Abstract
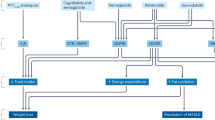
Peptide hormones, a class of pharmacologically active molecules, have a critical role in regulating energy homeostasis. Prohormone convertase 1/3 (also known as PCSK1/3) represents a key enzymatic mechanism in peptide processing, as exemplified with the therapeutic target glucagon-like peptide 1 (GLP-1)1,2. However, the full spectrum of peptides generated by PCSK1 and their functional roles remain largely unknown. Here we use computational drug discovery to systematically map more than 2,600 previously uncharacterized human proteolytic peptide fragments cleaved by prohormone convertases, enabling the identification of novel bioactive peptides. Using this approach, we identified a 12-mer peptide, BRINP2-related peptide (BRP). When administered pharmacologically, BRP reduces food intake and exhibits anti-obesity effects in mice and pigs without inducing nausea or aversion. Mechanistically, BRP administration triggers central FOS activation and acts independently of leptin, GLP-1 receptor and melanocortin 4 receptor. Together, these data introduce a method to identify new bioactive peptides and establish pharmacologically that BRP may be useful for therapeutic modulation of body weight.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in the manuscript and supporting files. The raw LC–MS data (raw.d mass spectrometry data) generated in this study have been deposited to Mendeley at https://doi.org/10.17632/y8m4669bkn.1, https://doi.org/10.17632/kphczhdcy8.1 and https://doi.org/10.17632/3x9yg2hzbx.1. Further information and requests for reagents should be directed to the corresponding author. Source data are provided with this paper.
Code availability
The code for the Peptide Predictor is freely available at https://github.com/Svensson-Lab/pro-hormone-predictor. There are no access restrictions.
References
Philippe, J. et al. A nonsense loss-of-function mutation in PCSK1 contributes to dominantly inherited human obesity. Int. J. Obes. 39, 295–302 (2015).
Sandoval, D. A. & D’Alessio, D. A. Physiology of proglucagon peptides: role of glucagon and GLP-1 in health and disease. Physiol. Rev. 95, 513–548 (2015).
Sobrino Crespo, C., Perianes Cachero, A., Puebla Jiménez, L., Barrios, V. & Arilla Ferreiro, E. Peptides and food intake. Front. Endocrinol. 5, 58 (2014).
Campbell, J. E. & Drucker, D. J. Pharmacology, physiology, and mechanisms of incretin hormone action. Cell Metab. 17, 819–837 (2013).
Muttenthaler, M., King, G. F., Adams, D. J. & Alewood, P. F. Trends in peptide drug discovery. Nat. Rev. Drug Discov. 20, 309–325 (2021).
Tatemoto, K., Carlquist, M. & Mutt, V. Neuropeptide Y–a novel brain peptide with structural similarities to peptide YY and pancreatic polypeptide. Nature 296, 659–660 (1982).
Lee, K. L., Middleditch, M. J., Williams, G. M., Brimble, M. A. & Cooper, G. J. S. Using mass spectrometry to detect, differentiate, and semiquantitate closely related peptide hormones in complex milieu: measurement of IGF-II and vesiculin. Endocrinology 156, 1194–1199 (2015).
Lee, J. E. Neuropeptidomics: mass spectrometry-based identification and quantitation of neuropeptides. Genomics Inform. 14, 12–19 (2016).
Fricker, L. D., Lim, J., Pan, H. & Che, F.-Y. Peptidomics: identification and quantification of endogenous peptides in neuroendocrine tissues. Mass Spectrom. Rev. 25, 327–344 (2006).
Muthusamy, B. et al. Plasma Proteome Database as a resource for proteomics research. Proteomics 5, 3531–3536 (2005).
Schwenk, J. M. et al. The Human Plasma Proteome Draft of 2017: building on the Human Plasma Peptide Atlas from mass spectrometry and complementary assays. J. Proteome Res. 16, 4299–4310 (2017).
Southey, B. R., Amare, A., Zimmerman, T. A., Rodriguez-Zas, S. L. & Sweedler, J. V. NeuroPred: a tool to predict cleavage sites in neuropeptide precursors and provide the masses of the resulting peptides. Nucleic Acids Res. 34, W267–W272 (2006).
Southey, B. R., Sweedler, J. V. & Rodriguez-Zas, S. L. A Python analytical pipeline to identify prohormone precursors and predict prohormone cleavage sites. Front. Neuroinformatics 2, 7 (2008).
Chance, R. E., Ellis, R. M. & Bromer, W. W. Porcine proinsulin: characterization and amino acid sequence. Science 161, 165–167 (1968).
Pollock, H. G., Hamilton, J. W., Rouse, J. B., Ebner, K. E. & Rawitch, A. B. Isolation of peptide hormones from the pancreas of the bullfrog (Rana catesbeiana). Amino acid sequences of pancreatic polypeptide, oxyntomodulin, and two glucagon-like peptides. J. Biol. Chem. 263, 9746–9751 (1988).
