Abstract
Parity and breastfeeding reduce the risk of breast cancer, particularly triple-negative breast cancer (TNBC)1,2, yet the immunological mechanisms underlying this protection remain unclear. Here we show that parity is associated with increased numbers of CD8+ T cells, including cells with a tissue-resident-memory-like phenotype within human normal breast tissue. In mouse models, pregnancy followed by lactation and involution drove the accumulation of CD8+ T cells in the mammary gland, coinciding with reduced tumour growth and increased intratumoural immune cell infiltration, effects that were abrogated by CD8+ T cell depletion. Importantly, this CD8+ T-cell-dependent tumour control was observed only after a complete cycle of lactation and involution. Consistent with this, primary triple-negative breast cancers from parous women exhibited greater T cell infiltration and improved clinical outcomes. Together, these findings, spanning preclinical models and over 1,000 patient samples, provide insights into how reproductive history shapes breast immunity, positioning CD8+ T cells as key mediators of parity-associated protection and informing strategies for both the prevention and the treatment of breast cancer.
Similar content being viewed by others
Main
It is widely recognized that parity and lactation are associated with a reduced long-term risk of breast cancer1,2. Parity is thought to protect by reshaping mammary epithelial cell differentiation and growth pathways that occur during pregnancy, lactation and involution, thereby reducing susceptibility to malignant transformation over time3,4,5. While parity has been associated with a decreased risk of hormone-receptor-positive breast cancer, breastfeeding appears to confer a more specific reduction in the risk of triple-negative breast cancer (TNBC)1,6. The cellular and molecular mechanisms underlying this association are unclear. Deeper understanding could enable new subtype-specific prevention and treatment strategies for breast cancer. This could be particularly relevant for women at higher risk of TNBC, such as those who carry germline pathogenic variants, or for groups with poorer outcomes even after adjusting for key prognostic factors, such as certain ethnic groups7.
The prognostic role of CD8+ tumour-infiltrating lymphocytes (TILs) is well established in both early- and advanced-stage TNBC8,9 and T cell checkpoint inhibitors are now part of standard-of-care therapy, underscoring the critical role of host immunity. Our previous research demonstrated that a subpopulation of CD8+ T cells with a tissue-resident-memory (TRM)-like phenotype mediates robust anti-tumour responses and is associated with reduced breast cancer recurrence8,10,11. We also showed that TRM-like cells were present in both breast tumours and cancer-unaffected normal breast tissue, implicating them in treatment response and long-term immune surveillance of the breast.
Post-partum, immune cell populations in mammary tissue undergo considerable modulation: pregnancy and lactation are followed by widespread apoptosis and tissue remodelling in the process known as involution12,13. Changes in immune subsets, such as increased natural killer T (NKT) cells in the post-involution mammary gland, suggest a parity-induced immune surveillance mechanism14. Furthermore, circulating T cells increases in parous women have been observed to persist for months post-partum and are thought to contribute to autoimmunity14,15,16,17,18. Collectively, these observations suggest long-lasting immunological breast remodelling after pregnancy, although the functional importance of these populations in tumour surveillance remains unclear. Here we tested the hypothesis that parity and post-lactational involution favourably reshape the mammary microenvironment by recruiting and retaining CD8+ T cells that persist long-term in the breast, to support immune surveillance and protect against tumorigenesis.
Parity associations with T cell quantity
To assess how reproductive history influences the immune landscape of normal breast tissue, we first compared the quantity and phenotype of CD8+ T cells in cancer-unaffected breast tissue from parous and nulliparous women11. We analysed four single-cell transcriptomic datasets comprising 985,662 single cells (including more than 35,000 CD8+ T cells) from normal breast tissue of 170 adult women at average breast cancer risk, collated from a previous study16. We observed that parous women (at least one full-term pregnancy) had a significantly higher proportions of CD3+, CD8+ and CD8+ TRM-like cells than nulliparous women (Fig. 1a). Other cell subsets analysed did not differ significantly (Extended Data Fig. 1a–e).
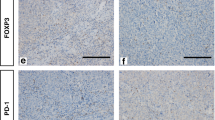
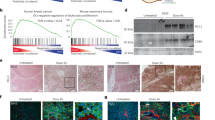
a, The immune cell abundance in normal breast from single-cell RNA-seq (scRNA-seq; y axis, cell type frequency) data, calculated as the indicated immune cells over total epithelial cells in women with average breast cancer risk. n = 170. The violin plots show the data distribution; the box plots show the interquartile range (IQR) extending from Q1 to Q3 (box limits) and the median (white bar), and the whiskers extend to the most extreme dot point within 1.5 × IQR and the points beyond show outliers. Statistical analysis was performed using two-sided Wilcoxon rank-sum tests; unadjusted P values are shown. Total cells, 985,662; CD45+, 120,467 from nulliparous (N, n = 53) versus parous (P, n = 117); CD3+ (n = 72,288 cells) in nulliparous (n = 53) versus parous (n = 115); CD8+ (n = 38,834 cells) in nulliparous (n = 53) versus parous (n = 114); and CD8+ TRM (n = 18,963 cells) in nulliparous (n = 53) versus parous (n = 114). b, The proportions of CD45+, CD3+ and CD8+ T cells of total cells and CD69+CD103– and CD69+CD103+ T cells of total CD45RA−CCR7−CD8+ T cells were determined by flow cytometry (nulliparous, n = 25; and parous, n = 65) analysis of normal breast in women with a high risk of breast cancer. Parous was defined as 1 or more full-term pregnancies (Methods). Statistical analysis was performed using two-sided Wilcoxon rank-sum tests, unadjusted for multiple comparisons. c, Representative multiplex OPAL images of nulliparous versus parous normal breast stained for DAPI, pan-cytokeratin (AE1/AE3), CD3, CD8, CD69 and CD103. Green indicates representative CD8+ T cell and white indicates merged CD69+CD103+CD8+CD3+ T cell (Methods). Scale bars, 100 μm. d, Persistence of CD69+CD103– versus CD69+CD103+CD8+ T cells over time (x axis; years from last live birth to human breast donation). n = 65. Datapoints are coloured according to age <50 years or >50 years for menopausal status. Unadjusted two-sided P values are shown. e, Volcano plot of breast CD69+CD103+CD8+ versus autologous circulating CD8+ T cells from non-cancerous parous women. n = 3. Significantly upregulated genes in CD69+CD103+CD8+ (limma test, false-discovery rate (FDR) < 1%, log[fold change (FC)] > 1) are referred to as the PB-TRM signature, f, Gene set enrichment analysis (GSEA) of the PB-TRM signature in normal breast (n = 109) from nulliparous women (n = 30) versus parous women (n = 79) from a previous study21. Exact P values are shown.
