Abstract
Chronic inflammation and tissue fibrosis are common responses that worsen organ function, yet the molecular mechanisms governing their cross-talk are poorly understood. In diseased organs, stress-induced gene expression changes fuel maladaptive cell state transitions1 and pathological interaction between cellular compartments. Although chronic fibroblast activation worsens dysfunction in the lungs, liver, kidneys and heart, and exacerbates many cancers2, the stress-sensing mechanisms initiating transcriptional activation of fibroblasts are poorly understood. Here we show that conditional deletion of the transcriptional co-activator Brd4 in infiltrating Cx3cr1+ macrophages ameliorates heart failure in mice and significantly reduces fibroblast activation. Analysis of single-cell chromatin accessibility and BRD4 occupancy in vivo in Cx3cr1+ cells identified a large enhancer proximal to interleukin-1β (IL-1β, encoded by Il1b), and a series of CRISPR-based deletions revealed the precise stress-dependent regulatory element that controls Il1b expression. Secreted IL-1β activated a fibroblast RELA-dependent (also known as p65) enhancer near the transcription factor MEOX1, resulting in a profibrotic response in human cardiac fibroblasts. In vivo, antibody-mediated IL-1β neutralization improved cardiac function and tissue fibrosis in heart failure. Systemic IL-1β inhibition or targeted Il1b deletion in Cx3cr1+ cells prevented stress-induced Meox1 expression and fibroblast activation. The elucidation of BRD4-dependent cross-talk between a specific immune cell subset and fibroblasts through IL-1β reveals how inflammation drives profibrotic cell states and supports strategies that modulate this process in heart disease and other chronic inflammatory disorders featuring tissue remodelling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All source data, including sequencing reads and single-cell expression matrices have been deposited in the NCBI’s Gene Expression Omnibus under accession number GSE221699. Source data are provided with this paper.
References
Badeaux, A. I. & Shi, Y. Emerging roles for chromatin as a signal integration and storage platform. Nat. Rev. Mol. Cell Biol. 14, 211–224 (2013).
Buechler, M. B. et al. Cross-tissue organization of the fibroblast lineage. Nature 593, 575–579 (2021).
Wang, N., Wu, R., Tang, D. & Kang, R. The BET family in immunity and disease. Signal Transduct. Target. Ther. 6, 23 (2021).
Alexanian, M. et al. A transcriptional switch governs fibroblast activation in heart disease. Nature 595, 438–443 (2021).
Filippakopoulos, P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067–1073 (2010).
Anand, P. et al. BET bromodomains mediate transcriptional pause release in heart failure. Cell 154, 569–582 (2013).
Spiltoir, J. I. et al. BET acetyl-lysine binding proteins control pathological cardiac hypertrophy. J. Mol. Cell. Cardiol. 63, 175–179 (2013).
Duan, Q. et al. BET bromodomain inhibition suppresses innate inflammatory and profibrotic transcriptional networks in heart failure. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aah5084 (2017).
Antolic, A. et al. BET bromodomain proteins regulate transcriptional reprogramming in genetic dilated cardiomyopathy. JCI Insight 5, e138687 (2020).
Auguste, G. et al. BET bromodomain inhibition attenuates cardiac phenotype in myocyte-specific lamin A/C-deficient mice. J. Clin. Invest. 130, 4740–4758 (2020).
Padmanabhan, A. et al. BRD4 (bromodomain-containing protein 4) interacts with GATA4 (GATA binding protein 4) to govern mitochondrial homeostasis in adult cardiomyocytes. Circulation 142, 2338–2355 (2020).
Korf-Klingebiel, M. et al. Myeloid-derived growth factor protects against pressure overload-induced heart failure by preserving sarco/endoplasmic reticulum Ca2+-ATPase expression in cardiomyocytes. Circulation 144, 1227–1240 (2021).
Reboll, M. R. et al. EMC10 (endoplasmic reticulum membrane protein complex subunit 10) is a bone marrow-derived angiogenic growth factor promoting tissue repair after myocardial infarction. Circulation 136, 1809–1823 (2017).
