Abstract
Infection by yellow fever virus (YFV), the prototype Orthoflavivirus, induces a febrile syndrome in humans that can progress to liver failure, haemorrhage and death1. Despite decades of study, the entry receptors for YFV remain unclear. Here, using a surface protein-targeted CRISPR–Cas9 screen, we identified LRP4, a low-density lipoprotein receptor (LDLR) family member, as a candidate entry receptor for YFV. Genetic ablation of LRP4 impaired YFV infection of cells and, reciprocally, complementation or ectopic expression of LRP4 increased infection. Related viruses in the YFV antigenic complex also showed LRP4-dependent infection. LRP4 promoted YFV entry into cells through LDLR type A (LA) domain binding to domain III of the YFV envelope protein. Soluble LRP4–Fc decoy receptors neutralized YFV infection in cell culture and reduced viral burden in vivo. As we observed residual YFV infection in LRP4-deficient cells, we evaluated whether other LDLR family members promote YFV entry. This approach identified LRP1 and VLDLR as additional receptors for YFV infection in cell culture. LRP1–Fc, LRP4–Fc and VLDLR–Fc decoys protected mice from YFV challenge, and LRP1–Fc decoys inhibited YFV infection and liver pathogenesis in mice engrafted with human hepatocytes. A genetic deficiency of LRP1 in primary human hepatocyte cultures also resulted in reduced YFV infection. Our findings establish a role for multiple LDLR family members in YFV entry, infection and pathogenesis, which has implications for receptor use and countermeasure development for multiple emerging orthoflaviviruses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data supporting the findings of this study are available within the Article and its Supplementary Information, or in files uploaded to the Washington University School of Medicine’s Bernard Becker Medical Library (https://doi.org/10.17632/xnpzsx3729). Reagents will be made available on request after completion of a materials transfer agreement. Raw CRISPR screen sequencing data have been uploaded to NCBI (BioSample: SAMN50433716. Accession: SRX29989461 and SRX29989462). Source data are provided with this paper.
Code availability
A Python script for analysis of SPR data has been uploaded to GitHub (https://github.com/kaszubat/YFV-LRP4-SPR-Analysis).
References
Pierson, T. C. & Diamond, M. S. The continued threat of emerging flaviviruses. Nat. Microbiol. 5, 796–812 (2020).
Fishburn, A. T., Pham, O. H., Kenaston, M. W., Beesabathuni, N. S. & Shah, P. S. Let’s get physical: flavivirus-host protein-protein interactions in replication and pathogenesis. Front. Microbiol. 13, 847588 (2022).
Anwar, M. N. et al. The interactions of flaviviruses with cellular receptors: Implications for virus entry. Virology 568, 77–85 (2022).
Pan, Y. et al. Flaviviruses: innate immunity, inflammasome activation, inflammatory cell death, and cytokines. Front. Immunol. 13, 829433 (2022).
Lopes, R. L. et al. Kidney involvement in yellow fever: a review. Rev. Inst. Med. Trop. Sao Paulo 61, e35 (2019).
Monath, T. P. & Vasconcelos, P. F. Yellow fever. J. Clin. Virol. 64, 160–173 (2015).
Collins, N. D. & Barrett, A. D. Live attenuated yellow fever 17D vaccine: a legacy vaccine still controlling outbreaks in modern day. Curr. Infect. Dis. Rep. 19, 14 (2017).
Gianchecchi, E., Cianchi, V., Torelli, A. & Montomoli, E. Yellow fever-origin, epidemiology, preventive strategies and future prospects. Vaccines 10, 372 (2022).
Zimmerman, O., Holmes, A. C., Kafai, N. M., Adams, L. J. & Diamond, M. S. Entry receptors—the gateway to alphavirus infection. J. Clin. Invest. 133, e165307 (2023).
Hastings, A. K. et al. TAM receptors are not required for Zika virus infection in mice. Cell Rep. 19, 558–568 (2017).