We next evaluated immune infiltration using flow cytometry in cancer-unaffected healthy breast tissue from women undergoing prophylactic mastectomy including parous (n = 65; >1 child) and nulliparous (n = 25) individuals at high risk of breast cancer (Supplementary Table 1). Compared with nulliparous women, the breast tissue of parous women showed increased proportions of CD45+, CD3+ and CD8+ lymphocytes and, notably, significantly increased proportions of CD8+ T cells expressing CD69 and CD103 canonical markers of tissue residency (TRM-like cells; Fig. 1b). Multiplex OPAL staining further confirmed increased numbers of CD3+ and CD8+ T cells, including a CD69+CD103+CD8+ TRM-like cell population in breast tissue from parous compared with nulliparous women, with localization predominantly in intraepithelial or periductal pan-cytokeratin-positive regions (Fig. 1c and Extended Data Fig. 2a,b). This dataset also enabled us to examine the long-term persistence of CD8+ TRM-like cells over a wide range of age intervals between last live birth and tissue donation, in both pre- and post-menopausal women. CD69+CD103+CD8+ T cells were stably maintained for over 30 years after pregnancy and showed superior long-term retention compared with their CD69+CD103−CD8+ T cell counterparts, consistent with previous observations that CD103+ TRM-like cells predominate over time19,20 (Fig. 1d).
To investigate the qualitative differences in breast-associated CD8+ T cells related to parity and lactation, we performed bulk RNA-sequencing (RNA-seq) analysis of TRM-like cells isolated from breast tissue from parous women unaffected by cancer who had breast fed for at least 6 months, comparing the gene expression profiles with those of autologous circulating CD8+ T cells. We identified 347 differentially expressed genes in the parous breast TRM-like cells (Supplementary Table 2), including upregulation of ITGAE, ITGA1 and CXCR6, hereafter the parous breast-associated-TRM (PB-TRM) signature (Fig. 1e). Analysis of an independent dataset comprising 109 normal breast tissue samples from cancer-unaffected women21 revealed significant positive enrichment of the signature in parous compared with nulliparous women, confirming parity-associated transcriptional differences (Fig. 1f). We also confirmed that the PB-TRM signature was significantly enriched in the CD8+ TRM compared with CD8+ TEM annotated single cells in the collated transcriptomic datasets16 (Extended Data Fig. 2c).
Increased CD8+ T cells in healthy mouse MFP
We next investigated immune cell populations in the mouse mammary fat pad (MFP) after pregnancy, lactation and involution in C57BL/6 mice under three conditions: (1) a complete cycle of natural lactation (21 days) followed by pup weaning, with tissue analysed at 28 days after involution when the mammary gland had returned to a pre-pregnancy state (hereafter, d28-inv mice); (2) early force-weaned (FW) involution, whereby pups were removed within 12–24 h of birth with minimal lactation and tissues were analysed 10 days after involution (d10-FW mice); and (3) respective age-matched controls to parous mice (virgin mice) (Fig. 2a).
a, Experimental schematic of parity. b, Immune cell frequencies (%) among CD45+ cells in the MFP were analysed using flow cytometry analysis of virgin (n = 9) and d28-inv (n = 12) C57BL/6 healthy mice; CD8α+CD44high T cells are highlighted in green. c, The numbers of the indicated immune cell subpopulations per gram of MFP from virgin (n = 9) and d28-inv (n = 12) C57BL/6 mice. d, The numbers of the indicated immune cell subpopulations per gram of MFP from virgin (n = 8) and d28-inv (n = 8) BALB/c mice. e, The numbers of the indicated immune cell subpopulations per gram of MFP from virgin (n = 8) and d10-FW (n = 8) C57BL/6 mice. f, Uniform manifold approximation and projection (UMAP) analysis of CD8α+ T cells generated from flow cytometry data from the MFP of C57BL/6 virgin (n = 8) and d28-inv (n = 9) mice; three enriched are clusters indicated. Each dot represents an individual cell. g, Feature plots of the indicated markers on CD8α+ T cells projected on UMAP. Data are mean ± s.e.m. The results represent two combined biologically independent experiments. Statistical significance was determined using two-sided (c–e) and one-sided (f) Mann–Whitney U-tests. Exact P values are shown, or P < 0.0001 in cases in which P values are very small.
Immune profiling of the MFP at these timepoints revealed a selective increase in the frequency of CD44highCD8+ T cells among CD45+ populations in d28-inv mice (Fig. 2b and Extended Data Fig. 3a). Total CD45+ and CD8+ T cell numbers were also significantly elevated compared with virgin mice, including CD8+ T cell subpopulations expressing CD69+CD103− and CD69+CD103+ (Fig. 2c and Extended Data Fig. 3b). A similar enrichment of CD8+ T cell subsets was observed in parous BALB/c mice at 28 days after involution, reinforcing that CD8+ T cell responsiveness is a strain-independent feature of the post-involution mammary gland (Fig. 2d). By contrast, CD8+ T cells and TRM-like cell populations were not significantly increased in d10-FW C57BL/6 mice compared with the virgin controls, and only modestly in BALB/c mice (Fig. 2e and Extended Data Fig. 3c). However, d10-FW mice showed a marked increase in ductal macrophages, consistent with previous reports13,22 (Fig. 2e).
