Abstract
Glycosylation is central to the localization and function of biomolecules1. We recently discovered that small RNAs undergo N-glycosylation2 at the modified RNA base 3-(3-amino-3-carboxypropyl) uridine (acp3U)3. However, the functional significance of N-glycosylation of RNAs is unknown. Here we show that the N-glycans on glycoRNAs prevent innate immune sensing of endogenous small RNAs. We found that de-N-glycosylation of cell-culture-derived and circulating human and mouse glycoRNA elicited potent inflammatory responses including the production of type I interferons in a Toll-like receptor 3- and Toll-like receptor 7-dependent manner. Furthermore, we show that N-glycans on cell surface RNAs prevent apoptotic cells from triggering endosomal RNA sensors in efferocytes, thus facilitating the non-inflammatory clearance of dead cells. Mechanistically, N-glycans conceal the hypermodified uracil base acp3U, which we identified as immunostimulatory when exposed in RNA. Consistent with this, genetic deletion of an enzyme (DTWD2) that synthesizes acp3U abrogated innate immune activation by de-N-glycosylated small RNAs and apoptotic cells. Furthermore, synthetic acp3U-containing RNAs are sufficient to trigger innate immune responses. Thus, our study has uncovered a natural mechanism by which N-glycans block RNAs from inducing acp3U-dependent innate immune activation, demonstrating how glycoRNAs exist on the cell surface and in the endosomal network without inducing autoinflammatory responses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-seq data have been deposited and are accessible at Gene Expression Omnibus number GSE291767. Source data are provided with this paper.
References
Varki, A. et al. Essentials of Glycobiology 3rd edn (Cold Spring Harbor Laboratory Press, 2015).
Flynn, R. A. et al. Small RNAs are modified with N-glycans and displayed on the surface of living cells. Cell 184, 3109–3124 (2021).
Xie, Y. et al. The modified RNA base acp3U is an attachment site for N-glycans in glycoRNA. Cell 187, 5228–5237.e12 (2024).
Alexopoulou, L., Holt, A. C., Medzhitov, R. & Flavell, R. A. Recognition of double-stranded RNA and activation of NF-kappaB by Toll-like receptor 3. Nature 413, 732–738 (2001).
Diebold, S. S., Kaisho, T., Hemmi, H., Akira, S. & Reis e Sousa, C. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science 303, 1529–1531 (2004).
Heil, F. et al. Species-specific recognition of single-stranded RNA via Toll-like receptor 7 and 8. Science 303, 1526–1529 (2004).
Greulich, W. et al. TLR8 is a sensor of RNase T2 degradation products. Cell 179, 1264–1275 (2019).
Asami, J. & Shimizu, T. Structural and functional understanding of the Toll-like receptors. Protein Sci. 30, 761–772 (2021).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat. Immunol. 5, 730–737 (2004).
Kato, H. et al. Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid-inducible gene-I and melanoma differentiation-associated gene 5. J. Exp. Med. 205, 1601–1610 (2008).
Sato, M. et al. Positive feedback regulation of type I IFN genes by the IFN-inducible transcription factor IRF-7. FEBS Lett. 441, 106–110 (1998).
Juang, Y. T. et al. Primary activation of interferon A and interferon B gene transcription by interferon regulatory factor 3. Proc. Natl Acad. Sci. USA 95, 9837–9842 (1998).
Huang, Y. et al. TLR7 promotes skin inflammation via activating NFκB-mTORC1 axis in rosacea. PeerJ 11, e15976 (2023).
Nainytė, M., Amatov, T. & Carell, T. Synthesis of an acp. Chem. Commun. 55, 12216–12218 (2019).
Ma, Y. et al. Spatial imaging of glycoRNA in single cells with ARPLA. Nat. Biotechnol. 42, 608–616 (2023).
Ren, Z. et al. Enzyme-mediated proximity labeling identifies small RNAs in the endoplasmic reticulum lumen. Biochemistry 62, 1844–1848 (2023).
