Abstract
The mammalian fatty acid synthase (FASN) enzyme is a dynamic multienzyme that belongs to the megasynthase family. In mammals, a single gene encodes six catalytically active domains and a flexibly tethered acyl carrier protein (ACP) domain that shuttles intermediates between active sites for fatty acid biosynthesis1. FASN is an essential enzyme in mammalian development through the role that fatty acids have in membrane formation, energy storage, cell signalling and protein modifications. Thus, FASN is a promising target for treatment of a large variety of diseases including cancer, metabolic dysfunction-associated fatty liver disease, and viral and parasite infections2,3. The multi-faceted mechanism of FASN and the dynamic nature of the protein, in particular of the ACP, have made it challenging to understand at the molecular level. Here we report cryo-electron microscopy structures of human FASN in a multitude of conformational states with NADPH and NADP+ plus acetoacetyl-CoA present, including structures with the ACP stalled at the dehydratase (DH) and enoyl-reductase (ER) domains. We show that FASN activity in vitro and de novo lipogenesis in cells is inhibited by mutations at the ACP–DH and ACP–ER interfaces. Together, these studies provide new molecular insights into the dynamic nature of FASN and the ACP shuttling mechanism, with implications for developing improved FASN-targeted therapeutics.
Similar content being viewed by others
Main
De novo lipogenesis (DNL) is essential in mammalian development to produce fatty acids for membrane formation, energy storage, cell signalling and protein modifications2. During diseases such as cancer and cancer metastasis, metabolic dysfunction-associated steatotic liver disease (MASLD) and viral and parasite infections, DNL is often upregulated along with the proteins involved in the pathway3,4. FASN is the primary enzyme in DNL that condenses cytosolic acetyl-CoA and malonyl-CoA into the 16-carbon saturated fatty acid palmitate1. The current leading inhibitor of FASN (denifanstat) is in various clinical trials (including phases I (NCT05835180), II (NCT03032484) and III (NCT05118776)) and has thus far shown promising clinical results for treatment of MASLD, cancer and acne vulgaris5,6. Many of the preclinical studies focused on targeting FASN to inhibit DNL and treat metabolic diseases have shown high efficacy; however, further studies have found that in certain contexts, full loss of DNL can cause unwanted effects such as decreased platelet production7 and learning deficits8, thus suggesting that further studies on the FASN mechanism and its role in health and disease are needed to harness the therapeutic potential of targeting FASN safely and effectively.
In mammals, the FASN enzyme is a large type I megasynthase that has six catalytic domains and a flexibly tethered ACP domain that shuttles reaction intermediates among them (Fig. 1a,b). A related family of megasynthases, the polyketide synthases (PKSs), also use a carrier protein system that shuttles covalently linked reaction intermediates between catalytic domains to coordinately synthesize a diverse set of biologically important compounds9. The PKS family includes a distinct set of genes across organisms, with type I iterative PKSs, and an exciting more recently characterized class of animal FASN-like PKSs10, showing the highest homology to mammalian FASN, even higher than yeast FASN enzymes11,12. In addition, the PKS family of enzymes synthesize an assortment of important natural products, including antibiotics, chemotherapy drugs and statins9. Current studies on PKSs have been working to harness the distinct chemistry across the family to create novel designer megasynthases that can synthesize a more diverse set of small molecules to expand compound libraries and discover new therapeutics13. These efforts have expanded to using domains from mammalian FASN to allow for incorporation of non-canonical starter and extender units, as the malonyl–acetyl transferase (MAT) domain of mammalian FASN has been shown to have the most promiscuous substrate specificity for loading the ACP in a megasynthase thus far14,15. Therefore, gaining functional insights into the molecular mechanism by which the ACP interacts with the various domains will provide important knowledge for ongoing work towards domain swapping parts of FASN with PKSs to produce novel compounds15,16.
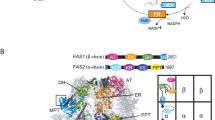
a, Diagram of the domains encoded in the human FASN gene. b, Schematic diagram of the ACP interactions, intermediates and movements during the mammalian FASN mechanism. c, Composite cryo-EM density of state 1 from the hFASN–NADPH dataset coloured by the corresponding domain. The colouring of each individual domain is kept consistent throughout the article unless otherwise stated.
Crystal structures of pig FASN have been determined to 4.5 Å (ref. 17) and 3.2 Å (ref. 18), elucidating an unexpected x-shaped domain architecture that diverges significantly from the more highly studied eukaryotic yeast FASN19,20. These structures and detailed biochemical studies21,22,23,24 provide a model for how mammalian FASN synthesizes palmitate from short-chain acyl-CoAs1. However, attempts to understand the ACP mechanism have been hindered by the transient nature of ACP interactions at the catalytic domains and technological limitations to capture the dynamic landscape of mammalian FASN25,26. Current models for the synthesis of palmitate by FASN propose that a combination of large structural rearrangements, and transient but specific domain–domain interactions (DDIs) made by the ACP domain, cooperate to shuttle intermediates to the appropriate catalytic domains. To our knowledge, this model, however, has not been resolved in a mammalian system26.
Here we used single-particle analysis cryo-electron microscopy (cryo-EM) to determine structures of full-length human FASN (hFASN) with the cofactor NADPH in various conformational states, and with NADP+ and acetoacetyl-CoA present. Our structures also resolve the ACP stalled at the DH and ER domains. Using in vitro biochemistry, we show that FASN activity can be inhibited by mutations at the ACP–DH and ACP–ER interfaces, thus providing new molecular insights into the ACP shuttling mechanism. Evaluations of these same mutants in mammalian cells reveal a defect in DNL and mimics FASN loss or small-molecule inhibition. Together, this work provides new molecular insights into the complex mechanism used by the multi-enzyme FASN to coordinate various active sites for palmitate biosynthesis, as well as a framework for improving next-generation FASN-targeted therapeutics.
FASN shows continuous heterogeneity
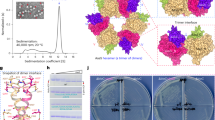
To facilitate a structure–function analysis of FASN, we prepared recombinant hFASN in insect cells for structural studies and mouse FASN (mFASN) in bacteria for in vitro biochemical work. The specific activity of mFASN had a 95% confidence interval of 292–339 nmol min−1 mg−1 for oxidation of NADPH. When titrating acetyl-CoA and malonyl-CoA in a 1:1 ratio up to 1 mM, we were able to observe a clear dose–response curve with no substrate inhibition, which is commonly seen with FASN when only one of the CoA derivatives is varied14 (Extended Data Fig. 1). The curve best fit to an allosteric sigmoidal model with a hill slope of 1.5, which suggests positive cooperativity. This has previously been reported for the β-ketosynthase (KS) domain when investigating the condensation reaction, but did not account for ACP shuttling in the context of a full reaction cycle27. Both proteins formed stable homodimers in solution (Extended Data Fig. 1). We mixed hFASN with NADPH (hFASN–NADPH) for single-particle analysis cryo-EM structure determination and obtained structures from the same cryo-EM sample in various conformations at 3.2–3.8 Å overall resolutions (Fig. 1c, Extended Data Table 1, Extended Data Figs. 2 and 3 and Supplementary Video 1). Similar to the crystal structure of pig FASN18, hFASN forms an x-shaped homodimer mediated by the KS, DH and ER domains. The structures provide strong cryo-EM density for five of the six catalytic domains (the KS, MAT, β-ketoreductase (KR), DH and ER domains), and the two pseudodomains (pseudo-KR (ΨKR) and pseudo-methyl transferase (ΨME)) in all conformations. In addition, we were able to resolve the ACP domain at the DH and ER domains in the modifying portion of hFASN with particle sets that share particles with each of the conformations. We were unable to resolve the C-terminal thioesterase domain in any of the structures, probably due to the flexible linker between the ACP and thioesterase domains and lack of stable interactions with other domains in the core of hFASN.