Dobaczewski, M., Xia, Y., Bujak, M., Gonzalez-Quesada, C. & Frangogiannis, N. G. CCR5 signaling suppresses inflammation and reduces adverse remodeling of the infarcted heart, mediating recruitment of regulatory T cells. Am. J. Pathol. 176, 2177–2187 (2010).
Dick, S. A. et al. Self-renewing resident cardiac macrophages limit adverse remodeling following myocardial infarction. Nat. Immunol. 20, 29–39 (2019).
Dick, S. A. et al. Three tissue resident macrophage subsets coexist across organs with conserved origins and life cycles. Sci. Immunol. 7, eabf7777 (2022).
Nahrendorf, M. et al. The healing myocardium sequentially mobilizes two monocyte subsets with divergent and complementary functions. J. Exp. Med. 204, 3037–3047 (2007).
Epelman, S. et al. Embryonic and adult-derived resident cardiac macrophages are maintained through distinct mechanisms at steady state and during inflammation. Immunity 40, 91–104 (2014).
Bajpai, G. et al. Tissue resident CCR2− and CCR2+ cardiac macrophages differentially orchestrate monocyte recruitment and fate specification following myocardial injury. Circ. Res. 124, 263–278 (2019).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Pinto, A. R. et al. An abundant tissue macrophage population in the adult murine heart with a distinct alternatively-activated macrophage profile. PLoS ONE 7, e36814 (2012).
Lee, M., Lee, Y., Song, J., Lee, J. & Chang, S.-Y. Tissue-specific role of CX3CR1 expressing immune cells and their relationships with human disease. Immune Netw. 18, e5 (2018).
Patel, B. et al. CCR2+ monocyte-derived infiltrating macrophages are required for adverse cardiac remodeling during pressure overload. JACC Basic Transl. Sci. 3, 230–244 (2018).
Revelo, X. S. et al. Cardiac resident macrophages prevent fibrosis and stimulate angiogenesis. Circ. Res. 129, 1086–1101 (2021).
Stratton, M. S. et al. Dynamic chromatin targeting of BRD4 stimulates cardiac fibroblast activation. Circ. Res. 125, 662–677 (2019).
Heidt, T. et al. Differential contribution of monocytes to heart macrophages in steady-state and after myocardial infarction. Circ. Res. 115, 284–295 (2014).
Hanna, A., Humeres, C. & Frangogiannis, N. G. The role of Smad signaling cascades in cardiac fibrosis. Cell. Signal. 77, 109826 (2021).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Link, V. M. et al. Analysis of genetically diverse macrophages reveals local and domain-wide mechanisms that control transcription factor binding and function. Cell 173, 1796–1809 (2018).
Kuppe, C. et al. Spatial multi-omic map of human myocardial infarction. Nature 608, 766–777 (2022).
Liu, T., Zhang, L., Joo, D. & Sun, S.-C. NF-κB signaling in inflammation. Signal Trans. Target. Ther. 2, 1–9 (2017).
Lighthouse, J. K. et al. Exercise promotes a cardioprotective gene program in resident cardiac fibroblasts. JCI Insight 4, e92098 (2019).
Pinteaux, E. et al. Cell-specific conditional deletion of interleukin-1 (IL-1) ligands and its receptors: a new toolbox to study the role of IL-1 in health and disease. J. Mol. Med. 98, 923–930 (2020).
Frangogiannis, N. G. Transforming growth factor-β in myocardial disease. Nat. Rev. Cardiol. 19, 435–455 (2022).
Amrute, J. M. et al. Targeting immune–fibroblast cell communication in heart failure. Nature https://doi.org/10.1038/s41586-024-08008-5 (2024).
Ridker, P. M. et al. Antiinflammatory therapy with canakinumab for atherosclerotic disease. New Engl. J. Med. 377, 1119–1131 (2017).
Everett, B. M. et al. Anti-inflammatory therapy with canakinumab for the prevention of hospitalization for heart failure. Circulation 139, 1289–1299 (2019).
Rhee, A. J. & Lavine, K. J. New approaches to target inflammation in heart failure: harnessing insights from studies of immune cell diversity. Annu. Rev. Physiol. 82, 1–20 (2020).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 1073–1079 (2013).
