Abstract
The composition of the intestinal microbiome varies considerably between individuals and is correlated with health1. Understanding the extent to which, and how, host genetics contributes to this variation is essential yet has proved to be difficult, as few associations have been replicated, particularly in humans2. Here we study the effect of host genotype on the composition of the intestinal microbiota in a large mosaic pig population. We show that, under conditions of exacerbated genetic diversity and environmental uniformity, microbiota composition and the abundance of specific taxa are heritable. We map a quantitative trait locus affecting the abundance of Erysipelotrichaceae species and show that it is caused by a 2.3 kb deletion in the gene encoding N-acetyl-galactosaminyl-transferase that underpins the ABO blood group in humans. We show that this deletion is a ≥3.5-million-year-old trans-species polymorphism under balancing selection. We demonstrate that it decreases the concentrations of N-acetyl-galactosamine in the gut, and thereby reduces the abundance of Erysipelotrichaceae that can import and catabolize N-acetyl-galactosamine. Our results provide very strong evidence for an effect of the host genotype on the abundance of specific bacteria in the intestine combined with insights into the molecular mechanisms that underpin this association. Our data pave the way towards identifying the same effect in rural human populations.
This is a preview of subscription content, access via your institution
Access options






Similar content being viewed by others
Data availability
All the 16S rRNA sequencing data, the metagenomics sequence data and the RNA-seq data were submitted to the GSA database under accession numbers CRA006230, CRA006239, CRA006240 and CRA006216. The genotype data were deposited at the GVM (http://bigd.big.ac.cn/gvm/getProjectDetail?project=GVM000310) under the GSA database under accession number GVM000310. The GWAS summary statistics are available at Figshare (https://doi.org/10.6084/m9.figshare.19313960). The whole-genome sequencing data of experimental pigs have been deposited in the GSA database (https://ngdc.cncb.ac.cn/gsa/browse/CRA006383) under accession number CRA006383. The source data are available at GitHub (https://github.com/yanghuijxau/Manuscript-microbiota-ABO).
Code availability
Codes to replicate the findings and the source data are available at GitHub (https://github.com/yanghuijxau/Manuscript-microbiota-ABO).
References
Kundu, P., Blacher, E., Elinav, E. & Pettersson, S. Our gut microbiome: the evolving inner self. Cell 171, 1481–1493 (2017).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
O’Hara, E., Neves, A. L. A., Song, Y. & Guan, L. L. The role of the gut microbiome in cattle production and health: driver or passenger? Annu. Rev. Anim. Biosci. 8, 199–220 (2020).
Schmidt, T. S. B., Raes, J. & Bork, P. The human gut microbiome: from association to modulation. Cell 172, 1198–1215 (2018).
Polderman, T. J. C. et al. Meta-analysis of the heritability of human traits based on 50 years of twin studies. Nat. Genet. 47, 702–709 (2015).
Polubriaginof, F. C. G. et al. Disease heritability inferred from familial relationships reported in medical records. Cell 173, 1692–1704 (2018).
Benson, A. K. et al. Individuality in gut microbiota composition is a complex polygenic trait shaped by multiple environmental and host genetic factors. Proc. Natl Acad. Sci. USA 107, 18933–18938 (2010).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Goodrich, J. K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
Blekhman, R. et al. Host genetic variation impacts microbiome compoistion across human body sites. Genome Biol. 16, 191 (2015).
Turpin, W. et al. Association of host genome with intestinal microbial composition in a large healthy cohort. Nat. Genet. 48, 1413–1417 (2016).
Bonder, M. J. et al. The effect of host genetics on the gut microbiome. Nat. Genet. 48, 1407–1412 (2016).
Wang, J. et al. Genome-wide association analysis identifies variation in vitamin D receptor and other host factors influencing the gut microbiota. Nat. Genet. 48, 1396–1406 (2016).
Hughes, D. A. et al. Genome-wide associations of human gut microbiome variation and implications for causal inference analyses. Nat. Microbiol. 5, 1079–1087 (2020).
Sankararaman, S. et al. The genomic landscape of Neanderthal ancestry in present-day humans. Nature 507, 354–357 (2014).
Patterson, N. et al. Genetic evidence for complex speciation of humans and chimpanzees. Nature 441, 1103–1108 (2006).
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2016).
Radjabzadeh, D. et al. Diversity, compositional and functional differences between gut microbiota of children and adults. Sci. Rep. 10, 1040 (2020).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743 (2016).
Cooling, L. Blood groups in infection and host susceptibility. Clin. Microbiol. Rev. 28, 801–870 (2015).
Rühlemann, M. C. et al. Genome-wide association study in 8,956 German individuals identifies influence of ABO histo-blood groups on gut microbiome. Nat. Genet. 53, 147–155 (2021).
Lopera-Maya, E. E. et al. Effect of host genetics on the gut microbiome in 7,738 participants of the Dutch microbiome project. Nat. Genet. 54, 143–151 (2022).
Qin, Y. et al. Combined effects of host genetics and diet on human gut microbiota and incident disease in a single population cohort. Nat. Genet. 54, 134–142 (2022).
Choi, M. K. et al. Determination of complete sequence information of the human ABO blood group orthologous gene in pigs and breed differences in blood type frequencies. Gene 640, 1–5 (2018).
Wang, S. et al. Design of glycosyl transferase inhibitors: serine analogues as pyrophosphate surrogates? ChemPlusChem 80, 1525–1532 (2015).
Ségurel, L. et al. The ABO blood group is a trans-species poilymorphism in primates. Proc. Natl Acad. Sci. USA 109, 18493–18498 (2012).
Groenen, M. A. M. A decade of pig genome sequencing: windo on pig domestication and evolution. Genet. Sel. Evol. 48, 23–32 (2016).