From a single dataset, we resolved the structure of hFASN in eight distinct conformational states, demonstrating large movements between the modifying (KR, DH, ER, ΨKR and ΨME domains) and condensing (MAT and KS domains) regions (Fig. 2 and Extended Data Figs. 2 and 3). The dimer forms two reaction chambers that each contain the entire catalytic machinery for synthesizing palmitate (Extended Data Fig. 4). We have designated our structure like the pig crystal structure as state 1 based on alignment to the deposited structure (Protein Data Bank (PDB) ID: 2VZ9), and subsequently numbered each conformation up to state 8 based on the swinging or rotation from state 1 (Extended Data Figs. 3 and 4). These conformations largely agree with the previous negative-stain electron microscopy study on the rat FASN homologue28. Except, in the negative-stain electron microscopy study, the authors reported large rearrangements in the modifying portion of rat FASN, but we do not see any major conformational changes in the modifying region or condensing regions relative to each other in any conformation lacking ACP density, with overall root mean square deviation < 0.6 Å for the modifying regions of states 2–8 relative to state 1, and a root mean square deviation < 0.5 Å for the condensing regions. The high agreement of the local structural elements suggests that the modifying and condensing regions act as rigid bodies that move relative to each other, but this motion is not allosterically controlled by rearrangements in the domains. We failed to resolve the glycine-serine-rich linker between the condensing and modifying portions of FASN, probably because of the inherent flexibility in the region that allows for the large rearrangements in solution. This prevented us from unambiguously identifying which KS domain is linked to which DH domain in the dimer, but previous biochemistry suggests that the MAT and KS domains can interact with the ACP in the same or opposite monomers24. We have denoted them as enclosing the same reaction chamber for clarity. The modifying and condensing regions were refined separately to obtain higher-resolution structures. The need to treat them as separate bodies to align to higher resolution suggests that the motion is continuous, and the sorted conformations represent snapshots along the trajectory. The eight conformations reported here sufficiently describe the motion, but do not show all possible conformations. Starting with state 1, states 1–4 represent a swinging motion of the condensing region relative to the modifying region all in the same plane, whereas states 5–8 represent a rotation out of this plane by the MAT of the condensing region. Comparison of the largest identified movements, states 1–3 show that a modest 5° swing moves the condensing region between the states, but comparing states 1–8 shows that the condensing region can rotate up to 90° relative to the modifying region, meaning that our reconstructions probably represent the 180° rotations of the condensing region relative to the modifying region that allow for the ACP to function with the KS and MAT domains from either monomer (reaction chamber)24 (Fig. 2). Together, these findings highlight the continuous motion between the rigid condensing and modifying regions of hFASN that the ACP coordinates fatty acid synthesis between.
a, Composite cryo-EM density of hFASN–NADPH in three states that highlight the swinging motion and rotation of the condensing region relative to the modifying region. b, Aligned models of state 1 and state 3 showing the modest 5° swinging in the condensing region that moves the MAT domains relative to the modifying region. c, Aligned models of state 1 and state 8 showing the large rotation in the condensing region that moves the MAT domains out of the plane with the modifying region by 90°.
NADPH oxidation state drives binding
In the structures of hFASN with NADPH, there is strong density for NADPH at the KR domains (Fig. 3a). The catalytic residues K1995, S2021 and Y2034 all sit near the nicotinamide ring on the NADPH that provides the hydride for reducing the keto group on the ACP (Fig. 3a). As we were unsuccessful in resolving the ACP at the KR domain in our original dataset, we also collected a dataset with acetoacetyl-CoA, to target the ACP to the KR domain, and NADP+ (hFASN–NADP+) to prevent turnover. Although we were unsuccessful in resolving the ACP at the KR domain, we determined structures of hFASN with a portion of NADP+ bound to the KR domain (Fig. 3 and Extended Data Fig. 5). In the crystal structure of pig FASN18, the amino acids 2067–2080 in a loop near the NADPH-binding site were ordered by the addition of NADP+ to the sample, but we did not resolve this loop in any of our structures (Fig. 3b). Structures of the modifying region of hFASN with NADPH and inhibitors that target the KR domain stabilize this loop and have suggested that it has a role in ACP binding, but its role remains elusive until the ACP–KR structure is elucidated29,30. The NADP+ molecules at the KR domain in the structure only resolve the adenine base, ribose ring and phosphate groups, and lack cryo-EM density for the nicotinamide ring (Fig. 3c). We observed no major structural rearrangements in the KR domain after the NADPH was oxidized (Fig. 3b). This suggests that the structural changes in the nicotinamide ring after oxidation are what drive cofactor release, as the NADPH-bound structure represents the priming of the KR domain for reducing the β-keto-ACP, and the NADP+-bound structure represents the release of NADP+ following catalysis.
Models of the hFASN modifying region bound to NADPH (centre top; NADPH in pink) and NADP+ (centre bottom; NADP+ in salmon) highlight the importance of the oxidation state and nicotinamide ring in facilitating binding at the KR and ER domains. a, NADPH bound to the KR domain (yellow) of hFASN with the nicotinamide ring near the known catalytic residues K1995, S2021 and Y2034. b, Comparison of the NADPH-bound and NADP+-bound KR domains from the two datasets. c, Cartoon representation of the KR domain (yellow) bound to NADP+. d, Atomic model of NADPH bound to the ER domain, with the nicotinamide ring at the active site near residues T1654, K1771, L1796 and D1797. e, Comparison of the ER domain when bound to NADPH (dark green) or no cofactor (dark grey). f, ER domain without cofactor bound.
In addition, the hFASN–NADP+ structure shows no density for NADP+ at the other NADPH-dependent active site in the ER domain, but all structures from the hFASN–NADPH dataset are fully occupied (Fig. 3d,f). NADPH binds to the ER domain with the nicotinamide ring near the catalytic residues T1654, K1771, L1796 and D1797 (Fig. 3d). The structures with NADPH bound to the ER domains and the ER domain without cofactor in the presence of NADP+ show no major structural changes in the ER domain (Fig. 3e), further implicating that the oxidation state and structural change in the nicotinamide ring of NADP(H) are what drives binding and release of the cofactor.
ACP shuttling revealed at the DH domain
In the hFASN–NADPH and hFASN–NADP+ datasets, we observed additional density at the DH domain of hFASN, which could be confidently modelled as the ACP domain (Fig. 4a,b). After 3D classification without alignments on the region, we obtained particle sets of approximately 10% of the final particles that had clear density at the DH domain when NADPH was present, and 18% of particles with NADP+ present (Extended Data Figs. 2 and 5). Although we do not have definitive evidence for the difference in the percentage of overall particles with the ACP stalled at the DH domain between the datasets, we hypothesized that the lack of turnover at the NADPH-dependent domains may have stalled more molecules primed for interactions with the DH domain. Focused refinement resulted in reconstructions of the modifying portion of hFASN with ACP density at nominal resolutions of 3.2 Å and 3.5 Å for the hFASN–NADPH and hFASN–NADP+ datasets, respectively. These structures show no major differences in the DH domain or the interface residues at the ACP–DH interface, and the overall maps agree well with a correlation of 0.97. Significantly, the phosphopantetheinyl (Ppant) arm on S2156 of the ACP domain is resolved and can be traced to the known catalytic residue in the DH domain H878 (Fig. 4c,d). Neither of the Ppant arms show signs of acylation, but this may be averaged out from containing a spectrum of intermediate chains, as we did not synchronize the ACP state in hopes of targeting it to each catalytic domain, or they may lack any acylation.
a, Cryo-EM density of the modifying region of hFASN with the ACP at the DH domain with NADPH bound to the KR and ER domains. b, Cryo-EM density of the modifying region of hFASN with the ACP at the DH domain with NADP+ bound to the KR domain. c, Model of the hFASN–NADPH ACP Ppant arm in the DH active site leading to H878 with the corresponding density. d, Model of the hFASN–NADP+ ACP Ppant arm in the DH active site leading to H878 with the corresponding density. e, Electrostatic surface potential of the ACP–DH interaction highlighting the charge complementarity. f, Interaction interface of the ACP and DH domain showing side chains of residues forming interactions. g, Overall in vitro activity of mFASN mutants affecting ACP–DH interactions compared with wild type (WT). The figure is numbered according to the human sequence, but correspond to residues R883, I924, R1106, L2151, M2152 and V2154 in the mouse protein. Data are shown as the average of n = 3 replicates, with error bars denoting s.d. h, Measurement of labelled palmitate in D42 FASN-knockout (KO) hepatocytes reconstituted with WT hFASN or indicated mutants, given [U-13C]-glucose for 48 h, shown as percent carbon enrichment. Data are presented as an average of n = 3 biological replicates, and error bars denote s.d.