Arandjelovic, S. & Ravichandran, K. S. Phagocytosis of apoptotic cells in homeostasis. Nat. Immunol. 16, 907–917 (2015).
Kawano, M. & Nagata, S. Efferocytosis and autoimmune disease. Int. Immunol. 30, 551–558 (2018).
Xie, Y. et al. Development and application of GlycanDIA workflow for glycomic analysis. Preprint at bioRxiv https://doi.org/10.1101/2024.03.12.584702 (2024).
Perr, J. et al. RNA-binding proteins and glycoRNAs form domains on the cell surface for cell-penetrating peptide entry. Cell https://doi.org/10.1016/j.cell.2025.01.040 (2025).
Kawai, T., Ikegawa, M., Ori, D. & Akira, S. Decoding Toll-like receptors: recent insights and perspectives in innate immunity. Immunity 57, 649–673 (2024).
Fisch, D. et al. Molecular definition of the endogenous Toll-like receptor signalling pathways. Nature 631, 635–644 (2024).
Godoy, P. M. et al. Large differences in small RNA composition between human biofluids. Cell Rep. 25, 1346–1358 (2018).
Srinivasan, S. et al. Small RNA sequencing across diverse biofluids identifies optimal methods for exRNA isolation. Cell 177, 446–462 (2019).
Brown, G. J. et al. TLR7 gain-of-function genetic variation causes human lupus. Nature 605, 349–356 (2022).
Satterthwaite, A. B. TLR7 signaling in lupus B cells: new insights into synergizing factors and downstream signals. Curr. Rheumatol. Rep. 23, 80 (2021).
Dorrity, T. J. et al. Long 3’UTRs predispose neurons to inflammation by promoting immunostimulatory double-stranded RNA formation. Sci. Immunol. 8, eadg2979 (2023).
Maharana, S. et al. SAMHD1 controls innate immunity by regulating condensation of immunogenic self RNA. Mol. Cell 82, 3712–3728 (2022).
Davis, P., Cunnington, P. & Hughes, G. Double-stranded RNA antibodies in systemic lupus erythematosus. Ann. Rheum. Disease. 34, 239 (1975).
Boada-Romero, E., Martinez, J., Heckmann, B. L. & Green, D. R. The clearance of dead cells by efferocytosis. Nat. Rev. Mol. Cell Biol. 21, 398–414 (2020).
Lee, S. J. et al. Transactivation of bad by vorinostat-induced acetylated p53 enhances doxorubicin-induced cytotoxicity in cervical cancer cells. Exp. Mol. Med. 46, e76 (2014).
Elliott, M. R. & Ravichandran, K. S. The dynamics of apoptotic cell clearance. Dev. Cell 38, 147–160 (2016).
Piva, T. J., Davern, C. M., Hall, P. M., Winterford, C. M. & Ellem, K. A. O. Increased activity of cell surface peptidases in HeLa cells undergoing UV-induced apoptosis is not mediated by caspase 3. Int. J. Mol. Sci. 13, 2650–2675 (2012).
Takakura, M., Ishiguro, K., Akichika, S., Miyauchi, K. & Suzuki, T. Biogenesis and functions of aminocarboxypropyluridine in tRNA. Nat. Commun. 10, 5542 (2019).
Zhang, N. et al. Cell surface RNAs control neutrophil recruitment. Cell 187, 846–860 (2024).
Liu, B. et al. Innate immune memory and homeostasis may be conferred through crosstalk between the TLR3 and TLR7 pathways. Sci. Signal. 9, ra70 (2016).
Sakaniwa, K. et al. TLR3 forms a laterally aligned multimeric complex along double-stranded RNA for efficient signal transduction. Nat. Commun. 14, 164 (2023).
Leonard, J. N. et al. The TLR3 signaling complex forms by cooperative receptor dimerization. Proc. Natl Acad. Sci. USA 105, 258–263 (2008).