Ravcheev, D. A. & Thiele, I. Comparative genomic analysis of the human gut microbiome reveals a broad distribution of metabolic pathways for the degradation of host-synthesized mucin glycans and utilization of mucin-derived monosaccharides. Front. Genet. 8, 111 (2017).
Tailford, L. A. et al. Mucin glycan foraging in the human gut microbiome. Front. Genet. 6, 81 (2015).
Lien, K. A., Sauer, W. C. & He, J. M. Dietary influences on the secretion into and degradation of mucin in the digestive tract of monogastric animals and humans. J. Anim. Feed Sci. 10, 223–245 (2001).
Brinkkötter, A. B., Klöss, H., Alpert, C.-A. & Lengeler, J. W. Pathways for the utilization of N-acetyl-galactosamine and galactosamine in Escherichia coli. Mol. Microbiol. 37, 125–135 (2000).
Rodionov, D. A. et al. Genomic encyclopedia of sugar utilization pathways in the Shewanella genus. BMC Genom. 11, 494 (2010).
Leyn, S. A., Gao, F., Yang, C. & Rodionov, D. A. N-acetylgalactosamine utilization pathway and regulon in proteobacteria. J. Biol. Chem. 287, 28047–28056 (2012).
Hu, Z., Patel, I. R. & Mukherjee, A. Genetic analysis of the roles of agaA, agaI, and agaS genes in the N-acetyl-d-galactosamine and d-galactosamine catabolic pathways in Escherichia coli strains O157:H7 and C. BMC Microbiol. 13, 94 (2013).
Bidart, G. N., Rodriguez-Diaz, J., Monedoro, V. & Yebra, M. J. A unique gene cluster for the utilization of the mucosal and human milk-associated glycans galacto-N-biose and lacto-N-biose in Lactobacillus casei. Mol. Microbiol. 93, 521–538 (2014).
Zhang, H. et al. Two novel regulators of N-acetyl-galactosamine utilization pathway and distinct roles in bacterial infections. Microbiol. Open 4, 983–1000 (2015).
Lawrence, J. Selfish operons: the evolutionary impact of gene clustering in prokrayotes and eukaryotes. Curr. Opin. Genet. Dev. 9, 642–648 (1999).
Koonin, E. V. Evolution of genome architecture. Int. J. Biochem. Cell Biol. 41, 298–306 (2009).
Lombard, V. et al. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Rahfeld, P. et al. An enzymatic pathway in the human gut microbiome that converts A to universal O type blood. Nat. Microbiol. 4, 1475–1585 (2019).
Rahfled, P. et al. Prospecting for microbial α-N-acetylgalactosaminidases yields a new class of GH31 O-glycanase. J. Biol. Chem. 294, 16400–16415.
Chen, Y. et al. ABO blood group and susceptibility to severe acute respiratory syndrome. JAMA 293, 1450–1451 (2005).
Ellinghaus, D. et al. The ABO blood group locus and a chromosome 3 gene cluster associate with SARS-CoV-2 respiratory failure in an Italian-Spanish genome-wide association analysis. Preprint at medRxiv https://doi.org/10.1101/2020.05.31.20114991 (2020).
Blancher, A. Evolution of the ABO supergene family. ISBT Sci. Ser. 8, 201–206 (2013).
Makivuokko, H. et al. Association between the ABO blood group and the human intestinal microbiota composition. BMC Microbiol. 12, 94 (2012).
Davenport, E. R. et al. ABO antigen and secretor statuses are not associated with gut microbiota composition in 1,500 twins. BMC Genom. 17, 941–955 (2016).
Kurilshikov, A. et al. Large-scale association analyses identify host factors influencing human gut microbiome composition. Nat. Genet. 53, 156–165 (2021).
Malmuthuge, N., Griebel, P. J. & Guan, L. L. Taxonomic identification of commensal bacteria associated with the mucosa and digesta throughout the gastrointestinal tracts of preweaned calves. Appl. Environ. Microbiol. 80, 2021–2028 (2014).
Hanson, M. E. B. et al. Population structure of human gut bacteria in a diverse cohort from rural Tanzania and Botswana. Genome Biol. 20, 16 (2019).
Warr, A. et al. An improved pig reference genome sequence to enable pig genetics and genomics research. Gigascience 9 (2019).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Rimmer, A. et al. Integrating mapping-, assembly- and haplotype-based approaches for calling variants in clinical sequencing applications. Nat. Genet. 46, 912–918 (2014).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Coppieters, W., Karim, L. & Georges, M. SNP-based quantitative deconvolution of biological mixtures: application to the detection of cows with subclinical mastitis by whole genome sequencing of tank milk. Genome Res. 30, 1201–1207 (2020).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahe, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Cheema, M. U. & Pluznick, J. L. Gut microbiota plays a central role to modulate the plasma and fecal metabolomes in response to angiotensin II. Hypertension 74, 184–193 (2019).
Zhou, X. & Stephens, M. Genome-wide efficient mixed-model analysis for association studies. Nat. Genet. 44, 821–824 (2012).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Visscher, P. M. et al. Assumption-free estimation of heritability from genome-wide identity-by-descent sharing between full siblings. PLoS Genet. 2, e41 (2006).
Ziyatdinov, A. et al. lme4QTL: linear mixed models with flexible covariance structure for genetic studies of related individuals. BMC Bioinform. 19, 68 (2018).
Haseman, J. K. & Elston, R. C. The investigation of linkage between a quantitative trait and a marker locus. Behav. Genet. 2, 3–19 (1972).
Aulchenko, Y. S., Ripke, S., Isaacs, A. & van Duijn, C. M. GenABEL: an R library for genome-wide association analysis. Bioinformatics 23, 1294–1296 (2007).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Vaser, R., Sovic, I., Nagarajan, N. & Sikic, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at arXiv https://arxiv.org/abs/1303.3997 (2013).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Harris, R. S. Improved Pairwise Alignment of Genomic DNA. PhD thesis, Pennsylvania State Univ. (2007).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Ai, H. et al. Adaptation and possible ancient interspecies introgression in pigs identified by whole-genome sequencing. Nat. Genet. 47, 217–225 (2015).