Vecchio, I., Tornali, C., Bragazzi, N. L. & Martini, M. The discovery of insulin: an important milestone in the history of medicine. Front. Endocrinol. 9, 613 (2018).
Tatemoto, K. & Neuropeptide, Y. Complete amino acid sequence of the brain peptide. Proc. Natl Acad. Sci. USA 79, 5485–5489 (1982).
Lovejoy, D. A. et al. Distinct sequence of gonadotropin-releasing hormone (GnRH) in dogfish brain provides insight into GnRH evolution. Proc. Natl Acad. Sci. USA 89, 6373–6377 (1992).
Steiner, D. F. The proprotein convertases. Curr. Opin. Chem. Biol. 2, 31–39 (1998).
Zheng, M., Streck, R. D., Scott, R. E., Seidah, N. G. & Pintar, J. E. The developmental expression in rat of proteases furin, PC1, PC2, and carboxypeptidase E: implications for early maturation of proteolytic processing capacity. J. Neurosci. 14, 4656–4673 (1994).
Lloyd, D. J., Bohan, S. & Gekakis, N. Obesity, hyperphagia and increased metabolic efficiency in Pc1 mutant mice. Hum. Mol. Genet. 15, 1884–1893 (2006).
Muhsin, N. I. A., Bentley, L., Bai, Y., Goldsworthy, M. & Cox, R. D. A novel mutation in the mouse Pcsk1 gene showing obesity and diabetes. Mamm. Genome 31, 17–29 (2020).
Burnett, L. C. et al. Deficiency in prohormone convertase PC1 impairs prohormone processing in Prader–Willi syndrome. J. Clin. Invest. 127, 293–305 (2017).
Lin, C. H. et al. An evaluation of liraglutide including its efficacy and safety for the treatment of obesity. Expert Opin. Pharmacother. 21, 275–285 (2020).
Wilding, J. P. H. et al. Once-weekly semaglutide in adults with overweight or obesity. N. Engl. J. Med. 384, 989–1002 (2021).
Kelly, A. S. et al. A randomized, controlled trial of liraglutide for adolescents with obesity. N. Engl. J. Med. 382, 2117–2128 (2020).
Peinado, J. R., Li, H., Johanning, K. & Lindberg, I. Cleavage of recombinant proenkephalin and blockade mutants by prohormone convertases 1 and 2: an in vitro specificity study. J. Neurochem. 87, 868–878 (2003).
Parvaz, N. & Jalali, Z. Molecular evolution of PCSK family: analysis of natural selection rate and gene loss. PLoS ONE 16, e0259085 (2021).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Mooney, C., Haslam, N. J., Pollastri, G. & Shields, D. C. Towards the improved discovery and design of functional peptides: common features of diverse classes permit generalized prediction of bioactivity. PLoS ONE 7, e45012 (2012).
Grønning, A. G. B., Kacprowski, T. & Schéele, C. MultiPep: a hierarchical deep learning approach for multi-label classification of peptide bioactivities. Biol. Methods Protoc. 6, bpab021 (2021).
Petryszak, R. et al. Expression Atlas update—an integrated database of gene and protein expression in humans, animals and plants. Nucleic Acids Res. 44, D746–D752 (2016).
Raffin-Sanson, M. L., de Keyzer, Y. & Bertagna, X. Proopiomelanocortin, a polypeptide precursor with multiple functions: from physiology to pathological conditions. Eur. J. Endocrinol. 149, 79–90 (2003).
Seizinger, B. R. et al. Isolation and structure of a novel C-terminally amidated opioid peptide, amidorphin, from bovine adrenal medulla. Nature 313, 57–59 (1985).
Ghatei, M. A. et al. Distribution, molecular characterization of pituitary adenylate cyclase-activating polypeptide and its precursor encoding messenger RNA in human and rat tissues. J. Endocrinol. 136, 159–166 (1993).
Foster, S. R. et al. Discovery of human signaling systems: pairing peptides to G protein-coupled receptors. Cell 179, 895–908.e21 (2019).
Eipper, B. A., Stoffers, D. A. & Mains, R. E. The biosynthesis of neuropeptides: peptide alpha-amidation. Annu. Rev. Neurosci. 15, 57–85 (1992).
Kim, K.-H. & Seong, B. L. Peptide amidation: production of peptide hormonesin vivo andin vitro. Biotechnol. Bioprocess Eng. 6, 244–251 (2001).
Mukai, H., Kawai, K., Suzuki, Y., Yamashita, K. & Munekata, E. Stimulation of dog gastropancreatic hormone release by neuromedin B and its analogues. Am. J. Physiol. 252, E765–E771 (1987).
Wettergren, A., Pridal, L., Wøjdemann, M. & Holst, J. J. Amidated and non-amidated glucagon-like peptide-1 (GLP-1): non-pancreatic effects (cephalic phase acid secretion) and stability in plasma in humans. Regul. Pept. 77, 83–87 (1998).
