Abstract
Across mammalian species, new mothers undergo behavioural changes to nurture their offspring and meet the caloric demands of milk production1,2,3,4,5. Although many neural circuits underlying feeding and parenting behaviours are well characterized6,7,8,9, it is unclear how these different circuits interact and adapt during lactation. Here we performed transcriptomic profiling of the arcuate nucleus (ARC) and the medial preoptic area (MPOA) of the mouse hypothalamus in response to lactation and hunger. Furthermore, we showed that heightened appetite in lactating mice was accompanied by increased activity of hunger-promoting agouti-related peptide (AgRP) neurons in the ARC (ARCAgRP neurons). To assess the strength of hunger versus maternal drives, we designed a conflict assay in which female mice chose between a food source or pups and nesting material. Although food-deprived lactating mothers prioritized parenting over feeding, hunger reduced the duration and disrupted the sequences of parenting behaviours in both lactating and virgin females. We found that ARCAgRP neurons inhibit bombesin receptor subtype 3 (BRS3) neurons in the MPOA (MPOABRS3 neurons), which become more active postpartum and govern parenting and satiety. Activation of this ARCAgRP-to-MPOABRS3 circuit shifted behaviours from parenting to food-seeking. Thus, hypothalamic networks are modulated by physiological states and work antagonistically during the prioritization of competing motivated behaviours.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data are provided in all main and extended figures and accompanying Source Data. Our raw sequencing data are deposited in the Gene Expression Omnibus (GEO) with the accession number GSE295610. Other public sequencing data analysed in our study are from GEO with the accession numbers GSE93374 and GSE113576. Our processed sequencing data are available at Zenodo (https://doi.org/10.5281/zenodo.15319501)68. Source data are provided with this paper.
Code availability
Code used to analyse scRNA-seq data was derived from https://satijalab.org/seurat/. Code used to analyse FED3 data was from https://github.com/earnestt1234/FED3_Viz. Custom code for our study is available at Zenodo (https://doi.org/10.5281/zenodo.15319501)68.
References
Prentice, A. M. & Prentice, A. Energy costs of lactation. Annu. Rev. Nutr. 8, 63–79 (1988).
Fleming, A. S. & Rosenblatt, J. S. Maternal behavior in the virgin and lactating rat. J. Comp. Physiol. Psychol. 86, 957–972 (1974).
Brunton, P. J. & Russell, J. A. The expectant brain: adapting for motherhood. Nat. Rev. Neurosci. 9, 11–25 (2008).
Numan, M. & Woodside, B. Maternity: neural mechanisms, motivational processes, and physiological adaptations. Behav. Neurosci. 124, 715–741 (2010).
Woodside, B., Budin, R., Wellman, M. K. & Abizaid, A. Many mouths to feed: the control of food intake during lactation. Front. Neuroendocrinol. 33, 301–314 (2012).
Sohn, J. W., Elmquist, J. K. & Williams, K. W. Neuronal circuits that regulate feeding behavior and metabolism. Trends Neurosci. 36, 504–512 (2013).
Ahn, B. H., Kim, M. & Kim, S. Y. Brain circuits for promoting homeostatic and non-homeostatic appetites. Exp. Mol. Med. 54, 349–357 (2022).
Alcantara, I. C., Tapia, A. P. M., Aponte, Y. & Krashes, M. J. Acts of appetite: neural circuits governing the appetitive, consummatory, and terminating phases of feeding. Nat. Metab. 4, 836–847 (2022).
Kohl, J. Parenting—a paradigm for investigating the neural circuit basis of behavior. Curr. Opin. Neurobiol. 60, 84–91 (2020).
Burnett, C. J. et al. Hunger-driven motivational state competition. Neuron 92, 187–201 (2016).
Burnett, C. J. et al. Need-based prioritization of behavior. eLife 8, e44527 (2019).
Sutton, A. K. & Krashes, M. J. Integrating hunger with rival motivations. Trends Endocrinol. Metab. 31, 495–507 (2020).
Barajas-Azpeleta, R., Tastekin, I. & Ribeiro, C. Neuroscience: How the brain prioritizes behaviors. Curr. Biol. 31, R1125–R1127 (2021).
Kovac, M. P. & Davis, W. J. Neural mechanism underlying behavioral choice in Pleurobranchaea. J. Neurophysiol. 43, 469–487 (1980).