Laureti, M., Narayanan, D., Rodriguez-Andres, J., Fazakerley, J. K. & Kedzierski, L. Flavivirus receptors: diversity, identity, and cell entry. Front. Immunol. 9, 2180 (2018).
Alen, M. M., Dallmeier, K., Balzarini, J., Neyts, J. & Schols, D. Crucial role of the N-glycans on the viral E-envelope glycoprotein in DC-SIGN-mediated dengue virus infection. Antiviral Res. 96, 280–287 (2012).
Perera-Lecoin, M., Meertens, L., Carnec, X. & Amara, A. Flavivirus entry receptors: an update. Viruses 6, 69–88 (2013).
Carnec, X. et al. The phosphatidylserine and phosphatidylethanolamine receptor CD300a binds Dengue virus and enhances infection. J. Virol. 90, 92–102 (2016).
Cordero-Rivera, C. D. et al. The importance of viral and cellular factors on flavivirus entry. Curr. Opin. Virol. 49, 164–175 (2021).
Lee, E. & Lobigs, M. E protein domain III determinants of yellow fever virus 17D vaccine strain enhance binding to glycosaminoglycans, impede virus spread, and attenuate virulence. J. Virol. 82, 6024–6033 (2008).
Nahain, A. A. et al. Antiviral activities of heparan sulfate mimetic RAFT polymers against mosquito-borne viruses. ACS Appl. Bio Mater. 7, 2862–2871 (2024).
Pönighaus, C. et al. Human xylosyltransferase II is involved in the biosynthesis of the uniform tetrasaccharide linkage region in chondroitin sulfate and heparan sulfate proteoglycans. J. Biol. Chem. 282, 5201–5206 (2007).
DePew, A. T. & Mosca, T. J. Conservation and innovation: versatile roles for LRP4 in nervous system development. J. Dev. Biol. 9, 9 (2021).
Hoffmann, H. H. et al. TMEM41B is a pan-flavivirus host factor. Cell 184, 133–148 (2021).
Lin, D. L. et al. The ER membrane protein complex promotes biogenesis of Dengue and Zika virus non-structural multi-pass transmembrane proteins to support infection. Cell Rep. 27, 1666–1674 (2019).
Marceau, C. D. et al. Genetic dissection of Flaviviridae host factors through genome-scale CRISPR screens. Nature 535, 159–163 (2016).
Ma, H. et al. A CRISPR-based screen identifies genes essential for West-Nile-virus-induced cell death. Cell Rep. 12, 673–683 (2015).
Khandia, R. et al. Modulation of Dengue/Zika virus pathogenicity by antibody-dependent enhancement and strategies to protect against enhancement in Zika virus infection. Front. Immunol. 9, 597 (2018).
Santos-Peral, A. et al. Prior flavivirus immunity skews the yellow fever vaccine response to cross-reactive antibodies with potential to enhance dengue virus infection. Nat. Commun. 15, 1696 (2024).
Qiu, X. & Bailey, A. L. Two mutations in NS2B are responsible for attenuation of the yellow fever virus (YFV) vaccine strain 17D. PLoS Pathog. 21, e1013373 (2025).
Zhang, J. et al. Amino acid changes in two viral proteins drive attenuation of the yellow fever 17D vaccine. Nat. Microbiol. 10, 1902–1917 (2025).
Abdullahi, I. N. et al. The interplay between environmental factors, vector competence and vaccine immunodynamics as possible explanation of the 2019 yellow fever re-emergence in Nigeria. N. Microbes N. Infect. 41, 100858 (2021).
Hobson-Peters, J. et al. A recombinant platform for flavivirus vaccines and diagnostics using chimeras of a new insect-specific virus. Sci. Transl. Med. 11, eaax7888 (2019).
Hardy, J. M. et al. A unified route for flavivirus structures uncovers essential pocket factors conserved across pathogenic viruses. Nat. Commun. 12, 3266 (2021).