Milbrandt, J. Nerve growth factor rapidly induces c-fos mRNA in PC12 rat pheochromocytoma cells. Proc. Natl Acad. Sci. USA 83, 4789–4793 (1986).
Gabellini, N., Minozzi, M. C., Leon, A. & Dal Toso, R. Nerve growth factor transcriptional control of c-fos promoter transfected in cultured spinal sensory neurons. J. Cell Biol. 118, 131–138 (1992).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Berkowicz, S. R., Featherby, T. J., Whisstock, J. C. & Bird, P. I. Mice lacking Brinp2 or Brinp3, or both, exhibit behaviors consistent with neurodevelopmental disorders. Front. Behav. Neurosci. 10, 196 (2016).
Wiggenhorn, A. L. et al. A class of secreted mammalian peptides with potential to expand cell-cell communication. Nat. Commun. 14, 8125 (2023).
Kim, J. T. et al. Cooperative enzymatic control of N-acyl amino acids by PM20D1 and FAAH. eLife 9, e55211 (2020).
Kim, J. T., Li, V. L., Terrell, S. M., Fischer, C. R. & Long, J. Z. Family-wide annotation of enzymatic pathways by parallel in vivo metabolomics. Cell Chem. Biol. 26, 1623–1629.e3 (2019).
Nair, A. B. & Jacob, S. A simple practice guide for dose conversion between animals and human. J. Basic Clin. Pharm. 7, 27–31 (2016).
Pedersen, K.-M. et al. Optimization of pig models for translation of subcutaneous pharmacokinetics of therapeutic proteins: liraglutide, insulin aspart and insulin detemir. Transl. Res. J. Lab. Clin. Med. 239, 71–84 (2022).
Camerlink, I. & Ursinus, W. W. Tail postures and tail motion in pigs: a review. Appl. Anim. Behav. Sci. 230, 105079 (2020).
Shaywitz, A. J. & Greenberg, M. E. CREB: a stimulus-induced transcription factor activated by a diverse array of extracellular signals. Annu. Rev. Biochem. 68, 821–861 (1999).
Sheng, M., McFadden, G. & Greenberg, M. E. Membrane depolarization and calcium induce c-fos transcription via phosphorylation of transcription factor CREB. Neuron 4, 571–582 (1990).
Keen, A. C. et al. OZITX, a pertussis toxin-like protein for occluding inhibitory G protein signalling including Gαz. Commun. Biol. 5, 256 (2022).
Peng, Q., Alqahtani, S., Nasrullah, M. Z. A. & Shen, J. Functional evidence for biased inhibition of G protein signaling by YM-254890 in human coronary artery endothelial cells. Eur. J. Pharmacol. 891, 173706 (2021).
Fukushima, N. et al. Melittin, a metabostatic peptide inhibiting Gs activity. Peptides 19, 811–819 (1998).
Borner, T. et al. GDF15 induces anorexia through nausea and emesis. Cell Metab. 31, 351–362.e5 (2020).
Tsai, V. W. W. et al. The anorectic actions of the TGFβ cytokine MIC-1/GDF15 require an intact brainstem area postrema and nucleus of the solitary tract. PLoS ONE 9, e100370 (2014).
Madsen, C. T. et al. Combining mass spectrometry and machine learning to discover bioactive peptides. Nat. Commun. 13, 6235 (2022).
Ma, J. et al. Improved identification and analysis of small open reading frame encoded polypeptides. Anal. Chem. 88, 3967–3975 (2016).
Donohue, M. J., Filla, R. T., Steyer, D. J., Eaton, W. J. & Roper, M. G. Rapid liquid chromatography–mass spectrometry quantitation of glucose-regulating hormones from human islets of Langerhans. J. Chromatogr. A 1637, 461805 (2021).
Paxinos, G. & Franklin, K. B. J. The Mouse Brain in Stereotaxic Coordinates (Academic Press, 2001).
Allen, W. E. et al. Thirst-associated preoptic neurons encode an aversive motivational drive. Science 357, 1149–1155 (2017).
Park, Y.-G. et al. Protection of tissue physicochemical properties using polyfunctional crosslinkers. Nat. Biotechnol. 37, 73–83 (2019).
Kim, S.-Y. et al. Stochastic electrotransport selectively enhances the transport of highly electromobile molecules. Proc. Natl Acad. Sci. USA 112, E6274–E6283 (2015).
Shelhamer, E., Long, J. & Darrel, T. Fully convolutional networks for semantic segmentation. Preprint at https://doi.org/10.48550/arXiv.1605.06211 (2016).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: convolutional networks for biomedical image segmentation. Preprint at https://doi.org/10.48550/arXiv.1505.04597 (2015).
Murray, E. et al. Simple, scalable proteomic imaging for high-dimensional profiling of intact systems. Cell 163, 1500–1514 (2015).