Hong, W., Kim, D. W. & Anderson, D. J. Antagonistic control of social versus repetitive self-grooming behaviors by separable amygdala neuronal subsets. Cell 158, 1348–1361 (2014).
Cheriyamkunnel, S. J. et al. A neuronal mechanism controlling the choice between feeding and sexual behaviors in Drosophila. Curr. Biol. 31, 4231–4245.e4 (2021).
Mei, L., Yan, R., Yin, L., Sullivan, R. M. & Lin, D. Antagonistic circuits mediating infanticide and maternal care in female mice. Nature 618, 1006–1016 (2023).
de Araujo Salgado, I. et al. Toggling between food-seeking and self-preservation behaviors via hypothalamic response networks. Neuron 111, 2899–2917.e6 (2023).
Marlin, B. J., Mitre, M., D’Amour J, A., Chao, M. V. & Froemke, R. C. Oxytocin enables maternal behaviour by balancing cortical inhibition. Nature 520, 499–504 (2015).
Stolzenberg, D. S. & Champagne, F. A. Hormonal and non-hormonal bases of maternal behavior: The role of experience and epigenetic mechanisms. Horm. Behav. 77, 204–210 (2016).
Ammari, R. et al. Hormone-mediated neural remodeling orchestrates parenting onset during pregnancy. Science 382, 76–81 (2023).
Matikainen-Ankney, B. A. et al. An open-source device for measuring food intake and operant behavior in rodent home-cages. eLife 10, e66173 (2021).
Campbell, J. N. et al. A molecular census of arcuate hypothalamus and median eminence cell types. Nat. Neurosci. 20, 484–496 (2017).
Krashes, M. J. et al. Rapid, reversible activation of AgRP neurons drives feeding behavior in mice. J. Clin. Invest. 121, 1424–1428 (2011).
Aponte, Y., Atasoy, D. & Sternson, S. M. AGRP neurons are sufficient to orchestrate feeding behavior rapidly and without training. Nat. Neurosci. 14, 351–355 (2011).
Takahashi, K. A. & Cone, R. D. Fasting induces a large, leptin-dependent increase in the intrinsic action potential frequency of orexigenic arcuate nucleus neuropeptide Y/Agouti-related protein neurons. Endocrinology 146, 1043–1047 (2005).
Mandelblat-Cerf, Y. et al. Arcuate hypothalamic AgRP and putative POMC neurons show opposite changes in spiking across multiple timescales. eLife 4, e07122 (2015).
Chen, Y., Lin, Y. C., Kuo, T. W. & Knight, Z. A. Sensory detection of food rapidly modulates arcuate feeding circuits. Cell 160, 829–841 (2015).
Betley, J. N. et al. Neurons for hunger and thirst transmit a negative-valence teaching signal. Nature 521, 180–185 (2015).
Padilla, S. L. et al. Agouti-related peptide neural circuits mediate adaptive behaviors in the starved state. Nat. Neurosci. 19, 734–741 (2016).
Alhadeff, A. L. et al. A Neural circuit for the suppression of pain by a competing need state. Cell 173, 140–152.e15 (2018).
Li, X. Y. et al. AGRP neurons project to the medial preoptic area and modulate maternal nest-building. J. Neurosci. 39, 456–471 (2019).
Numan, M. Medial preoptic area and maternal behavior in the female rat. J. Comp. Physiol. Psychol. 87, 746–759 (1974).
Numan, M., Numan, M. J., Marzella, S. R. & Palumbo, A. Expression of c-fos, fos B, and egr-1 in the medial preoptic area and bed nucleus of the stria terminalis during maternal behavior in rats. Brain Res. 792, 348–352 (1998).
Wu, Z., Autry, A. E., Bergan, J. F., Watabe-Uchida, M. & Dulac, C. G. Galanin neurons in the medial preoptic area govern parental behaviour. Nature 509, 325–330 (2014).
Kohl, J. et al. Functional circuit architecture underlying parental behaviour. Nature 556, 326–331 (2018).
Fang, Y. Y., Yamaguchi, T., Song, S. C., Tritsch, N. X. & Lin, D. A hypothalamic midbrain pathway essential for driving maternal behaviors. Neuron 98, 192–207.e10 (2018).