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: the genome analysis Toolkit best practices pipeline. Curr. Protoc. Bioinform. 43, 11.10.11–11.10.33 (2013).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984).
Nei, M. F-statistics and analysis of gene diversity in subdivided populations. Ann. Hum. Genet. 41, 225–233 (1977).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Hunt, M. et al. Circlator: automated circularization of genome assemblies using long sequencing reads. Genome Biol. 16, 294 (2015).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119 (2010).
Chen, S., Zhou, Y., Chen, Y. & Jia, G. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Li, D. et al. MEGAHIT v1.0: a fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102, 3–11 (2016).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP—a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Kanehisa, M., Sato, Y. & Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 428, 726–731 (2016).
Segata, N., Bornigen, D., Morgan, X. C. & Huttenhower, C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat. Commun. 4, 2304 (2013).
Li, M. et al. Aldolase B suppresses hepatocellular carcinogenesis by inhibiting G6PD and pentose phosphate pathways. Nat. Cancer 1, 737–747 (2020).
Nanchen, A., Fuhrer, T. & Sauer, U. Determination of metabolic flux rartios from 13C-experiments and gas chromatography-mass spectrometry data: protocol and principles. Methods Mol. Biol. 358, 177–197 (2007).
van Winden, W. A. et al. Correcting mass isoptopomer distributions for naturally occurring isotopes. Biotechnol. Bioeng. 80, 477–479 (2002).
Staley et al. Stable engraftment of human microbiota into mice with a single oral gavage following antibiotic conditioning. Microbiome 5, 87 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Momozawa, Y. et al. IBD risk loci are enriched in multigenic regulatory modules encompassing putative causative genes. Nat. Commun. 9, 2427 (2018).
Bushnell, B. BBMap: A Fast, Accurate, Splice-Aware Aligner (version 38.82) https://sourceforge.net/projects/bbmap/ (2014).
Köster, J. & Rahmann, S. Snakemake: a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Bolyen, E., Rideout, J. R., Dillon, M. R., Bokulich, N. A. & Caporaso, J. G. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Ghodsi, M., Liu, B. & Pop, M. DNACLUST: accurate and efficient clustering of phylogenetic marker genes. BMC Bioinform. 12, 271 (2011).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Srivastava, A. et al. Genomes of the mouse collaborative cross. Genetics 206, 537–556 (2017).
Yu, N. et al. Global patterns of human DNA sequence variation in a 10-kb region on chromosome 1. Mol. Biol. Evol. 18, 214–222 (2001).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Frantz, L. A. F. et al. Evidence of long-term gene flow and selection during domestication from analyses of Eurasian wild and domestic pig genomes. Nat. Genet. 47, 1141–1148 (2015).
Charlier, C. et al. NGS-based reverse genetic screen for common embryonic lethal mutations compromising fertility in livestock. Genome Res. 26, 1333–1341 (2016).
Georges, M., Charlier, C. & Hayes, B. Harnessing genomic information for livestock improvement. Nat. Rev. Genet. 20, 135–156 (2019).
Geraldes, A. et al. Inferring the history of speciation in house mice from autosomal, X-linked, Y-linked and mitochondrial genes. Mol. Ecol. 17, 5349–5363 (2008).
Huang, H. et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature 547, 173–178 (2017).
Suzuki, T. A. & Nachman, M. W. Spatial heterogeneity of gut microbioal composition along the gastrointestinal tract in natural populatiions of house mice. PLoS ONE 11, e0163720 (2016).
Vuik, F. E. R. et al. Composition of the mucosa-associated microbiota along the entire gastrointestinal tract of human individuals. UEG J. 7, 897–907 (2019).
Rowe, J. A. et al. Blood group O protects against severe Plasmodium falciparum malaria through the mechanism of reduced rosetting. Proc. Natl Acad. Sci. USA 104, 17471–17476 (2007).
Robinson, M. G., Tolchin, D. & Halpern, C. Enteric bacterial agents and the ABO blood groups. Am. J. Hum. Genet. 23, 135–145 (1971).
Camus, D., Bina, J. C., Carlier, Y. & Santoro, F. ABO blood groups and clinical forms of schistosomiasis mansoni. Trans. R. Soc. Trop. Med. Hyg. 71, 182 (1977).
Pereira, F. E. L., Bortolini, E. R., Carneiro, J. L. A., da Silva, C. R. M. & Neves, R. C. A, B, O blood groups and hepatosplenic form of schistosomiasis mansoni (Symmer’s fibrosis). Trans. R. Soc. Trop. Med. Hyg. 73, 238 (1977).
Ndamba, J., Gomo, E., Nyazema, N., Makaza, N. & Kaondera, K. C. Schistosomiasis infection in relation to the ABO blood groups among school children in Zimbabwe. Acta Trop. 65, 181–190 (1997).
Chaudhuri, A. & De, S. Cholera and blood groups. Lancet 2, 404 (1977).
Boren, T. et al. Attachment of Helicobacter pylori to human gastric epithelium mediated by blood group antigens. Science 262, 1892–1895 (1993).
Lindesmith, L. et al. Human susceptibility and resistance to Norwalk virus infection. Nat. Med. 9, 548–553 (2003).
Galili, U. in α-Gal and Anti-Gal (eds Galili, U. & Avila, J. L.) Vol. 32, 1–23 (Springer, 1999).
Prather, R. S., Shen, M. & Dai, Y. Genetically modified pigs for medicine and agriculture. Biotechnol. Genetic Eng. Rev. 25, 245–266 (2008).