To further characterize parity-associated CD8+ T cells, we next performed high-dimensional immunoproteomic analyses. This revealed three clusters that were significantly enriched in d28-inv mice compared with the virgin controls (Fig. 2f). All three clusters expressed canonical residency markers such as CD69, CD103 and CD49a along with differential expression of granzyme A, NK1.1 and CXCR3 (Fig. 2g and Extended Data Fig. 4a). Supporting the presence of TRM-like cells, the d28-inv MFP also showed increased expression of Tgfb2, Tnf and Cxcl16 (Extended Data Fig. 5a), factors that are associated with CD8+ TRM development23,24,25. Consistent with our human breast tissue findings, multiplex OPAL fluorescence microscopy revealed a significant increase in CD3+CD8+ TRM-like cells in the MFP of d28-inv mice compared with the virgin control mice (Extended Data Fig. 5b–d), with these cells located closer to E-cadherin+ epithelial cells (Extended Data Fig. 5e). Together, these data show that a complete cycle of lactation and involution promotes the accumulation and epithelial association of CD8+ T cells in mouse mammary tissue, paralleling observations in the human normal breast.
Lactation and involution restrain tumour growth
Accelerated tumour growth during early involution has been reported in preclinical models and associated with increased myeloid cell infiltration12,26. To further explore how different stages of involution affect tumour progression, we examined tumour growth in the MFP of d28-inv and d10-FW mice, compared with in the age-matched virgin controls using syngeneic orthotopic mouse models of TNBC. To this end, AT3-OVA TNBC cells were injected into the MFP of C57BL/6 mice at the timepoints described above (Fig. 2a). Tumour growth in d10-FW mice was comparable to virgin controls (Fig. 3a), consistent with previous reports12. By contrast, tumour growth was significantly reduced in d28-inv mice (Fig. 3b). This protective effect was also observed in BALB/c mice using the D2A1 breast cancer cells, which lack engineered antigens such as ovalbumin (OVA), providing an additional model of parity-induced tumour suppression (Fig. 3c,d).
a, Tumour growth (left) curves and end-point tumour volume (right) of AT3-OVA TNBC cells injected into the fourth MFP of d10-FW (n = 7) and age-matched virgin control (n = 7) C57BL/6 mice. b, Tumour growth (left) and end-point tumour volume (right) of AT3-OVA cells in d28-inv (n = 8) and age-matched virgin control (n = 8) C57BL/6 mice. c, Tumour growth (left) and end-point tumour volume (right) of D2A1 cells injected into d10-FW (n = 9) and age-matched virgin control (n = 10) BALB/c mice. d, Tumour growth (left) of D2A1 cells in d28-inv (n = 10) and age-matched virgin (n = 11) control mice and the end-point tumour volume (right) of d28-inv (n = 5) and age-matched virgin (n = 6) BALB/c mice. e, Immune cell frequencies (%) were analysed by flow cytometry among all CD45+ cells in AT3-OVA tumours from virgin (n = 9) and d28-inv (n = 10) mice 23–25 days after breast cancer tumour cell injection. f, The ratio of CD8+ to CD4+ T cells and the frequency of CD8+ T cells among all CD45+ cells AT3-OVA tumours from virgin (n = 9) and d28-inv (n = 10) mice. g, The numbers of the indicated immune cell subpopulations per gram of AT3-OVA tumour from virgin (n = 9) and d28-inv (n = 10) C57BL/6 mice. Data are mean ± s.e.m. Results represent two combined independent experiments. Statistical significance was determined using two-sided Mann–Whitney U-tests. Exact P values are shown, or P < 0.0001 in cases in which P values are very small.
Concomitant with reduced tumour outgrowth, we observed a significant increase in CD8+ T cells in d28-inv AT3-OVA tumours compared with in those from virgin mice (Fig. 3e,f), including OVA-specific tetramer+CD8+ T cells and CD69+CD103+ TRM-like subsets (Fig. 3g). Monocytes, CD4+ T helper type 1 cells, dendritic cells and XCR1+ type I dendritic cells were also elevated in d28-inv tumours, while B cells, natural killer (NK) cells and T cell receptor γδ (TCRγδ)+ cells were not significantly changed (Fig. 3g and Extended Data Fig. 6a). By contrast, CD8+ T cells were not increased in d10-FW AT3-OVA tumours compared with those from the virgin controls (Extended Data Fig. 6b). Similarly, in D2A1 tumours, CD45+ and CD8+ T cell numbers were higher in d28-inv mice, but not in d10-FW mice, relative to the virgin controls (Extended Data Fig. 6c,d).
CD8+ T cells mediate parity-induced protection
To investigate whether reduced tumour growth in parous mice was T cell mediated, we inoculated virgin and d28-inv Rag2−/−Il2rg−/−mice (lacking NK, T and B cells) with AT3-OVA cells and observed no difference in tumour growth (Fig. 4a). To assess whether CD8+ T cells could restore protection, we transferred naive CD8+ TCR transgenic OT-I cells (specific for OVA) into Rag2−/−Il2rg−/−mice before mating or into virgin controls. Tumour growth was significantly reduced in d28-inv mice compared with the virgin mice (Fig. 4b), but not when tumour cells were inoculated at 10 days after force-weaned involution (Extended Data Fig. 7a). Activated OT-I cells transferred before mating accumulated in greater numbers in the MFP at 28 days after involution compared with in virgin mice (Extended Data Fig. 7b), indicating that the post-lactation mammary microenvironment promotes the recruitment and differentiation of CD8+ TRM-like cells. In established tumours, CD45+ and TCR-vα2+CD8+ T cells were elevated in d28-inv mice compared with virgin mice, but not in d10-FW (Fig. 4c and Extended Data Fig. 7c). Adoptive transfer of activated gBT-I cells (irrelevant specificity) into Rag1−/− mice bearing AT3-OVA tumours had no effect on tumour growth (Extended Data Fig. 7d).