Batra, V. R. & Schrott, L. M. Acute oxycodone induces the pro-emetic pica response in rats. J. Pharmacol. Exp. Ther. 339, 738–745 (2011).
Acknowledgements
The authors thank the Stanford University Pathology Histology core facility; the Stanford Cardiovascular Institute; the Stanford Diabetes Research Center P30DK116074; the Pathology Department; the Long laboratory for helpful discussions; and the Stanford Behavioral and Functional Neuroscience Laboratory for behavioural testing, supported by the NIH S10 Shared Instrumentation for Animal Research (1S10OD030452-01). This research was mentored and financially supported by Stanford’s SPARK Translational Research Program. This work was funded by R01DK125260 (K.J.S.), P30DK116074 (K.J.S.), The Jacob Churg Foundation (K.J.S.), The SPARK Program at Stanford (K.J.S. and L.C.), Stanford Bio-X (K.J.S. and V.L.L.), Stanford Maternal and Child Health Research Institute (K.J.S.), American Heart Association 23IPA1042031 (K.J.S.), American Heart Association postdoctoral fellowship 1011077 (L.C.), American Heart Association postdoctoral fellowship 905674 (M.Z.), American Heart Association postdoctoral fellowship 1242818 (L.W.W.), K99/R00 NIH Pathway to Independence Award (K99AR081618) from NIAMS (M.Z.), Stanford School of Medicine Dean’s Fellowship Award 2022 (L.C.), Stanford School of Medicine Dean’s Fellowship Award 2023 (L.W.W.), Carlsberg Foundation (N.B.D.-S.), Fundacion Alfonso Martin Escudero postdoctoral fellowship (M.D.M.-G.), Wu Tsai Human, Performance Alliance postdoctoral fellowship (M.D.M.-G.), Stanford Maternal and Child Health, Research Institute Postdoctoral Fellowship (N.A.J.K.), Welcome 200837 (N.A.J.K.), National Institutes of Health GM113854 (V.L.L.) and HHMI (L.L.).
Author information
Authors and Affiliations
Contributions
Conceptualization: L.C. and K.J.S. Methodology: L.C., N.B.D.-S., Q.N., A.S., A.W., D.K., D.T., M.Z., E.B.-M., V.L.L., T.A., A.P., L.W.W., S.C.R., J.L., Y.W., J.J.K., I.L., M.D.M.-G., N.A.J.K., D.C.-H.W., L.L. and KJS. Investigation: L.C., N.B.D.-S., M.Z., E.B.-M., L.W.W., D.H.-C.C., D.C.-H.W. and K.J.S. Funding acquisition: K.J.S. Supervision: K.J.S. Writing—original draft: L.C., L.L. and K.J.S.
Corresponding author
Ethics declarations
Competing interests
K.J.S. and L.C. are named inventors on patents regarding BRP peptides for metabolic disorders. K.J.S. is a co-founder of Merrifield Therapeutics. All other authors declare no competing interests.
Peer review information
Peer review information
Nature thanks Richard DiMarchi and the other, anonymous, reviewer(s) for their contribution to the peer reviewof this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Peptide Predictor maps a large group of proteolytically processed human peptides.
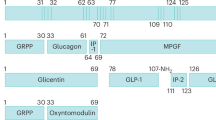
a, Number of precursor proteins with a minimum cleavage spacing of 3 amino acids between cleavage sites. b, Enriched biological pathways from the precursor proteins in the brain. c,d, Detected precursor proteins and the number of cleavage sites per precursors in pancreas (c) and pituitary (d). Blue: known prohormones. e, Validation of prediction method for pro-glucagon. f–p, Detected precursor proteins and the number of cleavage sites per precursors in brain (f), liver (g), blood (h), adipose tissue (i), adrenal gland (j), parathyroid gland (k), salivary gland (l), kidney (m), thyroid gland (n), skeletal muscle (o) and intestine (p). Blue: known prohormones. q, Bar diagram showing percent of peptides with a length between 5 and 25 amino acids (including the C-terminus cleavage site) per tissue (upper), and density plots of peptides within the same range (lower). Tics indicate individual peptides (n = 1–159).
Extended Data Fig. 2 BRP activates Fos expression and controls feeding without lowering blood glucose.