We can confidently trace the backbone and assign side chains throughout hFASN from our structures, but we were unable to confidently model many side chain rotamers at the interaction interface. Work on other ACP interactions has suggested that electrostatic charge complementarity is the driving factor for ACP interactions, so we searched the models for potential electrostatically matching patches. We found a negative patch on the ACP that interacts with a positive patch on the DH interface (Fig. 4e). Of note, charged residue R883 and the uncharged residue I924 on the DH domain and R1105 on the DH–ΨME linker are in proximity to the ACP when it is near the DH domain active site (Fig. 4f). On the basis of our model, these residues may interact with L2157, V2160 and M2158 on helix II of the ACP, respectively. We mutated the corresponding residues in the mFASN homologue and tested activity of the mutant enzymes in vitro by monitoring NADPH consumption. Single-point mutations R883A and I924A decreased overall FASN activity to approximately 30% and 40% of the wild-type level, respectively (Fig. 4g). A charge flipped R883E mutation further reduced the enzyme activity to only 20% of wild-type activity (Fig. 4g). Similarly, the R1105A mutant retained only 45% of wild-type activity, suggesting that the linker region to the catalytically incompetent ΨME domain also has a role in the ACP shuttling mechanism. The decreased activity in the alanine and charge flipped mutations suggest that a combination of hydrophobic, steric and charge complementarity at the ACP–DH interface is important for proper intermediate shuttling. We cannot rule out the possibility of certain degrees of freedom in the interaction that would allow for read-through when mutations are made as has been previously suggested for PKS enzymes31. Although the structures show interactions with L2157, this residue also lies in the consensus sequence for phosphopantetheinyl transferase (PPTase) enzymes32. Even though we used a promiscuous PPTase, we checked the phosphopantetheinylation status of the recombinant wild-type and L2157A mFASN by mass spectrometry and found that mutations to L2157 inhibit proper loading of the Ppant arm onto the ACP (Supplementary Table 1). The lack of the Ppant arm makes the loss of activity independent of ACP interactions at the domains, but highlights the essential nature of the Ppant arm for FASN function as the mutated enzyme retains essentially no activity. Mutations to the ACP interaction partners of the residues on the catalytic domain mutants M2158A and V2160A had little effect on overall FASN activity, with the M2158A and V2160A mutants both retaining 80% activity. The ACP must interact with each catalytic domain during palmitate synthesis, so the difference in activity is probably compounded by the interactions made with domains other than the DH. Together, these results suggest that the structurally observed ACP interaction interface on the DH domain is important for proper delivery of intermediates to the active site by the ACP in mammalian FASN.
To further evaluate the functional consequence of disrupted ACP shuttling and diminished FASN activity, we generated Fasn-knockout mouse hepatocellular carcinoma cells33 (D42-knockout cells; Extended Data Fig. 6) and re-expressed wild-type and mutant hFASN. The hepatocellular carcinoma cells have high levels of baseline DNL in culture and tolerate the knockout of Fasn, which allowed us to investigate FASN activity in a cellular context (Extended Data Fig. 6). Re-expression of hFASN wild type and mutants in the D42-knockout cells was confirmed by western blotting, and all mutants re-expressed at comparable levels (Extended Data Fig. 6). To measure DNL, cells were incubated with uniformly labelled carbon-13 glucose ([U-13C]-glucose) for 48 h, and palmitate labelling was measured by gas chromatography–mass spectrometry (GC–MS). We first measured the contribution of glucose to palmitate in the parental D42 cells and the hFASN-knockout cells and found that 40% of total palmitate carbons were derived from glucose in the parental cell line, but less than 1% of palmitate was labelled in the knockout cells, confirming that the knockout of FASN fully impaired the ability of the cells to synthesize palmitate (Extended Data Fig. 6b). We then compared the DNL level in the cells that re-expressed the wild-type and mutant hFASN. The expression of wild-type hFASN increased the percent enrichment of palmitate carbons from glucose to 14%; in agreement with our in vitro activity assays, the R883A, R883E and L2157A mutants had less than 1% enrichment in palmitate, like the knockout cells (Fig. 4h). In addition, the I924A and R1105A mutants rescued DNL to only 3% and 6% palmitate-labelled carbon enrichment, respectively, in line with the in vitro activity of each respective mutant (Fig. 4g,h). No significant differences were detected in saponified palmitate abundance between conditions (Extended Data Fig. 6c). Analysis of the isotopologue enrichment further supports that the wild-type cells are synthesizing palmitate from glucose, whereas the other cell lines all show little to no palmitate labelling (Extended Data Fig. 6d). This data further support the need for proper DDIs by the ACP to synthesize palmitate in a cellular context and suggest that the ACP-binding interface on the DH domain is a potentially targetable site for FASN inhibition.
ACP shuttling revealed at the ER domain
From the hFASN–NADPH dataset, we were also able to resolve the ACP docked at the NADPH-dependent ER domain to a nominal resolution of 3.2 Å (Fig. 5a). Even with extensive classification, we were unable to resolve the ACP at the ER domain in the hFASN–NADP+ dataset, potentially because the NADPH interacts with the Ppant arm (Fig. 5b). In the cryo-EM density, we could trace the Ppant arm into the ER domain active site, ending near the nicotinamide ring (Fig. 5b). In the structure of the ACP bound at the ER domain, the adenine ring of NADPH moves relative to the ACP-free structure, and residues 1573–1599 in the ER domain at the bottom of the active site move towards the Ppant arm (Fig. 5c). The residues of the ΨKR domain that interact with this region of the ER domain, 1535–1544, also shift towards the Ppant arm. We identified residues M1577, T1580, K1582 and R1841 on the ER domain that appear to be forming interactions with the Ppant arm or ACP itself (Fig. 5d). We made alanine mutations of these residues in mFASN or the corresponding interacting residues on the ACP (H2133 and E2161) and tested their effect on overall activity in vitro (Fig. 5e). Mutations of ER domain M1577A, which interacts with the Ppant arm and is shifted upon ACP binding, impaired overall FASN activity to roughly 25% of wild type. Similarly, an R1841A mutation at the surface of the ER domain, where the ACP interacts, reduced activity to 20% of wild type and was the only mutant at the ACP–ER interface that significantly decreased activity. R1841 does not appear to interact with helix II of the ACP, which is believed to be the predominant site of DDIs made by the ACP in other systems34 (Fig. 5d). This suggests that more than just helix II interactions are important for proper intermediate shuttling in mammalian FASN.
a, Cryo-EM map of the modifying region of hFASN with the ACP interacting with the ER domain. b, Model of the hFASN–NADPH ACP Ppant group and ER-bound NADPH with the corresponding densities. c, Comparison of the ER domain and bound NADPH molecule when the ACP is interacting (dark green) or not present (dark grey), showing the shift in the NADPH adenine base and ER loop. d, Interaction of the ER and ACP with the Ppant arm in the active site, highlighting potential side chain interactions. e, In vitro activity of mFASN mutants that were predicted to interrupt the interaction of the ACP with the ER domain. Data are shown as mean ± s.d. of n = 3 replicates. The figure is numbered according to the human sequence, with the residues corresponding to M1570, T1573, K1575, R1834, H2127 and E2155 in the mouse protein. f, Palmitate percent carbon labelling in FASN-knockout D42 hepatocellular carcinoma cells, reconstituted with hFASN WT and indicated mutants, given [U-13C]-glucose for 48 h. Data are plotted as the average, with error bars denoting the s.d. of n = 3 replicates. The WT and knockout data are equivalent to Fig. 4h but shown again for comparison.
