Abstract
Coronatine and related bacterial phytotoxins are mimics of the hormone jasmonyl-l-isoleucine (JA-Ile), which mediates physiologically important plant signalling pathways1,2,3,4. Coronatine-like phytotoxins disrupt these essential pathways and have potential in the development of safer, more selective herbicides. Although the biosynthesis of coronatine has been investigated previously, the nature of the enzyme that catalyses the crucial coupling of coronafacic acid to amino acids remains unknown1,2. Here we characterize a family of enzymes, coronafacic acid ligases (CfaLs), and resolve their structures. We found that CfaL can also produce JA-Ile, despite low similarity with the Jar1 enzyme that is responsible for ligation of JA and l-Ile in plants5. This suggests that Jar1 and CfaL evolved independently to catalyse similar reactions—Jar1 producing a compound essential for plant development4,5, and the bacterial ligases producing analogues toxic to plants. We further demonstrate how CfaL enzymes can be used to synthesize a diverse array of amides, obviating the need for protecting groups. Highly selective kinetic resolutions of racemic donor or acceptor substrates were achieved, affording homochiral products. We also used structure-guided mutagenesis to engineer improved CfaL variants. Together, these results show that CfaLs can deliver a wide range of amides for agrochemical, pharmaceutical and other applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Nucleotide sequences for the mutants generated as part of this study are available in Supplementary Information. Other nucleotide sequences for the enzymes used in this study were obtained from GenBank, and their accession numbers are provided within the paper or in Supplementary Information. The original materials and data that support the findings of this study are either available within the paper or are available from the corresponding author upon reasonable request. Crystallographic coordinates of wild type and mutant PbCfaL have been deposited in the Protein Data Bank as 7A9I (wild type) and 7A9J (R395G mutant).
References
Rangaswamy, V., Jiralerspong, S., Parry, R. & Bender, C. L. Biosynthesis of the Pseudomonas polyketide coronafacic acid requires monofunctional and multifunctional polyketide synthase proteins. Proc. Natl Acad. Sci. USA 95, 15469–15474 (1998).
Ullrich, M. & Bender, C. L. The biosynthetic gene cluster for coronamic acid, an ethylcyclopropyl amino acid, contains genes homologous to amino acid-activating enzymes and thioesterases. J. Bacteriol. 176, 7574–7586 (1994).
Staswick, P. E. & Tiryaki, I. The oxylipin signal jasmonic acid is activated by an enzyme that conjugates it to isoleucine in Arabidopsis. Plant Cell 16, 2117–2127 (2004).
Fonseca, S. et al. (+)-7-iso-Jasmonoyl-l-isoleucine is the endogenous bioactive jasmonate. Nat. Chem. Biol. 5, 344–350 (2009).
Westfall, C. S. et al. Structural basis for prereceptor modulation of plant hormones by GH3 proteins. Science 336, 1708–1711 (2012).
Winn, M., Richardson, S. M., Campopiano, D. J. & Micklefield, J. Harnessing and engineering amide bond forming ligases for the synthesis of amides. Curr. Opin. Chem. Biol. 55, 77–85 (2020).
Parry, R. J., Jiralerspong, S., Mhaskar, S., Alemany, L. & Willcott, R. Investigations of coronatine biosynthesis. Elucidation of the mode of incorporation of pyruvate into coronafacic acid. J. Am. Chem. Soc. 118, 703–704 (1996).
Tao, T. & Parry, R. J. Determination by enantioselective synthesis of the absolute configuration of CPE, a potential intermediate in coronatine biosynthesis. Org. Lett. 3, 3045–3047 (2001).
Strieter, E. R., Koglin, A., Aron, Z. D. & Walsh, C. T. Cascade reactions during coronafacic acid biosynthesis: elongation, cyclization, and functionalization during Cfa7-catalyzed condensation. J. Am. Chem. Soc. 131, 2113–2115 (2009).