a, AT3-OVA tumour growth (left) and end-point tumour burden (right) in the fourth MFP of Rag2−/−Il2rg−/− d28-inv (n = 13) and age-matched virgin control (n = 14) mice. b, AT3-OVA tumour growth (left) and end-point tumour burden (right) in the fourth MFP of Rag2−/−Il2rg−/− d28-inv (n = 11) and age-matched virgin control (n = 10) mice pre-inoculated with OT-I cells 7 days before mating. c, The numbers of the indicated immune cells per gram of AT3-OVA tumour from virgin (n = 6) and d28-inv (n = 6) Rag2−/−Il2rg−/− mice at 4 weeks after tumour development. d, AT3-OVA tumour growth in the fourth MFP of virgin (left) and d28-inv (right) C57BL/6 mice treated with anti-CD4 (virgin, n = 12; d28-inv, n = 13), anti-CD8α (n = 9), anti-CD8β (virgin, n = 12; d28-inv, n = 14) and combined anti-CD4/CD8α (n = 14) or isotype control (n = 16). e, End-point AT3-OVA tumour weight in virgin (left) and d28-inv (right) C57BL/6 mice depleted with anti-CD4 (virgin, n = 12; d28-inv, n = 13), anti-CD8α (n = 9), anti-CD8β (virgin, n = 12; d28-inv, n = 14) and combined anti-CD4/CD8α (n = 14) or isotype control (n = 16). f, The numbers of CD3+ T cells per μl blood in virgin and d28-inv C57BL/6 mice treated with FTY720 (n = 8) or vehicle (n = 8). g, AT3-OVA tumour growth after treatment with FTY720 at the indicated timepoints (virgin, n = 11; d28-inv, n = 15) or treatment with vehicle (virgin, n = 12; d28-inv, n = 15). h, End-point AT3-OVA tumour weight in virgin and d28-inv C57BL/6 mice treated with FTY720 (virgin, n = 11; d28-inv, n = 15) or vehicle (virgin, n = 12; d28-inv, n = 15). i, The number of OVA-tetramer+ T cells per gram of AT3-OVA tumour (virgin, n = 10; d28-inv, n = 13) or vehicle (virgin, n = 10; d28-inv, n = 15) analysed 23–25 days after breast cancer tumour cell injection. Data are mean ± s.e.m. from two combined independent experiments. Statistical analysis was performed using two-sided Mann–Whitney U-tests (a–c), two-way analysis of variance (ANOVA) at the end point (d and g), Kruskal–Wallis tests (f and i) and one-way ANOVA (e and h). Exact P values are shown, or P < 0.0001 in cases in which P values are very small.
We next tested the requirement for CD8+ T cells in parity-induced protection. For this, virgin and d28-inv mice were depleted of CD8α+, CD8β+ or CD4+ T cells before tumour inoculation and throughout the experiment (Extended Data Fig. 8a,b). As expected, isotype-treated d28-inv mice had lower tumour burdens than virgins, accompanied by increased CD8α+CD44high T cell numbers in tumours (Extended Data Fig. 8c,d). Critically, we found that tumour protection in d28-inv mice was significantly reduced by either CD8α or CD8β depletion alone, identifying CD8αβ+ as the key mediators and excluding a dominant role for CD8αα+ T cells (Fig. 4d,e). In virgin mice, only combined CD8α+ and CD4+ T cell depletion increased tumour growth. Notably, CD4+ T cell depletion alone enhanced tumour control in d28-inv mice. High-dimensional proteomic analysis of CD45+ cells from tumours revealed an increased abundance of two clusters (C1 and C2) enriched for CD8+ T cells and cDC1 markers in CD4+ depleted d28-inv mice (Extended Data Fig. 8e–k).
To assess whether local CD8+ T cells were sufficient for tumour control or whether circulating T cell recruitment was required, we treated virgin and d28-inv mice with the S1PR1 agonist FTY720 to block T cell egress before tumour inoculation (Fig. 4f). Tumour protection was lost in FTY720-treated d28-inv mice, accompanied by reduced tumour-specific CD8+ T cells (Fig. 4g–i). Collagen abundance did not differ in between virgin and d28-inv tumours (Extended Data Fig. 8l,m), suggesting that stromal differences are unlikely to explain T cell recruitment patterns. Together, these findings demonstrate that parity-induced tumour protection is mediated by conventional CD8αβ+ T cells. After lactation and involution, tumour challenge drives both expansion of the mammary-resident T cells and recruitment of circulating tumour-specific effectors, enhancing local immune surveillance and tumour clearance.
Breastfeeding and tumour immune infiltration
Having established that reduced tumour growth after lactation and involution in mice is dependent on CD8+ T cells, we next examined whether a similar association exists in human TNBC patients, in which high T cell infiltrate is well known to correlate with improved prognoses8. Epidemiological studies also suggest that, while parity may transiently increase breast cancer risk, breastfeeding attenuates the risk of developing TNBC1. To explore this, we examined the Malaysian MyBrCa cohort, which includes 934 women with early-stage breast cancer and detailed clinicopathological and genomic information. We focused on the subset of 734 patients with recorded parity status and 656 with breastfeeding history27,28. In this cohort, the median maternal age at birth of first child was 27 years (range, 14–43 years), the median number of children per woman was 3 (range, 1–11) and the median lifetime breastfeeding duration in women who breastfed at all was 4 months (range, 0.5–156; Supplementary Table 3). Using the gene-expression-derived ESTIMATE immune infiltrate score29, our analysis revealed significantly higher tumoural immune content in basal-like (TNBC) in parous women overall, compared with in the nulliparous women who subsequently developed breast cancer post-partum (Fig. 5a). A similar increase in the ESTIMATE score was seen in parous women who breastfed compared with nulliparous women (Fig. 5b). We next performed differential expression analysis between these basal-like tumours in parous versus nulliparous patients and parous-breastfeeding versus parous-non-breastfeeding patients. We found that the PB-TRM signature generated in Fig. 1f was highly enriched with both parity and breastfeeding status (Fig. 5c,d), highlighting the specificity of the PB-TRM signature to breast involution by parity and lactation. Furthermore, parous women who had breastfed had a significantly greater intratumoural CD8+ T cell density in basal-like tumours and when compared with other breast cancer subtypes (Fig. 5e and Extended Data Fig. 9f).