Li, D., Wu, J., Bai, Y., Zhao, X. & Liu, L. Isolation and culture of adult mouse cardiomyocytes for cell signaling and in vitro cardiac hypertrophy. J. Vis. Exp. https://doi.org/10.3791/51357 (2014).
Bhattacharyya, S., Sathe, A. A., Bhakta, M., Xing, C. & Munshi, N. V. PAN-INTACT enables direct isolation of lineage-specific nuclei from fibrous tissues. PLoS ONE 14, e0214677 (2019).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Padmanabhan, A. et al. A genome-wide CRISPR screen identifies BRD4 as a regulator of cardiomyocyte differentiation. Nat. Cardiovasc. Res. 3, 317–331 (2024).
Zhang, J. et al. Functional cardiac fibroblasts derived from human pluripotent stem cells via second heart field progenitors. Nat. Commun. 10, 2238 (2019).
Gonzalez-Teran, B. et al. Transcription factor protein interactomes reveal genetic determinants in heart disease. Cell 185, 794–814.e30 (2022).
Vagnozzi, R. J. et al. An acute immune response underlies the benefit of cardiac stem cell therapy. Nature 577, 405–409 (2020).
Liao, Y., Smyth, G. K. & Shi, W. The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 47, e47 (2019).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Winter, G. E. et al. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science 348, 1376–1381 (2015).
Acknowledgements
We thank the Srivastava and the Alexanian laboratories for discussions; R. Micheletti, M. G. Rosenfeld, K. Pelka, G. Andreoletti and A. Wong for feedback; K. N. Ivey and J. Yang for discussions and feedback; J. Qi, D. Li and L. Sigua for providing the JQ1 molecule; J. Srivastava and all of the members of the Gladstone Flow Cytometry Core; the staff at the Gladstone Animal Facility for support with mouse colony maintenance; E. Pinteaux for providing the Il1b conditional allele; K. Claiborn and F. Chanut for editorial guidance; and A. C. Silva for help with figure editing and design. M.A. was supported by the Swiss National Science Foundation (P400PM_186704), Longevity Impetus Grant and is the Dario and Irina Sattui Investigator at Gladstone. A. Padmanabhan. was supported by the Tobacco‐Related Disease Research Program (578649), A.P. Giannini Foundation (P0527061), Michael Antonov Charitable Foundation, Sarnoff Cardiovascular Research Foundation, Frank. A. Campini Foundation and National Institutes of Health/NHLBI (K08 HL157700); T.N. by the Japan Society for the Promotion of Science Overseas Research Fellowship; R.J. by the Burroughs Wellcome Fund and by National Institutes of Health/NHLBI (R35HL166663); B.G.T. by the American Heart Association Postdoctoral Fellowship (18POST34080175) and American Heart Association/Children’s Heart Foundation Fellowship (818798); R.J.V. by NIH R01 HL169578; T.A.M. and J.G.T. by the American Heart Association (16SFRN31400013); T.A.M. by NIH R01 HL116848, NIH R01 HL147558, NIH R01 DK119594 and NIH R01 HL150225; J.G.T. by the National Institutes of Health/NHLBI (K99 HL166708); P.F.P. and K.S.P. by National Institutes of Health P01 HL098707, U01 HL098179, Gladstone Institutes and the San Simeon Fund; D.S. by the National Institutes of Health/NHLBI (P01 HL146366, R01 HL057181, R01 HL015100, R01 HL127240), Roddenberry Foundation, L.K. Whittier Foundation, Dario and Irina Sattui, Younger Family Fund and Additional Ventures; and D.S. and T.A.M. by the National Institutes of Health R01 HL127240.