Parry, R. J., Lin, M. T., Walker, A. E. & Mhaskar, S. Biosynthesis of coronatine: investigations of the biosynthesis of coronamic acid. J. Am. Chem. Soc. 113, 1849–1850 (1991).
Vaillancourt, F. H., Yeh, E., Vosburg, D. A., O’Connor, S. E. & Walsh, C. T. Cryptic chlorination by a non-haem iron enzyme during cyclopropyl amino acid biosynthesis. Nature 436, 1191–1194 (2005).
Kelly, W. L. et al. Characterization of the aminocarboxycyclopropane-forming enzyme CmaC. Biochemistry 46, 359–368 (2007).
Rangaswamy, V. et al. Expression and analysis of coronafacate ligase, a thermoregulated gene required for production of the phytotoxin coronatine in Pseudomonas syringae. FEMS Microbiol. Lett. 154, 65–72 (1997).
Slawiak, M. & Lojkowska, E. Genes responsible for coronatine synthesis in Pseudomonas syringae present in the genome of soft rot bacteria. Eur. J. Plant Pathol. 124, 353–361 (2009).
Bignell, D. R. D. et al. Streptomyces scabies 87–22 contains a coronafacic acid-like biosynthetic cluster that contributes to plant-microbe interactions. Mol. Plant Microbe Interact. 23, 161–175 (2010).
Fyans, J. K., Altowairish, M. S., Li, Y. & Bignell, D. R. Characterization of the coronatine-like phytotoxins produced by the common scab pathogen Streptomyces scabies. Mol. Plant Microbe Interact. 28, 443–454 (2015).
Mitchell, R. E. & Frey, E. J. Production of N-coronafacoyl-L-amino-acid-analogs of coronatine by Pseudomonas syringae pv Atropurpurea in liquid cultures supplemented with L-amino acids. J. Gen. Microbiol. 132, 1503–1507 (1986).
Mitchell, R. E. & Ford, K. L. Chlorosis-inducing products from Pseudomonas syringae pathovars: new N-coronafacoyl compounds. Phytochemistry 49, 1579–1583 (1998).
Littleson, M. M. et al. Scalable total synthesis and comprehensive structure-activity relationship studies of the phytotoxin coronatine. Nat. Commun. 9, 1105 (2018).
Sabatini, M. T., Boulton, L. T., Sneddon, H. F. & Sheppard, T. D. A green chemistry perspective on catalytic amide bond formation. Nat. Catal. 2, 10–17 (2019).
Sabatini, M. T., Boulton, L. T. & Sheppard, T. D. Borate esters: simple catalysts for the sustainable synthesis of complex amides. Sci. Adv. 3, e1701028 (2017).
Krause, T., Baader, S., Erb, B. & Gooßen, L. J. Atom-economic catalytic amide synthesis from amines and carboxylic acids activated in situ with acetylenes. Nat. Commun. 7, 11732 (2016).
Stephenson, N. A., Zhu, J., Gellman, S. H. & Stahl, S. S. Catalytic transamidation reactions compatible with tertiary amide metathesis under ambient conditions. J. Am. Chem. Soc. 131, 10003–10008 (2009).
Allen, C. L., Atkinson, B. N. & Williams, J. M. J. Transamidation of primary amide with amines using hydroxylamine hydrochloride as an inorganic catalyst. Angew. Chem. Int. Edn 51, 1383–1386 (2012).
Al-Zoubi, R. M., Marion, O. & Hall, D. G. Direct and waste-free amidations and cycloadditions by organocatalytic activation of carboxylic acids at room temperature. Angew. Chem. Int. Edn 47, 2876–2879 (2008).
Noda, H., Furutachi, M., Asada, Y., Shibasaki, M. & Kumagai, N. Unique physicochemical and catalytic properties dictated by the B3NO2 ring system. Nat. Chem. 9, 571–577 (2017).
Scheidt, K. Amide bonds made in reverse. Nature 465, 1020–1022 (2010).
Goswami, A. & Van Lanen, S. G. Enzymatic strategies and biocatalysts for amide bond formation: tricks of the trade outside of the ribosome. Mol. Biosyst. 11, 338–353 (2015).
