Abstract
Immune checkpoint blockade (ICB) has transformed cancer therapy1,2. The efficacy of immunotherapy depends on dendritic cell-mediated tumour antigen presentation, T cell priming and activation3,4. However, the relationship between the key transcription factors in dendritic cells and ICB efficacy remains unknown. Here we found that ICB reprograms the interplay between the STAT3 and STAT5 transcriptional pathways in dendritic cells, thereby activating T cell immunity and enabling ICB efficacy. Mechanistically, STAT3 restrained the JAK2 and STAT5 transcriptional pathway, determining the fate of dendritic cell function. As STAT3 is often activated in the tumour microenvironment5, we developed two distinct PROTAC (proteolysis-targeting chimera) degraders of STAT3, SD-36 and SD-2301. STAT3 degraders effectively degraded STAT3 in dendritic cells and reprogrammed the dendritic cell–transcriptional network towards immunogenicity. Furthermore, STAT3 degrader monotherapy was efficacious in treatment of advanced tumours and ICB-resistant tumours without toxicity in mice. Thus, the crosstalk between STAT3 and STAT5 transcriptional pathways determines the dendritic cell phenotype in the tumour microenvironment and STAT3 degraders hold promise for cancer immunotherapy.
Similar content being viewed by others
Main
Immunotherapy, including ICB and T cell therapy, uses the immune system to eliminate cancer cells1,2,6. However, most patients are either poorly responsive to ICB or develop therapeutic resistance. Emerging evidence suggests that ICB does not trigger potent immune responses in patients with limited and impaired dendritic cells3 (DCs) in the tumour microenvironment (TME). DCs present tumour antigens to prime and activate T cells, thereby driving anti-tumour immunity4,7,8,9,10,11. In line with this, increased type 1 dendritic cell (DC1) infiltration correlates with prolonged survival in patients with cancers12,13,14. However, the TME can impede DC1 maturation and function, partly owing to disruptive environmental signals4,15,16,17. Thus, understanding how the DC phenotype is shaped in the TME and exploring strategies to enhance DC1 function are crucial for developing more effective cancer immunotherapy.
Alterations in STAT signalling pathways exert profound effects on anti-tumour responses in the TME18. For example, STAT3 is often activated and mediates immune inhibition in the TME5,19. STAT3 activation leads to the production of various pro-tumour factors, including VEGF and IL-6, impeding DC maturation and function20,21. Moreover, STAT3 signalling can inhibit T helper 1 (TH1)-type chemokine expression and subdue DC tumour trafficking, resulting in the exclusion of T cells from the TME21. By contrast, STAT5 is activated in response to cytokine signals, such as GM-CSF and IL-222, and has a positive role in the anti-tumour immune response23,24. On this basis, we hypothesized that the balance between the STAT5 and STAT3 transcriptional pathways in DCs may shape the fate of distinct immune responses in the TME, determining ICB responses in patients with cancer.
In this study, we analysed single-cell and bulk RNA-sequencing (RNA-seq) datasets from tumour tissues in patients with cancer who were receiving ICB. We found that the transcriptional pathways of STAT5 and STAT3 in DC1s correlated with T cell immunity and was associated with clinical outcome in patients treated with ICB. Moreover, genetic deletion and pharmacologic inhibition of STAT3 signalling led to DC1 activation and profound anti-tumour T cell immune responses. Furthermore, we provide proof of concept that pharmacological degradation of STAT3 can treat multiple tumour types as monotherapy and sensitize tumours to ICB.
Role of DC STAT5 and STAT3 in ICB
The STAT5 and STAT3 transcriptional pathways regulate tumour immune responses21,23,24. We thus reasoned that interplay between these two transcriptional pathways in immune cells, particularly in DCs, influences ICB responses in patients with cancer. To examine this possibility, we analysed RNA-seq data from tumour tissues collected before ICB treatment in a patient cohort (cohort 1) at the University of Michigan25 (Extended Data Table 1). As previously reported25, among these patients, 17% achieved a complete response to ICB, whereas 14% achieved partial response, 17% had stable disease and 51% had progressive disease according to RECIST (response evaluation criteria in solid tumours) criteria (Fig. 1a), and the complete response group exhibited the longest overall survival rate in cohort 1 (Fig. 1b). To determine whether the clinical response is associated with the effector T cell-mediated anti-tumour response, we evaluated outcomes stratified by expression of T effector cell signatures26. Patients with a high CD8+ effector T cell score exhibited improved overall survival (Fig. 1c). Furthermore, we calculated a DC1 maturation score27. We found that patients with a high DC1 maturation and function score demonstrated improvements in overall survival resembling those in patients with a high CD8+ effector T cell score (Fig. 1d). This similarity in survival was mirrored by a strong positive correlation between CD8+ effector T cell and DC1 maturation scores (Fig. 1e). Thus, DC-mediated antigen cross-presentation and T cell activation may drive ICB response in patients with cancer.
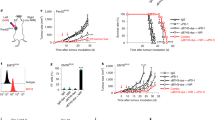
a–d, DC1 score and overall survival. a, Pie chart of number and percentages show RECIST-defined responses to ICB in cohort 1. CR, complete response; PD, progressive disease; PR, partial response; SD, stable disease. b–d, Kaplan–Meier survival curves were stratified by clinical response groups (b), cytotoxic T lymphocyte (CTL) score (c) and median DC1 maturation score (d). n = 92; log-rank test. e, Pearson correlation analysis shows a correction between CTL score and DC1 maturation score. The shaded area represents the 95% confidence interval. n = 92. f,g, Kaplan–Meier curves of overall survival in patients from cohort 1 were stratified by DC score and STAT5/STAT3 expression (f) or STAT5 transcriptional pathway/STAT3 expression (g). n = 92; P = 0.049 (f) and P = 0.00018 (g), log-rank test. h, Kaplan–Meier curves of overall survival in cohort 2 were stratified by DC score and the ratio of STAT5 versus STAT3. n = 114; P = 0.0041, log-rank test. i–l, STAT5 and STAT3 signalling signatures in DCs in cohort 3. i,j, STAT5 (i) and STAT3 (j) gene signatures in DCs were analysed in post-ICB samples of responders and the respective pre-ICB samples. k,l, STAT5 (k) and STAT3 (l) gene signatures within DCs were compared between non-responders and responders. In box plots, the central line represents the median, the box extends from the 25th to 75th percentiles, and whiskers span the smallest and largest values within 1.5× the interquartile range. Outliers are shown as individual points. Responders: pre- (332 DCs) and post- (170 DCs) treatment (4 patients). Non-responders: pre- (215 DCs), post- (194 DCs) treatment (6 patients). **P = 0.0012 (j), ***P = 0.00092 (k), **P = 0.0045 (l) and ****P < 0.0001 (i), Wilcoxon rank-sum test. P values are two-sided without correction for multiple comparisons.
To explore the role of the relationship between the STAT5 and STAT3 transcriptional pathways in the context of DC1 maturation and function, we stratified patients (cohort 1) by the median DC1 maturation score and the median ratio of STAT5/STAT3 expression into four groups: DC1hiSTAT5/STAT3hi, DC1hiSTAT5/STAT3low, DC1lowSTAT5/STAT3hi, and DC1lowSTAT5/STAT3low. Among the four patient groups, those classified as DC1hiSTAT5/STAT3hi had the longest overall survival, whereas the other groups exhibited similar survival trends to each other (Fig. 1f). We observed similar results when we stratified patients by the median ratio of STAT5 transcriptional pathway activation and STAT3 expression (Fig. 1g) and by excluding patients with tumours that were inherently unresponsive to ICB (Extended Data Fig. 1a). We analysed additional patients with melanoma28,29 (cohort 2), a type of cancer that is sensitive to anti-PD-1 and anti-CTLA-4 therapy (Extended Data Table 2). DC1hiSTAT5/STAT3hi in cohort 2 exhibited the longest overall survival, whereas the DC1lowSTAT5/STAT3low patients had the shortest (Fig. 1h and Extended Data Fig. 1b,c). Thus, the relationship between DC1 maturation and the STAT5 and STAT3 transcriptional pathways may affect responsiveness to ICB.
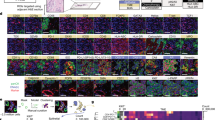
To solidify this finding, we analysed a single-cell RNA-sequencing (scRNA-seq) dataset from patients with triple-negative breast cancer (TNBC) (cohort 3), and examined DCs in tumour tissues collected before and after ICB treatment30. We conducted an analysis of the STAT5 and STAT3 transcriptional pathways in DC clusters. The analysis revealed a bimodal distribution of the STAT3 transcriptional pathway, with the second mode exhibiting a relatively high STAT3 signalling activity (Extended Data Fig. 1d). We conducted a detailed analysis of the STAT5 and STAT3 transcriptional pathways in DC clusters with relatively high STAT3 expression. Of note, STAT5 transcriptional signalling increased in post-ICB samples compared with pre-ICB samples among responders to ICB (Fig. 1i), but such an increase was not observed in non-responders (Extended Data Fig. 1e). Conversely, STAT3 transcriptional signalling showed a decrease in post-ICB samples of responders compared with their pre-ICB samples (Fig. 1j), with no significant change observed in non-responders (Extended Data Fig. 1f). Furthermore, post-ICB samples showed an increase in STAT5 transcriptional signalling and a reduction in STAT3 transcriptional signalling in responders relative to non-responders (Fig. 1k,l). We then examined the STAT5 and STAT3 transcriptional signalling pathways in specific DC subsets in these patients30 (cohort 3). Using the Louvain algorithm for shared nearest neighbours clustering and uniform manifold approximation and projection (UMAP) visualization to distinguish the cells, we conducted an analysis with two major distinct DC clusters, conventional DCs (cDCs) and monocyte-derived DCs (moDCs) (Extended Data Fig. 1g). In the cDC subset, the post-ICB samples from responders displayed increased STAT5 transcriptional signalling and decreased STAT3 transcriptional signalling compared with pre-ICB samples (Extended Data Fig. 1h,i). These changes were not discernible in the non-responder group (Extended Data Fig. 1j,k). In moDCs, an increase in STAT5 signalling was observed in the post-ICB samples from responders, whereas STAT3 transcriptional signalling remained unchanged compared with pre-ICB samples in both responders and non-responders (Extended Data Fig. 1l–o). Furthermore, in the post-ICB samples, we observed a noticeable increase in STAT5 transcriptional signalling and a corresponding decrease in STAT3 transcriptional signalling in the responders compared to the non-responders within cDCs (Extended Data Fig. 1p,q). In moDCs, neither STAT5 nor STAT3 transcriptional signalling showed significant differences between post-ICB samples from responders and non-responders (Extended Data Fig. 1r,s). Collectively, ICB treatment dynamically reprograms the STAT5 and STAT3 transcriptional signalling pathways in cDCs, affecting ICB efficacy.
STAT3 restrains STAT5-driven DC function
Given the negative effect of STAT3 transcriptional activity on DC maturation and ICB response, we examined whether alteration of STAT3 activity would affect DC phenotype. We used short hairpin RNAs (shRNAs) targeting Stat3 (shStat3) in JAWSII cells, a DC line31,32 (Fig. 2a). Knocking down STAT3 expression resulted in a robust increase in phosphorylation of STAT5 (pSTAT5) and a minor increase in pSTAT1, but had no effect on pSTAT6 and p-p65 in JAWSII cells cultured with GM-CSF (Fig. 2a). We crossed Stat3fl/fl mice with Xcr1cre mice (which express mCherry under the control of the Xcr1 promoter) and generated cDC1-specific STAT3 knockout (Stat3−/−) mice. We used Stat3fl/flXcr1+/+ mice as the Stat3+/+ controls, enabling us to use mCherry fluorescence to validate knockout specificity in the mouse offspring (Extended Data Fig. 2a). No mCherry expression was detected in other immune cells in Stat3−/− mice (Extended Data Fig. 2b). We confirmed the absence of STAT3 protein in cDC1s generated from Stat3−/− mice (Fig. 2b and Extended Data Fig. 2c). At 6 to 10 weeks of age, Stat3+/+ and Stat3−/− mice exhibited similar body size (Extended Data Fig. 2d), similar appearance of lymphoid organs (Extended Data Fig. 2e) and equivalent levels of CD8+ and CD4+ T cells in tissues (Extended Data Fig. 2f,g). Notably, we detected an increase in STAT5 activation in Stat3−/− cDC1s compared with Stat3+/+ cDC1s (Fig. 2b). An unbiased phospho-receptor kinase array showed higher levels of pSTAT5 in Stat3−/− cDC1s than Stat3+/+ cDC1s (Fig. 2c). Thus, STAT3 restrains STAT5 activation in DCs. Flow cytometry analysis revealed higher levels of typical DC maturation markers, including major histocompatibility complex class I (MHCI; Fig. 2d), major histocompatibility complex class II (MHCII; Fig. 2e), CD80 (Fig. 2f and Extended Data Fig. 2h) and CD86 (Fig. 2g and Extended Data Fig. 2h) in Stat3−/− cDC1s than in Stat3+/+ cDC1s.