Author information
Authors and Affiliations
Contributions
M.A., A. Padmanabhan. and D.S. conceived the study and interpreted the data. M.A. wrote the manuscript with input from A. Padmanabhan., S.M.H. and D.S. Y.H. performed heart surgeries and echocardiography. M.A., A. Padmanabhan., Y.H., L.Y., C.Y.L., N.S., K.A., A.Z., Y.A., I.C., C.A.E., A.J.C., M.W.C. and C.M. performed and interpreted flow cytometry analysis, and isolated cells for sc/snRNA-seq and ATAC–seq. M.A. and T.N. prepared chromium libraries. P.F.P. and K.S.P. developed computational methods for correlation and enhancer discovery analysis. M.A. analysed scRNA-seq. T.N. and P.F.P. analysed scATAC–seq. M.A., A. Pelonero., N.S. and B.G.T. performed and interpreted CUT&RUN. R.J., L.L. and Z.G. performed fibrosis analysis and quantification. A. Pelonero analysed bulk RNA-seq and CUT&RUN data. J.G.T., K.L.-C., R.J.V. and T.A.M. performed and interpreted the studies on BMDMs, collagen contractility and αSMA immunofluorescence. L.Y. generated enhancer KO clones, and performed ChIP–seq and luciferase experiments. M.A., L.Y., N.S., W.R.F. and C.K.-S.K. performed experiments on human iPSC-CFs.
Corresponding author
Ethics declarations
Competing interests
D.S. is scientific co-founder, shareholder and director of Tenaya Therapeutics. S.M.H. is an executive, officer and shareholder of Amgen and is a scientific co-founder and shareholder of Tenaya Therapeutics. T.A.M. received funding from Italfarmaco for an unrelated project. K.S.P. is a shareholder of Tenaya Therapeutics.
Peer review
Peer review information
Nature thanks Andres Hidalgo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Single-cell RNA-seq in heart failure with BET inhibition identifies a highly dynamic Cx3cr1-expressing monocyte/macrophage subpopulation.
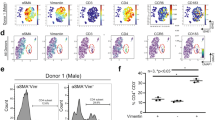
A. scRNA-seq expression Feature Plots of cell population markers in non-CM cells. B. Expression Dot Plot of the 24 most positively-correlated genes with LV ejection fraction in Sham, TAC, and TAC JQ1. C,D. UMAP plots of myeloid cells coloured by cluster (C) and sample identity (D). E. Expression Feature Plots monocyte-derived and resident macrophages in myeloid cells. F. Sample distribution within clusters in myeloid cells. G. Sample distribution of resident macrophages (defined as clusters 0,2,3,4,6) and Ccr2-positive cells (defined as clusters 1 and 5). H,I. Flow cytometry strategy (H) and number of immune cells, monocytes and macrophages (I) obtained from the ventricular tissues of TAC and TAC JQ1 at day 7 post-surgery. I, Data are mean ± s.d. Two-tailed Mann-Whitney U test.
Extended Data Fig. 2 BRD4 in infiltrated but not resident Cx3cr1-expressing cells contributes to heart failure pathogenesis.
A,B. Schematic of experimental settings of Brd4 deletion in Cx3cr1+ cells with continuous tamoxifen treatment (A), and Brd4 expression measured by qPCR in sorted CX3CR1+, fibroblasts (mEF-SK4+, CD45−, CD31−), endothelial cells (mEF-SK4−, CD45−, CD31+) and in unsorted CMs (B). C. Representative images of Sirius green staining (top) and fibrosis quantification (bottom) between TAC (red) and TAC Brd4-KO (blue). D,E. Schematic of experimental settings of pulsatile tamoxifen (TAM) treatment, which allows for Brd4 deletion specifically in Cx3cr1+ cardiac resident macrophages (D) and Brd4 expression measured by qPCR in blood and cardiac sorted CX3CR1+ cells (E). F. LV ejection fraction quantified by echocardiography in Brd4flox/flox and Cx3cr1CreERT2;Brd4flox/flox littermates. Only Cx3cr1+ resident macrophages have downregulation of Brd4 in this experiment. G. UMAP plot of CD45+ cells coloured by cluster identity. H. Expression Violin Plots of monocyte/macrophage markers in CD45+ cells. I. Expression Feature Plots of immune cell markers in CD45+ cells. J,K. Expression of Timd4 by clusters (J) and in UMAP plot (K) in monocytes/macrophages. L. UMAP plot of monocytes/macrophages coloured by cluster (left) and cluster composition within samples (right). Changes in cluster 4 are highlighted. M. Flow cytometry analysis of immune cells, monocytes and macrophages obtained from the ventricular tissues of Brd4flox/flox and Cx3cr1CreERT2;Brd4flox/flox littermates. N. Heatmap of expression depicting the top 50 genes upregulated in TAC versus TAC Brd4-KO in the monocyte/macrophage population. Data are mean ± s.e.m (B,C,E,F) and ± s.d. (M). Unpaired, two-tailed Student’s t-test (B,C,E), one-way ANOVA followed by Tukey post hoc test (F). Two-tailed Mann-Whitney U test (M).