Petchey, M. et al. The broad aryl acid specificity of the amide bond synthetase McbA suggests potential for the biocatalytic synthesis of amides. Angew. Chem. Int. Edn 57, 11584–11588 (2018).
Wood, A. J. L. et al. Adenylation activity of carboxylic acid reductases enables the synthesis of amides. Angew. Chem. Int. Edn 56, 14498–14501 (2017).
Pattabiraman, V. R. & Bode, J. W. Rethinking amide bond synthesis. Nature 480, 471–479 (2011).
Pérombelon, M. C. M. Potato diseases caused by soft rot Erwinias: an overview of pathogenesis. Plant Pathol. 51, 1–12 (2002).
Bell, K. S. et al. Genome sequence of the enterobacterial phytopathogen Erwinia carotovora subsp. atroseptica and characterization of virulence factors. Proc. Natl Acad. Sci. USA 101, 11105–11110 (2004).
Bottini, R., Fulchieri, M., Pearce, D. & Pharis, R. P. Identification of gibberellins A1, A3 and iso-A3 in cultures of Azospirillum lipoferum. Plant Physiol. 90, 45–47 (1989).
Schüler, G. et al. Coronalon: a powerful tool in plant stress physiology. FEBS Lett. 563, 17–22 (2004).
Shockey, J. M., Fulda, M. S. & Browse, J. Arabidopsis contains a large superfamily of acyl-activating enzymes. Phylogenetic and biochemical analysis reveals a new class of acyl-coenzyme A synthetases. Plant Physiol. 132, 1065–1076 (2003).
Stuhlsatz-Krouper, S. M., Bennett, N. E. & Schaffer, J. E. Substitution of alanine for serine 250 in the murine fatty acid transport protein inhibits long chain fatty acid transport. J. Biol. Chem. 273, 28642–28650 (1998).
Nocek, B. P. et al. Structural insights into substrate selectivity and activity of bacterial polyphosphate kinases. ACS Catal. 8, 10746–10760 (2018).
Sheldon, R. A. Cross-linked enzyme aggregates (CLEAs): stable and recyclable biocatalysts. Biochem. Soc. Trans. 35, 1583–1587 (2007).
Zhang, L. et al. α-Ketoamides as broad-spectrum inhibitors of coronavirus and enterovirus replication: structure-based design, synthesis, and activity assessment. J. Med. Chem. 63, 4562–4578 (2020).
Boras, B. et al. Discovery of a novel inhibitor of coronavirus 3CL protease for the potential treatment of COVID-19. Preprint at https://www.biorxiv.org/content/10.1101/2020.09.12.293498v3 (2020).
Zhou, H.-J. et al. Design and synthesis of an orally bioavailable and selective peptide epoxyketone proteasome inhibitor (PR-047). J. Med. Chem. 52, 3028–3038 (2009).
Chen, C.-S., Fujimoto, Y., Girdaukas, G. & Sih, C. J. Quantitative analyses of biochemical kinetic resolutions of enantiomers. J. Am. Chem. Soc. 104, 7294–7299 (1982).
Jervis, P. J., Amorim, C., Pereira, T., Martins, J. A. & Ferreura, P. M. T. Exploring the properties and potential biomedical applications of NSAID-capped peptide hydrogels. Soft Matter 16, 10001–10012 (2020).
Tiwari, A. D. et al. Microwave assisted synthesis and QSAR study of novel NSAID acetaminophen conjugates with amino acid linkers. Org. Biomol. Chem. 12, 7238–7249 (2014).
Andexer, J. N. & Richter, M. Emerging enzymes for ATP regeneration in biocatalytic processes. ChemBioChem 16, 380–386 (2015).
Strohmeier, G. A., Eiteljorg, I. C., Schwarz, A. & Winkler, M. Enzymatic one-step reduction of carboxylates to aldehydes with cell-free regeneration of ATP and NADPH. Chem. Eur. J. 25, 6119–6123 (2019).