a,b, Immunoblots of transcription factors in shStat3 JAWSII cells (a) and Stat3+/+ and Stat3−/− cDC1s (b). n = 3. c, Scatter plots of relative pixel density versus corresponding −log10(P value) for a phospho-antibody array incubated with lysate from LPS-stimulated Stat3+/+ and Stat3−/− cDC1s. n = 4. d,e, Fluorescence-activated cell sorting (FACS; left) and mean fluorescence intensity (MFI; right) of MHCI (d) and MHCII (e) in Stat3+/+ and Stat3−/− cDC1s. Data are mean ± s.e.m., n = 3; **P = 0.0084 (d) and *P = 0.0125 (e), unpaired two-tailed t-test. f,g, FACS and MFI of CD80 (f) and CD86 (g) in Stat3+/+ or Stat3−/− cDC1s. Data are mean ± s.e.m., n = 3; **P = 0.0098 (f) and *P = 0.0453 (g), unpaired two-tailed t-test. h,i, Immunoblots of transcription factors in JAWSII cells cultured with GM-CSF and transfected with shGmrβ (h) or shJak2 (i). n = 3. j,k, Immunoprecipitation with anti-GMRβ (j) or anti-JAK2 (k) shows the interaction between GMRβ and transcription factors in activated Stat3+/+ and Stat3−/− cDC1s. n = 2. l,m, FACS analysis of MHCII (l) and IL-12 enzyme-linked immunosorbent assay (ELISA; m) in Stat3+/+ and Stat3−/− cDC1s cultured with STAT5i. Data are mean ± s.e.m., n = 3, **P = 0.0079 (l); n = 4, ****P < 0.0001 (m); one-way ANOVA. n–q, Heat map comparing expression of DC maturation genes in Stat3+/+ versus Stat3−/− cDC1s as z-scores (n) or calculated with EdgeR (o), and GSEA shows differences in expression of genes associated with antigen processing and presentation (p) and IL-12 production (q). NES, normalized enrichment score. r,s, RNA-seq analysis of LPS-stimulated Stat3+/+ and Stat3−/− cDC1s shows changes in genes associated with multiple signalling pathways (r) and STAT5 signalling (s). t–w, ChIP–qPCR shows promoter occupancy of H2-D1 (t), H2-Eb1 (u), Cd80 (v) and Cd86 (w) in LPS-treated Stat3+/+ or Stat3−/− cDC1s. Data are mean ± s.e.m. fold change over control, n = 3; **P = 0.0017 (t), ***P = 0.0003 (u), *P = 0.0271 (v) and **P = 0.0074 (w); one-way ANOVA.
To explore how STAT5 activation is regulated in DCs, we assessed the expression profile of a range of cytokine receptors that are capable of activating STAT5, including the receptors for IL-2, IL-3, IL-5, IL-7, IL-15, IL-21 and GM-CSF, in cDC1s. Among these cytokine receptors, we detected higher transcript levels of GM-CSF receptors, Gmrα (also known as Csf2ra) and Gmrβ (also known as Csf2rb), compared with other cytokine receptors in cDC1s (Extended Data Fig. 2i). Knocking down GMRβ with specific shRNAs resulted in a decrease in phosphorylation of JAK2, STAT5 and STAT3 (Fig. 2h), but not of JAK1, TYK2, JAK3 and STAT1 in JAWSII cells cultured with GM-CSF (Extended Data Fig. 2j). Moreover, knocking down JAK2 caused a profound reduction of pSTAT5 and a moderate reduction of pSTAT3, but not pSTAT1, in JAWSII cells in response to GM-CSF (Fig. 2i). GM-CSF stimulation induced the phosphorylation of JAK2, but not JAK1, JAK3 or TYK2 in JAWSII cells (Extended Data Fig. 2k). Similar findings have been reported in human neutrophils33. Thus, GM-CSF and signalling pathways downstream of the GM-CSF receptor may regulate STAT5 activation via JAK2 in DCs.
We next investigated whether STAT3 is involved in the GMR–JAK2–STAT5 axis in DCs. We found that Stat3−/− and Stat3+/+ cDC1s expressed similar levels of GM-CSF (Extended Data Fig. 2l) and GMRβ (Extended Data Fig. 2m). Then, we performed a reciprocal co-immunoprecipitation assay with anti-GMRβ and anti-JAK2 in Stat3−/− and Stat3+/+ cDC1s. Co-immunoprecipitation experiments demonstrated the co-existence of GMRβ, JAK2 and STAT5 in activated cDC1s (Fig. 2j,k). Despite similar levels of GMRβ and JAK2 in the co-immunoprecipitation products of Stat3−/− and Stat3+/+ cDC1s (Fig. 2j,k), genetic deletion of STAT3 led to an increase in the interaction between GMRβ and STAT5 (Fig. 2j) and between JAK2 and STAT5 (Fig. 2k). shStat3 JAWSII cells (Fig. 2a) and Stat3−/− cDC1s (Fig. 2b) expressed higher levels of pSTAT5 than their counterparts. The data suggest that STAT3 may restrain the GMRβ–JAK2–STAT5 signalling pathway, thereby affecting DC phenotype and function in vitro (Fig. 2d–g and Extended Data Fig. 2h). Given that increased IL-6 and activated STAT3 are often observed in the TME, we assessed the expression of IL-6 and its receptors in human and mouse DCs. IL-6-expressing DCs represented a negligible fraction among IL-6-positive cells from patients with TNBC30 (cohort 3) (Extended Data Fig. 2n). Mouse Stat3+/+ and Stat3−/− cDC1s expressed low levels of IL-6 (Extended Data Fig. 2o,p). Moreover, cDC1s expressed similar amounts of IL-6 receptor transcripts (Il6ra and Il6st) (Extended Data Fig. 2q,r) and proteins (IL-6Rα and GP130) (Extended Data Fig. 2s) in Stat3+/+ and Stat3−/− cDC1s. These results suggest that a paracrine-activated IL-6–STAT3 pathway regulates the GMRβ–JAK2–STAT5 signalling pathway in DCs, affecting DC phenotype in the TME.
To validate this possibility in vivo and elucidate the biological involvement of JAK2, we treated B16F10 or MC38 colon carcinoma-bearing Stat3−/− and Stat3+/+ mice with the JAK2 inhibitor FLLL32. As expected, Stat3−/− cDC1s expressed higher levels of pSTAT5 (Extended Data Fig. 2t,u) and maturation markers compared with Stat3+/+ cDC1s (Extended Data Fig. 2v–x), whereas treatment with FLLL32 abolished this effect (Extended Data Fig. 2t–x). Then, we treated Stat3−/− and Stat3+/+ cDC1s with the STAT5 inhibitor STAT5i34. We observed higher expression levels of MHCII (Fig. 2l) and IL-12 (Fig. 2m) in Stat3−/− cDC1s than in Stat3+/+ cDC1s, and these effects were abolished by the STAT5i (Fig. 2l,m). Therefore, STAT3 may restrain JAK2-mediated STAT5 activation in DC1s and the functional maturation of these cells.
To complement these observations and systemically explore how STAT3 regulates DC phenotype, we conducted RNA-seq analysis of Stat3+/+ and Stat3−/− cDC1s. We observed a noticeable difference in the expression levels of maturation- and function-related genes between Stat3+/+ and Stat3−/− cDC1s (Fig. 2n,o). Gene set enrichment analysis (GSEA) also showed a positive enrichment of maturation and function signatures in differentially expressed genes in Stat3−/− cDC1s relative to Stat3+/+ cDC1s, including antigen processing and presentation (Fig. 2p), IL-12 production (Fig. 2q), CCR chemokine receptor binding (Extended Data Fig. 2y) and IFNγ-production pathways (Extended Data Fig. 2z). Consistently, the top upregulated canonical pathways in response to lipopolysaccharide (LPS) in Stat3−/− cDC1s compared with Stat3+/+ cDC1s included antigen presentation, DC maturation and interferon signalling (Fig. 2r). Differential gene expression analysis revealed that genes associated with the STAT5 pathway were upregulated with LPS treatment in Stat3−/− cDC1s compared with Stat3+/+ cDC1s (Fig. 2s). Furthermore, STAT5 chromatin immunoprecipitation followed by quantitative PCR (ChIP–qPCR) analysis revealed enhanced STAT5 binding at the promoter sites of H2-D1 (Fig. 2t), H2-Eb1 (Fig. 2u), Cd80 (Fig. 2v) and Cd86 (Fig. 2w) in Stat3−/− cDC1s compared with Stat3+/+ cDC1s. Thus, STAT3 inhibits DC1 maturation and function in a JAK2–STAT5-dependent transcriptional manner.
STAT3 dampens DC1-mediated tumour immunity
We next investigated the role of STAT3 in DC1-mediated anti-tumour response. We primed OT-I cells with Stat3+/+ and Stat3−/− cDC1s with the SIINFEKL peptide (Fig. 3a–d) or without antigen (Extended Data Fig. 3a–d), and subsequently assessed T cell response. We observed that Stat3−/− cDC1s induced increased proliferation (Fig. 3a), effector cytokine expression (Fig. 3b,c) and GZMB production (Fig. 3d) compared with Stat3+/+ cDC1s. These effects were not observed in the absence of SIINFEKL peptide-loaded cDC1s (Extended Data Fig. 3a–d).
a–d, OT-I cells were cultured with Stat3+/+ or Stat3−/− cDC1s in the presence of SIINFEKL peptides. a–d, FACS analysis of Ki-67+ (a), IFNγ+ (b), TNF+ (c) and GZMB+ (d) T cells. Data are mean ± s.e.m., n = 4; ****P < 0.0001 (a–c) and **P = 0.009 (d), unpaired two-tailed t-test. e–r, MC38 tumours were inoculated into Stat3+/+ and Stat3−/− mice. e–g, Tumour volume (e), images (f) and mass (g). h–r, FACS analysis of pSTAT5 (h–j), MHCII (k,l) and CD86 (m,n) in tumour-infiltrating cDC1s, and the percentage of tumour-infiltrating IFNγ+ (o,p) and GZMB+ (q,r) CD8+ T cells. One representative flow histogram or dot plot is shown (h,k,m,o,q). Data are mean ± s.e.m., n = 5 (e,g), n = 3 (i,j,r) and n = 4 (l,n,p); ****P < 0.0001 (e) **P = 0.0067 (g), *P = 0.0144 (i), *P = 0.0198 (j), *P = 0.0231 (l), **P = 0.003 (n), *P = 0.0223 (p) and *P = 0.0405 (r); two-way ANOVA (e), unpaired two-tailed t-test (g,i,j,l,n,p,r). s–z, B16F10 tumours were inoculated into Stat3+/+ and Stat3−/− mice. s,t, Tumour volume (s) and mass (t). u–z, FACS analysis of MHCII (u,v) and CD86 (w,x) in tumour-infiltrating cDC1s, and the percentage of tumour-infiltrating IFNγ+ (y) and GZMB+ (z) CD8+ T cells. One representative flow histogram or dot plot is shown (u,w,y,z). Data are mean ± s.e.m., n = 5 (s,t), n = 3 (v,x) and n = 4 (y,z); **P = 0.0018 (s), *P = 0.045 (t), *P = 0.0197 (v), **P = 0.0015 (x), ***P = 0.0005 (y) and **P = 0.0085 (z); two-way ANOVA (s), unpaired two-tailed t-test (t,v,x–z).
To explore the in vivo effect of STAT3 on DCs in response to tumour challenge, we inoculated MC38 cells subcutaneously into Stat3+/+ and Stat3−/− mice. Loss of STAT3 in cDC1s resulted in reduced tumour progression compared with Stat3+/+ mice, as indicated by tumour size (Fig. 3e,f) and mass (Fig. 3g). To examine whether STAT3 deficiency affected STAT5 activation in intratumoral cDC1s, we gated on tumour-infiltrating cDC1s and detected higher levels of STAT5 activation in Stat3−/− cDC1s than in Stat3+/+ cDC1s (Fig. 3h–j and Extended Data Fig. 3e). Genetic deletion of STAT3 in cDC1s led to a marked increase in the interaction between JAK2 and STAT5 in cDC1s in the TME (Extended Data Fig. 3f). Stat3−/− cDC1s in the TME (Fig. 3k–n) and tumour-draining lymph nodes (TDLNs) (Extended Data Fig. 3g–j) exhibited higher expression levels of MHCII and CD86 compared with Stat3+/+ cDC1s. This was accompanied by enhanced effector T cell responses in the TME and TDLN, as indicated by higher percentages of IL-2+ (Extended Data Fig. 3k–o), IFNγ+ (Fig. 3o,p and Extended Data Fig. 3p,q) and GZMB+ (Fig. 3q,r) CD8+ T cells in Stat3−/− mice compared with Stat3+/+ mice.
We extended our studies to B16F10 melanoma, a less immunogenic type of tumour. Again, B16F10 tumours progressed more slowly in Stat3−/− mice compared with Stat3+/+ mice (Fig. 3s,t). B16F10 tumour-infiltrating Stat3−/− cDC1s exhibited higher levels of maturation and function markers compared with Stat3+/+ cDC1s (Fig. 3u–x and Extended Data Fig. 3r–u). Moreover, we also observed higher percentages of IFNγ+ (Fig. 3y), GZMB+ (Fig. 3z), IL-2+ (Extended Data Fig. 3v,w) and TNF+ (Extended Data Fig. 3x,y) CD8+ T cells in the TME in Stat3−/− mice than in Stat3+/+ mice. Thus, STAT3 deficiency in DCs enhances T cell function, leading to improved tumour control in vivo.