Extended Data Fig. 3 Decreased profibrotic signature in fibroblasts with Brd4 deletion in Cx3cr1-expressing monocytes/macrophages.
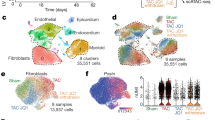
A. Expression Feature Plots of cell population markers in nuclei from cardiac tissue. B. Expression of Brd4 in TAC and TAC Brd4-KO across all clusters. C. UMAP plot of fibroblasts coloured by cluster sample identity. D. Expression Feature Plot of Brd4 in fibroblasts. E. UMAP plot of fibroblasts coloured by cluster identity. F. Expression Violin Plots of profibrotic markers across clusters in fibroblasts. G. Expression Feature Plots of profibrotic markers in fibroblasts. H. Sample distribution within fibroblast clusters. I. Heatmap of expression depicting the top 50 genes upregulated in TAC versus TAC Brd4-KO in fibroblasts. J. Heatmap indicating significance (−log10(p−val)) for indicated GO terms in genes upregulated in TAC vs TAC Brd4-KO (left, in red) or upregulated in TAC Brd4-KO vs TAC (right, in blue) in fibroblasts. K. Venn diagram comparing the genes upregulated in fibroblasts in TAC versus TAC + JQ1 (from Fig. 1 analysis) and those upregulated in TAC vs TAC Brd4-KO (from Fig. 2 analysis).
Extended Data Fig. 4 Changes in fibroblast chromatin accessibility with Brd4 deletion in Cx3cr1-expressing monocytes/macrophages.
A. Chromatin accessibility gene score of cell population markers across nuclei from cardiac tissue. B. UMAP plot (scATAC-seq) of fibroblasts coloured by cluster identity. C. Chromatin accessibility gene score of Comp, Cilp, Mfap4 and Thbs4 in fibroblasts. D. TF motif enrichment of fibroblast chromatin regions more accessible in TAC versus TAC Brd4-KO. Top 10 motifs are highlighted. A hypergeometric test was used to calculate p-values.
Extended Data Fig. 5 Identification of monocyte/macrophage population with single-cell ATAC-seq.
A. UMAP plot (scATAC-seq) of CD45+ cells coloured by cluster, the clusters encompassing the monocyte/macrophage populations are highlighted. B. Chromatin accessibility gene score of monocyte/macrophage markers in CD45+ cells. C. UMAP plot (scATAC-seq) of monocytes/macrophages coloured by cluster identity. D. Monocyte/macrophage cluster distribution within samples.
Extended Data Fig. 6 Chromatin accessibility and BRD4 occupancy in Cx3cr1-expressing cells identifies set of highly dynamic monocyte/macrophage distal elements in heart failure.
A. Distribution of accessibility in monocytes/macrophages in the TAC state identifies a class of distal regions (super-enhancers (SE)) for which the accessibility falls over the inflection point of the curve. B. Volcano plot depicting p65/RELA binding in all differentially expressed genes between TAC and TAC Brd4-KO (Log2FC > 0.125 or Log2FC < −0.125; FDR < 0.05) in monocytes/macrophages. C. Schematic of the targeting strategy to generate the Brd4 3xFlag mouse. D. Western blot showing WT and 3xFLAG knock-in bands in WT and Brd4flag/+ animals. E. Liver tissue western blot showing expression of endogenous long and short BRD4 isoforms treated with vehicle or with the small-molecule BET protein degrader dBET152. F. Coverage from anti-FLAG Cut&Run in Sham and TAC identifies regions enriched with BRD4 in stress (left) or at baseline (right). G. Genes that are upregulated in TAC Brd4-KO vs TAC in monocytes/macrophages (from Fig. 2 analysis) categorized by BRD4 binding in TAC. H. Examples of BRD4-dependent genes that display dynamic BRD4 recruitment in TAC in sorted Cx3cr1-positive cells. I. LogFC of BRD4 binding by CUT&RUN in TAC vs Sham at the super enhancers identified in the monocyte/macrophage population. J. Coverage from anti-FLAG CUT&RUN in Sham and TAC in selected ten super enhancer regions.