Barrat, F. J., Elkon, K. B. & Fitzgerald, K. A. Importance of nucleic acid recognition in inflammation and autoimmunity. Annu. Rev. Med. 67, 323–336 (2016).
Koffler, D., Agnello, V. & Kimkel, H. G. Polynucleotide immune complexes in serum and glomeruli of patients with systemic lupus erythematosus. Am. J. Pathol. 74, 109–124 (1974).
Jenks, S. A. et al. B cell subset composition segments clinically and serologically distinct groups in chronic cutaneous lupus erythematosus. Ann. Rheum. Dis. 80, 1190–1200 (2021).
Franceschini, F. & Cavazzana, I. Anti-Ro/SSA and La/SSB antibodies. Autoimmunity 38, 55–63 (2005).
Migliorini, P., Baldini, C., Rocchi, V. & Bombardieri, S. Anti-Sm and anti-RNP antibodies. Autoimmunity 38, 47–54 (2005).
Ah Kioon, M. D. et al. Modulation of plasmacytoid dendritic cells response in inflammation and autoimmunity. Immunol. Rev. 323, 241–256 (2024)
Barrat, F. J. & Su, L. A pathogenic role of plasmacytoid dendritic cells in autoimmunity and chronic viral infection. J. Exp. Med. 216, 1974–1985 (2019).
Reizis, B. Plasmacytoid dendritic cells: development, regulation, and function. Immunity 50, 37–50 (2019).
Kuriakose, J. et al. Patrolling monocytes promote the pathogenesis of early lupus-like glomerulonephritis. J. Clin. Invest. 129, 2251–2265 (2019).
Du, Y. et al. Chemokines form nanoparticles with DNA and can superinduce TLR-driven immune inflammation. J. Exp. Med. 219, e20212142 (2022).
Roberts, Z. J. et al. The chemotherapeutic agent DMXAA potently and specifically activates the TBK1-IRF-3 signaling axis. J. Exp. Med. 204, 1559–1569 (2007).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019)
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Frazee, A. C. et al. Ballgown bridges the gap between transcriptome assembly and expression analysis. Nat. Biotechnol. 33, 243–246 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Krämer, A., Green, J., Pollard, J. & Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 30, 523–530 (2014).
Acknowledgements
We thank the members of the Rathinam and Flynn laboratories for their constructive feedback. We also thank C. Bertozzi and N. Till for the synthesis of stereoisomerically pure acp3U, P. Sharp for helpful discussions, V. Hornung and M. Berouti for technical help and P. Hristov for chemical drawings. This work was supported by grants from Burroughs Wellcome Fund Career Award for Medical Scientists (R.A.F.), The Rita Allen Foundation (R.A.F.), The Sontag Foundation (R.A.F.), The Scleroderma Research Foundation (R.A.F. and F.J.B.) and National Institute of General Medical Sciences, National Institute of Diabetes and Digestive and Kidney Diseases, and the National Institute of Allergy and Infectious Diseases of the National Institutes of Health under grant award numbers F31AI178967 (V.R.G.), R35GM151157 (R.A.F.), R01AI119015 (V.A.R.), R01AI148491 (V.A.R.), R01AI132850 (S.K.V.), R01DK121805 (B.Z.) and R01HL172239 (B.Z.), the Deutsche Forschungsgemeinschaft (DFG) CA275/11-3 (ID 326039064), CRC1309 (ID 325871075, A4), CRC1361 (ID 393547839, P2) and TRR237 (ID 369799452, A27) and the European Research Council (ERC) under grant no. 741912 (EPiR) (T.C.).