Rey, F. A., Heinz, F. X., Mandl, C., Kunz, C. & Harrison, S. C. The envelope glycoprotein from tick-borne encephalitis virus at 2 A resolution. Nature 375, 291–298 (1995).
Volk, D. E. et al. Solution structure and antibody binding studies of the envelope protein domain III from the New York strain of West Nile virus. J. Biol. Chem. 279, 38755–38761 (2004).
Cao, D., Ma, B., Cao, Z., Zhang, X. & Xiang, Y. Structure of Semliki Forest virus in complex with its receptor VLDLR. Cell 186, 2208–2218 (2023).
Adams, L. J. et al. Structural and functional basis of VLDLR usage by Eastern equine encephalitis virus. Cell 187, 360–374 (2024).
Davis, C. W. et al. West Nile virus discriminates between DC-SIGN and DC-SIGNR for cellular attachment and infection. J. Virol. 80, 1290–1301 (2006).
Monath, T. P. & Barrett, A. D. Pathogenesis and pathophysiology of yellow fever. Adv. Virus Res. 60, 343–395 (2003).
Bailey, A. L. et al. Consumptive coagulopathy of severe yellow fever occurs independently of hepatocellular tropism and massive hepatic injury. Proc. Natl Acad. Sci. USA 117, 32648–32656 (2020).
Meier, K. C., Gardner, C. L., Khoretonenko, M. V., Klimstra, W. B. & Ryman, K. D. A mouse model for studying viscerotropic disease caused by yellow fever virus infection. PLoS Pathog. 5, e1000614 (2009).
Kafai, N. M. et al. Entry receptor LDLRAD3 is required for Venezuelan equine encephalitis virus peripheral infection and neurotropism leading to pathogenesis in mice. Cell Rep. 42, 112946 (2023).
Ma, H. et al. LDLRAD3 is a receptor for Venezuelan equine encephalitis virus. Nature 588, 308–314 (2020).
Johnson, E. B., Hammer, R. E. & Herz, J. Abnormal development of the apical ectodermal ridge and polysyndactyly in Megf7-deficient mice. Hum. Mol. Genet. 14, 3523–3538 (2005).
Weatherbee, S. D., Anderson, K. V. & Niswander, L. A. LDL-receptor-related protein 4 is crucial for formation of the neuromuscular junction. Development 133, 4993–5000 (2006).
Bhaskar, M., Satheesan, A. & Basu, A. Low-density lipoprotein receptor is an important host factor in flaviviral entry and replication in neurons. Biochem. Biophys. Res. Commun. 743, 151160 (2024).
Huerta, V. et al. The low-density lipoprotein receptor-related protein-1 is essential for dengue virus infection. Viruses 16, 1692 (2024).
Ganaie, S. S. et al. Lrp1 is a host entry factor for Rift Valley fever virus. Cell 184, 5163–5178 (2021).
Willnow, T. E., Armstrong, S. A., Hammer, R. E. & Herz, J. Functional expression of low density lipoprotein receptor-related protein is controlled by receptor-associated protein in vivo. Proc. Natl Acad. Sci. USA 92, 4537–4541 (1995).
Schwarz, M. M. et al. Oropouche orthobunyavirus infection is mediated by the cellular host factor Lrp1. Proc. Natl Acad. Sci. USA 119, e2204706119 (2022).
Clark, L. E. et al. VLDLR and ApoER2 are receptors for multiple alphaviruses. Nature 602, 475–480 (2022).
Li, W. et al. Shifts in receptors during submergence of an encephalitic arbovirus. Nature 632, 614–621 (2024).
Zhai, X. et al. LDLR is used as a cell entry receptor by multiple alphaviruses. Nat. Commun. 15, 622 (2024).
Palakurty, S. et al. The VLDLR entry receptor is required for the pathogenesis of multiple encephalitic alphaviruses. Cell Rep. 43, 114809 (2024).