Using the D42 FASN-knockout cells that we generated, we also tested M1577A-mutant and R1841A-mutant activity. Again, the hFASN mutants that perturbed the ACP–ER interaction and diminished mFASN activity in vitro prevented DNL in cells, with less than 1% palmitate carbon enrichment from 13C-glucose in both the M1577A-mutant and the R1841A-mutant cells (Fig. 5e). These findings further support the model that disrupting DDIs of the ACP with the catalytic domains prevent palmitate synthesis and mimics FASN loss or small-molecule inhibition.
FASN undergoes conformational dynamics
On the basis of the previous negative-stain electron microscopy structures of rat FASN at roughly 20 Å resolution28, the authors proposed that the conformational rearrangements made between the modifying and condensing regions drive asymmetry across the two reaction chambers, thus driving asynchrony in the FASN reaction cycle across the homodimer. Analysis of our structures extends these findings by visualizing the conformational states with the ACP at key steps. Although it might be hypothesized that each conformation in FASN may correlate with an individual ACP interaction and step in the synthesis cycle, our structures do not align with this model. We instead propose that the conformational dynamics provide a regulatory role in the hFASN condensing steps of the reaction cycle, as state 1 mimics the crystal structure, with the condensing region (MAT and KS domains) in range of the ACP, whereas swinging motions between states 1–4 cause asymmetry in which the MAT domain is accessible at reaction chamber 1 or chamber 2 exclusively (Extended Data Fig. 4). This suggests that ACP interactions are conformation dependent in hFASN in the condensing region. Our structures with the ACP at the DH and ER domains indicate, however, that the ACP interactions in the modifying portion do not correlate with the conformational state of the condensing region in relation to the modifying region. In support of this, in our consensus refinements of particles with the ACP at the DH or ER domains, we did not observe density for the MAT domains, and observed low-resolution density for the KS domain. Heterogenous refinement of the particles into eight classes that are seeded with a single initial model resulted in eight volumes with the condensing region in conformational states such as states 1–8; in all states, the ACP remains resolved at the modifying region in only one reaction chamber (Extended Data Fig. 7). This suggests that the ACPs in each reaction chamber work asynchronously, meaning the ACPs do not interact with the equivalent catalytic domain, nor are they necessarily loaded with the same intermediate at the same time, in each protomer.
Discussion
Here we have determined the structure of hFASN in multiple conformational states using cryo-EM, resolved the ACP in active conformations for the first time in a mammalian system at the DH and ER domains, and our structures support a model in which ACP–domain interactions occur independently of the conformational state.
We identified mutations at the ACP interface with the DH and ER domains that significantly decreased overall FASN activity both in vitro and in cells. This offers a new potential way to target FASN with small molecules that avoid the conserved active site regions. We observed that mutations to the DDI interface on the catalytic domain were more deleterious to overall FASN activity in vitro than the corresponding mutations on the ACP. This imbalance suggests that engineering the interaction interface on the catalytic domains in designer PKSs may be of greater importance than mutations on the ACP to improve product output.
In most prokaryotes, the catalytic enzymes for fatty acid synthesis are expressed as individual proteins, termed a type II system. This evolved into the megasynthases found in yeast and mammals that have the catalytic machinery in large complexes or type I systems. Yeast FASN expresses an α-chain and β-chain as separate genes that assemble into a large dodecameric cage-like structure that has six sets of catalytic domains for fatty acid synthesis. Although the architecture of each system varies greatly, the catalytic mechanism of each domain is largely conserved, and they all use ACPs for intermediate shuttling. The yeast system has an additional structural region linked to the canonical four α-helical bundle ACP (Extended Data Fig. 8a). Although our structures of hFASN revealed that the ACP–DH interaction is mostly mediated by helix II of the ACP and involves approximately 730 Å2 of buried solvent-accessible surface area, the Saccharomyces cerevisiae (yFASN) ACP–DH interaction involves approximately only 48 Å2 of buried solvent-accessible surface area, and only part of helix III (yeast equivalent to helix II) near the Ppant group interacts with the DH domain (Extended Data Fig. 8b). Likewise, the Escherichia coli homologue of the DH domain, FabA, has previously been crystallized with the AcpP in a trapped state using a mechanistic crosslinker35. The FabA–AcpP interaction involves approximately 600 Å2 of buried solvent-accessible surface area and involves significant contact by helix II of AcpP, making it a more similar interaction to the mammalian enzyme than yFASN (Extended Data Fig. 8c).
Interactions of the ACP with ER domain homologues are more poorly characterized, with the only previously available structures being from S. cerevisiae20. In yFASN, the ER domain uses a flavin mononucleotide cofactor in addition to the NADPH molecule. The structure of the yFASN ACP at the ER domain has an interaction interface with approximately 200 Å2 of buried solvent-accessible surface area, whereas the hFASN ER–ACP interaction consists of approximately 950 Å2. The yFASN interaction appears to be mostly mediated by helix III and charge complementarity (Extended Data Fig. 8d), like the hFASN interaction (Fig. 5). Although NADPH seems to aid in ACP interactions at the ER domain in hFASN, structures of yFASN show that NADPH and ACP binding at the ER domain are mutually exclusive20. This difference is probably driven by the flavin mononucleotide cofactor used by yFASN that makes co-binding of the two unnecessary. Overall, the ACP interactions in the mammalian system at the DH and ER domains diverge from the yFASN and FabA systems, but still share charge complementarity as driving factors.
From our cryo-EM data, we were able to resolve large conformational changes between the condensing and modifying region of hFASN (Fig. 2 and Extended Data Figs. 4 and 7). These conformational changes cause asymmetry across the homodimer as has been previously reported28. Our new data with the ACP at the DH and ER domain in one reaction chamber, although unresolved in the other, support a model in which hFASN asynchronously synthesizes palmitate across the dimer, which can be passed between the ACP and consequently the reaction chambers. This is consistent with previous biochemistry showing that mammalian FASN dimers that are catalytically inactive at the ACP and KS domains in opposite subunits retain partial activity24, and recent work on related modular PKSs that has also suggested asymmetric function across the dimers36,37. Although the asynchronous ACP shuttling observed in the cryo-EM might be considered inconsistent with the Hill slope of 1.5 (ref. 38), the meaning of such a Hill slope is convoluted in the complex case of FASN with six catalytic domains that also show changes in the reaction rate with different chain lengths on the ACP27. Yet, in the case of modular PKSs, which also work asynchronously, it is believed that the asymmetry may be important for ensuring proper modification and transfer of the intermediate to the next module9. In the case of hFASN, the polypeptide contains all the necessary catalytic functions for elongation and saturation of the growing acyl chain, and is more similar to iterative PKSs, but the recently structurally characterized iterative PKS lacks conformational heterogeneity39. Therefore, the ability of the FASN KS domain with the growing acyl chain to interact with either ACP is potentially advantageous for bypassing the need to correct the acyl group loaded on the ACP during synthesis when acyl chains other than malonyl groups are loaded to prevent stalling or off-loading of intermediates. These conformational rearrangements would allow for elongation by only a properly loaded malonyl-ACP and prevent or decrease stalling if the ACP in the specific reaction chamber is improperly loaded (Fig. 6). This may be specifically required in mammalian FASN to overcome the high promiscuity of the MAT domain that is not present in other systems23.