Acknowledgements
We thank Y. He, S. Xiao, W. Li, Y. Guo and Y. Xing for assistance in the construction of the experimental mosaic pig populations; Y. Su and J. Li for preparation of reagents and management of samples; Y. Momozawa, R. Mariman, M. Mni, L. Karim and M. Dekkers for generating the CEDAR-1 16S rRNA data; the staff at the Jiangxi Department for Education, the Ministry of Science and Technology of P. R. China, the Ministry of Agriculture and Rural Affairs of P. R. China, and Jiangxi department of Science and Technology for their long-term support of the swine heterogeneous stock project; and the members of the MIQUANT consortium for comments and discussions. L.H. is supported by The National Natural Science Foundation of China (31790410) and National pig industry technology system (CARS-35); C. Chen by the National Natural Science Foundation of China (31772579); H.Y. by the National Postdoctoral Program for Innovative Talent (no. BX201700102); L.S. by the FNRS IBD-GI-Seq project; M.G. by the Chinese Thousand Talents Program, the Belgian EOS ‘Miquant’ project and the FNRS (CDR ‘GEM’ project). C. Charlier is a senior research associate at the FNRS.
Author information
Authors and Affiliations
Contributions
H.Y. analysed the 16S rRNA sequence data, performed GWAS, meta-analyses and local association analyses, computed heritabilities of individual taxa, contributed to ABO genotyping and analysed the effect of the 2.3 kb deletion on taxa abundance. J.W. analysed the composition of the microbiome, including PCoA analyses, β- and α-diversity, correlations between kinship and microbiome dissimilarities, isolated the OTU476-like strains, performed the GalNAc feeding experiments, measured the concentrations of GalNAc in the caecal lumen, analysed the GalNAc import and use pathway in the MAGs, and contributed to ABO genotyping. X.H. participated in 16S rRNA sequencing (F6) and GWAS (F6). Y. Zhou performed metagenome sequencing analysis, analysed the GalNAc import and use pathway in MAGs, analysed the RNA-seq data from caecum samples and contributed to ABO genotyping. Y. Zhang participated in the preparation of the genotype data from whole-genome sequence information, participated in the computation of the genomic contribution of the different breeds in the F6 and F7 generation and the definition of expected mapping resolution, performed LD analyses, performed eQTL analysis for the ABO gene, participated in the characterization and sequence analysis of the ABO gene, including definition of the 2.3 kb deletion, and in the balancing selection and trans-species polymorphism analyses. M.L. assisted with the isolation of the OTU476-like strains, the GalNAc feeding experiments and genotyping of the ABO gene. Q.L. assisted with measuring the concentrations of GalNAc in caecal lumen. S.K., M.H., H.F., S.F., X.X., H.J., Z.C. and J.G. assisted with the experiments. Z.Z., X.T., Z.W., H.G. and Y.H. assisted with the preparation of genotype data from whole-genome sequencing data and conducted the analysis of the Nanopore data of the ABO region. J.M. assisted with the construction of the mosaic population. H.A. assisted with the bioinformatic analysis of the ABO region, de novo assembly of the A allele, and evolutionary analysis of the ABO alleles. L.S. analysed the effect of ABO genotype on intestinal microbiota composition in humans. W.C. assisted in the analysis of the sequencing data for the trans-species polymorphisms. C. Charlier supervised the characterization of the ABO gene and the 2.3 kb deletion and the corresponding haplotype structure in the F0, F6 and F7 population and for the trans-species polymorphism. B.Y. prepared the genotype data of whole-genome variants, assisted with raising the heterogeneous stock, and participated in the computation of the genomic contribution of the different breeds in the F6 and F7 generation and the definition of expected mapping resolution. M.G. supervised the bioinformatic and statistical analyses, performed bioinformatic and statistical analyses, and wrote the paper. C. Chen codesigned the study, supervised experiments, supervised bioinformatic and statistical analyses of gut microbiome, and wrote the paper. L.H. created the swine heterogeneous stock, designed the study, directed the project, supervised the experiments and analyses, and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Catherine Lozupone, Vincent Plagnol and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Generating a large mosaic pig population for genetic analysis of complex phenotypes.
(a) Rotational breeding design used for the generation of a large mosaic pig population for the genetic analysis of complex phenotypes, with sampling scheme for faeces (D25, D120, D240), luminal content of the ileum (IC) and caecum (CC), and mucosal scrapings in the ileum (IM) and caecum (CM). BX: Bamaxiang, EH: Erhualian, LA: Laiwu, TB: Tibetan, LW: Large White, LD: Landrace, PT: Piétrain, WD: White Duroc. (b) Average similarity (1 – π) between allelic sequences sampled within and between the eight founder breeds. The colour intensity ranges from black (breeds with lowest allelic similarity: BX vs WD, 1 - 4.3x10−3) to bright red (breed with highest allelic similarity: WD, 1 - 1.8x10−3). The acronyms for the breeds are as in (a). More than 30 million variants with MAF ≥ 3% segregate in this population, i.e. more than one variant every 100 base pairs. This is slightly lower than the 40 million high quality variants segregating in the mouse collaborative cross108. (c) Comparison of the average nucleotide diversity (π, i.e. the proportion of sites that differ between two chromosomes sampled at random in the population(s)) within and between European (Eur) and Asian (As) domestic pigs, and between modern European (HSEur), Asian humans (HSAs), Neanderthal (Neand) and Chimpanzee (Pan Trogl). The average nucleotide diversity within the four Chinese founder breeds was ~2.5x10−3 and within the four European founder breeds ~2.0x10−3. By comparison, π-values within African and within Asian/European human populations are ~9x10−4 and ~8x10−4, respectively109,110. Thus, against intuition (as domestication is often assumed to have severely reduced effective population size) the within population diversity is >2-fold higher in domestic pigs than in human populations, as previously reported111,112,113. Nucleotide diversities between Chinese founder breeds and between European founder breeds were ~3.6x10−3 and ~2.5x10−3, respectively, i.e. 1.44-fold and 1.25-fold higher than the respective within-breed π-values. These π-values are of the same order of magnitude as the sequence divergence between Homo sapiens and Neanderthals/Denosivans (~3x10−3, ref. 15). By comparison, π-values between Africans, Asians and Europeans are typically ≤ ~1x10−3 (ref. 109). The nucleotide diversity between Chinese and European breeds averaged ~4.3x10−3. This π-value is similar to the divergence