Erickson, A. K. & Pfeiffer, J. K. Spectrum of disease outcomes in mice infected with YFV-17D. J. Gen. Virol. 96, 1328–1339 (2015).
Ma, B., Huang, C., Ma, J., Xiang, Y. & Zhang, X. Structure of Venezuelan equine encephalitis virus with its receptor LDLRAD3. Nature 598, 677–681 (2021).
Shen, C., Xiong, W. C. & Mei, L. LRP4 in neuromuscular junction and bone development and diseases. Bone 80, 101–108 (2015).
Davis, C. W. et al. The location of asparagine-linked glycans on West Nile virions controls their interactions with CD209 (dendritic cell-specific ICAM-3 grabbing nonintegrin). J. Biol. Chem. 281, 37183–37194 (2006).
Muller, U. et al. Functional role of type I and type II interferons in antiviral defense. Science264, 1918–1921 (1994).
White, J. P. et al. Intestinal dysmotility syndromes following systemic infection by flaviviruses. Cell 175, 1198–1212 (2018).
Chen, R. E. et al. Implications of a highly divergent dengue virus strain for cross-neutralization, protection, and vaccine immunity. Cell Host Microbe 29, 1634–1648 (2021).
VanBlargan, L. A. et al. Broadly neutralizing monoclonal antibodies protect against multiple tick-borne flaviviruses. J. Exp. Med. 218, e20210174 (2021).
McArthur, M. A., Suderman, M. T., Mutebi, J. P., Xiao, S. Y. & Barrett, A. D. Molecular characterization of a hamster viscerotropic strain of yellow fever virus. J. Virol. 77, 1462–1468 (2003).
Goo, L., VanBlargan, L. A., Dowd, K. A., Diamond, M. S. & Pierson, T. C. A single mutation in the envelope protein modulates flavivirus antigenicity, stability, and pathogenesis. PLoS Pathog. 13, e1006178 (2017).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Yu, G. C., Smith, D. K., Zhu, H. C., Guan, Y. & Lam, T. T. Y. GGTREE: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Bodenhofer, U., Bonatesta, E., Horejs-Kainrath, C. & Hochreiter, S. msa: an R package for multiple sequence alignment. Bioinformatics 31, 3997–3999 (2015).
Sanson, K. R. et al. Optimized libraries for CRISPR-Cas9 genetic screens with multiple modalities. Nat. Commun. 9, 5416 (2018).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Miyoshi, H. & Stappenbeck, T. S. In vitro expansion and genetic modification of gastrointestinal stem cells in spheroid culture. Nat. Protoc. 8, 2471–2482 (2013).
Huch, M. et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160, 299–312 (2015).
Huch, M. et al. In vitro expansion of single Lgr5+ liver stem cells induced by Wnt-driven regeneration. Nature 494, 247–250 (2013).
Doyle, M. P. et al. Isolation of a potently neutralizing and protective human monoclonal antibody targeting yellow fever virus. mBio 13, e0051222 (2022).
Kim, A. S. et al. Pan-protective anti-alphavirus human antibodies target a conserved E1 protein epitope. Cell 184, 4414–4429 (2021).
Nelson, C. A., Lee, C. A. & Fremont, D. H. Oxidative refolding from inclusion bodies. Methods Mol. Biol. 1140, 145–157 (2014).
Oliphant, T. et al. Antibody recognition and neutralization determinants on domains I and II of West Nile Virus envelope protein. J. Virol. 80, 12149–12159 (2006).
BIAevaluation Version 3.0 Software Handbook (Biacore AB, 1997).
Acknowledgements
This research was funded by NIH grants (U01 AI073755 and U19 AI181960 to M.S.D., J.E.C. and D.H.F.) and contracts (75N93019C00062 and 75N93022C00035 to D.H.F.), as well as the NIH Director’s Early Independence Award (DP5 OD029608) to A.L.B. We thank N. Perlmutter for help in generating LRP4 orthologues and T. Pierson for providing the plasmids for production of YFV RVPs. Some graphics in the figures were made using BioRender, using an institutional license to Washington University in St. Louis.