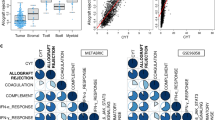
a, The effect of parity on subsequent tumour immune infiltration, as calculated using the RNA-seq-based ESTIMATE score in the MyBrCa cohort. Statistical analysis was performed using a linear model for ESTIMATE score with parity and sequencing batch as covariates. β is the estimated non-standardized coefficient for parity with 95% CI. b, The immune (ESTIMATE) score from the MyBrCa cohort with known breastfeeding (BF) status before breast cancer. Statistical analysis was performed using ANOVA with breastfeeding status and sequencing batch as covariates; two-sided, 95% CI; the mean difference was compared using Tukey’s honest significant difference test. P values were adjusted for intrasubtype comparisons only. c, GSEA of the PB-TRM signature in parous versus nulliparous women in the MyBrCa basal cohort. d, GSEA of the PB-TRM signature in breastfed compared with non-breastfed (no-BF) parous women in the MyBrCa basal cohort. e, The intratumoural T cell density in basal-like breast cancer was determined through immunohistochemistry analysis of CD8+ and CD3+ in the MyBrCa cohort, comparing nulliparous versus parous women by breastfeeding durations before breast cancer. The percentage CD8+ and CD3+ T cell density was quantified as the ratio of stain-positive pixels to all pixels within the tumour margins. Modelled with β regression adjusting for covariates of age at diagnosis and tumour grade. Unadjusted P values are shown with the average marginal effect (AME) for each group compared with nulliparous women. f, Kaplan–Meier survival analysis of OS in women (n = 270) diagnosed post-partum with primary hormone-receptor-negative breast cancer, stratified by breastfeeding status (yes versus no). Statistical analysis was performed using univariate Cox regression hazard ratio with 95% CIs. g, Quantification of stromal TILs on standard H&E-stained digital slides from primary hormone-receptor-negative women with high familial risk (n = 136). Age-adjusted percentage infiltration (y axis) by breastfeeding duration (x axis; Supplementary Table 4). The horizontal lines indicate the median values, and the vertical lines show the IQR. Group differences were tested using Kruskal–Wallis (K-W) tests. For a, b and e, the box plots show the median (horizontal line), IQR (box limits), the whiskers extend to the most extreme dot point within 1.5 × IQR and the points beyond show outliers. Exact P values are shown.
We further examined an independent of cohort of 270 Australian women who carry a germline pathogenic variant and had been diagnosed with hormone-receptor-negative early breast cancer after at least one full-term pregnancy, with available breastfeeding history and survival outcomes (Fig. 5f and Supplementary Table 4). After adjustment for known prognostic factors, we found that patients who breastfed exhibited significantly longer overall survival (OS) after breast cancer diagnosis compared with patients who did not breastfeed (hazard ratio = 0.39; 95% confidence interval (CI) = 0.19–0.79, P = 0.009; Extended Data Fig. 9a). Using an established method to assess immune infiltration on haematoxylin-and-eosin-stained (H&E) digital slides30, we found increased lymphocytic quantity in individuals with longer breastfeeding duration (Fig. 5g). Although precise separation of parity and breastfeeding effects in human datasets is challenging, our analyses of two independent cohorts suggest that breastfeeding may contribute additional immune-mediated protection beyond parity alone.
To further support the general clinical relevance of the PB-TRM signature, we examined associations of the PB-TRM signature with TILs and survival outcomes in the available early-stage breast cancer cohort31,32. We observed a strong correlation between increased TIL quantities30 and higher levels of the PB-TRM signature in patients with TNBC in the TCGA Breast Cancer dataset32 (Extended Data Fig. 9b,c). The PB-TRM signature was also enriched in basal-like tumours and associated with improved disease specific and OS in the METABRIC dataset (Extended Data Fig. 9d,e).
Discussion
The mammary gland undergoes profound structural and immunological changes across reproductive stages. While ductal macrophages in mammary gland biology are well studied, evidence increasingly supports a critical role for adaptive immunity, particularly during puberty and pregnancy13,33. Various immune cell subsets populate the ductal epithelium, and progesterone has been shown to influence T cell polarization during pregnancy12,33. Moreover, breast cancers that develop in the immediate post-partum period have a poorer prognosis34,35. However, the mechanisms by which parity and breastfeeding confer long-term protection from TNBC have remained unclear.
We show that normal breast tissue from parous women has significantly more CD8+ T cells, particularly those with a TRM-like phenotype, compared with nulliparous women across diverse backgrounds16. Given that CD8+ T cell quantity is associated with improved TNBC outcomes, we hypothesized that parity-driven CD8+ T cell responses could enhance long-term cancer immunosurveillance. Supporting this, CD8+ TRM-like cells in normal breast tissue from parous women were observed to persist for decades post-partum.
In mouse models, completion of a full reproductive cycle was associated with reduced tumour growth and a sustained increase in T cells in the MFP, including those with a TRM-like phenotype. These cells localized to the mammary epithelium under homeostatic conditions and were more frequently found within tumours after lactation, supporting their role in tumour immune surveillance. Depletion of conventional CD8αβ+ T cells abrogated parity-induced protection, confirming their essential functional role. Similarly, FTY720-mediated blockade of lymphocyte egress impaired tumour protection, suggesting that the parous mammary gland not only maintains resident CD8+ T cells, but also facilitates the recruitment of circulating tumour-reactive T cells from the periphery. This was accompanied by increased numbers of conventional type 1 dendritic cells the tumours of parous mice, consistent with enhanced CD8+ T cell priming and maintenance36. Previous studies reporting expansion of mammary intraepithelial lymphocytes and NKT cells through epithelial CD1d expression support the concept of immune remodelling after lactation, but our results indicate that these populations are not the primary mediators of tumour protection in our mouse models14,17,37. Instead, our data complement these studies by demonstrating a durable, CD8+ T-cell-mediated layer of tumour protection in the mammary tissue after lactation. These findings were paralleled in human TNBC, in which tumours from parous women exhibited greater T cell infiltration, which was also correlated with longer breastfeeding duration. Such associations may reflect cumulative antigen exposure, cytokine signalling, in addition to the involution-associated tissue changes that promote long-term T cell retention.
Given known population-level variation in immune infiltration and breast cancer risk, it is important to consider how these findings may generalize across populations. For example, higher TIL content in primary TNBCs from Asian compared with European populations has been previously reported, with exploratory substudies from phase III trials suggesting a numerically greater magnitude of benefit from checkpoint inhibition, although these comparisons remain hypothesis generating38. Although our findings focus on the potential preventive role of parity-induced CD8+ T cells that are resident in the breast, they suggest that the immune contexture of the parous breast could influence TNBC therapeutic responsiveness. This is particularly relevant given the increasing use of immune checkpoint blockade in early-stage disease. However, we acknowledge that this remains the subject of future investigation.