a, Peptide library design. 100 amidated peptides ranging from 5 to 25 amino acids length were synthesized with a C-terminal amide. b, Fos gene expression in NS-1 cells treated for 1 h with vehicle, NGF (50 ng/ml), BRP or scrambled BRP peptide (100 ug/ml). N = 3 biological replicates/group. c, Fos gene expression in Neuro2a cells treated with vehicle, or BRP peptide (50 µg/ml) for indicated times. N = 3 biological replicates/group. d, Fos gene expression in NS-1 cells treated with vehicle, or BRP peptide (50 µg/ml) for indicated times. N = 3 biological replicates/group. e, Fos gene expression in Neuro2a cells treated for 1 h with vehicle or BRP peptide every day for four days. N = 3 biological replicates/group. f, Fos gene expression in NS-1 cells treated for 1 h with vehicle, NGF (50 ng/ml), unmodified or amidated BRP (100 g/ml). N = 3 biological replicates/group. g, Fos gene expression in NS-1 cells treated for 24 h with vehicle or increasing doses of BRP peptide. N = 3 biological replicates/group. h, Fos gene expression in Neuro2a cells treated for 24 h with vehicle or increasing doses of BRP peptide. N = 3 biological replicates/group. i, Fos gene expression in Neuro2a cells treated for 1 h with vehicle or increasing doses of BRP peptide. N = 3 biological replicates/group. j, Fos gene expression in NS-1 cells treated for 1 h with vehicle or increasing doses of BRP peptide. N = 3 biological replicates/group. k, Fos gene expression in INS1 cells treated for 1 h with vehicle or increasing dose of GLP-1. N = 3 biological replicates/group. l, Pharmacokinetic levels of plasma and brain BRP after I.P. injection of 10 mg/kg BRP in lean C57BL/6 J male mice. N = 3 mice per group and per time point. m, Plasma levels of BRP, THRIL and LFNLC peptides after I.P. injection of 10 mg/kg BRP in lean C57BL/6 J male mice. N = 3 mice per group and per time point. n, Food intake measurement in lean C57BL/6 J male mice for up to 3 h after I.P. injection of vehicle, 5 mg/kg BRP or 5 mg/kg THRIL peptide. N = 3 independent experiments with 3 mice/group in each experiment. o, Food intake measurement in lean C57BL/6 J male mice for up to 3 h after I.P. injection of vehicle, 5 mg/kg BRP or 5 mg/kg LFNLC peptide. N = 3 independent experiments with 3 mice/group in each experiment. p, Food intake measurement in lean C57BL/6 J female mice for up to 3 h after I.P. injection of vehicle, 5 mg/kg BRP peptide, or 2 mg/kg GLP-1. N = 3 independent experiments with 3 mice/group in each experiment. q, Food intake measurement in C57BL/6 J DIO male mice for up to 3 h after I.P. injection of vehicle, 10 mg/kg BRP peptide, or 2 mg/kg GLP-1. N = 3 mice/group. r, Food intake measurement in lean C57BL/6 J male mice for up to 3 h after I.P. injection of vehicle or 5 mg/kg truncated BRP peptides. N = 3 independent experiments with 3 mice/group in each experiment. s, Fasting blood glucose levels in lean C57BL/6 J male mice for up to 3 h after I.P. injection of vehicle, 5 mg/kg peptides or 2 mg/kg GLP-1. N = 3 mice/group. t, Fasting insulin levels in lean C57BL/6 J male mice for up to 2 h after I.P. injection of vehicle or 5 mg/kg BRP. N = 3 mice/group. Data are presented as mean +/− S.E.M, P value = ns (non-significant), *P < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by one way Anova (b, f) or two-way Anova for multiple comparisons (c, d, e, m, n, o, p, q, r, s).
Extended Data Fig. 3 BRINP2 expression in humans, mice and pigs.
a, Brinp2 gene expression in tissues from WT C57BL/6 J mice. N = 5 biological replicates. b, Expression of BRINP2 across human tissues (Human Protein Atlas). c, Expression of BRINP2 across pig organs (data from the Pig RNA Atlas). Data are presented as mean +/− S.E.M (a).
Extended Data Fig. 4 BRP peptide is present in human cerebrospinal fluid and mouse brain.
a–h, Representative extracted ion chromatogram of the parent +3 ion at m/z = 533.3 (a), the MRM 533.3 to 178.1 y1+ ion transition (b), the MRM 799.5 to 239.1 b2+ ion transition (c), the MRM 533.3 to 239.1 b2+ ion transition (d), the MRM 799.5 to 791.2 z12 + + ion transition (e), the MRM 799.5 to 960.7 z7+ ion transition (f), the MRM 799.5 to 1186.7 z9+ ion transition (g), and the parent +2 ion at m/z = 799.5 (h) in human CSF at 15 min. Blue = synthesized amidated BRP standard. Black = amidated BRP peptide in human CSF. i, Concentration of BRP (533.3 to 239.1 b2+ ion transition) in human CSF. j, MS/MS mirror plot of BRP in human CSF and the synthesized BRP standard. k, LC-MS (MS1 spectra) demonstrating the detection of amidated BRP (+3 ion) in male and female mouse brain at 15 min. Blue = synthesized amidated BRP standard. Grey = BRP peptide in Pc1WT mice. Purple = BRP peptide in Pc1N22D/N222D mice. N = 5 pooled brain samples/group. l, LC-MS (MS2 spectra) demonstrating the detection of amidated BRP (538.6 to 178.1 y1+ ion transition) in male and female mouse brain at 15 min. Blue = synthesized amidated BRP standard. Grey = BRP peptide in Pc1WT mice. Purple = BRP peptide in Pc1N22D/N222D mice. N = 5 pooled brain samples/group. Data are presented as mean +/− S.E.M (i).