Author information
Authors and Affiliations
Contributions
Z.C. and M.S.D. designed the study. Z.C. performed the CRISPR screen and target gene validation. Z.C., S.P. and P.L. performed cell culture studies. S.H., C.A.N., D.A.P. and Z.C. generated recombinant proteins. Z.C., S.R., D.W.L., P.W.R., S.P.J.W., I.B.-P. and G.K.A. generated key plasmids and proteins. S.H., Z.C., P.D.H., T.K. and M.N.N. performed protein binding studies. J.E.C. provided purified recombinant antibodies. Z.C., A.S., X.Q. and A.L.B. performed in vivo studies. D.H.F. supervised the protein biophysical studies. Z.C. and M.S.D. wrote the initial draft with all of the other authors providing comments.
Corresponding author
Ethics declarations
Competing interests
M.S.D. is a consultant or on a scientific advisory board for Inbios, IntegerBio, Akagera Medicines, GlaxoSmithKline, Merck and Moderna. The Diamond laboratory has received unrelated funding support in sponsored research agreements from Vir Biotechnology, Bavarian Nordic and Moderna. J.E.C. is a former member of the scientific advisory boards of Gigagen (Grifols) and BTG International, has consulted for Moderna, is founder of IDBiologics and receives royalties from UpToDate. The Crowe laboratory received unrelated sponsored research agreements from IDBiologics during the conduct of the study.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
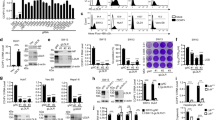
Extended Data Fig. 1 CRISPR–Cas9 screen identifies LRP4 as a factor required for efficient YFV infection.
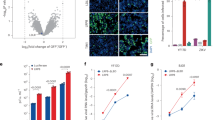
a, HAP1 ΔHS cells were transduced with the cell surface library containing 4,630 sgRNAs, selected with puromycin, and inoculated with YFV-17D at an MOI of 10. Surviving cells were expanded, and 3 weeks later, genomic DNA from cells was harvested and sequenced for sgRNA abundance. b, Next-generation sequencing confirmation of LRP4 gene editing in HAP1 ΔHS cells. Allele frequency is indicated next to the sequence. c, HAP1 ΔHS (control), HAP1 ΔHS ΔLRP4 (ΔLRP4) and LRP4-complemented HAP1 ΔHS ΔLRP4 (ΔLRP4 + LRP4) cells were analysed for surface expression of LRP4 by flow cytometry using an anti-Flag mAb. d, Next-generation sequencing confirmation of LRP4 gene editing in 293 T cells. Allele frequency is indicated next to the sequence. e, 293 T (control), ΔLRP4 and LRP4-complemented ΔLRP4 293 T (ΔLRP4 + LRP4) cells were inoculated with YFV-17D (MOI of 20) for 16 h, and infection was measured by flow cytometry (anti-E mAbs). f, Flag-LRP4 expression levels on the surface of control, ΔLRP4, and LRP4-complemented ΔLRP4 293 T (ΔLRP4 + LRP4) cells were analysed by flow cytometry (anti-Flag mAb). g-h, Control and Flag-LRP4 expressing K562 cells were analysed for surface expression of LRP4 by flow cytometry (g, anti-LRP4 mAb; h, anti-Flag mAb). i, ENTV (MOI of 5) was incubated with 200 ng/mL of E60 or isotype control mAb at for 1 h at 37 °C and then used to infect K562 cells. Infection levels were measured by flow cytometry at 24 h (anti-E mAbs). Data are mean ± s.d. (e and i) of 3 experiments, each performed in triplicate. Statistical analysis: one-way ANOVA with Dunnett’s post-test (e) or two-tailed unpaired t test (i). The diagram in a was created in BioRender. Diamond, M. (2025) https://BioRender.com/zw9zpr4.