Schematic representation of the proposed mechanism for how hFASN shuttles intermediates between active sites to synthesize palmitate.
Although our structures show that the interactions at the DH and ER domains are independent of the overall conformational state, interaction with the KS and MAT domains in the condensing region probably are specific to certain conformational states. On the basis of our structures, we hypothesize that states 1–4 are compatible with ACP interactions at the KS and MAT domains. These conformations leave the KS domain accessible in both reaction chambers, based on an estimated range of approximately 55 Å for the ACP to interact with the KS or MAT interface. The MAT is only accessible in a single reaction chamber in states 3 and 4, further supporting a potential regulatory role and asymmetric mechanism. Future investigations on ACP shuttling in mammalian FASN, with the technological bases established in this work, will help to complete our understanding of the FASN mechanism.
Methods
Cloning and protein expression
The plasmids pAR17_StrepI_hFASh_H8_pET22b and pAR18_StrepI_mFASm_H8_pET22b containing the genes for hFASN and mFASN respectively, were a gift from M. Grininger (Addgene plasmids #122846 and #122847)14,23. The coding region for hFASN was amplified from pAR17_StrepI_hFASh_H8_pET22b, and subcloned into a pFastBac-HTB vector containing an N-terminal Flag tag and a C-terminal 8×-histidine tag. The gene encoding the Holo-ACP synthase from Streptomyces coelicolor (2JBZ_A) was codon optimized for E. coli expression and inserted into a pET-28(a) vector by BioBasic and used as a co-expression with mFASN to yield functional protein from E. coli. hFASN was expressed in Sf9 cells that were grown to a cell density of 2 × 106 before infection with baculovirus containing the expression construct. The cells were harvested after 48 h growth at 27 °C, pelleted by ultracentrifugation and stored at −80 °C until purification. The mFASN construct was transformed into BL21-Rosetta (DE3) cells (Novagen), with Holo-ACP synthase, and expression was induced with isopropyl β-d-1-thiogalactopyranoside (IPTG) following the published protocol14.
Protein purification
For purification of mFASN, cells were resuspended in lysis buffer containing 40 mM HEPES (pH 7.6), 500 mM NaCl, 5 mM β-mercaptoethanol (βME), 1 mM PMSF, 5% glycerol, DNase, lysozyme, 0.05% Tween-20, 1 mM ATP, 1 mM MgCl2 and 20 mM imidazole. The cells were lysed by sonication on ice for 1 min, and the lysate was clarified by centrifugation at 50,000g for 60 min. The lysate was incubated with Ni-NTA resin (Thermo Fisher Scientific) that was equilibrated with lysis buffer for 1 h at 4 °C with slow rotation. The lysate was removed from the resin by flowing through a gravity column before washing with 50 ml of low-salt buffer (30 mM Tris (pH 8.0), 100 mM NaCl, 20 mM imidazole, 5 mM βME and 5% glycerol), and eluting with 50 ml of elution buffer (30 mM Tris (pH 8.0), 100 mM NaCl, 300 mM imidazole, 5 mM βme and 5% glycerol). The elution was immediately loaded onto a HiTrap Q column (Cytiva) and eluted with a slow gradient of high-salt buffer (1 M NaCl) to elute mFASN. The peak corresponding to mFASN was concentrated using a 100,000 MWCO spin concentrator (Amicon), diluted into low-pH buffer (25 mM Bis-Tris (pH 6.7), 150 mM NaCl, 5 mM TCEP and 5% glycerol) and incubated for 30 min. The concentrated sample was injected onto a Superose6 gel filtration column (Cytiva) equilibrated with a final buffer (25 mM HEPES (pH 7.0), 150 mM NaCl and 5 mM TCEP). The fractions containing mFASN were concentrated and used directly or flash frozen in liquid nitrogen and stored at −80 °C.
The hFASN construct was purified following a similar protocol, except the lysate was further clarified by vacuum filtration following centrifugation, and the N-terminal Flag tag was used for a second affinity purification step in place of the ion-exchange chromatography.
Cryo-EM sample preparation
Purified hFASN was diluted to 4 mg ml−1 and supplemented with indicated substrates (1 mM concentrations) and allowed to incubate for 10 min at room temperature before grid preparation. Quantifoil 300 Mesh Cu R0.6/1 grids were glow discharged for 1–2 min under vacuum at 0.25 mA. CHAPSO (Avanti) was spiked into the FASN sample to a final concentration of 8 mM immediately before applying 3 µl of sample to the grids. The grids were blotted using a VitroBot Mark IV (Thermo Fisher Scientific) and Whatman blotting paper for 2–3 s at a blot force of 10 in 100% humidity at 4 °C.
Cryo-EM data collection and data processing
All cryo-EM data were collected on a Titan Krios microscope (Thermo Fisher) equipped with a GatanK3 camera and an energy filter. The microscope was operated at 300 kV, a nominal magnification of ×81,000, and an energy filter slit width of 20 eV. Leginon software was used for automated data collection of 19,291 movies with an accumulated dose of 52.50 e− Å−2 across 40 frames40. A nominal defocus range of −0.8 to −3.0 µm was used.
The images were imported into RELION 3.1 (ref. 41) at a calibrated pixel size of 1.069 Å and motion corrected using MotionCor2 (ref. 42) with 5 × 5 patches and no binning. The motion-corrected images were imported into cryoSPARC (v4.2.1)43, and contrast transfer function (CTF) estimations were calculated using a wrapper for CTFFIND4.1 (ref. 44). Micrographs were filtered to remove any images with a CTF estimation worse than 6 Å, leaving a total of 15,231 micrographs for downstream processing. A total of 1,237 particles were manually picked from 23 micrographs and used to train a Topaz particle-picking model45. The trained model was used to pick 989,117 particles that were extracted with a box size of 360 pixels and classified into 300 classes in 2D with a maximum resolution of 12 Å. A subset of 271 classes (943,639 particles) were selected, and 100,000 of these particles were used for ab initio reconstruction into 4 classes. The class with the most features resembling mammalian FASN were used to seed a heterogenous refinement with ten identical classes. The outputs were used to seed a subsequent job, and this was repeated for a total of four rounds of heterogenous refinement. A total of eight classes remained that had FASN features, and two junk classes. The 8 FASN classes were used to pick particles on a subset of 5,000 micrographs to train a new Topaz model. The updated model was used to pick 1.5 million particles from the dataset. The processing steps were repeated through heterogenous refinement, and eight FASN-like classes remained again. Each class was individually refined to high resolution with homogenous refinement, NURefinement with CTF and defocus refinement, partial signal subtraction of the modifying and condensing regions independently, and local refinement of the remaining region. Each individual map was manually inspected, sharpened with DeepEMhancer for visualization, and run through the 3DFSC server to confirm 3D resolution and map sphericity before model building.
In addition, to elucidate the ACP at the catalytic domains, all the classes with FASN features were pooled and refined into a single map. A focused map was made for each of the active sites on the core (MAT, KS, KR, DH and ER) and used for focused classification using 3D classification in simple mode with ten classes. Classes that contained density for the ACP at the DH and ER domains were resolved, and the particles were refined to high-resolution maps using NURefinement with CTF and defocus refinement. The modifying regions with the ACP was well resolved, but the condensing region was unresolved, so it was subtracted, and local refinement was used to obtain a final high-resolution reconstruction of the modifying portion with the ACP. Each map was manually inspected, sharpened with DeepEMhancer46 and run through the 3DFSC47 server before model building.