Author information
Authors and Affiliations
Contributions
V.R.G., R.A.F. and V.A.R. conceived the project, designed experimental strategies, analysed the data, generated the figures and wrote the manuscript. V.R.G. and J.P. performed experiments. J.P. and C.G.L. performed rPAL analysis. P.C. generated DTWD2 KO cells. I.M. and T.C. designed and synthesized mononucleosides as well as unmodified and modified synthetic RNA. J.V.P. and M.R.W. collected and processed the human SLE samples. M.D.A.K. and F.J.B. isolated and performed human pDC and primary macrophage assays. A.J.M. and B.Z. performed RNA-seq and related analyses. A.G.H., M.S.S., S.S.W., J.P., P.W., S.K.V., R.R.-A. and X.W. aided in experimental design and helped to interpret results. V.R.G., P.W., S.K.V., B.Z., F.J.B., T.C., R.A.F. and V.A.R. acquired the funding for this work. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.A.F. is a stockholder of ORNA Therapeutics. R.A.F. is a board of directors’ member and stockholder of Chronus Health and Blue Planet Systems. V.R.G., R.A.F. and V.A.R. have filed a patent on the sequences and use of acp3U in various functional immune applications. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Bryan Dickinson, Nan Yan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 N-glycans shield small RNAs from innate immune recognition.
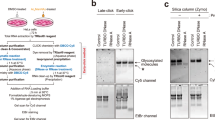
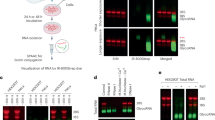
(A) Experimental design to test if the enzymatic removal of N-glycans from glycoRNAs induces a type I IFN response. (B) IFN-β ELISA of RAW macrophage supernatants 6 hours post-stimulation with 500 ng, 1 µg, or 2 µg of small RNAs untreated or treated with PNGase F, RNase cocktail, or PNGase F+RNase cocktail. (C) Immunoblotting for IRAK2, TIRAP and MyD88 in MyD88 immunoprecipitates and IRAK2, TIRAP, MyD88, and β-Actin in input lysates from macrophages stimulated as indicated. (D) TNF and IL-6 ELISA of supernatants from primary human macrophages stimulated with small RNAs treated with or without PNGase F (one dot = triplicate averages from one human donor). (E) IFN-α, TNF, IL-6 ELISA of supernatants from primary human plasmacytoid dendritic cells stimulated with small RNAs treated with or without PNGase F (one dot = triplicate averages from one human donor). (F) IFN-β ELISA of RAW macrophage supernatants 6 hours post treatment with 2 µg, 6 µg, or 10 µg of PNGase F alone or in combination with RNase. Pooled data from 3 (F) or 4 (B, D, E) independent experiments are shown as mean ± SEM (B, D–F) or one representative blot of two independent replicates (C). *, p < 0.05; ***, p < 0.001; ****, p < 0.0001; ns, not significant. Two-way ANOVA with Tukey’s test (B, F), One-way ANOVA with Šídák’s test (D, E). Schematic in a was created with BioRender (https://biorender.com).
Extended Data Fig. 2 GlycoRNAs are present in the circulation.
(A) Quantification of rPAL signal from glycoRNAs enriched from human and mouse sera RNA. (B) One dot equals one donor/mouse. rPAL signal intensity of individual RNA samples (quantified in A). (C) Blotting of glycoRNAs from HeLa cells and human sera samples normalized at 1 µg of RNA per well. (D) Quantification of glycoRNA signal intensity observed in (C). (E) rPAL signal intensity of normalized human and HeLa cell-derived RNA samples (quantified in D). (F) Blotting of glycoRNAs from healthy human donors or SLE patient sera samples normalized at 1 µg of RNA per well. Data are represented as mean ± SEM (A, C, and D). Pooled data from four (A–E) or 6 (F) independent sera samples are shown. ***, p < 0.001; ns, not significant. Unpaired two-tailed t-test (D, F).
Extended Data Fig. 3 De-N-glycosylation of cell surface RNAs makes apoptotic cells immunostimulatory.