STAT3 degraders for treatment of advanced tumours
STAT3 has been pursued as a cancer therapeutic target5,35. Small-molecule inhibitors or oligonucleotides have been developed to target STAT35, with limited clinical success35. An indirect strategy to target STAT3 is through inhibition of JAK2 activity, such as with pacritinib5. However, JAK2 inhibition may affect the positive effects of STAT5 on DC1s (as we show here) and T cells24. Thus, we used SD-36, a highly selective and effective PROTAC degrader of the STAT3 protein36 (Extended Data Fig. 4a). We treated cDC1s with SD-36 and observed a dose-dependent reduction in the amount of STAT3 protein (Fig. 4a). Next, we treated Stat3+/+ and Stat3−/− cDC1s and subsequently stimulated them with LPS. We observed enhanced STAT5 activation in Stat3+/+ cDC1s treated with SD-36 (Fig. 4b). Knocking out Stat3 led to increased STAT5 activation, and the addition of SD-36 did not further increase STAT5 activation in Stat3−/− cDC1s (Fig. 4b). To pharmacologically validate our genetic studies on the role of STAT3 in DCs (Fig. 2 and Extended Data Fig. 2), we performed a reciprocal co-immunoprecipitation assay with anti-GMRβ and anti-JAK2 in cDC1s treated with SD-36. Pharmacological degradation of STAT3 by SD-36 led to a dose-dependent increase in the interaction between GMRβ and STAT5 (Fig. 4c) and between JAK2 and STAT5 (Fig. 4d). Moreover, DCs treated with SD-36 exhibited increased expression of MHCI (Fig. 4e,f), MHCII (Fig. 4g,h), CD80 (Fig. 4i,j) and CD86 (Fig. 4k,l) compared with the control group. As genetically knocking down STAT3 caused a slight increase in pSTAT1 in JAWSII cells (Fig. 2a), we tested whether STAT1 is involved in SD-36-mediated modification of DC phenotype. Knocking down STAT1 (Extended Data Fig. 4b) had no effect on the expression of MHCI (Extended Data Fig. 4c), MHCII (Extended Data Fig. 4d) or CD80 (Extended Data Fig. 4e) compared with the control group, and SD-36 remained equally effective in enhancing the expression of these DC maturation molecules in shStat1 JAWSII cells and wild-type JAWSII cells (Extended Data Fig. 4c–e). Genetic deletion of STAT1 (Extended Data Fig. 4f) had no effect on STAT5 activation in cDC1s (Extended Data Fig. 4g), as shown in Stat1+/+ and Stat1−/− cDC1s. Stat1+/+ and Stat1−/− cDC1s expressed similarly high levels of MHCI, MHCII and CD80 in response to SD-36 (Extended Data Fig. 4h–j). Thus, depletion of STAT3 improves DC function via STAT5 activation.
a, Immunoblot showing STAT3 expression in mouse cDC1s treated with different concentrations of SD-36 in vitro. b, Immunoblot showing expression of pSTAT5 and STAT5 in cDC1s from Stat3+/+ and Stat3−/− mice treated with SD-36 (200 nM) for 48 h and LPS (20 ng ml−1) for 1 h. In a,b, one of three experiments is shown. c,d, cDC1s were treated with different doses of SD-36 for 24 h and were immunoprecipitated (IP) with anti-GMRβ (c) or anti-JAK2 (d). The immunoprecipitation shows the interaction between GMRβ, JAK2, STAT3 and STAT5 in cDC1s. One of two experiments with repeats is shown. e–l, FACS analysis of MHCI (e,f), MHCII (g,h), CD80 (i,j) and CD86 (k,l) on cDC1s treated with SD-36 (e–h) and SD-36 plus LPS (i–l). Representative histograms are shown. Data are mean ± s.e.m., n = 4; *P = 0.0241 (h), *P = 0.023 (j), *P = 0.0329 (l) and **P = 0.0015 (f), unpaired two-tailed t-test. m–p, Mice were inoculated with 4T1 (m), MC38 (n), ID8 (o) or LLC (p) cells and were treated with SD-36 (20 mg kg−1) every 3 days, and tumour volumes were monitored. Data are mean ± s.e.m.; n = 6 (m,p) and n = 5 (n,o); **P = 0.0021 (p) and ****P < 0.0001 (m–o), two-way ANOVA. q–r. Mice bearing CT26 tumours (500 mm3 per tumour) were treated with SD-36 or vehicle and tumour volumes (q) and mouse survival (r) were monitored. Data are mean ± s.e.m.; n = 8; *P = 0.0111 (r), log-rank test. s,t, Luciferase radiance images (s) and total fluxes of metastatic 4T1 tumours (t) from mice treated with SD-36. Data are mean ± s.e.m.; vehicle: n = 5, SD-36: n = 6; *P = 0.0348, two-way ANOVA.
We examined the role of SD-36 in the tumour immune response in vivo. We inoculated B16F10 melanoma into C57BL/6J mice and treated them with different doses of SD-36. SD-36 at 10 mg kg−1 moderately but significantly slowed down B16F10 tumour progression compared with controls (Extended Data Fig. 4k). We observed a marked inhibitory effect of SD-36 at 20 mg kg−1 on tumour growth in mice bearing 4T1 mammary carcinoma (Fig. 4m), MC38 colon carcinoma (Fig. 4n and Extended Data Fig. 4l,m), ID8 ovarian cancer (Fig. 4o) or Lewis lung carcinoma (LLC) (Fig. 4p). Consistently, we observed that mice treated with SD-36 at 100 mg kg−1 also exhibited a slower rate of progression of B16F10, CT26 and EMT6 tumours compared with the control group (Extended Data Fig. 4n–p) and had no obvious effect on the body weight of these mice (Extended Data Fig. 4q–s). Thus, SD-36 monotherapy attenuates the growth of multiple tumours including LLC, an ICB-resistant tumour37. Large, metastatic cancers are often resistant to ICB38,39. We initiated the treatment with SD-36 in CT26 tumour-bearing mice with tumour volumes greater than 500 mm3. Of note, SD-36 remained therapeutically effective at this late stage (Fig. 4q and Extended Data Fig. 4t,u) as shown by increased mouse survival compared with the control group (Fig. 4r). Finally, we treated 4T1 metastatic tumour-bearing mice with SD-36. This treatment effectively reduced metastatic 4T1 tumour volume (Fig. 4s,t). Thus, SD-36 monotherapy is effective in treating advanced, metastatic and ICB-resistant cancers.
STAT3 degradation in DCs boosts immunity
We next studied whether the anti-tumour effect of SD-36 was immune- and/or DC-dependent. To this end, we treated MC38 tumour-bearing NOD.SCID γcnull (NSG) mice with a low dose of SD-36 (20 mg kg−1). This dose of SD-36 had no effect on tumours, as shown by tumour volume (Fig. 5a) and mass (Fig. 5b) in NSG mice. A high dose of SD-36 (100 mg kg−1) had a modest but significant inhibitory effect on tumour progression in NSG mice (Extended Data Fig. 5a). Given that both innate and adaptive immunity are deficient in NSG mice, we treated MC38-bearing Rag1−/− mice, which have functional innate immunity, with SD-36. Similarly, a low dose of SD-36 (20 mg kg−1) did not have any effect on tumour volume (Fig. 5c), weight (Fig. 5d) or appearance (Fig. 5e) in Rag1−/− mice. Thus, the therapeutic effect of SD-36 is dependent on an intact adaptive immune system. We treated MC38-bearing wild-type mice with anti-CD8 antibody to deplete CD8+ T cells (Extended Data Fig. 5b). CD8+ T cell depletion resulted in the loss of SD-36 therapeutic effectiveness (Fig. 5f). Indeed, SD-36 treatment increased the fraction of CD8+ T cells that express TNF (Fig. 5g,h), IFNγ (Fig. 5i,j) and GZMB (Fig. 5k,l) in 4T1 tumours and TDLNs (Extended Data Fig. 5c–f) and LLC (Extended Data Fig. 5g–r). Thus, STAT3 degradation enhances CD8+ T cell-mediated anti-tumour efficacy.
a–e, Tumours in MC38-bearing NSG (a,b) and Rag1−/− mice (c–e) treated with SD-36 were monitored. n = 5 (a,b) and n = 6 (c,d); two-way ANOVA (a,c) and unpaired two-tailed t-test (b,d). Scale bar, 1 cm. NS, not significant. f, Tumour volume was monitored in MC38-bearing mice treated as indicated. Data are mean ± s.e.m., n = 6; **P = 0.0018, two-way ANOVA. g–l, FACS analysis of 4T1-infiltrating TNF+ (g,h), IFNγ+ (i,j) and GZMB+ (k,l) CD8+ T cells from mice treated with SD-36. Data are mean ± s.e.m., n = 6; *P = 0.0175 (h), *P = 0.0232 (j) and *P = 0.0404 (l), unpaired two-tailed t-test. m,n, Tumour volumes were monitored in Batf3−/− and wild-type mice bearing MC38 (m) or B16F10 (n) tumours and treated with SD-36. Data are mean ± s.e.m.; n = 5 (m), and n = 8 (n); *P = 0.0158 (m) and *P = 0.0126 (n), two-way ANOVA. o, FACS analysis of STAT3 expression in different cell types from LLC-bearing mice. Data are mean ± s.e.m., n = 4; ****P < 0.0001, one-way ANOVA. p–r, FACS analysis of pSTAT3 (p,q) and pSTAT5 (r) in MC38-infiltrating cDC1s from mice treated with SD-36. Data are mean ± s.e.m., n = 5; *P = 0.0335 and ***P = 0.0002, unpaired two-tailed t-test. s–u, FACS analysis of MHCI (s), MHCII (t) and CD80 (u) in B16F10-infiltrating cDC1s from mice treated with SD-36. Data are mean ± s.e.m., n = 5; **P = 0.0021 (s), **P = 0.0035 (t) and **P = 0.0023 (u), unpaired two-tailed t-test. v, Tumour volume in B16F10-bearing Stat3+/+ and Stat3−/− mice treated with SD-36. Data are mean ± s.e.m., n = 7; **P = 0.0067 and ***P = 0.0006, two-way ANOVA. w, B16F10-bearing Batf3−/− mice were transferred with Stat3+/+ and Stat3−/− cDC1s and treated with SD-36, and tumour volume was monitored. Data are mean ± s.e.m., n = 7; **P = 0.0094, two-way ANOVA. x, Stat5+/+ and Stat5−/− cDC1s were transferred into MC38-bearing Batf3−/− mice, treated with SD-36, and tumour volume was monitored. Data are mean ± s.e.m., n = 7; ****P < 0.0001, two-way ANOVA. y,z, Tumour volume was monitored in mice bearing MC38 (y) and B16F10 (z) tumours and treated as indicated. Data are mean ± s.e.m.; n = 6 (y) and n = 7 (z); **P = 0.0094 (z) and ***P = 0.0009 (y), two-way ANOVA.
To investigate whether DC1s were involved in the therapeutic efficacy of SD-36, we treated tumour-bearing Batf3−/− mice lacking cDC1s40. The anti-tumour efficacy of SD-36 therapy was absent in Batf3-deficient mice bearing MC38 (Fig. 5m), B16F10 (Fig. 5n) or LLC (Extended Data Fig. 5s) tumours. STAT3 was highly expressed in intratumoral DCs in wild-type mice bearing LLC tumours (Fig. 5o), and SD-36 treatment resulted in a rapid and efficient degradation of STAT3 protein in DCs compared with tumour cells (Extended Data Fig. 5t). Of note, the intracellular concentration of SD-36 was four times higher in DCs than in tumour cells (Extended Data Fig. 5u). These results suggest that SD-36 effectively induces degradation of STAT3 in DCs to achieve its therapeutic efficacy in vivo.
Next, we found that SD-36 treatment reduced pSTAT3 and increased pSTAT5 in MC38 tumour-infiltrating cDC1s (Fig. 5p–r). We also observed enhanced STAT5 activation in DCs isolated from the spleens of MC38-bearing mice after SD-36 treatment (Extended Data Fig. 5v). SD-36 treatment increased DC maturation and function markers in cDC1s in B16F10 tumours (Fig. 5s–u).
We examined whether STAT3 in DCs is a direct functional target of SD-36 therapy in vivo in tumour-bearing mice. We treated B16F10-bearing Stat3+/+ and Stat3−/− mice with SD-36. SD-36 treatment inhibited tumour growth in Stat3+/+ mice, whereas this effect was abolished in Stat3−/− mice (Fig. 5v). Similar results were observed in MC38 tumour-bearing mice (Extended Data Fig. 5w). We next transferred Stat3+/+ or Stat3−/− cDC1s into B16F10 or MC38 tumour-bearing Batf3−/− mice and treated these mice with SD-36. Again, SD-36 improved the anti-tumour effect in Batf3−/− mice that received Stat3+/+ cDC1s, but not in those that received Stat3−/− cDC1s (Fig. 5w, Extended Data Fig. 5x).
We used STAT5i34 to assess the interplay between STAT3 and STAT5 in DCs (Fig. 2l,m). STAT5i can have off-target effects. To genetically validate whether STAT5 in DCs contributes to SD-36-mediated anti-tumour immunity, we transferred Stat5b−/− cDC1s or Stat5b+/+ cDC1s into MC38 tumour-bearing Batf3−/− mice and treated these mice with SD-36. SD-36 effectively controlled tumour progression in mice receiving Stat5b+/+ cDC1s, but not Stat5b −/− cDC1s (Fig. 5x). Therefore, SD-36 reprograms key transcription factors in cDC1s by degrading STAT3 and enhancing STAT5 signalling, thereby inducing potent anti-tumour immunity in vivo.
