Data Fig. 3 Interrupted coarsening stabilizes Turing foam on spherical surfaces.
Polyhedral MinE meshes (translucent cyan on a translucent black background; MinD not shown) with an increasing number of faces (foam domains) form in simulations as the sphere radius R is increased. An equatorial branch forms for 2R = 10 μm (a), a tetrahedral foam for 2R = 15 μm (b), a polyhedron with 4- and 5-sided domains at 2R = 20 μm (c), and a dodecahedral foam for 2R = 25 μm (d). The simulations are not drawn to scale. The model equations and parameters are given in Methods Sec. A 2.
Supplementary information
Supplementary Information (download PDF )
Supplementary Discussion including Figs. 1–5.
Supplementary Video 1 (download MP4 )
Emergence of two-dimensional Turing foam from random initial conditions in simulations of the Min system.
Supplementary Video 2 (download MP4 )
Interrupted coarsening in simulations of the Min system.
Supplementary Video 3 (download MP4 )
Domain splitting in simulations of the Min system.
Supplementary Video 4 (download MP4 )
Evolution of the Turing foam in in vitro experiments with the Min system.
Supplementary Video 5 (download MP4 )
Evolution of the Turing foam in simulations of the ferrocyanide–iodate–sulfite (FIS) reaction–diffusion system.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Weyer, H., Roth, T.A. & Frey, E. Protein pattern morphology and dynamics emerging from effective interfacial tension. Nat. Phys. 22, 94–102 (2026). https://doi.org/10.1038/s41567-025-03101-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41567-025-03101-6