between M. domesticus and M. castaneus114, and close to halve the ~1% difference between chimpanzee and human16. Note that Chinese and European pig breeds are derived from Chinese and European wild boars, respectively, which are thought to have diverged ~1 million years ago27, while M. domesticus and M. castaneus are thought to have diverged ≤ 500,000 years ago114. (d) Autosome-specific estimates of the genomic contributions of the eight founder breeds in the F6 and F7 generation. We used a linear model incorporating all variants to estimate the average contribution of the eight founder breeds in the F6 and F7 generation at genome and chromosome level56. At genome-wide level, the proportion of the eight founder breed genomes ranged from 11.2% (respectively 11.5%) to 14.1% (14.7%) in the F6 (F7) generations. At chromosome-specific level, the proportion of the eight founder breeds ranged from 6.7% (respectively 4.9%) to 20.7% (22.1%) in the F6 (F7) generations. The genomic contribution of the eight founder breeds in the F6 and F7 generation is remarkably uniform and close to expectations (i.e. 12.5%) both at genome-wide and chromosome-wide level, suggesting comparable levels of genetic diversity across the entire genome. This does not preclude that more granular examination may reveal local departures from expectations, or under-representation of incompatible allelic combinations at non-syntenic loci. (e-f) Indicators of achievable mapping resolution in the F6 generation: (e) Frequency distribution (density) of the number of variants in high LD (r2 ≥ 0.9) with an “index” variant (was computed separately for all variants considered sequentially as the “index”), corresponding to the expected size of “credible sets” in GWAS115. The red vertical line corresponds to the genome-wide median. The green vertical line corresponds to the mapping resolution achieved in this study for the ABO locus (see hereafter). (f) Frequency distribution (density) of the maximum distance between an index variant and a variant in high LD (r2 ≥ 0.9) with it, defining the spread of credible sets. Red and green vertical lines are as in (D).
Extended Data Fig. 2 Characterizing the age- and location-specific composition of the intestinal microbiome of the healthy pig.
(a) Definition of a core intestinal microbiome of the pig. A total of 58 OTUs that were annotated to 21 taxa were identified in >95% of day 120 and 240 faeces and caecum content samples of both F6 and F7 generations, hence defined as core bacterial taxa. (b) The compositions of the porcine and human intestinal microbiota are closer to each other than either is to that of the mouse. Boxplots are as is Fig. 1c. The number of samples available for analysis were 1281 pigs, 106 humans and 6 mice. (c) Abundances (F6-F7 averages when available) of the 43 families represented in Fig. 1b in the seven sample types relative to the sample type in which they are the most abundant (red – blue scale). The families are ordered according to the sample type in which they are the most abundant. The colour-code for phyla is as in Fig. 1b. Columns are added for comparison with mouse and human. Mouse data are from Fig. 1 in Suzuki & Nachman116, and human data from Fig. 6 in Vuik et al117. P_I: proximal ileum, D_IL: distal ileum, C: caecum, CO: colon, RE: rectum, F: faeces. The families differing the most with regards to location-specific distribution between species include Helicobacteriaceae, Veillonellaceae, Lactobacillaceae and Streptocaccaceae.
Extended Data Fig. 3 Evaluating the heritability of intestinal microbiota composition in the mosaic pig population.
Correlation between heritability estimates of taxa/OTUs in F6 and F7 generation by sample type (D25, D120, D240, CC and IC). Correlation coefficients (r) and associated p-values (p) were computed using heritability estimates that were pre-corrected for bacterial abundance (residuals of linear model). Heritability estimates indeed tend to slightly increase with taxa abundance. Yet, results show that this effect cannot account for the observed correlations between F6 and F7 estimates in D120, D240 and CC, hence pointing towards genuine genetic effects. The shaded areas correspond to the 95% confidence region for the regression fit. Correlation coefficients and two-sided p-values were computed using Spearman’s rank-based method. Reported p-values are nominal (i.e. uncorrected for multiple testing).
Extended Data Fig. 4 Identifying a microbiota QTL (miQTL) with major effect on the abundance of Erysipelotrichaceae species by whole genome sequence based GWAS.
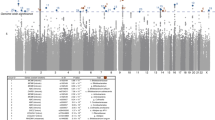
(a) Schematic illustration of the samples and SNPs used for the two types of analyses (abundance and presence/absence) performed for miQTL mapping. (b) (Upper) Distribution of log(1/p) values for 1,527 sets of 11 p-values obtained in 11 data-series for a SNP x taxon x analysis model combination that yielded a genome-wide significant signal (p < 5 x 10−8) in the 12th data-series. (Lower) Distribution of log(1/p) values for 1,527 sets of 11 p-values obtained in the same data-series and with the same analysis model as in (upper) but with randomly selected SNP x taxon combinations matching the ones in (upper) for MAF and taxa abundance. Log(1/p) values were computed using GenABEL as described in Methods. Corresponding p-values are nominal and two-sided. (c) Correlation between the average (F6 and F7) taxon heritability, and the average (F6 and F7) number of genome-wide significant (p ≤5 × 10−8) miQTL for D240 faecal samples. The shaded area corresponds to the 95% confidence region for the regression fit. Correlation coefficient and associated p-values are Spearman’s. (d) QQ plot for 1,527 (number of signals (SNP x taxon x model x one data series in one cohort) exceeding the genome-wide log(1/p) threshold value of 7.3) sets of ≤ 5-7 p-values (same SNP x taxon x model, all data series in the other cohort) for real SNPs (Blue: quantitative model; Green: binary model), and matched sets of ≤ 5-7 p-values corresponding to randomly selected SNP x taxon combinations matched for MAF and abundance or presence/absence rate (Brown: quantitative model; Yellow: binary model). Log (1/p) values were computed using GenABLE as described in Methods. Corresponding p-values are nominal and two-sided. (e) Same QQ plot as in (C) after removal of all SNPs in the chromosome 1: 272.8-273.1Mb interval. Log(1/p) values were computed using GenABLE as described in Methods. Corresponding p-values are nominal and two-sided. (f) Distribution of the association log(1/p) values and corresponding signed z-scores for SNP 1_272907239 and 31 p-75-a5 OTUs (red) and 83 Erysipelotrichaceae (yellow) OTUs, showing an enrichment of effects with same sign as for OTU476 and OTU327. Log(1/p) values were computed using Metal (v3.0) as described in Methods. Corresponding p-values are nominal and two-sided. See also Supplemental discussion 1.