Finally, we examined whether SD-36 could synergize with ICB. To this end, we treated mice bearing MC38 or B16F10 tumours with PD-L1 monoclonal antibody, SD-36 or both. Anti-PD-L1 inhibited MC38 tumour growth but had no effect on B16F10 tumours, whereas SD-36 monotherapy efficiently slowed down MC38 and B16F10 tumour growth. The combined treatment profoundly inhibited progression of MC38 and B16F10 tumours (Fig. 5y,z). We further tested SD-36 in human DC-mediated T cell activation in the context of ICB. We isolated DCs from fresh human ovarian cancer tissues and treated these DCs with SD-36 and/or anti-human PD-L1 monoclonal antibody, and co-cultured these DCs with T cells. Treatment with SD-36 or anti-PD-L1 increased the numbers of polyfunctional human T cells, as shown by GZMB+IFNγ+ (Extended Data Fig. 6a). The combined treatment additionally upregulated the levels of polyfunctional T cells (Extended Data Fig. 6a,b).
To further investigate degradation of STAT3 as a potential immunotherapeutic strategy, we have developed SD-2301, another highly potent and selective STAT3 degrader (Extended Data Fig. 6c). In contrast to SD-36, which engaged the cereblon–cullin 4A complex for induced STAT3 degradation, SD-2301 used a high-affinity VHL ligand to recruit VHL–cullin 2 for STAT3 degradation. SD-2301 effectively degraded STAT3 in DCs in vitro (Extended Data Fig. 6d). Treatment with SD-2301 resulted in a potent STAT3 degradation in DCs in vivo in B16F10 tumour-bearing mice (Extended Data Fig. 6e). Of note, SD-2301 exhibited four- to five-fold improved potency compared with SD-36 towards STAT3 degradation in DCs (Extended Data Fig. 6f). An efficacy experiment showed that 5 mg kg−1 SD-2301 was effective in reducing B16 tumour progression and was four times more potent than SD-36 (Extended Data Fig. 6g). Treatment with SD-2301 also increased the number of effector CD8+ T cells, as shown by IFNγ (Extended Data Fig. 6h,i) and GZMB (Extended Data Fig. 6j,k) expression. In line with this, SD-2301 treatment increased DC maturation and function markers in cDC1s in B16F10 tumours (Extended Data Fig. 6l–n). Treatment with SD-2301 had no effect on CD31+ blood vessel densities in B16F10 (Extended Data Fig. 6o) and MC38 (Extended Data Fig. 6p) tumour tissues and did not affect the body weight of mice bearing B16F10 tumours (Extended Data Fig. 6q).
In addition, whereas anti-PD-L1 monotherapy had no effect on B16F10 tumour growth, combined SD-2301 and anti-PD-L1 synergistically inhibited tumour growth (Extended Data Fig. 6r). In the MC38 tumour model, anti-PD-L1 or SD-2301 monotherapy effectively inhibited tumour growth, and the combined therapy profoundly suppressed MC38 tumour progression (Extended Data Fig. 6s). Accordingly, SD-2301 treatment increased the proportion of MC38 tumour neoantigen-specific (KSPWFTTL) tetramer+CD8+ T cells compared with the control treatment (Extended Data Fig. 6t,u). These results highlight that SD-2301 may enhance DC-mediated antigen specific T cell priming and presentation in vivo.
Next, we explored whether DC1s and STAT3 are direct cellular and molecular targets of SD-2301 therapy in vivo. To address this, we treated B16F10-bearing Stat3+/+ and Stat3−/− mice with SD-2301. As expected, SD-2301 treatment inhibited tumour growth in Stat3+/+ mice, whereas this effect was abolished in Stat3−/− mice (Extended Data Fig. 6v). The anti-tumour efficacy of SD-2301 therapy was absent in Batf3-deficient mice bearing MC38 tumours (Extended Data Fig. 6w). These results indicate that cDC1s have a key role in the therapeutic efficacy of SD-2301 in vivo.
Pharmacokinetic evaluation showed that SD-2301 had an excellent pharmacokinetic profile in mice, as characterized by a slow clearance and high plasma exposure (Extended Data Fig. 6x). Moreover, SD-2301 was highly potent and selective in inducing degradation of STAT3 over all other STAT proteins in human peripheral blood monocyte cells in vitro (Extended Data Fig. 6y). Together, our results suggest that STAT3 serves as a brake to restrain JAK2-mediated STAT5 activation in DCs and STAT3 degradation in DCs elicits potent anti-tumour immunity, thereby being a promising approach for cancer immunotherapy (Extended Data Fig. 6z).
Discussion
This study offers support for several undocumented perspectives in the field of tumour immunology and immunotherapy. First, STAT3 counteracts STAT5 activation, which restrains DC maturation in the TME. Second, STAT3 degradation is a potential immunotherapeutic modality for treating patients with advanced and ICB-resistant cancer. Third, STAT3 degraders work in an DC-dependent manner, supporting STAT5-mediated DC maturation and function.
DCs prime and activate T cells against cancer cells3,41,42. We found that ICB can reprogram the STAT5 and STAT3 transcriptional pathways in DCs, thereby normalizing DC function in the TME. It has long been thought that DCs are potential immunotherapeutic targets. However, it is challenging to therapeutically manipulate DCs owing to their numeric scarcity, subset diversity and lack of specific molecular targets. Our results suggest that STAT3 is a druggable molecular target in DC1s using our potent and highly selective PROTAC STAT3 degraders.
Despite the identification of STAT3 and STAT5 many years ago20,43,44,45, the interplay between STAT3 and STAT5 signalling pathways in specific immune cells remain poorly understood. STAT3 is frequently activated in the TME owing to abundant stimulating factors, including IL-6, IL-10 and VEGF20,21, whereas STAT5 is often activated by T cell-derived cytokines, such as GM-CSF and IL-2. Given that T cells are limited or dysfunctional in the TME, the balance between STAT5 and STAT3 in immune cells, including DCs, may be biased towards STAT3 overactivation. In support of this possibility, we demonstrate that there is an imbalance between these two transcriptional pathways in DC1s in the TME, tilting towards an activation in STAT3 over STAT5. At the molecular level, DC1s express high levels of GM-CSF receptor and the STAT3 transcriptional signalling pathway restrains the GMR–JAK2–STAT5 signalling axis in DCs in the TME. The interaction between GMR and JAK2 appears to regulate both the STAT3 and STAT5 transcriptional pathways in DC1s. In support of this possibility, we observed that GM-CSF stimulation or genetically knocking down GM-CSF receptor alters the phosphorylation of JAK2, STAT3 and STAT5, but not JAK1, JAK3, TYK2 and STAT1 in cDC1s. Of note, targeting STAT3 has no effect on the expression levels of GM-CSF and GMRβ but enhances the interaction of GMRβ and JAK2 with STAT5, leading to the phosphorylation of STAT5 and DC maturation. Therefore, STAT3 restrains JAK2-mediated STAT5 activation in DCs, thereby impairing DC1-mediated T cell immunity.
To rescue the STAT3-impaired DC phenotype, we developed two PROTAC molecules (SD-36 and SD-2301) that target STAT3 protein for degradation. There are several key functional characteristics of our STAT3 degraders. First, they are highly selective and efficacious in degrading STAT3. Second, low doses of SD-36 and SD-2301 effectively target DCs. Furthermore, low doses of SD-36 and SD-2301 mediate an immune-dependent anti-tumour effect, whereas high doses of SD-36 can directly control tumour progression in the absence of a functional immune system36. Intriguingly, SD-36 or SD-2301, as a monotherapy, exhibit remarkable efficacy in treating large, advanced tumours and ICB-resistant tumours. Given that the anti-tumour effect of SD-2301 is four to five times more potent than that of SD-36 in degrading STAT3 and it has excellent pharmacokinetics, we are evaluating SD-2301 as a potential drug candidate for clinical development. Notably, although STAT3 degraders efficaciously target DCs, the involvement of STAT3 in other cell types may also influence immune responses in the TME. Together, our study provides strong support for potential human clinical trials of STAT3 degraders in treating patients with large and metastatic tumours and/or ICB-resistant tumours.
Methods
Antibodies, plasmids and reagents
Western blot analyses were performed using primary antibodies at a dilution of 1:1,000 unless otherwise specified. Anti-STAT1 (D1K9Y, 14994), anti-STAT2 (D9J7L, 72604), anti-STAT3 (Rabbit, 79D7, 4904; Mouse, 124H6, 9139), anti-STAT4 (C46B10, 2653), anti-STAT5 (D2O6Y, 94205), anti-STAT6 (D3H4, 5397), anti-JAK1 (6G4, 3344), anti-JAK2 (D2E12, 3230), anti-JAK3 (D7B12, 8863), anti-TYK2 (E9H4T, 35615), anti-p65 (D14E12, 8242), anti-CD80 (E6J6N, 54521), anti-CD86 (E5W6H, 19589), anti-IL-6Rα (E7H4J, 39837), anti-GP130 (3732), anti-pSTAT1 (58D6, 9167), anti-pSTAT3 (D3A7, 9145), anti-pSTAT5 (D47E7, 4322), anti-pSTAT6 (D8S9Y, 56554), anti-pJAK1 (D7N4Z, 74129), anti-pJAK2 (C80C3, 3776), anti-pJAK3 (D44E3, 5031), anti-pp65 (93H1, 3033) and anti-β-actin (4967) were from Cell Signaling Technology. Anti-JAK2 (C-10, sc-390539), anti-GMRβ (F-12, sc-393281) and anti-β-actin (C4, sc-47778) were from Santa Cruz Biotechnology. Anti-pTYK2 (PA5-37762) was from Invitrogen. HRP horse anti-mouse IgG antibody (PI-2000, 1:5,000) and HRP goat anti-rabbit IgG (PI-1000, 1:5,000) were from Vector Laboratories.
For the in vivo experiments, anti-mouse CD8 (clone YTS 169.4, BE0117), anti-mouse PD-L1 (clone 10F.9G2, BE0101) and anti-IgG2b (clone LTF-2, BE0090) were from BioXcell. SD-36 and SD-2301 were designed and synthesized in our laboratory. FLLL32 (JAK2 inhibitor X, 5301530001) and STAT5 inhibitor (573108) were purchased from Sigma-Aldrich. Plasmids expressing shRNAs targeting Stat3 (TRCN0000071453, TRCN0000301946), Stat1 (TRCN0000054924), Jak2 (TRCN0000023649, TRCN0000023651, TRCN0000023652) and Gmrβ (TRCN0000067025, TRCN000067026) were from Sigma-Aldrich.
Cell culture
JAWSII, 293T, B16F10, CT26, EMT6, LLC and 4T1 cells were purchased from the American Type Culture Collection (ATCC). MC38 mouse colon cancer cell line was obtained from the University of Texas Southwestern Medical Center (Y.-X. Fu). Ovarian cancer cell line luciferase-ID8 cells were previously reported24. All cell lines were tested for mycoplasma contamination by MycoAlert Mycoplasma Detection kit and confirmed negative for Mycoplasma.
The 293T cells were transfected with packaging plasmids and non-targeting lentivirus vector, or a lentivirus vector encoding shRNAs targeting Stat3, Stat1, Jak2 or Gmrβ. The virus-containing supernatant was collected 48 h after transfection. JAWSII cells were transfected with the lentivirus. Then the cell lysates were analysed by immunoblotting.
Bone marrow-derived dendritic cells
Bone marrow was collected from the femurs and tibias of mice. Red blood cells were lysed using ACK lysis buffer. Bone marrow-derived DCs (BMDCs) were generated with GM-CSF (20 ng ml−1) and FLT3 ligand (100 ng ml−1) in IMDM (Iscove’s Modified Dulbecco’s Medium; Gibco, 12440-053) supplemented with 10% FBS, 1% penicillin-streptomycin and 55 μΜ β-mercaptoethanol for 7–10 days. BMDCs were either sorted as cDC1s (CD11c+XCR1+) using a FACSAria flow cytometry sorter (BD Biosciences) or purified using biotin anti-mouse XCR1 antibody (BioLegend,148212) and anti-biotin microbeads (Miltenyi Biotec, 130-090-485) following the manufacturer’s protocols for subsequent experiments. Primary cell cultures were tested to be mycoplasma-free by MycoAlert Mycoplasma Detection kit.
ELISA
Culture media were collected from Stat3+/+ and Stat3−/− cDC1s and centrifuged at 8,000g for 5 min. GM-CSF (MGM00) and IL-6 (DY406) were detected in the culture supernatants using the ELISA kit.
CD8 depletion
CD8+ T cells were depleted with anti-CD8 (YTS 169.4, BioXCell) antibodies. Anti-CD8 (100 μg per mouse) antibodies were injected intraperitoneally at the beginning of tumour inoculation and continuously administered every three days.
Drug cellular uptake
DCs and LLC tumour cells were treated with 1 μM SD-36 for 12 h. Then, DCs and LLC (106 each) were centrifuged at 1,500 rpm for 5 min at 4 °C and washed 3 times with ice-cold PBS. The pellets were resuspended in 100 μl of a 50% (v/v) water/methanol solution and subjected to 3 freeze–thaw cycles. After the final thaw cycle, these samples were centrifuged at 14,000 rpm, 4 °C for 20 min, and the supernatant were collected. All samples were submitted to the pharmacokinetics core (University of Michigan) for analysis.