Brown, R. S. E. et al. Prolactin action in the medial preoptic area is necessary for postpartum maternal nursing behavior. Proc. Natl Acad. Sci. USA 114, 10779–10784 (2017).
Yoshihara, C. et al. Calcitonin receptor signaling in the medial preoptic area enables risk-taking maternal care. Cell Rep. 35, 109204 (2021).
Atasoy, D., Betley, J. N., Su, H. H. & Sternson, S. M. Deconstruction of a neural circuit for hunger. Nature 488, 172–177 (2012).
Betley, J. N., Cao, Z. F., Ritola, K. D. & Sternson, S. M. Parallel, redundant circuit organization for homeostatic control of feeding behavior. Cell 155, 1337–1350 (2013).
Moffitt, J. R. et al. Molecular, spatial, and functional single-cell profiling of the hypothalamic preoptic region. Science 362, eaau5324 (2018).
Guenthner, C. J., Miyamichi, K., Yang, H. H., Heller, H. C. & Luo, L. Permanent genetic access to transiently active neurons via TRAP: targeted recombination in active populations. Neuron 78, 773–784 (2013).
DeNardo, L. A. et al. Temporal evolution of cortical ensembles promoting remote memory retrieval. Nat. Neurosci. 22, 460–469 (2019).
Cazzulino, A. S., Martinez, R., Tomm, N. K. & Denny, C. A. Improved specificity of hippocampal memory trace labeling. Hippocampus 26, 752–762 (2016).
McHenry, J. A. et al. Hormonal gain control of a medial preoptic area social reward circuit. Nat. Neurosci. 20, 449–458 (2017).
Ohki-Hamazaki, H. et al. Mice lacking bombesin receptor subtype-3 develop metabolic defects and obesity. Nature 390, 165–169 (1997).
Guan, X. M. et al. Antiobesity effect of MK-5046, a novel bombesin receptor subtype-3 agonist. J. Pharmacol. Exp. Ther. 336, 356–364 (2011).
Pinol, R. A. et al. Preoptic BRS3 neurons increase body temperature and heart rate via multiple pathways. Cell Metab. 33, 1389–1403.e6 (2021).
Motta, S. C. et al. Ventral premammillary nucleus as a critical sensory relay to the maternal aggression network. Proc. Natl Acad. Sci. USA 110, 14438–14443 (2013).
Donato, J. Jr. et al. Lesions of the ventral premammillary nucleus disrupt the dynamic changes in Kiss1 and GnRH expression characteristic of the proestrus-estrus transition. Neuroscience 241, 67–79 (2013).
Donato, J. Jr. et al. The ventral premammillary nucleus links fasting-induced changes in leptin levels and coordinated luteinizing hormone secretion. J. Neurosci. 29, 5240–5250 (2009).
Mei, L., Osakada, T. & Lin, D. Hypothalamic control of innate social behaviors. Science 382, 399–404 (2023).
Chen, P., Li, C., Haskell-Luevano, C., Cone, R. D. & Smith, M. S. Altered expression of agouti-related protein and its colocalization with neuropeptide Y in the arcuate nucleus of the hypothalamus during lactation. Endocrinology 140, 2645–2650 (1999).
Phillips, C. T. & Palmiter, R. D. Role of agouti-related protein-expressing neurons in lactation. Endocrinology 149, 544–550 (2008).
Suzuki, Y. et al. Changes in mRNA expression of arcuate nucleus appetite-regulating peptides during lactation in rats. J. Mol. Endocrinol. 52, 97–109 (2014).
Catalbas, K. et al. Hypothalamic AgRP neurons regulate the hyperphagia of lactation. Mol. Metab. 86, 101975 (2024).
Szymczak-Workman, A. L., Vignali, K. M. & Vignali, D. A. Verification of 2 A peptide cleavage. Cold Spring Harb. Protoc. 2012, 255–257 (2012).
Raymond, C. S. & Soriano, P. High-efficiency FLP and PhiC31 site-specific recombination in mammalian cells. PLoS ONE 2, e162 (2007).
Mickelsen, L. E. et al. Neurochemical heterogeneity among lateral hypothalamic hypocretin/orexin and melanin-concentrating hormone neurons identified through single-cell gene expression analysis. eNeuro https://doi.org/10.1523/ENEURO.0013-17.2017 (2017).