Philpott, H. K., Thomas, P. J., Tew, D., Fuerst, D. E. & Lovelock, S. L. A versatile biosynthetic approach to amide bond formation. Green Chem. 20, 3426–3431 (2018).
Hara, R., Hirai, K., Suzuki, S. & Kino, K. A chemoenzymatic process for amide bond formation by an adenylating enzyme-mediated mechanism. Sci. Rep. 8, 2950 (2018).
Acknowledgements
We thank the BBSRC (grants BB/K002341/1 and BB/N023536/1) and Syngenta for funding. F.W. was supported by the China Scholarship Council (grant no. 201806155100) and L.B. was funded by the Deutsche Forschungsgemeinschaft (DFG, grant BE 7054/1). The Michael Barber Centre for Collaborative Mass Spectrometry provided access to MS instrumentation. We also thank J. Vincent and N. Mulholland (Syngenta) for helpful discussions in the early stages of the project, and N. J. Turner (University of Manchester) for kindly providing the CHU plasmid. We also thank Diamond Light Source for beamtime access on i03 and i04-1 (proposal mx17773-56 and 76).
Author information
Authors and Affiliations
Contributions
M.W. and J.M. designed experiments; M.W., M.R., F.W., L.B. and D.F. carried out the experiments and provided additional experiment design; C.L. performed crystallographic studies. M.W. and J.M. wrote the manuscript. J.M. led the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Francesca Paradisi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Melting point determination of CfaL enzymes.
Melting point temperatures (Tm) of the CfaLs in this study, obtained using a fluorescence-based assay conducted in a Bio-Rad CFX Connect qPCR machine. Higher Tm indicates improved thermal stability. The Tm is calculated as the lowest point when plotting the negative derivative of RFU (relative fluorescence units) as a function of temperature (dT), versus the temperature (degrees Celsius).
Extended Data Fig. 2 Rational mutagenesis of PbCfaL.
a, Structural comparison between PbCfaL (left) and the mutant PbCfaL(R395G) (right). R395 (circled) of PbCfaL (PDB ID 7A9I) is in the hinge region between the N-terminal domain (blue) and the flexible C-terminal domain (red). In PbCfaL(R395G) (PDB ID 7A9J) this large arginine residue is replaced by a much smaller glycine (circled) that is found in the other members of the CfaL family and many other similar ANL ligases. The overall structure of this mutant exhibits no other substantial structural difference from that of the wild type. b, Overlay of PbCfaL with three published ATP-dependent ligase structures (in ellipse) showing the conserved ATP binding location. When superimposed, PbCfaL (PDB ID 7A9I, blue), McbA (AMP bound, PDB ID 6SQ8, red), GrsA (ATP bound, PDB ID 1AMU, green) and AuaEII (anthranoyl-AMP bound, PDB ID 4WV3, light brown) show the conserved location of ATP binding. The corresponding loop in PbCfaL (inset, arrowed) is larger than in the other structures which may affect ATP binding. The location of this region within the structure of PbCfaL (grey) is also shown for reference. Structural alignment was performed using Chimera (version 1.14) MatchMaker.
Extended Data Fig. 3 Synthesis and NMR spectra of crude product 64.
a, Preparative-scale synthesis of 64 from 10 and l-Ile catalysed by PbCfaL(R395G/A294P) (lysate), with either the addition of ATP (87% isolated yield) or recycling the endogenous ATP present in the lysate using the kinase (CHU) and polyphosphate (polyP) (52% isolated yield). b, 1H-NMR spectrum of crude product 64. c, 13C-NMR spectrum of crude product 64.
Extended Data Fig. 4 Catalysed conversion of 10 and l-Ile to give 64.