Extended Data Fig. 2 Cell surface expression of LRP4 variants and orthologs in K562 cells.
a, K562 cells ectopically expressing indicated LRP4 variants were measured by flow cytometry (anti-Flag mAb). b, Sequence alignment of LRP4 LBD from indicated species. c, K562 cells ectopically expressing LRP4 orthologs were measured by flow cytometry (anti-Flag mAb). d, MFI of Flag-LRP4 expression levels in (c).
Extended Data Fig. 3 Binding of LRP4 to YFV.
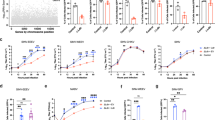
a, SDS-PAGE gel with Coomassie staining of purified human LRP4-LBD-Fc and LDLRAD3-LA1-Fc proteins. b-c, SDS-PAGE gel with Coomassie staining of YFV-M185D/BinJ (fraction in lane 2 was used for BLI experiments) (b), YFV-ES504/BinJ (fraction in lane 3 was used for BLI experiments) and YFV-Asibi/BinJ (c) (fraction in lane 7 was used for BLI experiments). d, Micrographs of YFV-Asibi/BinJ and YFV-M185D/BinJ chimeric virions show mature and immature particles of approximately 50 nm in diameter. Scale bar, 100 nM. e-f, Inhibition of YFV-17D infection with indicated concentrations of LRP4-LBD-Fc fusion protein in 293 T (e) or Vero (f) cells was measured by flow cytometry (anti-E mAbs). g-h, Inhibition of YFV-17D infection with indicated concentrations of LRP4-Fc fusion proteins containing only one LA domain (g) or two LA domains (h) in HAP1 cells was measured by flow cytometry (anti-E mAbs). i-k, Infection of control K562 cells or those expressing indicated LRP4 variants with YFV-17D RVPs at 24 h was measured by flow cytometry. l, K562 cells ectopically expressing indicated LRP4 variants in (h-j) were measured by flow cytometry (anti-Flag mAb). Data (a-d) are representative of 2 experiments. Data (e-f and i-k) are mean ± s.d. from 3 experiments, each performed in duplicate (e-f) or triplicate (g-k). Data (g-h) are mean values from 3 experiments, each performed in triplicate. Source gel data images (a-c) are in Supplementary Fig. 1.
Extended Data Fig. 4 Contacts between YFV E-DIII and antibody.
a, Alignment of YFV-Asibi and YFV-17D E-DIII proteins with other YFV strains and YFV complex viruses. Red dots underneath indicate mutated residues in panels b-c. b, Binding of YFV-Asibi E-DIII mutant proteins to anti-YFV Adi-49147 mAb by BLI. Biosensors were coated with Adi-49147 mAb following incubation with YFV-Asibi E-DIII mutant proteins in solution. c, Infection of YFV-17D RVPs with indicated mutations in E-DIII mutant was measured by flow cytometry on Raji-DC-SIGNR cells (GFP expression). Data (b, c) are representative of 2 experiments. The diagram in b was created in BioRender. Diamond, M. (2025) https://BioRender.com/d5zz6nu.
Extended Data Fig. 5 LRP4 mRNA expression pattern in humans and mice.
a-b, Human LRP4 mRNA levels (a) or protein levels (b) are showed in indicated tissues. c, Mouse Lrp4 mRNA levels are showed from indicated tissues. Data were obtained from Human Protein Atlas database (https://www.proteinatlas.org/ENSG00000134569-LRP4/tissue) (a-b) or BioGPS database (http://biogps.org/#goto=genereport&id=228357) (c).