(A) Efferocytosis of CellBright Red-labeled live or apoptotic HeLa cells treated with or without PNGase F, or PNGase F+RNase cocktail by CD11b+ peritoneal macrophages 2 hours post-feeding as assessed via flow cytometry. (B) IFN-β ELISA of RAW macrophage supernatants 24 hours post-stimulation with apoptotic HeLa cells treated with or without PNGase F or PNGase F+RNase cocktail at indicated macrophage:HeLa cell ratios. (C) IL-6 ELISA of peritoneal macrophage supernatants 24 hours post-stimulation with live or apoptotic HeLa cells treated with or without PNGase F or PNGase F+RNase cocktail at indicated macrophage:HeLa cell ratios. (D) TNF ELISA of peritoneal macrophage supernatants 24 hours post-stimulation with live or apoptotic HeLa cells treated with or without PNGase F or PNGase F+RNase cocktail at indicated macrophage:HeLa cell ratios. Data are represented as mean ± SEM (A–D). Pooled data from 3 independent experiments (A–D) are shown. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; ns, not significant. Two-way ANOVA with Tukey’s test (A–D).
Extended Data Fig. 4 The modified RNA base acp3U stimulates innate immune responses.
(A) IFN-β ELISA of peritoneal macrophage supernatants 6 hours post-transfection with small RNA derived from WT and two separate single cell clones of DTWD2 KO U2OS cells treated with or without PNGase F, RNase cocktail, or PNGase F+RNase cocktail. (B) IFN-β ELISA of peritoneal macrophage supernatants 24 hours post-stimulation with apoptotic WT, DTWD2 KO1 and DTWD2 KO2 U2OS cells treated with or without PNGase F or PNGase F with RNase cocktail at 1:2 macrophage:U2OS cell ratio. (C) TNF and IL-6 ELISAs of supernatants of THP1 macrophages 6 hours post-transfection of small RNAs harvested from DTWD2 KO cells treated with or without PNGase F or in combination with RNase. (D) TNF and IL-6 ELISAs of supernatants of THP1 macrophages 6 hours post-stimulation with apoptotic DTWD2 KO cells treated with or without PNGase F or in combination with RNase. (E) IFN-β ELISA of RAW macrophage (left panel) and peritoneal macrophage (right panel) supernatants 6 hours post-transfection with small RNA treated with or without Endo-F2/F3 or Endo-F2/F3+RNase cocktail (This experiment was performed at the same time as Fig. 1F). (F) IFN-β ELISA of peritoneal macrophage supernatants 24 hours post-stimulation with apoptotic HeLa cells treated with or without PNGase F or Endo-F2/F3 at 1:4 macrophage:HeLa cell ratio. Data are represented as mean ± SEM (A–F). Pooled data from 3 independent experiments (A–F) are shown. **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; ns, not significant. Two-way ANOVA with Tukey’s test (A–E) or One-way ANOVA with Šídák’s test (F).
Extended Data Fig. 5 The modified RNA base acp3U stimulates innate immune responses.
(A and B) TNF ELISA of THP-1 macrophage supernatants 6 hours post-stimulation (A) or post-transfection (B) with 10 nm, 100 nm, or 250 nm of synthetic unmodified RNA or synthetic RNA modified with acp3U (acp3U-8C and acp3U-8D). (C and D) Volcano plot of differentially expressed genes in THP1 macrophages stimulated with acp3U-8C-RNA (C) or PNGase F-treated small RNAs (D) compared to THP1 macrophages stimulated with unmodified RNA (C) or untreated small RNA (D), respectively, as revealed by RNA-seq. Each dot represents a gene (Total of 32,389 genes tested). A cutoff of absolute value fold-change of 0.5 was used to select significantly differentially expressed genes for gene ontology, yielding 5107 genes in PNGase F-treated small RNAs vs. untreated small RNAs and 6472 genes in acp3U-8C RNA vs. unmodified RNA. (E) Heat map of the genes represented in the indicated immune pathways in THP1 macrophages stimulated as in C and D. Rows are samples grouped by condition; Columns are genes grouped by pathways; Color is the scaled expression of genes in each sample. Data are represented as mean ± SEM (A and B). Pooled data from 3 (A and B) or 2 (C–E) independent experiments are shown. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; ns, not significant. Two-way ANOVA with Tukey’s test (A and B).