Tumour-extrinsic factors, such as the mammary microbiome and dietary influences, may further modulate immune infiltration and tumour development39. Moreover, post-partum tissue remodelling, including clearance of mutant clones and epigenetic changes3,40, might act synergistically with CD8+ T-cell-mediated surveillance to eliminate premalignant cells. Similar immune-mediated protective mechanisms have been observed in other tissues, such as commensal skin human papilloma viruses promoting local CD8+ T cell populations; immune-mediated regression of early lung neoplastic conditions; and HER2-specific T cells linked to parity-associated protection of ductal carcinoma in situ41,42,43. These examples underscore the importance of physiological immune health in normal tissue as a mechanism for cancer prevention—a concept that we demonstrate here specifically for TNBC.
Our study has limitations. Although we focused on TNBC, reproductive history might also influence immune-mediated protection in other breast cancer subtypes, although this remains to be determined. We recognize that we did not assess hormone-receptor-positive models due to limitations in preclinical systems, even though parity and breastfeeding is associated with reduced risk of all breast cancer subtypes. Oestrogen signalling has been shown to have diverse effects on immune cell function, with context- and receptor-specific outcomes. Although oestrogen can suppress certain immune responses through ERα signalling, ERβ activation in immune cells has been associated with enhanced anti-tumour activity44. Moreover, our human cohorts varied in reproductive histories, population backgrounds, time from tissue donation and last child, potentially introducing heterogeneity. Lastly, antigen specificity of the CD8+ T cells identified remains undefined, warranting future research.
In conclusion, our findings reveal that lactation-associated remodelling of the mammary gland confers durable CD8+ T-cell-mediated protection against TNBC. By integrating data from mouse models and diverse human populations, this work provides a new framework for understanding how reproductive history shapes long-term immune surveillance and influences breast cancer risk. These insights support the need for tailored prevention and treatment strategies that consider parity status and the importance of systematically capturing reproductive history in future immunotherapy clinical trials. This is particularly critical given the rising global burden of breast cancer45.
Methods
Patient samples
This project was approved by the Human Research Ethics Committee of the Peter MacCallum Cancer Centre (project approval number ‘SEGMENT’ 13/123 and 97/27 for the Kathleen Cuningham Foundation Consortium for Research into Familial Breast Cancer (kConFab) with project approval numbers 129 and 150). All of the participants provided written informed consent before tissue and blood collection. Ethics approvals for the human MyBrCa cohort can be found in the original publication28.
Tissue processing of human normal breast
Cancer-unaffected normal breast tissues were collected from prophylactic mastectomies from women from high-risk familial cancer families. Mammary tissues were divided into segments and either placed in neutral-buffered formalin for processing to formalin-fixed paraffin-embedded (FFPE) blocks, or a single-cell suspension was created for further processing. In brief, adipose tissue was removed from the mammary glands and the associated connective tissues were then finely diced into smaller fragments in RPMI1640 containing 1 mg ml−1 collagenase type 4 (Worthington Biochemical), 30 U ml−1 DNase (Roche Diagnostics) and incubated for 30 min at 37 °C. Digested tissue fragments were teased through a 70-μm sterile cell strainer, the sieve was irrigated with Dulbecco’s PBS and the homogenized cells were collected into multiple 50 ml conical tubes. Pelleted cells were stained with monoclonal antibodies for immunophenotyping and resuspended in 2% paraformaldehyde (PFA) buffer for flow cytometry spectral analysis.
Antibody labelling of human cells for FACS
Homogenized cells in suspension were labelled with monoclonal antibodies for 30 min at 4 °C in fluorescence-activated cell sorting (FACS) buffer (2% FBS in Dulbecco’s PBS), and washed twice in FACS wash buffer. The antibody panel included CD45 and HLA-ABC to discriminate between lymphocytes from other stromal and epithelial cell compartments in suspension, T cell markers CD3, CD4 and CD8, and the T cell differentiation markers CCR7, CD45RA, CD69 and CD103. Viable cells were revealed using the fixable zombie red, PE-Tex-red or fixable yellow viability dyes (BioLegend). Compensation controls were prepared for each antibody using UltraComp beads (BD Biosciences). Multiparameter flow cytometry data were acquired on the BD FACS Symphony A5 or LSR Fortessa X-20 instrument (BD Biosciences) and data were analysed using FlowJo v.10 (BD Biosciences). A representative flow cytometry gating strategy of immune cells in human normal breast tissue is shown in Extended Data Fig. 10a.
Human normal breast data analysis
Two-sided Wilcoxon rank-sum tests were used to compare parous and nulliparous groups across different T cell subpopulations. β regression was used to model the interval from last birth to tissue donation and how this affected CD8+CD103− and CD8+CD103+ T cells, as implemented in the betareg R package (v.3.2-1). Regression lines were fitted using β regression, controlling for age at tissue donation, with the 95% CIs shaded. The AME for trend over time, representing the change in the percentage abundance per 1 year increase in birth to donation interval (Fig. 1d). Age at tissue donation was included as a continuous covariate. The AME of the interval since last pregnancy was estimated using the ‘margins’ function from the margins R package (v.0.3.28), interpreted as the average change in cell proportion per year increase in the interval, averaged over the empirical distribution of age at tissue donation. To represent model fit on plot and calculate CIs, the fitted β-regression model was supplied to the ggpredict function of the ggeffects R package (v.2.2.1).
Mouse tissue processing and FACS
Spleens were mashed through a 70-μm cell strainer and incubated with red blood cell lysis buffer for 5 min at room temperature before staining. Mouse mammary tissue with the lymph nodes removed or tumour were minced into fragments and incubated in RPMI1640 containing 1 mg ml−1 collagenase type 4 and DNase II at 0.2 mg ml−1 for 30 min at 37 °C with shaking (300 rpm). Cells were serially passed and washed through 70-μm strainers followed by viability staining in PBS then Fc receptor (FcR) blockade antibody (clone 2.4G2) at 1:500 dilution for 10 min at room temperature. Flow cytometry staining was then performed with cell surface antibodies in FACS wash buffer (4% FCS, 2 mM EDTA) containing BD Horizon Brilliant Stain Buffer Plus (BD). After staining with an antibody cocktail for 30 min at room temperature, the samples were washed twice with FACS buffer and fixed with FOXP3/transcription factor staining buffer set (Invitrogen) according to the manufacturer’s instructions. The samples were either resuspended in 2% PFA or underwent intracellular staining in FOXP3/transcription factor permeabilization buffer followed for 30 min at room temperature. Intracellular stained samples were washed twice and resuspended in 2% PFA. The samples were analysed on the BD Symphony FACS analyser or 5-laser Cytek Aurora. Data were analysed using FlowJo v.10 or OMIQ cloud-based platform.