Immunoprecipitation and immunoblot analysis
Cells were lysed in immunoprecipitation lysis buffer (50 mM Tris-HCl pH 7.4, 120 mM NaCl, 1 mM EDTA and 0.5% NP-40) and supplemented with Halt Protease and Phosphatase Inhibitor Cocktail (Thermo Fisher Scientific). Cells were repeatedly passed through a 21-gauge needle with sonication. Then 1,000 µg total cell lysates were incubated with the appropriate antibody (2 µg), with rotation overnight at 4 °C, followed by a 3-h incubation with Protein A/G Sepharose beads (Santa Cruz Biotechnology). Immune complexes were washed three times with wash buffer (20 mM Tris-HCl pH 7.4, 100 mM NaCl, 1 mM EDTA and 0.2% NP-40); then the immunoprecipitated proteins were denatured by the addition of sample buffer (Bio-Rad) and boiled for 10 min, resolved by SDS–PAGE, and immunoblotted with indicated antibodies.
Cells were lysed in RIPA buffer (Thermo Fisher Scientific) supplemented with Halt Protease and Phosphatase Inhibitor Cocktail (Thermo Fisher Scientific). The protein concentrations of cell lysates were determined by BCA protein assay kit (Thermo Fisher Scientific). Equivalent amounts of total cellular protein were separated by SDS–PAGE, transferred to polyvinylidene fluoride membranes, and immunoblotted with the indicated antibodies.
For phospho-antibody array cDC1s were treated with LPS (20 ng ml−1) for 3 h and lysed for protein extraction. Lysates were analysed using the Proteome Profiler Phospho-Kinase Array Kit (R&D Systems, number ARY003C) according to the manufacturer’s instructions. Western blotting data were processed and analysed by image lab 6.1 (Bio-Rad) and ImageJ 1.51n (NIH).
Immunohistochemistry staining
For histological analysis, tissue samples were collected, fixed in 10% formalin (Sigma) and processed for formalin-fixed paraffin-embedded tissue analysis. Four-micrometre paraffin sections underwent heat-induced epitope retrieval. Staining was performed with anti-mouse CD31 antibody (Cell Signaling Technology, D8V9E, 77699, 1:100) using an automated immunostainer (Biocare Intellipath, Biocare Medical). Sections were counterstained with haematoxylin (Biocare Medical), and slides were scanned with the Aperio AT2 Scanner (Leica Biosystems Imaging). Quantification was performed with QuPath v0.5.1.
Quantitative PCR
Total RNA was isolated from cells by column purification (RNeasy Micro Kit, Qiagen) with DNase treatment. cDNA was synthesized using the RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher Scientific) with random hexamer primers. Quantitative PCR was performed on cDNA using Fast SYBR Green Master Mix (Thermo Fisher Scientific) on a QuantStudio 3 Real-Time PCR System (Thermo Fisher Scientific). Gene expression was quantified using specific primers (Extended Data Table 3). Fold changes in messenger RNA expression were calculated with the ΔΔCt method using Actb as an endogenous control. The results are expressed as fold changes normalized to the controls.
Synthesis of SD-2301 and its chemical data
The synthesis of SD-2301 is outlined in scheme 1 (Supplementary Fig. 1). It began with a known compound, 1, which was converted into compound 2 through a two-step process. First, compound 1 was reacted with hept-6-ynoic acid to form an amide, which was then saponified using LiOH in aqueous media to yield acid intermediate 2 in 65% overall yield. Azide 6 was synthesized in 80% yield from commercially available amine 5 using a previously reported method. A click reaction between alkyne acid 2 and azide 6, with sodium ascorbate and CuSO4, resulted in acid intermediate 3 in 78% yield. Amine 7 was prepared from Boc-l-glutamine through a two-step procedure. Initially, Boc-l-glutamine was coupled with substituted benzyl amine in the presence of HATU (hexafluorophosphate azabenzotriazole tetramethyl uronium) and DIPEA (N,N-diisopropylethylamine) in DMF (N,N-dimethylformamide) to form a Boc-protected amide, which was then Boc-deprotected using 4 (N) HCl in DCM to give amine 7 in 70% yield over two steps. The coupling of acid 3 with amine 7 in the presence of HATU and DIPEA in DMF produced a Boc-protected intermediate. This intermediate was then Boc-deprotected using 4 (N) HCl in dioxane to yield amine intermediate 4 in 75% yield over two steps. Finally, compound 4 was converted into SD-2301 in 86% yield through an amide formation reaction with literature known intermediate 8, using HOBt and DIPEA in DMF.
The purity of SD-2301 was confirmed by ultra-performance liquid chromatography (UPLC) to be >99%. UPLC−MS (ESI) m/z: calculated, 683.7 for C64H80N13O15PS2 [M + 2H]+/2; found, 684.09 (Supplementary Fig. 2). Proton nuclear magnetic resonance (1H-NMR) and carbon nuclear magnetic resonance (13C-NMR) spectroscopies were performed on Bruker Advance 400 NMR spectrometers, and chemical shifts are reported in parts per million (ppm) relative to an internal standard (Supplementary Figs. 3 and 4).
In vivo mouse experiments
Animal studies were approved by the Institutional Animal Care and Use Committee at the University of Michigan. All mice were maintained under specific pathogen-free housing (about 22 °C with approximately 40% humidity) on a 12-h dark:12-h light cycle. The following mice (at 6–8 weeks of age) (The Jackson laboratory) were used for this study: C57BL/6J, BALB/cJ, NOD-scid IL2Rγ null (NSG), Rag1tm1Mom (Rag1–/–), C57BL/6-Tg(TcraTcrb)1100Mjb/J (OT-I), B6.129S(C)-Batf3tm1Kmm/J (Batf3−/−), B6.129S(Cg)-Stat1tm1Dlv/J (Stat1−/−), B6(129S4)-Xcr1tm1.1(cre) Kmm/J (Xcr1cre), B6.129S1-Stat3tm1Xyfu/J (Stat3fl/fl) mice and CD-1 mice were obtained from Charles River Laboratories. Stat5b–/– C57BL/6J mice were from the National Institutes of Health (Warren Leonard). Stat3fl/fl mice were crossed with Xcr1cre mice to obtain specific STAT3 deficiency in DC1s (Stat3−/− mice). In-house littermates were used in the control arm when specific mouse strains were generated in-house.
MC38 (2 × 106), CT26 (105), EMT6 (105), B16F10 (2 × 105) and LLC (2 × 105) cells were injected subcutaneously into syngeneic mice. Luciferase-expressing ID8 cells (2 × 106) were injected into the peritoneal cavity of female mice. Luciferase-expressing 4T1 cells (105) were injected into female mouse mammary fat pad. From day 5–7, SD-36 (low doses: 10 or 20 mg kg−1; high dose: 100 mg kg−1) or SD-2301 (5 mg kg−1) was administered intravenously every 3 days. In 4T1 tumour metastasis model, treatment with SD-36 (20 mg kg−1) started on day 14, followed by every 3 days. In some cases, peritoneal tumour-bearing mice were similarly treated with SD-36 (20 mg kg−1) or SD-2301 (5 mg kg−1) or were injected intraperitoneally with FLLL32 (30 mg kg−1) for 24 h. For the ICB experiments, PD-L1 monoclonal antibody or isotype antibody (BioXcell, 200 μg) were administrated intraperitoneally into tumour-bearing mice, starting on day 3, then repeated this treatment every 3 days. For DC1 transfusion experiments, bone marrow-derived cDC1s (2 × 106) were sorted and injected intravenously into tumour-bearing Batf3−/− mice on day −2 and day 8. SD-36 (20 mg kg−1) was administered intravenously every three days, starting on day 5. Tumour inoculation time was considered day 0. Tumour cells were inoculated into age-, sex- and source-matched mice. Tumour size was measured using calipers with a Vernier scale and calculated as previously46.
To evaluate tolerability and toxicity, tumour-bearing C57BL/6J mice received intravenous injections of SD-2301 (5 mg kg−1) every 3 days for 18 days, with body weights were monitored throughout.
For pharmacokinetic evaluation, female CD-1 mice were administered a single intravenous dose of SD-2301 at 5 mg kg−1 with 25% of PCP as the dosing vehicle. Then, 250–300 μl blood samples were collected from 5 min to 24 h. Following centrifugation at 15,000 rpm for 10 min, at least 100 μl of plasma was collected. All samples were analysed by the Pharmacokinetic and Mass Spectrometry core at the University of Michigan. Compound concentrations in plasma were determined using a validated LC-MS/MS method with an internal control. Chromatographic separation was achieved with a Waters XBridge-C18 column (5 cm × 2.1 mm, 3.5 μm) on a Shimadzu HPLC system. Detection was performed on an AB Sciex QTrap 5500 mass spectrometer in positive-ion MRM mode. The mobile phases were 0.1% formic acid in water (A) and in acetonitrile (B). The gradient for B was: 10% (0–0.3 min), increased to 95% at 0.7 min, held for 2.3 min, and returned to 10% for a 2-min re-equilibration. Flow rate was 0.4 ml min−1. Pharmacokinetic parameters were calculated using noncompartmental methods in WinNonlin v.3.2 (Pharsight).
Human studies
All clinical records used in this study were acquired and used with the approval of Institutional Review Boards. Patients (cohort 1) who underwent ICB therapy were recruited from the University of Michigan Hospital, Ann Arbor, MI, USA. Inclusion in the analysis applied to those who were enroled in the Michigan Center for Translational Pathology’s (MCTP) continuous comprehensive clinical sequencing programme39,47,48,49, MI-Oncoseq, and who possessed sequenced libraries of pre-treatment tumours. From the initiation of therapy, overall survival times were measured. Treatment responses were determined according to RECIST1.1 criteria50 and imRECIST51. Integrative clinical sequencing was conducted using standard protocols approved for use in MCTP’s Clinical Laboratory Improvement Amendments-compliant sequencing laboratory, as previously described47,48,52. After purification with the AllPrep DNA/RNA/miRNA kit (Qiagen), total RNA was sequenced using the exome-capture transcriptome platform on an Illumina HiSeq 2000 or HiSeq 2500 in paired-end mode. Quality control, alignment, and expression quantification was executed through the standard clinical RNA-seq pipeline, CRISP53. Read count tables were then normalized into fragments per kilobase million (FPKM) then transcripts per million (TPM) using the edgeR 4.2.2 package54. Gene scores in RNA-seq data were generated using the rank-based inverse normal transformation (INT) as demonstrated previously55. The CTL score and DC1 maturation score were generated using the following respective gene sets: (CD8A, CXCL10, CXCL9, GZMA, GZMB, PRF1, IFNG and TBX21) and (CD40, CD80, CD86, HLA-DQA1, XCR1, CLEC9A, IL12A, IL12B, HLA-DRA and IDO1). STAT5/STAT3 expression was calculated as the ratio of combined TPM of STAT5A and STAT5B (STAT5AB) to STAT3 TPM for all patients (Extended Data Table 1). Stratification was performed at the median STAT5AB/STAT3 TPM ratio and at the median DC1 maturation score. This was performed for cohort 2 (Extended Data Table 2) in each dataset, before combining group-wise. STAT5 pathway INT score was calculated using the GM-CSF reactome gene set (R-HSA-512988). STAT5 pathway/STAT3 was calculated by dividing the INT score by the STAT3 TPM in each respective sample, this value was stratified by the median.
The single-cell RNA-seq analysis of TNBC (cohort 3) (dataset GSE169246) was conducted in R using Seurat (v.4.3.0), ssGSEA, and standard data wrangling and plotting packages. All analyses were carried out after subsetting the data for immunotherapy treatment. DCs were extracted based on the original authors’ annotations. Only cells expressing 200 or more genes in at least 3 cells were included in the analysis, and cells with mitochondrial reads greater than 10% were excluded. The STAT5 signalling score for each cell was calculated by applying the GM-CSF reactome gene set (R-HSA-512988) as input to the enrichIt function from the escape (v.2.0.0) package, which implements ssGSEA. Similarly, the STAT3 signalling score was determined using the same approach. The genes used for the IL-10-driven STAT3 signalling score were obtained from previously reported gene orthologues in murine DCs56. This gene set comprised: RAP1GAP, CAMKK1, CASR, KIF1A, HPCAL4, DRAM1, RAMP2, MT2A, IGFBP6, GATA3, CXCR5, GDA, FAM65C (also known as RIPOR3), PLET1, SOCS3, MUC1 and GBP5. The DC maturation score was computed using the same method with genes canonically associated with DC maturation, including CD40, CD80, CD83, CD86, LAMP3, CCL19, IL12A, IL12B, CCL5, CCL22, CXCL9, CXCL10, NFKB1, NFKB2, NFKBIA, NFKBIB, FSCN1 and CCR7. The binary division of high and low STAT3 signalling scores was determined by visually identifying two distinct clusters. The monocyte-DCs versus conventional DC distinction was determined through a combination of iterative clustering and expression of CD14. The Wilcoxon signed-rank test was used to compare the ssGSEA scores using the stat_compare_means function from the ggpubr (v.0.6.0) package.
For the in vitro human ICB study, lin−CD45+CD11c+MHCII+ DCs were enriched and sorted from fresh cancer tissues of patients with high grade serous ovarian carcinoma. DCs (106 per ml) were pretreated with SD-36 (1 μM), washed, and co-cultured with normal peripheral blood T cells (2 × 106 per ml) in the presence of anti-human PD-L1 (10 μg ml−1), anti-CD3 (2 μg ml−1), and anti-CD28 (1 μg ml−1) for 3 days. T cell cytokine profile was analysed by FACS. For detecting the effect of SD-2301 on STAT members in human immune cells in vitro, normal human peripheral blood mononuclear cells (PBMCs) were treated with SD-2301 for 14 h, followed by immunoblotting to analyse STAT protein levels. Normal human immune cells were collected from commercial buffy coats (Carter BloodCare).