Mickelsen, L. E. et al. Cellular taxonomy and spatial organization of the murine ventral posterior hypothalamus. eLife 9, e58901 (2020).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030.e16 (2018).
Friard, O. & Gamba, M. BORIS: a free, versatile open-source event-logging software for video/audio coding and live observations. Methods Ecol. Evol. 7, 1325–1330 (2016).
Gao, C. et al. Molecular and spatial profiling of the paraventricular nucleus of the thalamus. eLife 12, e81818 (2023).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinformatics 22, 433 (2021).
Alcantara, I. C., Papas, B. N., & Krashes, M. J. Single-cell RNA-sequencing of the ARC and MPOA across satiety and lactation states in female mice. Zenodo https://doi.org/10.5281/zenodo.15319501 (2025).
Acknowledgements
The authors thank A. Fleischmann, W. Li, W. S. Young, and members of the M.J.K. and A. Lutas laboratories for discussions and support; G. Parham for assisting with coding and data analysis; and A. Franks for assisting with mouse husbandry. This research was funded by the Intramural Research Program of the National Institutes of Health, the National Institute of Diabetes and Digestive and Kidney Diseases (ZIA-DK075087, ZIA-DK075088) and the National Institute of Environmental Health Sciences (ZIC-ES102545).
Author information
Authors and Affiliations
Contributions
I.C.A. and M.J.K. conceived the project and designed the experiments. I.C.A. and C.L. performed stereotaxic surgeries. I.C.A., S.R.G., A.I.G. and E.O.K. performed behavioural experiments and fibre photometry recordings. R.A.P. performed cold exposure experiments paired fibre photometry recordings. C.L. performed slice electrophysiological recordings. I.C.A. analysed behavioural data. I.C.A., C.L. and R.A.P. analysed neuronal activity data. I.C.A., L.E.M., C.M.M. and G.C. collected samples for scRNA-seq. I.C.A., C.G., B.N.P. and J.-L.L. analysed the scRNA-seq data. I.C.A., C.G. and I.M.C. performed RNAscope and analysed the data. I.C.A., A.I.G. and M.J.K. performed histology and antibody staining. A.I.G. performed cell counting and axonal projection density analysis. C.X. and M.L.R. generated the Brs3-T2A-FlpO mouse line. I.C.A., C.L., B.N.P., A.I.G. and I.d.A.S. prepared the figures. I.C.A. wrote the manuscript, which all authors reviewed and edited.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Dayu Lin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Lactating mice exhibit increased appetite during dark and light cycles.
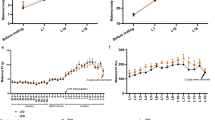
a, mean daily food intake of control (n = 8) and lactating (n = 14) mice during baseline, pregnancy, lactation, and post-weaning. b-c, daily food intake of representative lactating mouse with 9 pups (b) or 3 pups (c). d-i, daily food intake measurement using FED3 in the dark cycle (d-e) and light cycle (f-g) of control (n = 6) and lactating (n = 5) mice during baseline, pregnancy, lactation, and post-weaning. h-i, same data as e and g but replotted to compare light vs. dark cycle food intake in control (h) and lactating (i) mice. Plots show mean ± s.e.m. Data analyzed using two-way RM-ANOVA with Šidák’s multiple comparisons test (a, e, g, h, i). Statistical details in Source Data.
Extended Data Fig. 2 Agrp+ neuronal cluster in the ARC exhibits the most changes in gene expression upon fasting.
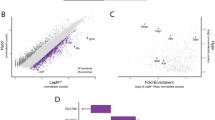
a, heat map of relative expression of marker genes in each ARC neuronal cluster with dendrogram (top) showing similarities between clusters. b, comparison of our ARC clusters (columns) with ARC clusters from a previous scRNA-seq publication (rows). c, comparison of fasted virgin vs. fed virgin groups showing the contribution of each cluster to the first 8 principal components (PCs), highlighting the top contributor, cluster 0. Dashed red line shows contribution if all cluster contributed equally. d, FeaturePlot showing ARC neurons and clusters in UMAP space with Agrp and Npy expression.