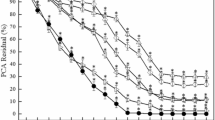
a, Catalysed by PbCfaL (R395G/A294P) CLEAs. Reactions (100 mM Tris-HCl, 10 mM MgCl2, 10 mM ATP, 1 mM 10, 3 mM l-isoleucine, 50 ml total volume) were run for 24 h; the CLEA (cross-linked enzyme aggregate) was then removed, washed, and reintroduced to an identical reaction. While activity was seen to reduce over the 5 days, the CLEA still retained high levels of productivity even after 5 recycles over 5 days, whereas cell lysates generally precipitated and lost all activity within 12 h. Although CfaL undergoes extensive conformational changes during catalysis, encapsulating it within CLEAs shows the potential of immobilization to extend the functional lifespan of the CfaL. More sophisticated immobilization techniques may have the potential to further retain activity. Conversion values were calculated from HPLC peak area ratios of product and starting materials, and represent means where n = 5, error bars denote s.d. b, Catalysed by purified PbCfaL(R395G/A294P), showing percentage conversions of 10 and l-Ile in the presence of various solvents and at different concentrations. Conversion values were calculated from HPLC peak area ratios of product and starting materials and represent means where n = 3, error bars denote s.d.
Extended Data Fig. 5 LCMS analysis and extracted-ion chromatograms (EICs) of fragments of telaprevir and bortezomib synthesized by PbCfaL(R395G/A294P).
a, Proposed route towards the antiviral agent telaprevir by CfaL (see reaction at top). The expected product of the reaction, 110, was detected by LCMS (top trace, expected m/z 262.1197, observed 262.1193 [M-H]−). Additional peaks consistent with dipeptide 111, formed from condensation of two cyclohexylglycines, 58, were also detected (bottom trace, 111a and 111b, expected m/z 295.2027, observed 295.2036 [M-H]−). Although CfaL are highly selective for l-amino acid substrates, the appearance of two products of the same mass suggests formation of diastereomers, which may be due to a lack of enantioselectivity in the adenylation step forming the acyl donor when racemic cyclohexylglycine (58) is used. This indicates that 58 can function as both a carboxylic acid and an amine donor. b, Proposed route towards anti-cancer agent bortezomib via the synthesis of 112 by CfaL (see reaction at top). The expected product of the reaction was detected by LCMS (top trace, expected m/z 270.0884, observed 270.0878 [M-H]−). An additional peak consistent with an l-Phe dipeptide (113) was also detected (bottom trace, expected m/z 311.1401). This indicates that l-Phe can function as both acyl donor and amine acceptor. c, The reaction between carboxylic acid substrate (9), which is a good substrate for the enzyme, and cyclohexylglycine (58) gives only the desired product (114, top trace, expected m/z 274.1449, observed 274.1460 [M-H]−). No cyclohexylglycine homodimer (dipeptide 111) was evident in this case, indicating that homocoupling of 58 only takes place when carboxylic acid (acyl donor) substrates that are not well accepted by CfaL are used.
Supplementary information
Supplementary Information (download PDF )
This file contains supplementary methods and compound characterization, NMR Spectra and supplementary references.
Supplementary Figures (download PDF )
This file contains supplementary text, supplementary figures 1 – 21, supplementary tables 1 – 4 and supplementary references.
Rights and permissions
About this article
Cite this article
Winn, M., Rowlinson, M., Wang, F. et al. Discovery, characterization and engineering of ligases for amide synthesis. Nature 593, 391–398 (2021). https://doi.org/10.1038/s41586-021-03447-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-021-03447-w
This article is cited by
-
Accelerated enzyme engineering by machine-learning guided cell-free expression
Nature Communications (2025)
-
Biocatalytic kinetic resolution for efficient and enantioselective synthesis of δ-haloalcohols and tetrahydrofurans
Communications Chemistry (2025)
-
Enzymatic combinatorial synthesis of E-64 and related cysteine protease inhibitors
Nature Chemical Biology (2025)
-
Biocatalytic enantioselective formation and ring-opening of oxetanes
Nature Communications (2025)
-
Biocatalytic Synthesis of N-trans-feruloyltyramine Using an Amide Bond Synthetase with an ATP Recycling
Applied Biochemistry and Biotechnology (2025)