The dataset of hFASN with NADP+ and acetoacetyl-CoA was processed in a similar manner, except the maps for the individual conformations were not taken to high resolution, and 3D classification only resolved the ACP at the DH domain. Local resolution estimates were made for all maps using cryoSPARC with an FSC cut-off of 0.143. UCSF’s ChimeraX48 was used for all visualizations of cryo-EM maps and models.
Atomic model building
All maps with resolution of less than 4 Å were manually fitted with atomic models. For the condensing region, PDB 3HHD was manually fit into the density using ChimeraX and further refined using a combination of Coot49 and Phenix50. For the modifying portion, two copies of the AlphaFold2 (refs. 51,52) prediction (AF-P49327-F1) residues 855–2112 were manually fit into the density in ChimeraX and further refined using a combination of Coot and Phenix. The NADP(H) molecules were built into the density using Coot and the available coordinates from the PDB. Maps, which had density for the ACP, were additionally fit with the crystal structure (PDB: 2CG5) in ChimeraX. The Ppant arm was built into the models in Coot using the previously reported coordinates for 4HH in the PDB.
Structure analysis
Analysis and visualization of all models were done using ChimeraX, including calculations of buried solvent-accessible surface area, interactions and root mean square deviation.
Enzyme kinetics
Aliquots of mFASN protein were thawed under running water and centrifuged at 16,000g for 10 min to remove any precipitate. The sample concentrations were individually determined using a Nanodrop with an extinction coefficient of 252,000, and stocks at 1 mg ml−1 were made by diluting into fresh buffer. The enzymatic assays were assembled to a final concentration of 200 nM FASN, 0.005% Tween-20, 50 mM HEPES (pH 7.0), 150 mM NaCl and 5 mM TCEP. To determine the specific activity and generate a Michaelis–Menten curve, acetyl-CoA and malonyl-CoA were titrated in a 1:1 ratio from 4 to 1,000 µM in the presence of 500 µM NADPH and activity was monitored continuously by absorbance of NADPH at 340 nM wavelength in a Tecan Spark plate reader. The enzyme velocity was calculated using a standard curve of NADPH in the same buffer to calculate NADPH consumption. Prism was used to fit the curve to an allosteric model that was the best fit compared with the Michaelis–Menten and substrate inhibition models with an R2 value of 0.9755. On the basis of the Michaelis–Menten curve, conditions were chosen for comparing mutant to wild-type activity. The relative activity assays were activated with 500 µM acetyl-CoA and malonyl-CoA and 500 µM NADPH. The activity of all mutants was calculated relative to a wild-type control in the same plate.
Knockout cell line generation
The D42 murine hepatocellular carcinoma cells used in this study were previously generated and described33. FASN-knockout D42 cells were generated using CRISPR–Cas9 with single guide RNA targeting exon 2 of mFASN (aaacCTTGGGTAATCCATAGAGCC) designed with Benchling (Biology Software, 2023). The method previously used for CRISPR–Cas9 knockout in the D42 cells was used to generate the FASN-knockout cells33, and knockout was confirmed by western blot with antibodies to FASN (C20G5, Cell Signaling; rabbit monoclonal antibody) at a 1:1,000 dilution and actin (8H10D10, Cell Signaling; mouse monoclonal antibody) at a 1:2,000 dilution.
Fatty acid methyl-ester measurements by GC-MS
Tracing of [U-13C]-glucose into palmitate was done for each cell line using a fatty acid methyl-ester derivatization and measurement by GC-MS as previously described33. In short, the cells were plated and grown in Dulbecco’s modified eagle medium (DMEM) supplemented with calf serum overnight. The medium was exchanged for medium containing 10 mM [U-13C]-glucose or unlabelled glucose and 10% charcoal-dextran-treated serum for 48 h before harvesting the cells. Fatty acids were extracted with a mixture of methanol and chloroform, and 10 µl of 1 mM heptadecanoic acid in methanol was added as an internal standard before drying under nitrogen. Dried lipids were first resuspended in a mixture of toluene (10 ml), methanol (50 ml) and butylated hydroxytoluene (5 mg), and 2 µl of acetyl chloride was then added. Reaction mixture was heated to 95 °C for 1 h to derivatize fatty acids to methyl-esters. The lipophilic portion was separated and loaded into the GC-MS for analysis. Analysis was performed on an Agilent GC-MS 7890A/5975A with a DB-5MS 5% phenyl methyl Silox column (Agilent). Carbon enrichment into palmitate was determined using FluxFix53 utilizing unlabelled glucose samples as controls. Relative quantification of palmitate was performed by normalizing the sum of the area under the curve of all isotopologues to the sum of the area under the curve of all isotopologues of heptadecanoic acid. Each sample was then further normalized to its protein quantification calculated using the Pierce BCA protein assay.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Atomic models and cryo-EM maps have been deposited into the PDB and Electron Microscopy Data Bank (EMDB) for hFASN–NADPH state 1 (EMD-43337 and PDB 8VLE), hFASN–NADPH state 2 (EMD-43340 and PDB 8VLO), hFASN–NADPH state 3 (EMD-43341 and PDB 8VLP), hFASN–NADPH state 4 (EMD-43350 and PDB 8VM0), hFASN–NADPH state 5 (EMD-43352 and PDB 8VM5), hFASN–NADPH state 6 (EMD-43355 and PDB 8VMC), hFASN–NADPH state 7 (EMD-43354 and PDB 8VM7), hFASN–NADPH state 8 (EMD-43353 and PDB 8VM6), hFASN–NADPH DH–ACP (EMD-43199 and PDB 8VG4), hFASN–NADPH ER–ACP (EMD-43187 and PDB 8VF7) and hFASN–NADP+ DH–ACP (EMD-43356 and PDB 8VMD). Any additional data supporting this publication will be provided by the corresponding authors on request. Source data are provided with this paper.
References
Paiva, P. et al. Animal fatty acid synthase: a chemical nanofactory. Chem. Rev. 121, 9502–9553 (2021).
Ameer, F., Scandiuzzi, L., Hasnain, S., Kalbacher, H. & Zaidi, N. De novo lipogenesis in health and disease. Metabolism 63, 895–902 (2014).
Batchuluun, B., Pinkosky, S. L. & Steinberg, G. R. Lipogenesis inhibitors: therapeutic opportunities and challenges. Nat. Rev. Drug Discov. 21, 283–305 (2022).
Ferraro, G. B. et al. Fatty acid synthesis is required for breast cancer brain metastasis. Nat. Cancer 2, 414–428 (2021).
Falchook, G. et al. First-in-human study of the safety, pharmacokinetics, and pharmacodynamics of first-in-class fatty acid synthase inhibitor TVB-2640 alone and with a taxane in advanced tumors. EClinicalMedicine 34, 100797 (2021).
Loomba, R. et al. TVB-2640 (FASN inhibitor) for the treatment of nonalcoholic steatohepatitis: FASCINATE-1, a randomized, placebo-controlled phase 2a trial. Gastroenterology 161, 1475–1486 (2021).
Kelly, K. L. et al. De novo lipogenesis is essential for platelet production in humans. Nat. Metab. 2, 1163–1178 (2020).
Bowers, M. et al. FASN-dependent lipid metabolism links neurogenic stem/progenitor cell activity to learning and memory deficits. Cell Stem Cell 27, 98–109.e111 (2020).
Grininger, M. Enzymology of assembly line synthesis by modular polyketide synthases. Nat. Chem. Biol. 19, 401–415 (2023).