Extended Data Fig. 5 The chromosome 1 miQTL is caused by a 2.3 kb deletion in the orthologue of the human ABO gene.
(a) Breakpoints of the 2.3 kb deletion showing the role of a duplicated SINE sequence in mediating an intra-chromosomal recombination. (b) Illustrative example of allelic balance for the cG146C SNP in an AA homozygote and of allelic imbalance for the same SNP in an AO heterozygote. (c) (Upper) eQTL analysis for the porcine AO gene maximizing at the exact position of the 2.3 kb deletion (p = 1.9x10−43) and showing the additive effect of the A allele increasing transcript levels ~3-fold (inset; FPKM: Fragments Per Kilobase of transcript per Million mapped reads). The “n’s” correspond to the number of animals of each genotype available for analysis. Boxplots are as in Fig. 1c. (Lower) Genome wide eQTL scan for the porcine ABO gene showing the strong cis-eQTL signal on chromosome 1. eQTL analysis was conducted with GEMMA (v0.97)64. Reported log-transformed p-values are nominal and two-sided. (d) Effect of N-acetyl-galactosaminyl transferase genotype (AA, AO or OO) on abundance of OTU327 and p-75-a5 in the twelve data series. Absence of an effect of N-acetyl-galactosaminyl transferase genotype (AA, AO or OO) on abundance of E. coli in the twelve data series. Sample sizes are as in STable 4.1. Boxplots are as in Fig. 3d. (e) Abundance of OTU476, OTU327 and p-75-a5 in the twelve data series. Violin plots with indication of the median. Numbers (n’s) are as in STable 4.1. See also Supplemental discussion 2.
Extended Data Fig. 6 cis-eQTL analyses in the vicinity of the chromosome 1 miQTLK supports the causality of the 2.3 kb deletion.
(a) Cis-eQTL analysis for the porcine N-acetyl-galactosaminyl transferase (“ABO”), GBTG1, LCN1 (=OBP2B), MED22 and SURF6 genes in caecum. The blue triangle corresponds to the top SNP for the miQTL. The red triangles correspond to the top SNPs for the respective cis-eQTL. Only for N-acetyl-galactosaminyl transferase are blue and red variants the same. eQTL analyses were conducted with GEMMA (v0.97)64. Reported log-transformed p-values are nominal and two-sided. (b) Effect of AO genotype on the expression levels of the corresponding genes in caecum. There was no evidence for an effect of AO genotype on the expression of any of these genes other than ABO. The number of AA, AO and OO samples available for cis-eQTL analysis for each gene are given (n). Boxplots are as in Fig. 1c. We tested the difference in gene expression level between pairs of genotype classes using a two-sided t-test. (c) Effect of the top cis-eQTL SNPs (blue triangles in A) on OTU476 abundance. Only the top cis-eQTL SNPs for ABO has an effect on OTU476 abundance. The number of AA, AO and OO samples available for miQTL analysis for each gene are given (n). Boxplots are as in Fig. 1c. We tested the difference in bacterial abundance between pairs of genotype classes using a two-sided t-test.
Extended Data Fig. 7 The 2.3 kb deletion in the orthologue of the human ABO gene is 3.5 million years old and under balancing selection.