Extended Data Fig. 5 The anorexigenic effects of BRP are comparable to GLP-17–37.
a–d, Analysis using metabolic cages for respiratory exchange ratio (RER) (a), oxygen consumption VO2 (b), carbon dioxide production VCO2 (c), and ambulatory activity (d) in lean C57BL/6 J male mice after a single I.P. injection of vehicle or 5 mg/kg scrambled BRP peptide. N = 12 mice/group. e–h, Analysis using metabolic cages for respiratory exchange ratio (RER) (e), oxygen consumption VO2 (f), carbon dioxide production VCO2 (g), and ambulatory activity (h) in lean C57BL/6 J male mice after a single I.P. injection of vehicle or 2 mg/kg GLP-1. N = 8 mice/group. Data are presented as mean +/− S.E.M, P value = ns (non-significant), *P < 0.05, **P < 0.01, ***P < 0.001 by two-way Anova for multiple comparisons (a, b, c, d, e, f, g, h).
Extended Data Fig. 6 BRP, but not scrambled BRP, reduces food intake without affecting energy expenditure.
a–e, Analysis using metabolic cages for food intake (a,b), meal frequency (c), time between meals (d), and meal size (e) in lean C57BL/6 J male mice after two consecutive days of I.P. injection of vehicle (N = 8 mice) or 5 mg/kg BRP (N = 7 mice). Injections were done at time 0 and 24 h. f–l, Analysis using metabolic cages for respiratory exchange ratio (RER) (f, g), oxygen consumption VO2 (h, i), carbon dioxide production VCO2 (j, k), and ambulatory activity (l) in lean C57BL/6 J male mice after two consecutive days of I.P. injection of vehicle (N = 8 mice) or 5 mg/kg BRP (N = 7 mice). Injections were done at time 0 and 24 h. Bar graphs indicate the mean of RER (g), VO2 (i), and VCO2 (k) the first three hours after injection. m–q, Analysis using metabolic cages for food intake (m, n), meal frequency (o), meal size (p), and time between meals (q) in lean C57BL/6 J male mice after two consecutive days of I.P. injection of vehicle or 5 mg/kg scrambled BRP. Injections were done at time 0 and 24 h. N = 8 mice/group. r–u, Analysis using metabolic cages for respiratory exchange ratio (RER) (r), oxygen consumption (VO2) (s), carbon dioxide production (VCO2) (t), and ambulatory activity (u) in lean C57BL/6 J male mice after two consecutive days of I.P. injection of vehicle or 5 mg/kg scrambled BRP. N = 8 mice/group. Injections were done at time 0 and 24 h. Data are presented as mean ± S.E.M. P value = ns (non-significant), *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 by two-tailed Student’s t-test (b, c, d, n, o, q), or two-way Anova (a, e, f, g, h, i, j, k, l, m, p, r, s, t, u).
Extended Data Fig. 7 BRP does not induce anxiety-like behaviour in minipigs.
a, Ethogram used for recording tail postures and motions. Coordinates for valence (x value in red) and arousal (y value in blue) were given for each tail posture in circles. The ethogram was adapted and re-drawn from ref. 50. b, Representative photos of the tail postures of the Yucatan minipigs 1 h after I.M. injection of vehicle, 2 mg/kg BRP peptide or 8 µg/kg Liraglutide. The circled numbers represent the recorded tail postures according to the ethogram. c, Scores for valence and arousal for the pigs used in this study. All the pigs were scored with a tail posture of 2, 4, 5 or 6. N = 20 photos (Vehicle), N = 29 photos (BRP), and N = 15 photos (Liraglutide) (N = 4 pigs/group).
Extended Data Fig. 8 GLP1R, leptin and MC4R are dispensable for the anti-obesity and anorexigenic action of BRP.