Extended Data Fig. 3 Hunger delays and reduces parenting behaviors in virgin females and lactating mothers in the dark cycle.
a, schematic of the timing of fasting, refeeding, and conflict assay for fasted and refed groups. b, percent of animals that either performed pup retrieval or feeding as their first behavior (nvirgin = 7 mice, nlactating = 6 mice). c-f, probability density of pup retrieval and feeding events throughout the conflict assay (all trials). 1 total pellet consumed by 1 virgin female (refed) at time = 32.65 min is not shown to avoid skewing the visualization. g, transition probabilities between pup retrieval and feeding events in fasted virgin females and fasted lactating mothers. Only the feeding events prior to the completion of pup retrieval were used for the calculation. h, food intake. i, number of pups retrieved. j, latency to retrieve the first pup. k, nest score. l, preference index. m, food intake (# pellets) vs. % time in parenting chamber of virgin and lactating females in the fasted condition. Bar graphs show mean ± s.e.m. Box plots show median, upper/lower quartiles, and upper/lower extremes. Data analyzed using two-way RM-ANOVA with Šidák’s multiple comparisons test (h-l) or simple linear regression (m). Statistical details in Source Data.
Extended Data Fig. 4 Photostimulation of ARCAgRP projections to MPOA expressing ChR2 or eYFP.
a-i, ARCAgRP-ChR2. a, schematic of stimulation paradigms in fed mice. Created in BioRender. Alcantara, I. (2025) https://BioRender.com/09eqaav. b, 30 min food intake across different stimulation paradigms at 10 and 1 mW light intensity; “no stim” data are identical for 10 and 1 mW comparisons (n = 10 virgin females). c-d conflict assay without food (n = 8 virgin females). c, number of pups retrieved. d, number of pups attacked. e, nest score. f-i, conflict assay in Npy-KO background (n = 6 virgin females). f, food intake. g, number of pups retrieved. h, number of pups attacked. i, nest score. j-o, ARCAgRP-eYFP. j, histology. k, food intake. l-o, conflict assay. l, food intake. m, nest score. n, number of pups retrieved. o, preference index. Scale bars = 200 μm. Bar graphs show mean ± s.e.m. Box plots show median, upper/lower quartiles, and upper/lower extremes. Data analyzed using one-way RM-ANOVA with Šidák’s multiple comparisons test (b), one-sided paired t-test (c-e, g-h), two-sided paired t-tests (f, i, k) or one-way RM-ANOVA with Tukey’s multiple comparisons test (l-o). Statistical details in Source Data.
Extended Data Fig. 5 Inhibition of parenting neurons in MPOA reduces parenting in fed lactating mothers.
a, number of pups retrieved. b, nest score. c-d, food intake over time in Parent-TRAP (c) and Negative-TRAP (d) animals. e, total food intake at 40 min time point. f-g, preference index over time in Parent-TRAP (f) and Negative-TRAP (g) animals (n = 7 Parent-TRAP lactating mothers, n = 6 Negative TRAP lactating mothers). Bar and line graphs show mean ± s.e.m. Box plots show median, upper/lower quartiles, and upper/lower extremes. Data analyzed using two-way RM-ANOVA with Šidák’s multiple comparisons test. P-values in c-d, f-g indicate the treatment effect of CNO. Statistical details in Source Data.
Extended Data Fig. 6 MPOA neuronal clusters co-express several genetic markers for parenting-promoting neurons.
a, heat map of relative expression of marker genes in each MPOA neuronal cluster with dendrogram (top) showing similarities between clusters. b, comparison of our MPOA clusters (columns) with MPOA clusters from a previous scRNA-seq publication (rows). c, comparison of fed lactating vs. fed virgin groups showing the contribution of each cluster to the first 9 PCs. d, comparison of fasted virgin vs. fed virgin groups showing the contribution of each cluster to the first 10 PCs. e, DotPlot of clusters 3, 5, and 14 showing expression of Slc17a6 (Vglut2), Slc32a1 (Vgat), top 5 enriched genes in cluster 3 (Pde1c, Ptprk, Esr1, Dgkb, Grm8), parenting markers (Gal, Calcr, Prlr, Pgr, Brs3), and NPY receptors (Npy1r, Npy2r). f, fractions of neurons that are Brs3+, i.e. contains at least 1 unique molecular identifier (UMI), per MPOA cluster. g, left: co-expression matrix showing fraction of Gene A+ MPOA neurons that are also Gene B+, highlighting co-expression with Brs3 (e.g., 0.38 of Brs3+ neurons are Slc17a6+); right: fractions of MPOA neurons that are positive for select genes.