Lin, Z., Li, F., Krug, P. J. & Schmidt, E. W. The polyketide to fatty acid transition in the evolution of animal lipid metabolism. Nat. Commun. 15, 236 (2024).
Jenke-Kodama, H., Sandmann, A., Müller, R. & Dittmann, E. Evolutionary implications of bacterial polyketide synthases. Mol. Biol. Evol. 22, 2027–2039 (2005).
Nivina, A., Yuet, K. P., Hsu, J. & Khosla, C. Evolution and diversity of assembly-line polyketide synthases. Chem. Rev. 119, 12524–12547 (2019).
Fischer, M. & Grininger, M. Strategies in megasynthase engineering — fatty acid synthases (FAS) as model proteins. Beilstein J. Org. Chem. 13, 1204–1211 (2017).
Rittner, A., Paithankar, K. S., Drexler, D. J., Himmler, A. & Grininger, M. Probing the modularity of megasynthases by rational engineering of a fatty acid synthase type I. Protein Sci. 28, 414–428 (2019).
Rittner, A. et al. Chemoenzymatic synthesis of fluorinated polyketides. Nat. Chem. 14, 1000–1006 (2022).
Buyachuihan, L., Reiners, S., Zhao, Y. & Grininger, M. The malonyl/acetyl-transferase from murine fatty acid synthase is a promiscuous engineering tool for editing polyketide scaffolds. Commun. Chem. 7, 187 (2024).
Maier, T., Jenni, S. & Ban, N. Architecture of mammalian fatty acid synthase at 4.5 Å resolution. Science 311, 1258–1262 (2006).
Maier, T., Leibundgut, M. & Ban, N. The crystal structure of a mammalian fatty acid synthase. Science 321, 1315–1322 (2008).
Leibundgut, M., Jenni, S., Frick, C. & Ban, N. Structural basis for substrate delivery by acyl carrier protein in the yeast fatty acid synthase. Science 316, 288–290 (2007).
Singh, K. et al. Reconstruction of a fatty acid synthesis cycle from acyl carrier protein and cofactor structural snapshots. Cell 186, 5054–5067.e16 (2023).
Witkowski, A., Joshi, A. K. & Smith, S. Characterization of the β-carbon processing reactions of the mammalian cytosolic fatty acid synthase: role of the central core. Biochemistry 43, 10458–10466 (2004).
Witkowski, A., Joshi, A. K. & Smith, S. Mechanism of the β-ketoacyl synthase reaction catalyzed by the animal fatty acid synthase. Biochemistry 41, 10877–10887 (2002).
Rittner, A., Paithankar, K. S., Huu, K. V. & Grininger, M. Characterization of the polyspecific transferase of murine type I fatty acid synthase (FAS) and implications for polyketide synthase (PKS) engineering. ACS Chem. Biol. 13, 723–732 (2018).
Rangan, V. S., Joshi, A. K. & Smith, S. Mapping the functional topology of the animal fatty acid synthase by mutant complementation in vitro. Biochemistry 40, 10792–10799 (2001).
Brink, J. et al. Quaternary structure of human fatty acid synthase by electron cryomicroscopy. Proc. Natl Acad. Sci. USA 99, 138–143 (2002).
Buyachuihan, L., Stegemann, F. & Grininger, M. How acyl carrier proteins (ACPs) direct fatty acid and polyketide biosynthesis. Angew. Chem. Int. Edn https://doi.org/10.1002/anie.202312476 (2023).
Rittner, A., Paithankar, K. S., Himmler, A. & Grininger, M. Type I fatty acid synthase trapped in the octanoyl‐bound state. Protein Sci. 29, 589–605 (2020).
Brignole, E. J., Smith, S. & Asturias, F. J. Conformational flexibility of metazoan fatty acid synthase enables catalysis. Nat. Struct. Mol. Biol. 16, 190–197 (2009).
Hasan, S. M. N., Lou, J. W., Keszei, A. F. A., Dai, D. L. & Mazhab-Jafari, M. T. Atomic model for core modifying region of human fatty acid synthase in complex with denifanstat. Nat. Commun. 14, 3460 (2023).
Hardwicke, M. A. et al. A human fatty acid synthase inhibitor binds β-ketoacyl reductase in the keto-substrate site. Nat. Chem. Biol. 10, 774–779 (2014).
Klaus, M. et al. Solution structure and conformational flexibility of a polyketide synthase module. JACS Au 1, 2162–2171 (2021).
Bunkoczi, G. et al. Mechanism and substrate recognition of human holo ACP synthase. Chem. Biol. 14, 1243–1253 (2007).
Izzo, L. T. et al. Acetylcarnitine shuttling links mitochondrial metabolism to histone acetylation and lipogenesis. Sci. Adv. 9, eadf0115 (2023).
Chan, D. I. & Vogel, H. J. Current understanding of fatty acid biosynthesis and the acyl carrier protein. Biochem. J. 430, 1–19 (2010).
Nguyen, C. et al. Trapping the dynamic acyl carrier protein in fatty acid biosynthesis. Nature 505, 427–431 (2014).
Bagde, S. R., Mathews, I. I., Fromme, J. C. & Kim, C.-Y. Modular polyketide synthase contains two reaction chambers that operate asynchronously. Science 374, 723–729 (2021).
Cogan, D. P. et al. Mapping the catalytic conformations of an assembly-line polyketide synthase module. Science 374, 729–734 (2021).
Monod, J., Wyman, J. & Changeux, J.-P. On the nature of allosteric transitions: a plausible model. J. Mol. Biol. 12, 88–118 (1965).
Wang, J. et al. Structural basis for the biosynthesis of lovastatin. Nat. Commun. 12, 867 (2021).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Tan, Y. Z. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat. Methods 14, 793–796 (2017).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2021).
Trefely, S., Ashwell, P. & Snyder, N. W. FluxFix: automatic isotopologue normalization for metabolic tracer analysis. BMC Bioinformatics 17, 485 (2016).
Acknowledgements
We thank S. Molugu and S. Steimle from the Electron Microscopy Resource Lab and Beckman Center for Cryo-EM from the Institute of Structural Biology at the University of Pennsylvania for support with cryo-EM grid preparation and screening (RRID: SCR_022375); C. Freeman and D. Schultz from the High Throughput Screening Core (RRID: SCR_022379) at the University of Pennsylvania for production of hFASN in Sf9 cells; Z. Arany for access to a GC-MS instrument for measurement of fatty acids; and E. Eng, E. Chua and K. Neselu for help with data collection performed at the National Center for CryoEM Access and Training and the Simons Electron Microscopy Center located at the New York Structural Biology Center, supported by the NIH Common Fund Transformative High Resolution Cryo-Electron Microscopy program (U24 GM129539), and by grants from the Simons Foundation (SF349247) and NY State Assembly. Structural visualization and analysis were done with UCSF ChimeraX, which was developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco. ChimeraX was developed with support from US National Institutes of Health (NIH) R01-GM129325 and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases. We thank H.-Y. Tang and T. Beer from The Wistar Institute Proteomics and Metabolomics Facility, supported in part by a Cancer Center Support Grant awarded by the National Cancer Institute to the Ellen and Ronald Caplan Cancer Center, for mass spectrometry confirmation of post-translational modifications. NIH grants R35 GM118090 to R.M., R01 CA262055 to R.M. and K.E.W., K00 CA245802 to P.C.-P., T32 DK007314 and a Diversity Supplement of R01 CA262055 to L.V.P., and T32 GM133398 to K.S. supported this work.