Extended Data Fig. 6 LRP4-Fc decoy molecules serum levels and activity.
a, AG129 mice were given 500 μg of LRP4-LBD-Fc, LRP4-LA (6-8)-Fc or LDLRAD3-LA1-Fc protein by i.p. at 4 h before infection. Mice were then inoculated with 105 PFU of YFV-17D by intraperitoneal injection. At 5 days post-infection, viral RNA levels in indicated tissues were measured by qRT-PCR (n = 7, 7, and 9, respectively). b, Indicated LRP4-Fc decoy neutralization activity in vitro and serum levels at day +1 after injection. c, Binding of YFV-M185D/BinJ to indicated LRP4 single LA domain Fc fusion proteins by BLI. Biosensors were coated with indicated Fc-fusion proteins following incubation with YFV-M185D/BinJ. d, Inhibition of YFV-17D infection with indicated concentrations of LRP4-LA (2-4)-3X-Fc fusion protein in HAP1 cells was measured by flow cytometry (anti-E mAbs). e, 500 μg of LRP4-LA (2-4)-Fc or LDLRAD3-LA1-Fc protein were incubated with 105 PFU of YFV-17D at 37 °C for 1 h and then inoculated into AG129 mice via intraperitoneal injection. Mice were also given 500 μg of LRP4-LA (2-4)-Fc or LDLRAD3-LA1-Fc protein at 1- and 2-days after infection (n = 12 and 8, respectively). At 3 d.p.i., viral RNA levels in indicated tissues were measured. f, Wild-type and Lrp4hypo/hypo mice were inoculated in the footpad with 102 PFU of VEEV ZPC-738. At 4 d.p.i., tissues were collected, and viral RNA levels tissues were measured (n = 7 and 11, respectively). Data in (a, e, and f) are pooled from 2 experiments. Bars indicate median values. Data in (c) are representative of 2 experiments. Data in (d) are from 3 experiments, each performed in triplicate. Statistical analysis: one-way ANOVA with Dunnett’s post-test (a, c); two-tailed Mann-Whitney test (e), ns, not significant. The diagram in c was created in BioRender. Diamond, M. (2025) https://BioRender.com/v82694b.
Extended Data Fig. 7 Ectopic expression of LDLR family members.
a, Diagram of LDLR family members. The indicated LBD domains were expressed ectopically using lentiviruses. b, Expression of indicated LDLR family members in K562 cells was measured by flow cytometry (anti-Flag mAb). The diagram in a was created in BioRender. Diamond, M. (2025) https://BioRender.com/xga4f13.
Extended Data Fig. 8 LRP1 and VLDLR mRNA expression pattern in humans and mice.
a-b, human LRP1 (a) or mouse Lrp1 (b) mRNA levels are showed in indicated tissues. Data were obtained from Human Protein Atlas database (https://www.proteinatlas.org/ENSG00000123384-LRP1/tissue) (a) or BioGPS database (http://biogps.org/#goto=genereport&id=16971) (b). c-d, human VLDLR (c) or mouse Vldlr (d) mRNA levels are showed in indicated tissues. Data were obtained from Human Protein Atlas database (https://www.proteinatlas.org/ENSG00000147852-VLDLR/tissue) (c) or BioGPS database (http://biogps.org/#goto=genereport&id=22359) (d).
Extended Data Fig. 9 LRP1 and VLDLR interact with YFV virions.