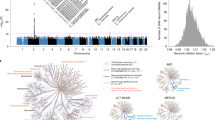
(a) UPGMA tree based on nucleotide diversities between 14 AA and 34 OO animals in windows of increasing size (0.5 to 40 kb) centred on the 2.3 kb deletion in the porcine N-acetyl-galactosaminyl transferase gene (porcine O allele). PA: Phacochaerus Africanus, SC: Sus cebifrons, SV: Sus verrucosus, SU: Sus scrofa vittatus, CB: Chinese wild boar, RB: Russian wild boar, EB: European wild boar, ERH: Erhualian, BX: Bamaxiang, T: Tibetan, LA: Laiwu, LR: Landrace, LW: Large White, PI: Piétrain, WD: White Duroc. Context: To gain additional insights in the age of the porcine O allele, we generated phylogenetic trees of the A and O alleles of 14 AA and 34 OO animals including domestic pigs, wild boars, Visayan and Javanese warty pigs, and common African warthog. Examination of their local SNP genotypes (50K window encompassing the ABO gene) reveals traces of ancestral recombinations between O and A haplotypes as close as 300 and 800 base pairs from the proximal and distal deletion breakpoints, respectively, as well as multiple instances of homoplasy that may either be due to recombination, gene conversion or recurrent de novo mutations. On their own, these signatures support the old age of the O allele. We constructed UPGMA trees based on nucleotide diversity for windows ranging from 500 bp to 40 kb centred on the 2.3 kb deletion. Smaller windows have a higher likelihood to compare the genuine ancestral O versus A states, yet yield less robust trees because they are based on smaller number of variants. Larger windows will increasingly be contaminated with recombinant A-O haplotypes blurring the sought signal. Indeed, for windows ≥ 20 kb or more, the gene tree corresponds to the species tree, while for windows ≤ 15 kb the tree sorts animals by AA vs OO genotype. For all windows ≤ 15 kb the Sus cebifrons O allele maps outside of the Sus scrofa O allele supporting a deep divergence (rather than hybridization) and hence the old age of the O allele. Of note, for windows ≤1.2 kb, the warthog A allele is more closely related to the Sus A alleles than to the Sus O alleles (ED7A). This suggests that the O allele may be older than the divergence of the Phacochoerus and Sus A alleles, i.e. > 10 MYA. It will be interesting to study larger numbers of warthog to see whether the same 2.3 kb deletion exists in this and other related species as well. (b) Alignment of ~900 base pairs of the O alleles of domestic pigs (Bamaxian), European and Asian wild boars, and Sus cebufrons demonstrating that these are identical-by-descent. The SINE element that is presumed to have mediated the recombinational event that caused to 2.3 kb deletion is highlighted in red. Context: To further support their identity-by-descent we aligned ~900 base pairs (centred on the position of the 2.3 kb deletion) of the O alleles of domestic pig, European and Asian wild boars and Sus cebifrons. The sequences were nearly identical further supporting our hypothesis. It is noteworthy that the old age of the “O” allele must have contributed to the remarkable mapping resolution (≤3 kb) that was achieved in this study. In total, 42 variants were in near perfect LD (r2 ≥ 0.9) with the 2.3 kb deletion in the F0 generation, spanning 2,298 bp (1,522 on the proximal side, and 762 on the distal side of the 2.3 kb deletion). This 2.3 kb span is lower than genome-wide expectations (17th percentile), presumably due to the numerous cross-overs that have accrued since the birth of the 2.3 kb deletion that occurred in the distant past. Yet the number of informative variants within this small segment is higher than genome-wide average of (57% percentile) also probably due at least in part to the accumulation of numerous mutations since the remote time of coalescence of the A and O alleles (see Fig. 1d in main text). (c) QQ plots for the effect of AO genotype on 150 phenotypes pertaining to meat quality, growth, carcass composition, hematology, health, and other phenotypes in the F6 and F7 generation. P-values were obtained using a mixed model followed by meta-analysis (weighted Z score) across the F6 and F7 generations as described in Methods. log-transformed p-values used for the QQ plot are nominal and two-sided. Context: Our findings in suidae are reminiscent of the trans-species polymorphism of the ABO gene in primates attributed to balancing selection26. The phenotype driving balancing selection remain largely unknown yet a tug of war with pathogens is usually invoked: synthesized glycans may affect pathogen adhesion, toxin binding or act as soluble decoys, while naturally occurring antibodies may be protective20,44. In humans, the O allele may protect against malaria118, E. Coli and Salmonella enteric infection119, SARS-CoV-142, SARS-CoV-243 and schistosomiasis120,121,122, while being a possible risk factor for cholera123, H. pylori124 and norovirus infection125. Whatever the underlying selective force, it appears to have operated independently in at least two mammalian branches (primates and suidae), over exceedingly long periods of time, and over broad geographic ranges, hence pointing towards its pervasive nature. To gain insights in what selective forces might underpin the observed balanced polymorphism, we tested the effect of porcine AO genotype on >150 traits measured in the F6 and F7 generations pertaining to carcass composition, growth, meat quality, hematological parameters, disease resistance and behaviour. No significant effects were observed when accounting for multiple testing, including those pertaining to immunity and disease resistance. (d) Expression profile of the AO gene in a panel of adult and embryonic porcine tissues (own RNA-Seq data).
Extended Data Fig. 8 The chromosome 1 miQTL affects caecal N-acetyl-D-galactosamine (GalNAc) concentrations which are correlated with the abundance of Erysipelotyrichaceae species within AO genotype: theory.
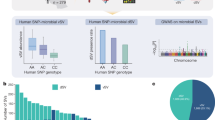
(a) ABO and α-gal epitopes in pigs and human. The glycosyltransferase gene located on 9q34.2 and underpinning the human ABO blood group is characterized in most human populations by three major alleles: (i) IA encoding a α-3-N-acetyl-D-galactosaminyltransferase that is adding GalNAc to H and Lewis antigens (yielding the A antigen) on various glycoproteins including mucins secreted in the intestinal lumen, (ii) IB encoding a α-3-D-galactosyltransferase that is adding galactose to the same antigens (yielding the B antigen), and (iii) the inactive IO null allele that precludes expression of either the A and/or the B antigen. Mutations in the fucosyltransferase 2 gene (FUT2) preclude formation of the H antigen on secreted proteins and hence the detection of A and B antigens in secretions20. The pig orthologue of the human ABO glycosyltransferase gene is located on the telomeric end of porcine chromosome 1q, and is characterized by two major alleles: (i) the A allele, encoding a α-3-N-acetyl-D-galactosaminyltransferase that is adding GalNAc to H and Lewis antigens, similar to the human IA allele, and (ii) the O allele corresponding to a null allele as a result of a 2.3 kb deletion similar to the human IO allele24. Thus, the B antigen (Galα1-3(Fucα1-2)Galβ1-4GlcNAc-R) is not observed in pig populations. However, what is found abundantly on the surface of cells in many tissues is the so-called “α-gal epitope” (Galα1-3Galβ1-4GlcNAc-R), which results from the addition of a galactose to the Galβ1-4GlcNAc-R precursor by a α1,3galactosyltransferase encoded by the GGTA1 gene. The orthologue of the GGTA1 gene is non-functional in human and Old World non-human primates, which, however, have high titers of circulating anti-α-gal antibodies contributing to acute rejection of xenografts126,127. (b) Identifying whether changes in GalNAc concentration are the cause of the observed changes in abundance of Erysipelotrichaceae species by searching for a correlation between the two phenotypes “within AO genotype”. (b1) If AO genotype is associated with the abundance of Erysipelotrichaceae species and GalNAc concentrations by virtue of different molecular mechanisms (for instance because they involved distinct causative mutations albeit in linkage disequilibrium, or because the gene has an as of yet unknown other activity that is causing the change in bacterial abundance, independently of its glycosyltransferase activity), there is no reason to expect a correlation between bacterial abundance and GalNAc concentration within AO genotype (red horizontal lines in the dotted circles). There is of course a correlation across genotypes that is due to the fact that AO genotype has a (direct or indirect) effect on both phenotypes. (b2) If, on the other hand, AO genotype causes the change in GalNAc concentration (which is very likely given its known enzymatic activity) which then causes the change in the abundance of Erysipelotrichaceae species, one can expect that bacterial abundance and GalNAc concentration will be correlated, also within AO genotype, as indicated by the sloped red lines within the dotted ellipses. This is what is observed with the real data.