OMIQ was used for high dimensional analysis. Data were scaled using hyperbolic arcsine (asinh) transformation and clustered using Phenograph. For high dimensional analysis of immune cells in the MFP, TCRγδ+ and CD1d+ cells were excluded and CD45+TCRβ+CD8α+ cells were clustered based on the expression of granzyme B, CD38, CD62L, TCF-1, CD49a, NK.1, CD8β, SLAMF6, CD103, CD44, Ly6C, KLRG1, CXCR3, CD39, PD1, CD244, TBET, CD69, CXCR6, CD8α and granzyme A. For high-dimensional analysis of immune cells in AT3-OVA tumours, CD45+ cells were clustered based on the expression of CD19, NK1.1, CD3, TCRb, TCRγδ, CD8α, CD8β, CD4, CD44, CD62L, CD69, CD103, CD49a, CD11b, CD11c, F480, MHC-II, SIGLEC-F, XCR1, SIRPα, CX3CR1, CD64, CD39, OVA-tetramer, PD-1, PD-L1, LAG3, KLRG1, CD38, FOXP3, TBET, TCF-1 and CTLA-4.
Bulk RNA-seq gene expression analysis
Cancer-unaffected normal-breast-tissue-associated CD69+CD103+CD8+ T cells and autologous blood derived CD8+ circulating T cells (5 × 103–1 × 104 cells) were FACS purified under sterile conditions using the BD FACSAria II Cell Sorter (BD Biosciences). For reanalysis of isolated CD8+ T cell subpopulations after FACS, each individual sample was assessed for >90% purity. Total RNA was extracted from the indicated T cell subpopulations from n = 3 pooled cancer-unaffected normal breast tissue samples using the RNeasy mini kit (Qiagen) according to the manufacturer’s instructions. RNA Tape Station (Agilent) analysis was performed according to the manufacturer’s instructions to assess the quantity and quality of RNA present in the sample. Then, 2–5 ng was used for RNA library preparation according to the manufacturer’s instructions with ribo-depletion NEB. The library was then amplified with 3′ PCR primers containing sample indices and the Illumina clustering guides. 10× lysis Buffer is a cell lysis buffer that can be used together with the SMARTer Ultra Low Input RNA Kit for Sequencing v3 and the SMART-Seq v4 Ultra Low Input RNA Kit for Sequencing. Indexed libraries were pooled and sequenced on the Illumina NextSeq HO 75SE (Illumina) system. Two to five million single-end reads were generated as an output. Adaptor trimming was performed, and reads were aligned to the mm10 reference genome using HISAT2 (v.2.2)46. Aligned reads were quantified using HTSeq (v.2.0.3). Counts were normalized and unwanted variance that was introduced by library size differences was removed using RUV-III (v.0.9.7.1) in R. Differential expression analysis was performed using limma (v.3.60.3). Volcano plots were produced with custom code using ggplot2 (v.3.5.1). The PB-TRM signature was derived from differential expression analysis by filtering the genes by FDR < 1% and log[FC] > 1. A gene expression dataset of premenopausal parous and nulliparous human breast tissue was accessed at Gene Expression Omnibus (GEO: GSE112825). We used limma47 to analyse the previously normalized expression and perform differential expression analysis of parous against nulliparous women. To detect enrichment of PB-TRM gene signature, we performed GSEA using the differential expression results with the fgsea (v.1.30.0) library in R (v.4.4.1)48.
Single-cell cohort
The processed and annotated single-cell normal breast tissue atlas presented previously16 was used; only cells annotated as ‘not sorted’ or ‘live sorted’ were collated from several datasets of normal human breast tissue. We used samples from women annotated as ‘average risk’ or ‘unknown risk’ of breast cancer, that is, normal breast samples from reduction mammoplasties and not cancer mastectomies or prophylactic mastectomies, and where parity status was known.
To calculate and compare the abundance of normal breast cell types between parous and nulliparous women, we calculated ratios of CD3+ T cells, CD8+ T cells and CD8+ TRM-like cells over total epithelial cells per sample, compared using unpaired Wilcoxon tests; Violin plots shown were produced in R (v.4.4.1). We conducted a focused analysis on the CD8+ T cells, performing integration with Harmony (v.1.2.3)49 using the default parameters (using dataset and donor as covariates), and dimensionality reduction and visualization with Seurat (v.5.2.1)50. We used the scaled gene counts and calculated single-cell signature enrichment of the PB-TRM signature using the AUCell (v.1.28.0) package in R51. We compared single-cell enrichments of two different previously annotated CD8+ T cell subpopulations (TEM and TRM-like phenotype) using the unpaired Wilcoxon rank-sum test.
Mouse cell lines
AT3-OVA cells were provided by P. Darcy. D2A1 cells52 were provided by K. Britt. AT3-OVA TNBC cells were generated by transducing the parental AT3 cell line with a retroviral vector pMIG/MSCV-IRES-eGFP plasmid encoding membrane-bound chicken OVA cDNA (model antigen) protein, tagged with GFP as previously described53. Both cell lines were well established from Peter MacCallum Cancer Centre laboratory stocks and published. Breast cancer cells were cultured with complete DMEM medium supplemented with 10% FCS. For in vivo experiments, 2.5 × 105 AT3-OVA or 1 × 104 D2A1 cells at an early passage state were resuspended in 50 µl PBS solution at neutral pH and injected orthotopically into the right fourth MFP of mice. Cell lines were verified to be mycoplasma negative at the Victorian Infectious Diseases References Lab and were maintained at 37 °C in a humidified incubator under 5% CO2.
Mouse tumour and parity models
All animal experiments conducted in this study were approved by the relevant Peter MacCallum Cancer Centre Animal Experimentation Ethics Committee (E648, 2025-05) or by The University of Melbourne Animal Ethics Committee (21938) and conducted in accordance with the National Health and Medical Research Council Australian Code of Practice for the Care and Use of Animals for Scientific Purposes.