a, Food intake per mouse per day in C57BL/6 J DIO male mice during 14 days of daily treatment with vehicle, 100 µg/kg Liraglutide, or 5 mg/kg BRP. N = 10 mice/group. b, Body weight in C57BL/6 J DIO male mice during 14 days of daily treatment with vehicle, 100 µg/kg Liraglutide, or 5 mg/kg BRP. N = 10 mice/group. c, Fasting insulin levels in C57BL/6 J DIO male mice after 14 days of daily treatment with vehicle, 100 µg/kg Liraglutide, or 5 mg/kg BRP. N = 9 mice/group. d, Fasting glucagon levels in C57BL/6 J DIO male mice after 14 days of daily treatment with vehicle, 100 µg/kg Liraglutide, or 5 mg/kg BRP. N = 9 mice/group. e, Fasting leptin levels in C57BL/6 J DIO male mice after 14 days of daily treatment with vehicle, 100 µg/kg Liraglutide, or 5 mg/kg BRP. N = 8 mice/group. f, Fasting ghrelin levels in C57BL/6 J male mice 30 and 60 min after I.P. injection of vehicle (N = 4 mice) or 5 mg/kg BRP peptide (N = 5 mice). g, Fasting leptin levels in C57BL/6 J male mice 30 and 60 min after I.P. injection of vehicle (N = 4 mice) or 5 mg/kg BRP peptide (N = 5 mice). h, Food intake in C57BL/6 J male mice for up to 3 h after I.P. injection of vehicle, 2 mg/kg GLP-1 or 5 mg/kg BRP peptide or GLP-1 and BRP combined. N = 3 independent experiments with 3 mice/group in each experiment. i, Food intake in C57BL/6 J male mice for up to 3 h after I.P. injection of vehicle, 2 mg/kg GLP-1 or 5 mg/kg BRP peptide or both in combination with 100 µg/kg Exendin(9-39)amide. N = 3 independent experiments with 3 mice/group in each experiment. j, Body weight in WT C57BL/6 J and ob/ob male mice. N = 9 mice/group. k, Fasting blood glucose levels in WT and ob/ob C57BL/6 J male mice. N = 9 mice/group. l, Food intake in WT and ob/ob C57BL/6 J male mice. N = 6 mice/group. m, Food intake in C57BL/6 J ob/ob male mice for up to 3 h after I.P. injection of vehicle, 2 mg/kg GLP-1 or 5 mg/kg BRP peptide. N = 3 independent experiments with 3 mice/group in each experiment. n, Food intake in C57BL/6 J ob/ob male mice during 14 days of daily I.P. injection of vehicle (N = 4 mice), or 5 mg/kg BRP (N = 5 mice) followed by a 7-day washout period. o, Accumulated food intake in C57BL/6 J ob/ob mice during 14 days of daily I.P. injection of vehicle (N = 4 mice), or 5 mg/kg BRP (N = 5 mice). p, Body weight in C57BL/6 J ob/ob male mice during 14 days of daily I.P. injection of vehicle (N = 4 mice), or 5 mg/kg BRP (N = 5 mice) followed by a 7-day washout period. N = 4-5 mice/group. q, Body weight change in C57BL/6 J ob/ob male mice during 14 days of daily I.P. injection of vehicle, or 5 mg/kg BRP followed by a 7-day washout period. r, Fasting blood glucose levels in WT and MC4R-KO C57BL/6 J male mice. N = 9 mice/group. s, Food intake in WT and MC4R-KO C57BL/6 J male mice. N = 9 mice/group. t, Body weight in WT C57BL/6 J and MC4R-KO mice. N = 9 mice/group. u, Food intake in MC4R-KO mice for up to 3 h after I.P. injection of vehicle, 2 mg/kg GLP-1 or 5 mg/kg BRP peptide. N = 3 independent experiments with 3 mice/group in each experiment. v, Food intake in MC4R-KO male mice during 7 days of daily I.P. injection of vehicle, or 5 mg/kg BRP followed by a 5-day washout period. N = 4 mice/group. w, Accumulated food intake in MC4R-KO mice during 7 days of daily I.P. injection of vehicle, or 5 mg/kg BRP. N = 4 mice/group. x, Body weight in MC4R-KO male mice during 7 days of daily I.P. injection of vehicle, or 5 mg/kg BRP followed by a 5-day washout period. N = 4 mice/group. y, Body weight change in MC4R-KO mice during 7 days of daily I.P. injection of vehicle, or 5 mg/kg BRP followed by a 5-day washout period. N = 4 mice/group. Data are presented as mean +/− S.E.M. P value = ns (non-significant), *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 by by two-tailed Student’s t-test (j, k, l, r, s, t), one-way Anova (c, d, e) or two-way Anova for multiple comparisons (a, b, f, g, h, i, m, n, o, p, q, u, v, w, x, y).
Extended Data Fig. 9 BRP activates a central CREB/Fos pathway.