Extended Data Fig. 7 Inhibition of Esr1, Calcr, Crh, or MC4R neurons in MPOA with hM4Di.
a-d, conflict assay: histology (scale bars = 200 μm) (1), food intake (2), number of pups retrieved (3), nest score (4), and preference index (5) of animals expressing hM4Di in MPOAEsr1 (a, n = 8 virgin females), MPOACalcr (b, n = 10 virgin females), MPOACrh (c, n = 6 virgin females), or MPOAMC4R (d, n = 6 lactating mothers). Bar graphs show mean ± s.e.m. Box plots show median, upper/lower quartiles, and upper/lower extremes. Data analyzed using two-way RM-ANOVA with Fisher’s LSD test. Statistical details in Source Data.
Extended Data Fig. 8 Photometry MPOABRS3 expressing eYFP or GCaMP6s during parenting, feeding, and cold exposure.
a-e, MPOABRS3-eYFP fiber photometry of virgin females (n = 5). a, histology (scale bar = 200 μm). b-d, signal in response to pup (b), object (c), or food (d). e, mean response to all 3 stimuli. f-n, MPOABRS3-GCaMP6s fiber photometry in lactating mothers (n = 6). f, signal in response to pup or object in the fed condition. g, mean response to pup or object. h, signal aligned to pup presentation. i, mean response to pup in the fasted state. j, signal aligned to food presentation. k, mean response to food. l-m, representative traces in the fed (l) and fasted (m) states aligned to behaviors. n, mean signal during behaviors. o-q, MPOABRS3-GCaMP6s neurons in virgin females (n = 7) during cold exposure. o, experimental setup. Created in BioRender. Alcantara, I. (2025) https://BioRender.com/09eqaav. p, GCaMP6s signal (bottom) aligned to changes in ambient temperature (top). q, mean signal at 30 °C or 15 °C at time = 5 to 30 min. Bar and line graphs show mean ± s.e.m. Box plots show median, upper/lower quartiles, and upper/lower extremes. Data analyzed using two-way RM-ANOVA with Šidák’s multiple comparisons test (e, n) or two-tailed paired t-tests (g, i, k, q). P-values in n indicate the effect of hunger states and behavior types to the variance by two-way RM-ANOVA. Statistical details in Source Data.
Extended Data Fig. 9 Generation of the Brs3-T2A-FlpO line.
a, schematic of the Brs3-T2A-FlpO locus with genotyping strategy by PCR (n = 3 FlpO/+ mice); x475 = wild-type forward primer, x803 = FlpO forward primer, x564 = reverse primer. b, sequences of guide RNAs and long single-stranded DNA template. c, schematic of Flp-dependent GCaMP6s virus injection in MPOA of Brs3-T2A-FlpO line (n = 3 mice). d, representative confocal images from RNAscope experiment showing expression of Flp-dependent GCaMP mRNA and colocalization with Brs3 mRNA in MPOA (scale bar = 50 μm). e, representative images from n = 3 mice showing expression pattern of tdTomato in Brs3-T2A-FlpO mouse line crossed with RC::FLTG reporter along anterior/posterior axis relative to Bregma (scale bars = 200 μm).
Extended Data Fig. 10 Intrinsic properties of MPOABRS3 neurons in virgin females and lactating mothers measured by whole-cell patch clamp.
a, proportion of cells in depolarization block (DB) in virgin and lactating mice (nvirgin = 46 cells, nlactating = 40 cells, n = 6 mice each group). b, current required to hold cells at −70 mV. c, rheobase at −70 mV. d, capacitance. e, resistance. f, inter-spike membrane potential. Data analyzed using two-way RM-ANOVA with Šidák’s multiple comparisons test (b-f). Statistical details in Source Data.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1 and Supplementary Table 1
Source data
Rights and permissions
About this article
Cite this article
Alcantara, I.C., Li, C., Gao, C. et al. A hypothalamic circuit that modulates feeding and parenting behaviours. Nature 645, 981–990 (2025). https://doi.org/10.1038/s41586-025-09268-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09268-5