Extended Data Fig. 7 BRD4-dependent regulation of the Il1b super enhancer in heart failure.
A. Schematic of the Il1b super enhancer locus displaying the 7 distal regions (Peak1 to Peak7) called as accessible peaks in scATAC-seq in monocytes/macrophages. B. Chromatin accessibility trend between samples (mean and 95% confidence interval) in all identified Il1b super-enhancer peaks. C. Schematic of the Il1b locus displaying the location of the gRNAs used to delete the 7 distal regions (Peak1 to Peak7). D,E. Agarose gel electrophoresis to assess distal peak deletions in Il1b locus (D) and unaffected region around Il1b promoter (E) (1813bp) in WT and KO clones.
Extended Data Fig. 8 Chromatin accessibility in human myeloid cells.
A. Chromatin accessibility gene score of TIMD4, LYVE1 and CCR2 in human cardiac myeloid cells30. B. Dot Plot indicating significance (−log10(p−val)) for indicated GO terms across human myeloid cell clusters.
Extended Data Fig. 9 IL1B synergizes with TGFB to drive profibrotic function of human induced pluripotent stem cell (iPSC) derived cardiac fibroblast.
A. UMAP plot (snRNA-seq) of nuclei from cardiac tissue coloured by cluster identity. B. Expression Feature Plot of Il1r1 in nuclei from cardiac tissue. Fibroblasts are highlighted. C. Protocol to generate human induced pluripotent cardiac fibroblasts (iPSC-CFs). D. Immunofluorescence staining of αSMA (left) in Unstimulated iPSC-CFs or treated with IL1B, TG-B or TGFB + IL1B. Nuclei are marked by Hoechst. Scale bars, 200 μm. Right, quantification of αSMA staining. E. Images (left) and quantification (right) of iPSC-CFs seeded on compressible collagen gel matrices in unstimulated or with IL1B, TGFB or TGFB + IL1B treatments. F. IL1R1 expression by qPCR in Unstimulated human iPSC-CFs with control or IL1R1-targeting siRNAs. G–J. Representative images (G) and quantification of human iPSC-CFs seeded on compressible collagen gel matrices comparing unstimulated cells with IL1B-treated (H), TGFB-treated (I) or TGFB + IL1B-treated (J) using a control or IL1R1-targeting siRNAs. K. Il1b expression by qPCR in Unstimulated and LPS treated RAW264-7 macrophages. L. MEOX1 expression by qPCR in Unstimulated human iPSC-CFs with control or MEOX1-targeting siRNAs. M–O. Representative images (M) and quantification of human iPSC-CFs seeded on compressible collagen gel matrices comparing unstimulated cells with IL1B-treated (N) and TGFB-treated (O) using a control or MEOX1-targeting siRNAs. P. Human MEOX1 locus showing H3K27Ac in unstimulated iPS-CFs. The syntenic region of the mouse Meox1 Peak9/10 regulatory element4 is highlighted and the six p65/RELA motifs within the region indicated. Q. MEOX1 expression by qPCR in Unstimulated iPSC-CFs or treated with IL1B, TGFB, or TGFB + IL1B with vehicle or JQ1. R. p65/RELA expression by qPCR in Unstimulated iPSC-CFs or treated with IL1B, TGFB or TGFB + IL1B with control or p65/RELA-targeting siRNAs. D-F,H-L,N,O,Q,R. Data are mean ± s.e.m. One-way (D,Q,R) and two-way (E,H-J,N,O) ANOVA followed by Tukey post hoc test. Unpaired, two-tailed Student’s t-test (F,K,L).
Extended Data Fig. 10 In vivo antibody-mediated IL1B neutralization decreases tissue fibrosis and profibrotic gene expression in fibroblasts.