a, Western blot of pCREBS133, total CREB, β-actin, or tubulin in metabolic tissues 15 min after a single intravenous injection of vehicle or 10 mg/kg BRP in lean C57BL/6 J male mice. N = 3 mice/group. b, Quantification of protein expression of pCREBS133/total CREB by western blot quantified from figure a. N = 3 mice/group. c,d, Heatmaps showing mean Fos+ cell density on Allen Mouse Brain Atlas planes (650 microns apart) in lean C57BL/6 J male mice 1 h after I.P. injection of vehicle (c) or 10 mg/kg BRP (d). N = 3 mice/group. e, Heatmaps showing subtracted (BRP-Vehicle treated mice) mean Fos+ cell density on Allen Mouse Brain Altas planes (7000 microns). The subpanel on the right is a higher magnification heatmap of the hypothalamic area outlined in black. N = 3 mice/group. f, Heatmaps showing subtracted (BRP-Vehicle treated mice) mean Fos+ cell density on Allen Mouse Brain Altas planes (5400 microns). The subpanel on the right is a higher magnification heatmap of the hypothalamic area outlined in black. N = 3 mice/group. g, Heatmaps and representative images of Fos staining in the preoptic nucleus of the hypothalamus 1 h after I.P. injection of vehicle (upper panel) or 10 mg/kg BRP (lower panel) in lean C57BL/6 J male mice. N = 3 mice/group. The subpanels on the right are higher magnification (3×) images of the area outlined in white. Scale bars: 1000 μm (left), 250 μm (right). h, Representative images (N = 4-5 sections per three biological replicates) of Fos staining in arcuate nucleus 1 h after a single intravenous injection of vehicle, 10 mg/kg BRP, or 100 μg/kg Liraglutide in lean C57BL/6 J male mice. N = 3 mice/group. i, Quantification of the number of Fos positive cells in arcuate nucleus 1 h after a single intravenous injection of vehicle, 10 mg/kg BRP, or 100 μg/kg Liraglutide in lean C57BL/6 J male mice (N = 3 mice/group with 4 sections per biological replicate). j, Representative images (N = 4 sections per three biological replicates) of Fos and Pomc staining in arcuate nucleus 1 h after a single intravenous injection of vehicle or 10 mg/kg BRP in lean C57BL/6 J male mice. N = 3 mice/group. k, Quantification of the percentage of Pomc cells expressing Fos in arcuate nucleus 1 h after a single intravenous injection of vehicle or 10 mg/kg BRP in lean C57BL/6 J male mice (N = 3 mice/group with 4 sections per biological replicate). Data are presented as mean +/− S.E.M. P value = ns (non-significant), *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 by two-tailed Student’s t-test (k), or one-way Anova (b, i).
Extended Data Fig. 10 Leu8 is necessary for the signalling and anorexigenic action of BRP.
a,b, Fos gene expression in NS-1 (a) or Neuro2a (b) cells treated for 1 h with vehicle, 100 µg/ml BRP, or 100 µg/ml alanine substituted BRP peptides. N = 3 biological replicates/group. c, Food intake measurement in lean C57BL/6 J male mice for up to 3 h after I.P. injection of vehicle, 5 mg kg−1 BRP or 5 mg kg−1 alanine substituted BRP peptides. N = 3 independent experiments with 3 mice/group in each experiment. d, Fos gene expression in Neuro2a cells treated for 1 h with vehicle or 100 µg/ml alanine substituted BRP peptides (y axis) and the percentage of food intake at 30 min relative to vehicle after I.P. injection of 5 mg kg−1 alanine substituted BRP peptides in lean C57BL/6 J male mice (x axis). e, Food intake measurement in lean C57BL/6 J male mice for up to 3 h after I.P. injection of vehicle, 5 mg kg−1 BRP or 5 mg kg−1 BRPL8A peptide. f, Predicted structure and lipophilicity of BRP and BRPL8A mutant (top ranked prediction out of a total of five predictions per structure for BRP and the BRPL8A mutant). Surface is coloured by hydrophobicity score. g, cAMP reporter assay in Neuro2a cells treated with vehicle (N = 25 cells), BRP (N = 13 cells), scrambled BRP peptide (N = 25 cells) or BRPL8A mutant (100 µg/ml) (N = 25 cells). h, Representative western blot (N = 3 in total) of pCREBS133, total CREB, and β-actin in Neuro2a cells treated with vehicle, BRP, scrambled BRP peptide or BRPL8A mutant (100 µg/ml) for 15 min. i, Quantification of protein expression of pCREBS133/total CREB quantified from three independent experiments. N = 3 biological replicates/group. j, Fos gene expression in Neuro2a cells treated for 1 h with vehicle, BRP, scrambled BRP peptide or BRPL8A mutant (100 µg/ml). N = 3 biological replicates/group. k, Fos gene expression in Neuro2a cells treated for 1 h with vehicle or BRP (100 µg/ml) in combination with different doses of BRPL8A mutant. N = 3 biological replicates/group. Data are presented as mean +/− S.E.M, P value = ns (non-significant), *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 by one-way Anova (a, b, i, j, k) or two-way Anova for multiple comparisons (c, e, g).
Supplementary information
Supplementary Information (download PDF )
Supplementary data scans for Fig. 4a,b,d,f,h,j,n and Extended Data Figs. 9a and 10h
Supplementary Data (P values) (download XLSX )
Source data (P values)
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Coassolo, L., B. Danneskiold-Samsøe, N., Nguyen, Q. et al. Prohormone cleavage prediction uncovers a non-incretin anti-obesity peptide. Nature 641, 192–201 (2025). https://doi.org/10.1038/s41586-025-08683-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-08683-y
This article is cited by
-
Non-incretin peptide curbs appetite
Nature Reviews Drug Discovery (2025)
-
Microbial unusual gene clusters without prominent core enzymes: natural products, biosynthesis and genome mining
The Journal of Antibiotics (2025)
-
A novel anti-obesity peptide that targets the hypothalamus
Nature Reviews Endocrinology (2025)