Author information
Authors and Affiliations
Contributions
K.S., P.C.-P., L.V.P., K.E.W. and R.M. conceptualized the study. K.S., L.G., P.C.-P., L.V.P., J.R.-S., S.M.G., K.E.W. and R.M. generated the methodology. K.S., L.G., P.C.-P., L.V.P., J.R.-S. and S.M.G. performed the investigation. K.S., P.C.-P. and L.V.P. conducted formal analysis. K.S. wrote the original draft of the manuscript. K.S. and L.G. performed the visualization. K.S., L.G., P.C.-P., L.V.P., J.R.-S., S.M.G., K.E.W. and R.M. reviewed and edited the manuscript. K.E.W. and R.M. acquired funding. K.E.W. and R.M. provided resources. K.E.W. and R.M. supervised the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Martin Grininger and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Human and mouse FASN samples.
a, Coomassie stained SDS-PAGE gel of hFASN with a single band corresponding to the protein near 270 kDa (representative of n = 5 purifications). For gel source data, see Supplementary Fig. 3. b, Size exclusion chromatogram of hFASN on a Superose 6 column with a peak corresponding to hFASN at 14 mL. c, Mass photometry data show a primary peak for hFASN at 550 kDa, confirming that it is a stable homodimer in solution in our conditions (representative of n = 3 experiments with a gaussian curve fit to the histogram and SD shown). The peak near 30 kDa is high background from the TCEP in the sample. d, Coomassie stained SDS-PAGE gel of WT mouse FASN purified from bacteria. The band and purity are similar to hFASN. For gel source data, see Supplementary Fig. 3. e, Superose 6 chromatogram of mouse FASN with a peak like hFASN near 14 mL. f, Mass photometer data showing that mFASN primarily forms a stable homodimer in the conditions at 550 kDA. While we observe a small monomer peak at 272 kDa, the dimer is the primary species in solution (representative of n = 3 experiments with a gaussian curve fit to the histogram and SD shown). g, Michaelis-Menten curve of NADPH consumption by mFASN when titrating acetyl-CoA and malonyl-CoA in a 1:1 ratio (n = 3 replicates). The curve was fit with an allosteric sigmoidal model in Prism. h, Table of values generated from the allosteric sigmoidal model of fitting to the kinetics of mFASN with n = 3 replicates. Both the best fit and 95% confidence interval are reported.
Extended Data Fig. 2 Cryo-EM workflow for NADPH dataset.
a, Description of the data processing workflow used to determine the structures of hFASN with NADPH. b, Representative micrograph from the hFASN + NADPH dataset (representative of the 15,201 micrographs used for particle picking). Micrograph has been lowpass filtered to 10 Å. c, Representative 2D classes showing multiple views of hFASN. Classes were chosen from the 2D classification job before ab intio reconstruction. d,e, Unsharpened cryo-EM density of the modifying region of hFASN with the ACP docked at the DH or ER domain respectively, with corresponding 3D FSC curve.
Extended Data Fig. 3 Cryo-EM maps of hFASN/NADPH States 2-8.
a–h, Unsharpened composite cryo-EM densities of hFASN in States 1-8, respectively, that are coloured by local resolution.
Extended Data Fig. 4 Conformational dynamics of hFASN.
a, Models of States 1-8 from the hFASN/NADPH dataset coloured by domain. A centroid centred at A2112 with a diameter of 55 Å is shown to represent reaction chamber 1 (teal) and reaction chamber 2 (magenta) that represent the range that each ACP can access to interact with the catalytic sites at each domain. The MAT is labelled as rotating into the plane of the viewer in states 5-8, although the direction of rotation is ambiguous from the data.
Extended Data Fig. 5 Cryo-EM workflow for the hFASN/NADP+ dataset.
a, Detailed data processing workflow used to determine the structure of the modifying region of hFASN in the presence of acetoacetyl-CoA and NADP+ with the ACP at the DH domain. b, Representative micrograph of the 19,528 micrographs from the dataset that has been lowpass filtered to 10 Å resolution. c, Representative 2D classes from a 2D classification job before ab initio reconstruction. d, Unsharpened cryo-EM density coloured by local resolution with the corresponding 3D FSC curve and azimuth plot.
Extended Data Fig. 6 D42 FASN KO cell generation and re-expression of WT and mutant enzymes.
a, Western blot analysis of FASN levels in FASN KO cells, and re-expressed WT and mutant human FASN enzymes, with actin blotted as a loading control (representative of n = 2 experiments). The MW marker in the FASN section is 240 kDa and the corresponding marker in the actin blot is 42 kDa. For blot source data, see Supplementary Figs. 4 and 5. b, Comparison of palmitate synthesis from [U-13C]-glucose in D42 parental cells and the FASN KO clone. The average of n = 3 replicates is plotted with error bars representing the SD. c, Relative abundance of palmitate measured by GC-MS. Data are normalized to heptadecanoic acid as the internal standard and μg protein, and the average abundance of the WT cells set to 1. Each biological replicate (n = 3) is shown, with the mean plotted and error bars for the SD. d, Isotopologue analysis of heavy carbon atoms incorporated into palmitate in each cell line, with the mean of n = 3 replicates shown and error bars of the SD. Isotopologue enrichment was calculated using FluxFix.science.
Extended Data Fig. 7 Conformational dynamics of the condensing region during acyl chain modification.
a, Output from a heterogenous refinement job initially seeded with the map and particles providing the hFASN/NADPH DH-ACP structure. All 8 classes are coloured according to domain and show structural heterogeneity similar to the entire dataset. Labelling is changed to Class A-H in place of State 1-8 to avoid confusion with the structures that lack ACP density. b, Classes from the 4th round of heterogenous refinement on the particles from the hFASN/NADPH ER-ACP structure, with the density coloured by domain.
Extended Data Fig. 8 Comparison of hFASN ACP interactions with yeast and bacterial systems.
a, Alignment of the hFASN ACP with the yeast ACP and bacterial AcpP. b, Structure of the yFASN ACP-DH interaction showing the minor contacts made by Helix III and the DH surface. c, Crystal structure of the bacterial FabA homologue of the DH domain with the ACP crosslinked. The FabA is shown as a surface and AcpP in cartoon form to highlight the extensive interaction made by Helix II at the FabA interface. d, yFASN ACP-ER interaction structure highlighting the broad contacts made by the ACP at the ER interface.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–5: Figure S1 shows the 3D-FSC curves for States 1-8 in the hFASN/NADPH dataset. Figure S2 shows the Azimuth plots for each map for States 1-8 in the hFASN/NADPH dataset. Figure S3 is the source image for Extended Data Fig. 1a and d SDS-PAGE. Figure S4 and 5 are the source data for Extended Data Fig. 6a.
Supplementary Table 1 (download XLSX )
Mass spectrometry analysis of phosphopantetheinylation of mouse FASN proteins. The table includes details methods describing how the phosphopantetheinylation was detected on the purified recombinant wild-type and L2151A (mouse numbering) proteins and the corresponding peptides detected
Supplementary Video 1 (download MOV )
Video summarizing the structures from the hFASN/NADPH dataset. The video shows the structures describing the large conformational changes between the modifying and condensing regions of hFASN, and the ACP interactions at the DH and ER domains.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Schultz, K., Costa-Pinheiro, P., Gardner, L. et al. Snapshots of acyl carrier protein shuttling in human fatty acid synthase. Nature 641, 520–528 (2025). https://doi.org/10.1038/s41586-025-08587-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-08587-x
This article is cited by
-
Navigating the Lipid Landscape: The Role of Fatty Acid Synthase in Neural Stem Cell Fate and Central Nervous System Function
Molecular Neurobiology (2026)
-
Downregulation of FASN in granulosa cells and its impact on ovulatory dysfunction in PCOS
Journal of Ovarian Research (2025)
-
A lipid metabolism related gene signature predicts postoperative recurrence in pancreatic cancer through multicenter cohort validation
Scientific Reports (2025)
-
Visualizing acyl carrier protein interactions within a crosslinked type I polyketide synthase
Nature Communications (2025)