a, LRP1, LRP4 and VLDLR KO efficiency (as indicated by sequences of wild-type allele) in HepG2 cells was analysed by next-generation deep sequencing. b, Diagram of LRP1-CL-I-Fc, VLDLR-LBD-Fc and VLDLR-LA (1-2)-Fc proteins. c, SDS-PAGE gel with Coomassie staining of purified human LRP1-CL-I-Fc and VLDLR-Fc proteins. d, Binding of YFV virions to VLDLR-LBD-Fc fusion protein by BLI. Biosensors were coated with VLDLR-LBD-Fc fusion protein and incubated with YFV-M185D/BinJ virions in solution. e, Inhibition of YFV-17D infection with indicated concentrations of VLDLR-LBD-Fc fusion protein in HepG2 cells was measured by flow cytometry (anti-E mAbs). f, Infection of control K562 cells or those expressing indicated VLDLR variants with YFV-17D RVPs was measured by flow cytometry at 24 h. g, Expression of indicated VLDLR tandem LA domain repeats in (f) was measured by flow cytometry (anti-Flag mAb). h, 500 μg of LRP1-CL-I-Fc or LDLRAD3-LA1-Fc protein was incubated with 105 PFU of YFV-17D at 37 °C for 1 h and then inoculated into anti-IFNAR1 antibody treated C57BL/6J mice via intraperitoneal injection. Mice were also given 500 μg doses of LRP1-CL-I-Fc or LDLRAD3-LA1-Fc protein at 1 and 2 dpi (n = 10 and 7, respectively). At 3 d.p.i., viral RNA levels in indicated tissues were measured. i, 500 μg of LDLRAD3-LA1-Fc or VLDLR-LA (1-2)-Fc proteins were incubated with 105 PFU of YFV-M185D at 37 °C for 1 h and then inoculated into anti-IFNAR1 antibody treated C57BL/6J mice via intraperitoneal injection. Mice were also given 500 μg of the same Fc proteins at 1 and 2 dpi (n = 10 and 8, respectively). At 3 d.p.i., viral RNA levels in indicated tissues were measured. j, 500 μg of LDLRAD3-LA1-Fc or 1:1:1 mixture (167 μg of each) of LRP1-CL-I-Fc, LRP4-LA (2367)-Fc and VLDLR-LA (1-2)-Fc proteins were incubated with 105 PFU of YFV-M185D at 37 °C for 1 h and then inoculated into anti-IFNAR1 antibody treated C57BL/6J mice via intraperitoneal injection. Animals also were given 500 μg of the same Fc proteins at 1 and 2 dpi (n = 10 and 9, respectively). At 3 d.p.i., viral RNA levels in indicated tissues were measured. Data (c, d) are representative of 2 experiments. Data (e-f) are from 3 experiments, each performed in triplicate. Data (h-j) are pooled from 2 experiments, and bars indicate median values. Statistical analysis: two-tailed Mann-Whitney test (h-j), ns, not significant. Source gel data image (c) is in Supplementary Fig. 1. The diagrams in b,d were created in BioRender. Diamond, M. (2025) https://BioRender.com/psypg0x; Diamond, M. (2025) https://BioRender.com/v82694b.
Extended Data Fig. 10 LRP1-CL-I-Fc decoy protects hFRG mice against YFV infection in vivo.
a, Serum levels of VLDLR-LA (1-2)-Fc and LRP1-CL-I-Fc proteins were measured at 1 day post i.p. injection by ELISA. b, Individual hFRG mouse weight change in LDLRAD3-LA1-Fc or LRP1-CL-I-Fc treated groups after YFV infection. c, LRP1 KO efficiency (as indicated by sequences of wild-type allele) in PHH was analysed by next generation deep sequencing. Data (b) are from 2 experiments. d, Flow cytometry contour plots showing YFV infection in control and LRP1 KO PHH.
Supplementary information
Supplementary Figure 1 (download TIF )
Original source data of SDS–PAGE gels. The gels in a–c correspond to those in Extended Data Fig. 3a–c. The gel in d corresponds to data in Extended Data Fig. 9c. The red boxes indicate the gel regions that were used to create the Extended Data Figure panels.
Supplementary Table 1 (download XLSX )
sgRNA sequences in surfaceome library for used for gene editing.
Supplementary Table 2 (download XLSX )
MAGeCK analysis of genes in CRISPR–Cas9 screen.
Supplementary Table 3 (download XLSX )
Nucleotide sequence of LDLR family members used for expression and targeted screen in K562 cells.
Supplementary Table 4 (download XLSX )
Unnormalized infection data of control cells.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chong, Z., Hui, S., Qiu, X. et al. Multiple LDLR family members act as entry receptors for yellow fever virus. Nature 649, 173–182 (2026). https://doi.org/10.1038/s41586-025-09689-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09689-2
This article is cited by
-
Unlocking entry for yellow fever virus
Nature Reviews Microbiology (2026)