Bulk RNA-seq analysis in mouse DCs
Data processing
cDC1s were purified from BMDCs as described above. The RNA was isolated by column purification (Qiagen, 217048) with DNase treatment. The RNA-seq was conducted by BGI Genomics. Bulk RNA-seq analysis was performed using the edgeR (v.4.2.2) and limma (v.3.60.6) package workflows. Preliminary quality control measures were performed including the filtering of lowly expressed genes, and the calculation and assessment of library sizes across samples. The raw counts were then transformed to log counts per million (CPM) values.
Differential expression analysis
A design matrix was created containing cell population information (Stat3−/− versus Stat3+/+) and then contrasts for pairwise comparisons between Stat3−/− and Stat3+/+ were determined in limma using the makeContrasts function. As an additional normalization measure, removal of heteroscedasticity from the normalized counts data was achieved using the voom function. To identify differentially expressed genes, the mean-variance relationship of the normalized counts was assessed and then the normalized data were modelled as a linear combination of factors and covariates. The cutoff for the P value was P = 0.05. Differential expression analysis was performed on non-treated Stat3−/− versus Stat3+/+ cDC1s and LPS-treated Stat3−/− versus Stat3+/+ cDC1s, respectively.
Gene signature analysis
To assess the DC maturation signature, the log fold change and P values of DC maturation genes reported in the literature including (Cd40, Cd80, Cd83, Cd86, Ido1, Irf7, Tnfsf8, H2-Aa, H2-ab1, Tlr4, Il12b, Il12rb1, Il12rb2 and Cxcl9) was observed from the differential expression analysis results. To assess the significantly upregulated DC maturation pathways, gene set enrichment analysis (GSEA) was performed on the Stat3−/− versus Stat3+/+ differentially expressed genes in R using the gseGO function. To assess the significantly upregulated DC activation pathways in the DCs treated with LPS, GSEA was performed on the LPS-treated Stat3−/− versus Stat3+/+ differentially expressed genes in R using the gseGO function.
Data visualization
The z-score of the normalized counts were computed and plotted as a heat map using GraphPad Prism 10.2.2. DC maturation genes were selected from the differential expression analysis results and the log fold change and P values were plotted using GraphPad Prism 10.2.2. The GSEA plots were generated in R using the gseaplot function. The GSEA dot plot was generated by plotting key pathways from the GSEA results using the dotplot function from clusterProfiler 4.9.2 package. The volcano plot was generated by first performing differential expression analysis of LPS-treated Stat3−/− versus Stat3+/+ cDC1s and plotting log fold changes and P values of genes from the STAT5 pathway curated in the Hallmark pathway Molecular Signature Database (MSigDB).
Chromatin immunoprecipitation
Chromatin immunoprecipitation was performed using the SimpleChIP Enzymatic Chromatin IP Kit (Cell Signaling Technology, 9003s). Sheared chromatin was then immunoprecipitated with STAT5 (Cell Signaling Technology, 94205s) and IgG (Cell Signaling Technology, 2729) antibodies. SYBR green master mix (Applied Biosystems) was used to measure amplification of DNA using QuantStudio 3 Fast Real-Time PCR system (Applied Biosystems). The promoters of gene were quantified using the specific primers (Extended Data Table 3). After normalization to the Input DNA, the amount of output DNA of each target protein was calculated by subtracting that of the IgG control.
Flow cytometry analysis
Single-cell suspensions were prepared from fresh mouse tumour tissues or spleen, and lymphocytes were enriched by density gradient centrifugation. To assess intracellular cytokine production, cells were cultured for 4 h in the presence of phorbol myristate acetate (5 ng ml−1; Sigma-Aldrich), ionomycin (500 ng ml−1; Sigma-Aldrich) and brefeldin A (1: 1000, BD Biosciences). Cells were fixed and permeabilized with the Transcription factor staining buffer set (Invitrogen, 00-5523-00). Cellular phenotypes were assessed multi-parameter flow cytometry panels. Data were acquired on a BD LSRFortessa. Antibodies against the following mouse antigens were used: CD45 (30-F11, HI30), CD90 (53-2.1, 30-H12), CD3 (17A2), CD45R/B220 (RA3-6B2), CD4 (RM4-5, GK1.5), CD8 (53−6.7), IFNγ (XMG1.2), granzyme B (GB11), IL-2 (JES6-5H4), Ki-67 (B56), I-A/I-E (M5/114.15.2), CD80 (16-10A1), CD86 (GL1), H-2Kb (AF6-88.5.5.3), H-2 (M1/42), XCR1 (ZET), TNF (MP6-XT22), CD11c (HL3), phospho-Tyr694-STAT5 (SRBCZX), phospho-Tyr705-STAT3 (13A3-1). Additionally, antibodies against the following human antigens were used: CD8 (RPA-T8), IFNγ (B27) and TNF (MAb11). The strategies for DC gating and in vitro cultures and tissues are shown in Extended Data Figs. 2c and 3e, respectively. The strategy for T cell gating is presented in Extended Data Fig. 3k.
Statistics and reproducibility
Sample sizes were determined based on previously published studies to ensure appropriate statistical power. In vitro experiments were performed in at least two independent replicates. For in vivo studies, each group consisted of a minimum of five mice, which was considered sufficient to detect meaningful biological differences with good reproducibility, and mice were randomized into treatment groups at the start of the experiment. Experiments were not performed in a blinded manner, as knowledge of the treatment groups were necessary to carry out the study. t-tests were used to assess differences between two independent experimental groups, and one-way analysis of variance (ANOVA) was used to compare three or more groups. Two-way ANOVA was applied to compare tumour growth curves. Data are presented as mean ± s.e.m., and statistical significance was defined as P < 0.05 for all tests. All statistical analyses related to these experiments were performed using GraphPad Prism software v.10.2.2.
Kaplan–Meier analysis was conducted to estimate overall survival, with differences between groups assessed using log-rank tests. Cox proportional hazards models were used for multivariate survival analysis. Pearson’s correlation coefficient was applied to evaluate linear correlations between variables, and the Wilcoxon rank-sum test was performed to assess group differences. All P values are two-sided and not adjusted for multiple comparisons. The statistical analyses were conducted using R packages.
Ethics oversight
All human studies were conducted under the oversight and approval of the Institutional Review Board at the University of Michigan Medical School.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Clinical sequencing data are publicly available with raw data available upon request from dbGaP phs000673.v5.p1 (refs. 39,47). RNA-seq data that were newly generated in this study for in vitro analysis have been deposited at the Gene Expression Omnibus (GEO) at NCBI under accession GSE289764. Single-cell RNA-seq data that support the findings of this study were downloaded from GEO accession GSE169246. The two original melanoma RNA-seq datasets were deposited in the European Nucleotide Archive (ENA) under accession PRJEB23709 and in dbGaP under accession phs000452.v2.p1. The processed data for these two melanoma datasets can be found at https://ngdc.cncb.ac.cn/icb/resources. All materials are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Sharma, P. & Allison, J. P. The future of immune checkpoint therapy. Science 348, 56–61 (2015).
Zou, W., Wolchok, J. D. & Chen, L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: mechanisms, response biomarkers, and combinations. Sci. Transl. Med. 8, 328rv324 (2016).
Wculek, S. K. et al. Dendritic cells in cancer immunology and immunotherapy. Nat. Rev. Immunol. 20, 7–24 (2020).
Spranger, S., Dai, D., Horton, B. & Gajewski, T. F. Tumor-residing Batf3 dendritic cells are required for effector T cell trafficking and adoptive T cell therapy. Cancer Cell 31, 711–723 (2017).
Johnson, D. E., O’Keefe, R. A. & Grandis, J. R. Targeting the IL-6/JAK/STAT3 signalling axis in cancer. Nat. Rev. Clin. Oncol. 15, 234–248 (2018).
June, C. H. & Sadelain, M. Chimeric antigen receptor therapy. New Engl. J. Med. 379, 64–73 (2018).
Inaba, K., Young, J. W. & Steinman, R. M. Direct activation of CD8+ cytotoxic T lymphocytes by dendritic cells. J. Exp. Med. 166, 182–194 (1987).
Everts, B. et al. Migratory CD103+ dendritic cells suppress helminth-driven type 2 immunity through constitutive expression of IL-12. J. Exp. Med. 213, 35–51 (2016).
Maier, B. et al. A conserved dendritic-cell regulatory program limits antitumour immunity. Nature 580, 257–262 (2020).
Anderson, D. A. 3rd, Dutertre, C. A., Ginhoux, F. & Murphy, K. M. Genetic models of human and mouse dendritic cell development and function. Nat. Rev. Immunol. 21, 101–115 (2021).
Duong, E. et al. Type I interferon activates MHC class I-dressed CD11b+ conventional dendritic cells to promote protective anti-tumor CD8+ T cell immunity. Immunity 55, 308–323 (2022).
Ganesan, A. P. et al. Tissue-resident memory features are linked to the magnitude of cytotoxic T cell responses in human lung cancer. Nat. Immunol. 18, 940–94 (2017).
Bottcher, J. P. et al. NK cells stimulate recruitment of cDC1 into the tumor microenvironment promoting cancer immune control. Cell 172, 1022–1037 (2018).
Gerhard, G. M., Bill, R., Messemaker, M., Klein, A. M. & Pittet, M. J. Tumor-infiltrating dendritic cell states are conserved across solid human cancers. J. Exp. Med. 218, e20200264 (2021).
Salmon, H. et al. Expansion and activation of CD103+ dendritic cell progenitors at the tumor site enhances tumor responses to therapeutic PD-L1 and BRAF inhibition. Immunity 44, 924–938 (2016).
Qiao, J. et al. Targeting tumors with IL-10 prevents dendritic cell-mediated CD8+ T cell apoptosis. Cancer Cell 35, 901–915 (2019).
Guo, C. S. et al. SLC38A2 and glutamine signalling in cDC1s dictate anti-tumour immunity. Nature 620, 200–20 (2023).
Li, Y. J., Zhang, C., Martincuks, A., Herrmann, A. & Yu, H. STAT proteins in cancer: orchestration of metabolism. Nat. Rev. Cancer 23, 115–134 (2023).
Fu, X. Y. STAT3 in immune responses and inflammatory bowel diseases. Cell Res. 16, 214–219 (2006).
Zhong, Z., Wen, Z. & Darnell, J. E. Jr. Stat3: a STAT family member activated by tyrosine phosphorylation in response to epidermal growth factor and interleukin-6. Science 264, 95–98 (1994).
Yu, H., Kortylewski, M. & Pardoll, D. Crosstalk between cancer and immune cells: role of STAT3 in the tumour microenvironment. Nat. Rev. Immunol. 7, 41–51 (2007).
Liao, W., Lin, J. X., Wang, L., Li, P. & Leonard, W. J. Modulation of cytokine receptors by IL-2 broadly regulates differentiation into helper T cell lineages. Nat. Immunol. 12, 551–U247 (2011).
Eddy, W. E. et al. Stat5 is required for CD103+ dendritic cell and alveolar macrophage development and protection from lung injury. J. Immunol. 198, 4813–4822 (2017).
Zhou, J. et al. The ubiquitin ligase MDM2 sustains STAT5 stability to control T cell-mediated antitumor immunity. Nat. Immunol. 22, 460–470 (2021).
Choi, J. E. et al. PIKfyve, expressed by CD11c-positive cells, controls tumor immunity. Nat. Commun. 15, 5487 (2024).
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
Yuan, X. et al. Vitamin E enhances cancer immunotherapy by reinvigorating dendritic cells via targeting checkpoint SHP1. Cancer Discov. 12, 1742–1759 (2022).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Gide, T. N. et al. Distinct immune cell populations define response to anti-PD-1 monotherapy and anti-PD-1/anti-CTLA-4 combined therapy. Cancer Cell 35, 238–255 (2019).
Zhang, Y. et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell 39, 1578–1593 (2021).
Theisen, D. J. et al. WDFY4 is required for cross-presentation in response to viral and tumor antigens. Science 362, 694–69 (2018).
Choi, H. W. et al. Perivascular dendritic cells elicit anaphylaxis by relaying allergens to mast cells via microvesicles. Science 362, 656–65 (2018).
Al-Shami, A., Mahanna, W. & Naccache, P. H. Granulocyte-macrophage colony-stimulating factor-activated signaling pathways in human neutrophils—selective activation of Jak2, Stat3, and Stat5B. J. Biol. Chem. 273, 1058–1063 (1998).
Riggan, L. et al. The transcription factor Fli1 restricts the formation of memory precursor NK cells during viral infection. Nat. Immunol. 23, 556–55 (2022).
Huynh, J., Chand, A., Gough, D. & Ernst, M. Therapeutically exploiting STAT3 activity in cancer - using tissue repair as a road map. Nat. Rev. Cancer 19, 82–96 (2019).
Bai, L. et al. A potent and selective small-molecule degrader of STAT3 achieves complete tumor regression in vivo. Cancer Cell 36, 498–511 (2019).
Lin, H. et al. Host expression of PD-L1 determines efficacy of PD-L1 pathway blockade-mediated tumor regression. J. Clin. Invest. 128, 805–815 (2018).
Nishinol, M., Giobbie-Hurder, A., Ramaiyal, N. H. & Hodit, F. S. Response assessment in metastatic melanoma treated with ipilimumab and bevacizumab: CT tumor size and density as markers for response and outcome. J. Immunother. Cancer 2, 40 (2014).
Yu, J. et al. Liver metastasis restrains immunotherapy efficacy via macrophage-mediated T cell elimination. Nat. Med. 27, 152–164 (2021).