A. Representative images of Sirius green staining (left) and fibrosis quantification (right) between IgG Ab (red) and Anti-IL1B Ab (blue). B. UMAP plot of non-cardiomyocyte cells coloured by cluster. C. Expression Feature Plots of cell population markers in non-cardiomyocyte cell population. D. Expression Feature Plots of Comp, Cilp, Mfap4 and Thbs4 in fibroblasts. E. Expression Violin Plots of Comp, Cilp, Mfap4 and Thbs4 across clusters in fibroblasts. F. Heatmap indicating significance (−log10(p−val)) of top GO terms for each fibroblast cluster. G. Expression Violin Plots of Comp, Cilp, Mfap4 and Thbs4 across samples in fibroblasts. H. Heatmap of expression of the top 50 genes upregulated in TAC IgG Ab versus TAC Anti-IL1B Ab in fibroblasts. I. Heatmap indicating significance (−log10(p−val)) for indicated GO terms in genes upregulated in IgG vs Anti-IL1B (red) or upregulated in Anti-IL1B vs IgG (blue) in fibroblasts. J. Heatmap indicating significance (−log10(p−val)) for indicated GO in genes upregulated or downregulated in fibroblasts clusters 3&5 vs all other clusters. A, Data are mean ± s.e.m. Unpaired, two-tailed Student’s t-test.
Extended Data Fig. 11 Decreased profibrotic signature in fibroblasts with Il1b deletion in Cx3cr1-expressing myeloid cells.
A. Schematic of experimental settings for scRNA-seq from sorted cardiac fibroblasts at day 10 post TAC of Il1bflox/flox (TAC) and Cx3cr1CreERT2;Il1bflox/flox (TAC Il1b-KO) littermates (left). Il1b expression measured by qPCR in cardiac sorted CX3CR1+ cells (right). B. UMAP plot of fibroblast, endothelial, pericytes and smooth muscle cell markers in sorted fibroblasts at day 10 post TAC. C. Expression Violin Plots of profibrotic markers across clusters in sorted fibroblasts. D. Heatmap indicating significance (−log10(p−val)) of top GO terms for each fibroblast cluster. E. Expression Feature Plots of profibrotic markers in sorted fibroblasts. F. Expression Violin Plots profibrotic markers across samples in sorted fibroblasts. G. Heatmap of expression of the top 50 genes upregulated in TAC versus TAC Il1b-KO in sorted fibroblasts. H. Heatmap indicating significance (−log10(p−val)) for indicated GO terms in genes upregulated in TAC vs TAC Il1b-KO (red) or upregulated in TAC Il1b-KO vs TAC (blue) in sorted fibroblasts. A, Data are mean ± s.e.m. Unpaired, two-tailed Student’s t-test.
Extended Data Fig. 12 Comparison of cardiac fibroblast transcription with systemic IL1B inhibition and Il1b deletion in Cx3cr1-expressing cells after TAC.
A,B. UMAP plot generated by merging the fibroblasts from TAC IgG/TAC-Anti IL1B and TAC/TAC Il1b-KO datasets displaying clusters (A) and sample composition (B). C. Sample distribution within clusters in fibroblasts. D,E. Expression Feature plot (D) and violin plot across clusters (E) of Postn and Meox1 expression. F. Heatmap indicating significance (−log10(p−val)) of top GO terms for each fibroblast cluster. G. Sample distribution of fibroblasts categorized as transcriptionally positive or negative for Postn and Meox1.
Supplementary information
Supplementary Tables (download ZIP )
Supplementary Tables 1–13.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Alexanian, M., Padmanabhan, A., Nishino, T. et al. Chromatin remodelling drives immune cell–fibroblast communication in heart failure. Nature 635, 434–443 (2024). https://doi.org/10.1038/s41586-024-08085-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-024-08085-6
This article is cited by
-
Design of RGD-functionalized GSH-responsive pegylated polymeric protacs for selective BRD4 degradation and EndMT-driven cardiac fibrosis inhibition
Journal of Nanobiotechnology (2026)
-
Cardiac epigenome in heart development and disease
Nature Reviews Cardiology (2026)
-
Location and aetiology are determinants of fibroblast activation and heterogeneity in the failing human heart
Genome Medicine (2025)
-
The role of periodontitis as a modifier of diabetes mellitus in older patients with atrial fibrillation
BMC Endocrine Disorders (2025)
-
CD248+ fibroblasts drive cardiac fibrosis by interacting with immune cells
Nature Cardiovascular Research (2025)