Extended Data Fig. 6 RNA with the modified base acp3U stimulates innate immune responses.
(A and B) IFN-β ELISA of peritoneal macrophage supernatants 6 hours post-stimulation (A) or post-transfection (B) with 50 µM, 125 µM, 250 µM of uridine, acp3U, or uridine+acp3U (C and D) IL-6 ELISA of THP-1 macrophage supernatant 6 hours post-stimulation (C) or post-transfection (D) with 50 µM, 125 µM, 250 µM of uridine, acp3U, or uridine+acp3U. (E and F) TNF ELISA of THP-1 macrophage supernatant 6 hours post-stimulation (E) or post-transfection (F) with 50 µM, 125 µM, 250 µM of uridine, acp3U, or uridine+acp3U. (G) IL-6 and TNF ELISA of THP-1 macrophage supernatant 6 hours post-addition or post-transfection with 50 µM of uridine, acp3U, acp3U-modified with an amide (acp3U amide), acp3U+uridine or acp3U amide+uridine. (H) IL-6 ELISA of THP-1 macrophage supernatant 6 hours post-addition or post-transfection with 50 µM of uridine, acp3U, acp3U-modified with a GlcNAc (acp3U-GlcNAc), acp3U+uridine or acp3U GlcNAc+uridine. Data are represented as mean ± SEM (A–H). Pooled data from 3 independent experiments (A–H) are shown. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; ns, not significant. Two-way ANOVA with Tukey’s test (A–H).
Extended Data Fig. 7 TLR3 and TLR7 sense de-N-glycosylated glycoRNAs.
(A) IFN-β ELISA of supernatants from WT, TLR3 KO, and TLR7 KO peritoneal macrophages stimulated for 24 hours with small RNAs from HeLa cells that were treated with or without PNGase F or in combination with RNase and then column purified to remove the associated N-glycans and enzymes. Poly(I:C), CL097, R837 and LPS were used as controls. (B) TNF and IL-6 ELISA of supernatants from THP1 macrophages stimulated with small RNAs treated with or without PNGase F, or in combination with RNase, unmodified or acp3U-containing synthetic RNAs, as well as poly(I:C), CL097, ssRNA40, 2’,3’-cGMP, R837, and TL8. (C) IFN-β ELISA of supernatants of RAW macrophages 24 hours post-electroporation with free uridine, acp3U, uridine+acp3U in combination or the synthetic 18-nucleotide unmodified or acp3U containing RNAs and poly(dA:dT). Pooled data from 3 independent experiments (A–C) are shown. **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; ns, not significant. Two-way ANOVA with Tukey’s test (A) or One-way ANOVA with Šídák’s test (B–C).
Supplementary information
Supplementary Information (Fig. 1) (download PDF )
Supplementary Information related to Fig. 1.
Supplementary Information (Fig. 2) (download PDF )
Supplementary Information related to Fig. 2.
Supplementary Information (Extended Data Fig. 1) (download PDF )
Supplementary Information related to Extended Data Fig. 1.
Supplementary Information (Extended Data Fig. 2) (download PDF )
Supplementary Information related to Extended Data Fig. 2.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Graziano, V.R., Porat, J., Ah Kioon, M.D. et al. RNA N-glycosylation enables immune evasion and homeostatic efferocytosis. Nature 645, 784–792 (2025). https://doi.org/10.1038/s41586-025-09310-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09310-6
This article is cited by
-
Long noncoding RNA regulation of immunity
Nature Immunology (2026)
-
GlycoRNA complexed with heparan sulfate regulates VEGF-A signalling
Nature (2026)
-
Lectin-based detection and expression profiling of native glycoRNAs
Scientific Reports (2026)
-
The IRF transcription factor family in type I interferon-mediated antiviral immunity and associated diseases
Immunity & Inflammation (2026)