Hildner, K. et al. Batf3 deficiency reveals a critical role for CD8α+ dendritic cells in cytotoxic T cell immunity. Science 322, 1097–1100 (2008).
Durai, V. & Murphy, K. M. Functions of murine dendritic cells. Immunity 45, 719–736 (2016).
Murphy, T. L. & Murphy, K. M. Dendritic cells in cancer immunology. Cell. Mol. Immunol. 19, 3–13 (2022).
Fu, X. Y. & Zhang, J. J. Transcription factor p91 interacts with the epidermal growth factor receptor and mediates activation of the c-fos gene promoter. Cell 74, 1135–1145 (1993).
Sadowski, H. B., Shuai, K., Darnell, J. E. Jr. & Gilman, M. Z. A common nuclear signal transduction pathway activated by growth factor and cytokine receptors. Science 261, 1739–1744 (1993).
Wakao, H., Gouilleux, F. & Groner, B. Mammary gland factor (MGF) is a novel member of the cytokine regulated transcription factor gene family and confers the prolactin response. EMBO J. 13, 2182–2191 (1994).
Wang, D. et al. Controllable synthesis of dual-MOFs nanostructures for pH-responsive artemisinin delivery, magnetic resonance and optical dual-model imaging-guided chemo/photothermal combinational cancer therapy. Biomaterials 100, 27–40 (2016).
Robinson, D. R. et al. Integrative clinical genomics of metastatic cancer. Nature 548, 297–303 (2017).
Wu, Y. M. et al. Inactivation of CDK12 delineates a distinct immunogenic class of advanced prostate cancer. Cell 173, 1770–1782 (2018).
Wang, W. et al. CD8+ T cells regulate tumour ferroptosis during cancer immunotherapy. Nature 569, 270–274 (2019).
Watanabe, H. et al. New response evaluation criteria in solid tumours-revised RECIST guideline (version 1.1). Gan To Kagaku Ryoho 36, 2495–2501 (2009).
Hodi, F. S. et al. Immune-modified Response Evaluation Criteria In Solid Tumors (imRECIST): refining guidelines to assess the clinical benefit of cancer immunotherapy. J. Clin. Oncol. 36, 850–858 (2018).
Parolia, A. et al. Distinct structural classes of activating FOXA1 alterations in advanced prostate cancer. Nature 571, 413–418 (2019).
Cieslik, M. et al. The use of exome capture RNA-seq for highly degraded RNA with application to clinical cancer sequencing. Genome Res. 25, 1372–1381 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Yang, J. et al. FTO genotype is associated with phenotypic variability of body mass index. Nature 490, 267–272 (2012).
Chrisikos, T. T. et al. STAT3 inhibits autocrine IFN signaling in type i conventional dendritic cells. J. Immunol. 209, 1286–1299 (2022).
Acknowledgements
The authors thank J. Deng for providing Stat1–/– C57BL/6J mice (originally from The Jackson Laboratory); M. Wang, B. Wen and D. Sun for the pharmacokinetics study; and our laboratory members for their intellectual input. This work was supported in part by research grants from the Breast Cancer Research Foundation, NIH/NCI R01 grants (CA217648, CA123088, CA099985, CA193136, CA152470 and CA244509), and the NIH/NCI through the University of Michigan Rogel Cancer Center Grant (CA46592). J.-X.L. and W.J.L. are supported by the Division of Intramural Research, the National Heart, Lung and Blood Institute, the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
J.Z. conceived the study, designed experiments, conducted experiments and wrote and revised the manuscript. K.T. performed human bioinformatic analyses and wrote and revised the manuscript. H.Z. developed SD-36. L.B. developed SD-36 and SD-2301 and conducted in vitro human PBMC studies with SD-2301. R.K.A. developed SD-2301. D.M. assisted with in vivo experiments and supervised the use of SD-36 and SD-2301. H.M. provided assistance with in vivo experiments. Y.W. conducted the pharmacokinetics study. M.P. performed mouse bioinformatic analyses. J.E.C. assisted with human bioinformatic analyses. L.V. performed flow cytometry sorting of samples. P.L. designed ChIP–qPCR primers. J.Y. assisted in preparing single-cell suspensions from mouse tumour tissues. H.L. assisted with antigen presentation assays. L.J. assisted with the identification of Stat3+/+ and Stat3−/− mice. S. Wei identified and generated Stat3+/+ and Stat3−/− mice and guided the protocol for experimental animals. X.G. assisted with phospho-kinase array. S.G. generated Stat3+/+ and Stat3−/− mice. A.P. supervised bioinformatic analyses. M.C. assisted in acquiring human tumour samples and provided bioinformatic support. I.K. conducted in vitro human studies, assisted with the analysis of flow cytometry data, provided scientific input, engaged in discussions, revised the manuscript and supervised the study. M.D.G. contributed to the acquisition of human tumour samples and oversaw bioinformatic support. J.-X.L. provided Stat5b−/− mice. A.M.C. had a role in acquiring of clinical samples and supervised bioinformatic support. W.J.L. provided Stat5b−/− mice, provided scientific input, participated in discussions and revised the manuscript. S. Wang conceived the study, developed SD-36 and SD-2301, designed experiments, provided scientific input, participated in discussions, revised the manuscript, supervised the study and acquired funding. W.Z. conceived the study, designed experiments, composed the manuscript, provided scientific input, participated in discussions, wrote and revised the manuscript, supervised the study and acquired funding.
Corresponding authors
Ethics declarations
Competing interests
W.Z. serves as a scientific advisor or consultant for Cstone, Hanchor Bio and NextCure and was a consultant for Oncopia. S. Wang was a co-founder and a paid consultant for Oncopia Therapeutics and owned equity in Oncopia, which was acquired by Roivant Sciences and Proteovant Therapeutics. S. Wang was a paid consultant for Proteovant Therapeutics and Roivant Sciences and owned stock in Roivant Sciences. S. Wang was the principal investigator for a research contract between Oncopia/Proteovant and the University of Michigan. S. Wang is a co-founder, paid consultant, and member of the board of directors of Ascentage Pharma Group International and owns stock in Ascentage. A.M.C. was a co-founder and a paid consultant for Oncopia and owned equity in Oncopia and was a paid consultant for Proteovant Therapeutics. The University of Michigan owned stock in Oncopia and Ascentage. The University of Michigan has filed multiple patents on STAT3 degraders, which have been licensed by Oncopia/Proteovant (now SK Life Science Labs) for clinical development. S. Wang, H.Z., L.B., R.K.A., D.M. and H.M. are inventors on SD-36 and/or SD-2301 and have received royalty payments from the University of Michigan. All other authors declare no competing interests.
Peer review
Peer review information
Nature thanks David Frank, Ignacio Melero and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Role of DC STAT5 and STAT3 in ICB.
a. The Kaplan Meier (KM) survival curve was plotted to visualize the difference in survival probability between patient stratifications. Cohort 1 patients with ICB unresponsive tumors (sarcoma, lymphoma, prostate cancer, and others) were excluded for analysis. n = 69, p = 0.0374, the CoxPH model calculates the effect of the STAT5 pathway/STAT3 DC maturation (high/high group) on survival outcome relative to STAT5 pathway/STAT3 DC maturation (high/low group) when tumor type is controlled. b-c. KM curves of OS in melanoma patients (cohort 2) treated with ICB, by low or high DC score and the ratio of STAT5 versus STAT3 (DC1HiSTAT5/STAT3Hi, DCHiSTAT5/STAT3Lo, DCLoSTAT5/STAT3Hi, DCLoSTAT5/STAT3Lo). n = 72 (b), n = 42 (c), p = 0.038 (b), p = 0.051 (c) by log-rank test. d. High and low STAT3 gene signaling scores in patients with TNBC (cohort 3). Based on the STAT3 ssGSEA score, two distinct clusters were resolved. e-f. STAT5 and STAT3 gene signatures in DCs in non-responders (cohort 3). STAT5 (e) and STAT3 (f) gene signatures were analyzed in DCs in the post-ICB samples of non-responders compared to their pre-ICB samples. NS, not significant, Wilcoxon Rank-sum test. g. UMAP representations were used to identify different DC subtypes from the integrated scRNA-seq datasets of TNBC (cohort 3). The cells were then color-coded according to their respective DC subtypes. h-s. STAT5 and STAT3 gene signatures in cDCs and moDCs during ICB treatment (cohort 3). STAT5 (h, j, l, m, p, and r) and STAT3 (i, k, n, o, q, and s) gene signatures in cDCs and moDCs were compared between responders and non-responders after ICB treatment in TNBC. *p = 0.036 (m), *p = 0.034 (p), ****p < 0.0001 (h, i, l, q), Wilcoxon Rank-sum test. the central line represents the median. The lower and upper hinges of the box correspond to the 25th and 75th percentiles. The whiskers extend to the smallest and largest values no further than 1.5 * IQR from the hinges. Outliers are plotted as individual points.
Extended Data Fig. 2 STAT3 restrains STAT5-driven DC function.
a-b. Xcr1 Cre knock-in mice express an mCherry fluorescent protein under the control of the Xcr1 promoter in cDC1s. mCherry expression and XCR1-APC-Cy7 staining in DCs from Stat3+/+ and Stat3−/− mice were analyzed by FACS (a). mCherry expression and XCR1-APC-Cy7 staining in DCs (CD11c), B cells (B220), T cells (CD3 and CD90), macrophages (F4/80) from Stat3−/− mice were analyzed by FACS (b). One of four mice is shown. c. Gating strategy for cDC1s (CD11c+XCR1+) in BMDCs. d-g. Phenotype of Stat3−/− mice. The representative images show mouse body size (d), lymphoid organs (e), and T cell subset distribution (f-g) in Stat3+/+ and Stat3−/− mice. h. Western blotting shows CD80 and CD86 expression in Stat3+/+ and Stat3−/− cDC1s. One of three experiments is shown. i. Real time qPCR shows the expression of cytokine receptor mRNAs in mouse cDC1s. mean ± SEM, n = 4, ****p < 0.0001, one-way ANOVA. j. Effect of GMRβ knockdown on the JAK-STAT signaling pathway. JAWSII cells were transfected with shRNAs against Gmrβ or control shRNA and cultured with GM-CSF. Cell lysates were subjected to analysis by immunoblots. One of three experiments is shown. k. Effect of GM-CSF on JAWSII cells. The immunoblots show the expression levels of JAK family members in JAWSII cells. One of three experiments is shown. l-m. Effect of STAT3 deletion on GM-CSF and GMRβ expression in cDC1s. GM-CSF was detected by ELISA in culture supernatants in Stat3+/+ and Stat3−/− cDC1s on day 5 (l). n = 3, NS, not significant, unpaired two-tailed t-test. GMRβ was determined by Western blotting (m). One of three experiments is shown. n. The fractions of IL-6+ DCs (in red) and non-DCs are depicted in the TNBC patient dataset30. o. RNA-seq analysis (from Fig. 2n) shows the expression of Il6 by TPM (Transcripts Per Million) in mouse Stat3+/+ and Stat3−/− cDC1s. n = 4, NS, not significant, unpaired two-tailed t-test. p. Effect of STAT3 deletion on IL-6 expression in cDC1s. IL-6 was detected by ELISA in culture supernatants in Stat3+/+ and Stat3−/− cDC1s on day 4. n = 3, NS, not significant, unpaired two-tailed t-test. q-r. RNA-seq analysis (from Fig. 2n) shows the expression of IL-6 receptors (Il6ra, Il6st) by TPM (Transcripts Per Million) in mouse Stat3+/+ and Stat3−/− cDC1s. n = 4, NS, not significant, unpaired two-tailed t-test. s. Western blot shows expression of IL-6Rα and GP130 in Stat3+/+ and Stat3−/− cDC1s. One of three experiments is shown. t-x. Effect of FLLL32 on STAT5 activation and DC maturation in vivo. Stat3+/+ and Stat3−/− mice bearing peritoneal B16F10 (t) and MC38 (u-x) tumors were treated with FLLL32 on day 9 for 24 h. FACS shows expression of pSTAT5 (t, u), MHC-I (v), MHC-II (w), and CD80 (x) in peritoneal cDC1s. MFI, mean fluorescence intensity. mean ± SEM, n = 3. ****p < 0.0001 (t-w), ***p = 0.0001 (x), one-way ANOVA. y-z. GSEA analysis reveals significant differences in the CCR chemokine receptor binding (y) and interferon gamma production (z) between Stat3+/+ and Stat3−/− cDC1s. Source data contained unprocessed WB.