Extended Data Fig. 9 The chromosome 1 miQTL affects caecal N-acetyl-D-galactosamine (GalNAc) concentrations which are correlated with the abundance of Erysipelotyrichaceae species within AO genotype: results.
(a) Positive correlation between caecal GalNAc concentrations and bacterial abundance (upper panels: p-75-a5; lower panels: OTU327) “within AO genotype”. GalNAc concentrations and bacterial abundances were corrected for batch effects and AO genotype and scaled between 0 and 1 to equalize residual variance. Correlations were computed using all samples jointly and Spearman’s rank-based test; corresponding p-values (nominal; two-sided) are given (left panels). Regression lines are shown for the different AO genotypes separately (right panels); all of them are positive. Note that the scatter plots for p-75-a5 are not identical but very similar to those for OTU476 (Fig. 5b, c). This is because OTU476 accounts for most of the p-75-a5 genus in caecum content (see also Extended Data Fig. 5). These data can therefore not be considered to be independent. The shaded areas correspond to the 95% confidence regions for the regression fit. (b) Comparison of the free GalNAc concentrations in caecal content of OO, AO and AA pigs as well as in caecal content of germ-free mice gavaged with 200mg/kg GalNAc. Concentrations were determined in freeze-dried caecal content powder using LC-MS/MS. Number of analyzed samples are given (n). Boxplots are as in Fig. 1c.
Extended Data Fig. 10 The chromosome 1 miQTL affects bacteria with a functional GalNAc import and catabolic pathway.
Presence anywhere in the genome (green), presence in close proximity to agaS (red), or absence (black) of the orthologues of 24 genes implicated in the GalNAc TR/CP pathway in the genome of (i) two OTU476 like strains (4-15-1 and 4-8-110), (ii) 248 MAGs assigned to the Erysipelotrichaceae family, and (iii) 2,863 MAGs assigned to other bacterial families. The two lanes on the right of the three panels correspond to the Regulon (red) and Pathway (green) score respectively. Both scores range from 0 (black) to 6 (bright red or green). Means (range) for the corresponding dataset are given on top. P-values (nominal, two-sided, uncorrected) of the pathway and regulon scores were computed using a linear model described in Methods.
Extended Data Fig. 11 Different GalNAc operon structure and transcriptome response in miQTL-sensitive versus -insensitive GalNAc utilizing bacteria.
Maps of GalNAc “operons” in one of the two OTU476-like strains (NB: The organization of the GalNAc gene cluster was identical in both 4-15-1 and 4-8-110 strains), and six MAGs assigned respectively to an Erysipelotrichaceae, E. coli (an Enterobacteriaceae), a Collinsella (a Coriobacteriaceae), a Fusobacteriaceae, a Firmicutes and a Clostridium. Identified Open Reading Frames (ORFs) are represented as coloured boxes. Genes implicated in GalNAc import and catabolism are in red if they are part of the cluster and in green if located elsewhere in the genome. Genes with a known function unrelated to GalNAc are in blue. ORFs with uncharacterized gene product in gray. Gene acronyms are given next to the corresponding boxes. ORFs transcribed from the top (respectively bottom) strand are above (below) the dotted line. The respective transcriptional directions are marked by the arrows. The source of information used to confirm the map order is given (finished genome, multiple MAGs, single contig).
Extended Data Fig. 12 No effect of ABO genotype on intestinal Erysipelotrichaceae abundance in human.
Volcano and QQ plots for 43 (V1-V2), 20 (V3-V4) and 9 (V5-V6) OTUs classified as Erysipelotrichaceae for the contrasts (a) [AA, AO and AB] versus [BB, BO and OO], (b) [BB, BO and AB] versus [AA, AO and OO], and (c) [OO] versus [all others]. The shaded areas correspond to the 95% confidence intervals of the spread of the QQ plot under the null hypothesis of no QTL. The actual points are always within these intervals precluding us to reject the null hypothesis. P-values (nominal, two-sided) were computed using the linear model described in Methods and hereafter. See also Supplemental discussion 3.
Supplementary information
Rights and permissions
About this article
Cite this article
Yang, H., Wu, J., Huang, X. et al. ABO genotype alters the gut microbiota by regulating GalNAc levels in pigs. Nature 606, 358–367 (2022). https://doi.org/10.1038/s41586-022-04769-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-022-04769-z
This article is cited by
-
Genomics of host–microbiome interactions in humans
Nature Reviews Genetics (2026)
-
Maternal intestinal L. vaginalis facilitates embryo implantation and survival through enhancing uterine receptivity in sows
Microbiome (2025)
-
Gut microbiota and metabolites in lipid metabolism and intramuscular fat deposition: mechanisms and implications for meat quality
Journal of Animal Science and Biotechnology (2025)
-
Deciphering the coordinated roles of the host genome, duodenal mucosal genes, and microbiota in regulating complex traits in chickens
Microbiome (2025)
-
Unraveling the composition and function of pig gut microbiome from metagenomics
Animal Microbiome (2025)