Extended Data Fig. 3 STAT3 dampens DC1-mediated immunity.
a-d. Effect of STAT3 deletion on DC-mediated T cell activation. OT- I cells were cultured with Stat3+/+ or Stat3−/− cDC1s without loading SIINFEKL peptides. FACS shows the percentage of Ki67+ (a), IFNγ+ (b), TNFα+ (c), and GZMB+ (d) OT- I cells. e. Gating strategy for cDC1s (CD11c+XCR1+) in MC38 tumor bearing mouse model. f. Effect of STAT3 deficiency on the interaction between JAK2, STAT3, and STAT5 in DCs in vivo. Lin− CD11c+XCR1+ cDC1s were enriched and sorted from the TME in Stat3+/+ and Stat3−/− mice bearing peritoneal MC38 tumors. Co-immunoprecipitation (IP) was performed with anti-JAK2 in DC lysates. One of two experiments with repeats is shown. g-j. MC38 tumors were inoculated into Stat3+/+ and Stat3−/− mice. MHC-II (g, h), and CD86 (i, j) expression in cDC1 from MC38 tumor-draining lymph nodes (TDLNs) were analyzed by FACS. mean ± SEM, n = 4, **p = 0.0062 (h), *p = 0.0335 (j), unpaired two-tailed t-test. k. Gating strategy for CD8+ T cells in MC38 tumor bearing mouse model. l-q. Effect of STAT3 deletion on MC38 tumor progression. FACS shows the percentages of MC38 tumor infiltrating IL-2+CD8+ T cells (l, m), and the percentages of TDLN IL-2+ (n, o), and IFNγ+ (p, q) CD8+ T cells. For l, n, and p, one representative dot plot is shown in each panel. mean ± SEM, n = 3 (m), n = 4 (o, q), *p = 0.0141 (m), **p = 0.006 (o), **p = 0.0021 (q), unpaired two-tailed t-test. r-y. Effect of STAT3 deletion on the phenotype of DCs and T cells in B16F10 bearing mice. FACS shows the expression of MHC-I (r, s) and CD80 (t, u) in tumor-infiltrating cDC1s, and the percentages of tumor infiltrating IL-2+ (v, w) and TNFα+ (x, y) CD8+ T cells. For r, t, v, and x, one representative dot plot is shown in each panel. MFI, mean fluorescence intensity. mean ± SEM, n = 3 (s, u), n = 4 (w, y), *p = 0.038 (s), *p = 0.0363 (u), *p = 0.0109 (w), *p = 0.0172 (y), unpaired two-tailed t-test. Source data contained unprocessed WB.
Extended Data Fig. 4 STAT3 degrader treats advanced tumors.
a. SD-36 structure is shown. b. Immunoblotting shows STAT1 in shStat1 JAWSII cells. c-e. Effect of STAT1 on SD-36-regulated DC phenotype. FACS shows the expression of MHC-I (c), MHC-II (d), and CD80 (e) in shStat1 JAWSII cells treated with SD-36. MFI, mean fluorescence intensity. means ± SEM, n = 3, *p = 0.0117 (c), *p = 0.0195 (d), ***p = 0.0006 (e), one-way ANOVA. f. Immunoblotting shows STAT1 in bone marrow derived DC1s from Stat1+/+ and Stat1−/− mice. One of three experiments is shown. g. Effect of STAT1 on STAT5 activation in cDC1s. FACS shows the expression of pSTAT5 in bone marrow derived Stat1+/+ and Stat1−/− cDC1s. n = 3, NS, not significant, unpaired two-tailed t-test. h-j. Effect of STAT1 on SD-36-regulated DC maturation. Stat1+/+ and Stat1−/− DCs were treated with SD-36 for 12 h. FACS shows the expression of MHC-I (h), MHC-II (i), and CD80 (j) in cDC1s. MFI, mean fluorescence intensity. means ± SEM, n = 3, *p = 0.0314 (i), *p = 0.0138 (j), **p = 0.0059 (h), one-way ANOVA. k. Effect of SD-36 on B16F10 tumor progression. B16F10 tumor bearing C57/B6 mice were treated with 10 mg/kg SD-36. Tumor volumes were shown. mean ± SEM, n = 7, *p = 0.0114, two-way ANOVA. l-m. Effect of SD-36 on MC38 tumor progression. MC38 tumor bearing C57/B6 mice were treated with SD-36. Tumor weight (l) and image (m) were shown. mean ± SEM, n = 5, ***p = 0.0004, unpaired two-tailed t-test. n-s. Effect of SD-36 on tumor progression (n-p) and mouse body weights (q-s). Mice bearing B16F10 (n, q), CT26 (o, r), and EMT6 (p, s) were treated with SD-36 (100 mg/kg) three times a week when tumor volume reached 100 mm³. Tumor volumes and mouse body weights were monitored. means ± SEM are shown, n = 8 (n, q), n = 10 (o, p, r, s), ***p = 0.0005 (n), ****p < 0.0001(o, p), two-way ANOVA. t-u. Effect of SD-36 on large tumor progression. Mice bearing CT26 tumors (500 mm3/tumor) were treated with vehicle (t) or SD-36 (u). Tumor volumes were monitored. Source data contained unprocessed WB.
Extended Data Fig. 5 STAT3 degradation in DCs boosts immunity.
a. Effect of SD-36 on tumor growth in NSG mice. MC38 tumor-bearing NSG mice were treated with SD-36 (100 mg/kg) or vehicle. Tumor volume was monitored. mean ± SEM, *p = 0.0326, two-way ANOVA. b. Efficacy of anti-CD8 mAb in CD8+ T cell depletion. MC38 bearing were treated with anti-CD8 mAb. FACS shows CD8+ T cells in the tumor draining lymph node (TDLN). One of 5 is shown. c-f. Role of SD-36 in CD8+ T cells in 4T1 tumor draining lymph nodes (TDLN). 4T1 tumor bearing mice were treated with SD-36. FACS shows the percentages of TDLN TNFα+ (c, d), and IFNγ+ (e, f) CD8+ T cells. For c, e, one representative dot plot is shown in each panel. mean ± SEM, n = 4, *p = 0.0184 (f), **p = 0.0063 (d), unpaired two-tailed t-test. g-r. Role of SD-36 in CD8+ T cells in LLC tumor bearing mice. FACS shows the percentages of TNFα+ (g, h), IFNγ+ (i, j), and GZMB+ (k, l) CD8+ T cells from tumors (g-l) and tumor-draining lymph nodes (m-r). For g, i, k, m, o, and q, one representative dot plot is shown in each panel. mean ± SEM, n = 6 (h, j, l), n = 4 (n, p, r), *p = 0.0224 (p), *p = 0.0294 (r), **p = 0.0072 (h), **p = 0.0071 (n), ***p = 0.0007(j), ***p = 0.0008 (l), unpaired two-tailed t-test. s. Effect of SD-36 on tumor growth in Batf3−/− mice. Batf3−/− mice were inoculated with LLC cells and with SD-36 (n = 8) or vehicle (n = 11). Tumor volumes were monitored. t. Effect of SD-36 on STAT3 protein in DCs and LLC tumor cells. Immune blotting shows the levels of STAT3 in DCs and LLC tumor cells treated with SD-36 for 12 h, and the ratio of STAT3 to actin expression. u. DCs and LLC tumor cells were treated with 1 μM SD-36 for 12 h. Intracellular SD-36 was quantified by LC-MS/MS. mean ± SEM, n = 4, **p = 0.0011, unpaired two-tailed t-test. v. Role of SD-36 in DCs. MC38 tumor-bearing mice were treated with SD-36 or vehicle. Immune blotting shows key transcriptional factors in DCs isolated from spleens. One of three experiments is shown. w. Effect of cDC1 STAT3 in SD-36-regulated tumor progression. MC38 tumor bearing Stat3+/+ and Stat3−/− mice were treated with SD-36 or vehicle. Tumor volume was determined. mean ± SEM, n = 7, **p = 0.0056, two-way ANOVA. x. Role of cDC1 STAT3 in SD-36-regulated anti-tumor immunity. Stat3+/+ and Stat3−/− cDC1s were transferred into MC38-bearing Batf3−/− mice. These mice were treated with SD-36. Tumor volume was monitored. mean ± SEM, n = 5, **p = 0.002, two-way ANOVA. Source data contained unprocessed WB.
Extended Data Fig. 6 STAT3 degradation synergizes ICB.
a-b. Effect of SD-36 on ICB efficacy in humans in vitro. DCs from human ovarian cancer tissues were pretreated with SD-36, washed, and co-cultured with T cells in the presence of anti-human PD-L1 for 3 days. FACS shows the percentage of GZMB+IFNγ+ (a) and IL-2+TNFα+ (b) CD8+ T cells, mean ± SEM, n = 4, *p = 0.0237 (a, control vs. SD-36), *p = 0.0135 (a, control vs. Anti-PD-L1), *p = 0.0302 (b), ****p < 0.0001 (a), one-way ANOVA. c. Chemical structure of SD-2301 structure. d. Effect of SD-2301 on STAT3 expression in DCs in vitro. DCs were treated with SD-2301 (50 nM) for 24 h. Immune blotting shows STAT3 protein. One of 3 is shown. e. Effect of SD-2301 on STAT3 expression in DCs in vivo. Wild type mice bearing peritoneal B16F10 tumors were treated SD-2301 (5 mg/kg). Immune blotting shows STAT3 expression in peritoneal DCs. n = 3/group. f. Comparison of the effect of SD-2301 and SD-36 on STAT3 degradation in mouse DCs. DCs were treated with SD-2301 and SD-36 for 24 h. Immune blotting shows STAT3 expression in DCs. One of three experiments is shown. g-n. Comparison of the anti-tumor effect of SD-2301 and SD-36. B16F10 bearing mice were treated with SD-2301 (5 mg/kg) or SD-36 (20 mg/kg) every 3 days. Tumor volume was monitored (g). FACS shows the expression of IFNγ (h, i) and GZMB (j, k) in B16F10 infiltrating CD8+ T cells, and the expression of MHC-I (l), MHC-II (m), and CD80 (n) in tumor infiltrating cDC1s. MFI, mean fluorescence intensity. means ± SEM, n = 7 (g), n = 4 (i, k, l, m, n), **p = 0.0023 (g), ***p = 0.0003 (g), two-way ANOVA (g); **p = 0.0014 (i), **p = 0.0015 (l), **p = 0.0046 (n), ***p = 0.0001 (k), ***p = 0.0001 (m), unpaired two-tailed t-test (i, k, l, m, n). o-p. Effect of SD-2301 on blood vessels in tumors. B16-F10 and MC38 bearing mice were treated with SD-2301. CD31+ vessel density was quantified in B16-F10 (o) and MC38 (p) tumors. Representative immunohistochemical staining images and the quantification of CD31+ vessel densities are shown. n = 4 tumors, NS, not significant, unpaired two-tailed t-test. q. Effect of SD-2301 on mouse body weights. B16F10 bearing mice were treated with SD-2301 (5 mg/kg) every 3 days for 18 days. Mouse body weights were monitored. r-s. Effect of SD-2301 on ICB efficacy in vivo. B16F10 (r) or MC38 (s) bearing mice were treated with SD-2301, anti-PD-L1, and their combination. The group of B16F10 bearing mice (r) treated with SD-2301 is identical to that shown in Extended Data Fig. 6g. Tumor volumes were monitored. mean ± SEM, n = 7, *p = 0.0106, ***p = 0.0009, ****p < 0.0001, two-way ANOVA. t-u. Effect of SD-2301 on neoantigen specific CD8+ T cell response. MC38 bearing mice were treated with SD-2301 or vehicle. FACS shows the percentage of tumor infiltrating KSPWFTTL tetramer+CD8+ T cells. means ± SEM, n = 6, *p = 0.0216, unpaired two-tailed t-test. v. Effect of SD-2301 on the role of STAT3 in DC-regulated tumor progression. B16F10 tumor bearing Stat3+/+ and Stat3−/− mice were treated with SD-2301 or vehicle. Tumor volumes were monitored. mean ± SEM, n = 7, ****p < 0.0001, Two-way ANOVA. w. Effect of SD-2301 on cDC1-regulated tumor progression. MC38 bearing Batf3+/+ and Batf3−/− mice were treated with SD-2301 or vehicle. The group of control mice treated with SD-2301 is identical to that shown in Extended Data Fig. 6s. Tumor volumes were monitored. mean ± SEM, n = 7, ****p < 0.0001, Two-way ANOVA. x. Pharmacokinetics (PK) of SD-2301 in mice. Female CD-1 mice (n = 3) were administered with SD-2301 (5 mg/kg). Blood samples were collected for PK analysis. Cl, clearance; Vss, volume of distribution at steady state; T1/2, elimination half-life; Cmax, maximum drug concentration; AUC, area-under-the-curve. y. Effect of SD-2301 on STAT3 expression in human immune cells. Human peripheral blood mononuclear cells were treated with different concentrations of SD-2301 for 14 h. The immunoblots show the expression levels of STATs. One of two repeats. z. The schematic diagram shows that STAT3 restrains the JAK2-STAT5 signaling pathway, thereby reducing STAT5-dependent DC maturation and lessening T cell immunity and immunotherapy effectiveness. Source data contained unprocessed WB.
Supplementary information
Supplementary Information (download PDF )
This file contains experimental procedure for SD-2301, and Supplementary Figs. 1–4. Supplementary Fig. 1: Synthesis of SD-2301. Supplementary Fig. 2: UPLC spectrum of SD-2301. Supplementary Fig. 3: 1H-NMR Spectrum of SD-2301 (400 MHz, Methanol-d4). Supplementary Fig. 4: 13C-NMR Spectrum for SD-2301 (101 MHz, Methanol-d4).
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, J., Tison, K., Zhou, H. et al. STAT5 and STAT3 balance shapes dendritic cell function and tumour immunity. Nature 643, 519–528 (2025). https://doi.org/10.1038/s41586-025-09000-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09000-3
This article is cited by
-
Targeting inflammatory microenvironments: overcoming therapy resistance and immunosuppression
Molecular Cancer (2026)
-
Pharmacological Inhibition of JAK/STAT-IL2 Axis Alleviated Cisplatin-Induced Ototoxicity
Molecular Neurobiology (2026)
-
Discovery of a potent STAT3 inhibitor WR-S-647 for the treatment of ADPKD
Archives of Pharmacal Research (2026)
-
Recent advances in targeting protein degradation for tumor immunotherapy
Journal of Hematology & Oncology (2025)
-
The emerging role of dendritic cells in the tumor microenvironment: from antigen presentation to targeted immunotherapy
Cell Death & Disease (2025)