C57BL/6J wild type (WT), BALB/c WT, Rag2−/−Il2rg−/−, Rag1−/−, gBT-I and OT-I CD45.2 WT female mice were obtained from the Walter and Eliza Hall Institute of Medical Research or bred in-house (Victoria, Australia). Mice were allocated to experiments at 6–8 weeks of age. For tumour models, volume (length × width2 × 0.5) was calculated by vernier callipers after development of palpable tumours, and measurements were taken every 3–4 days. Mice were euthanized when MFP tumours reached an ethical limit of up to 1,500 mm3, or if the animals showed any signs of adverse health indications as per Peter MacCallum cancer centre institutional guidelines. For parity experiments, dams were whitened before the introduction of stud males to establish breeding pairs/trios. Pregnancy was confirmed by plug formation and studs were removed. After littering, dams underwent complete lactation and with pups for 21 days and, for forced wean experiments, the pups were removed at birth within 12–24 h from the dams post-partum and euthanized (d10-FW timepoints). Dams that underwent early pup removal were immediately co-housed with other female mice and did not exhibit changes in social behaviour. Housing conditions for the mice were followed according to institutional guidelines, including dark–light cycle, ambient temperature and humidity conditions.
Adoptive transfer of transgenic T cells
OT-I cells were isolated using the negative immunomagnetic cell-separation method, MACS (Stem Cell Technologies), from the spleen of WT OT-I CD45.2 donor mice. A total of 2.5 × 105 OT-I cells was intravenously transferred to recipient Rag2−/−Il2rg−/− mice, 7 days before mating and in house breeding began. OT-I cells were immunophenotyped by flow cytometry to assess the purity of OT-I cell isolation (>90–95%) with the use of CD45.2, TCR-vα2, CD8α, CD44 and CD62L cell surface markers. For experiments with effector OT-I cells, T cells were activated in culture for 4 days with OVA257–264 (SIINFEKL) peptide-pulsed, irradiated (50 Gy) splenocytes in the presence of recombinant human IL-2 (25U ml−1; PeproTech) at 37 °C and 5% CO2 and 1 × 104 cells were injected intravenously per mouse.
For the gBT-I experiment, HSV-specific gBT-I transgenic T cells were activated in culture for 4 days with gB498–505 (SSIEFARL) peptide-pulsed, irradiated (50 Gy) splenocytes in the presence of recombinant human IL-2 (25U ml−1; PeproTech) at 37 °C and 5% CO2. The 2 × 107 activated gBT-I cells were injected intravenously into naive Rag1−/− mice. Then, 56–62 days after gBT-I cell transfer, mice were subsequently inoculated orthotopically with 5 × 105 AT3-OVA cells into the fourth MFP as described above.
T cell depletion experiments
C57BL/6 mice were mated as described above before being allocated to tumour experiments. Seven days before AT3-OVA tumour challenge, non-randomized mice were injected intraperitoneally with 200 µg IgG2 isotype control combined with 200 µg anti-CD4 (GK1.5, Bio X Cell Leganon), anti-CD8α (YTS, Bio X Cell), anti-CD8b (53-5.8, Bio X Cell) or 400 µg IgG2 isotype control (Bio X Cell) twice then once weekly.
Fingolimod (FTY720) in vivo treatments
C57BL/6 mice were mated as described above before being allocated to tumour experiments. In brief, 3 days before AT3-OVA challenge, the mice were injected intraperitoneally with 1 mg per kg FTY720 compound reconstituted in 2% 2-hydroxypropyl-β-cyclodextrin (Merck) or vehicle control (2% 2-hydroxypropyl-β-cyclodextrin) with daily injections separated by one day breaks every 5 days.
Mouse histology and OPAL analysis
Intact fourth MFPs from C57BL/6 mice were fixed in 10% neutral-buffered formalin for 24 h and then processed and embedded into FFPE blocks. Then, 4-µm-thick sections were cut from blocks and mounted onto SuperFrost+ slides. m-IHC staining using OPAL fluorophores was then performed according to the manufacturer’s instructions (Akoya Bioscience). In brief, the reagents were loaded into the Bond Rx Autostainer (Leica Biosystems) system for an entire cycle of approximately 12 h. The tissue sections were baked, dewaxed and incubated in a 3% hydrogen peroxide (H2O2) solution (Merck EMSURE) to block endogenous peroxidases in the tissues. Subsequently, the slides were incubated with 1% BSA in Tris-buffered saline to block non-specific background binding. Slides were incubated iteratively with the primary antibodies including CD8 (1:4,000, pH 9), CD3e (1:600, pH 6) and CD103 (1:2,000 pH 9), for 30 min, the secondary antibody (OPAL polymer HRP mouse + rabbit) for 10 min, and the tyramide-conjugated OPAL fluorophores (570 and 690 (1:150) and 780 (1:25)) for 10 min. Finally, the slides were incubated with spectral DAPI for 10 min. All antigen retrieval was performed at 97 °C for 20 min and all of the other steps were performed at ambient temperature. After the completion of staining, the slides were briefly rinsed in milliQ water and then manually cover slipped using Prolong Glass Antifade Mountant (Invitrogen).
The stained 4-plex OPAL m-IHC slides were imaged on the Phenoimager HT imaging system (AKOYA Biosciences). The whole tissue section was spectrally imaged with a ×20 objective. The optimal exposure settings were manually set for each tissue. Whole-slide images were then unmixed to subtract the autofluorescence signal and images were exported as component .tiff files using InForm software (v.2.6.0, AKOYA Biosciences). In brief, the spectral library was created using multispectral images obtained from single stained slides for each marker and the associated fluorophore (DAPI, OPAL 570, 690 and 780) on InForm software (v.2.6.0, AKOYA Biosciences) for downstream image analysis workflow. For Masson’s trichrome staining, 4-µm-thick sections were cut from FFPE blocks and incubated in Bouin’s fixative for 60 min at 60 °C, washed in water and stained with haematoxylin. The slides were washed and stained with 1% Ponceau 2R and 1% acid fucshin for 2 min. Slides were washed with water stained with 2% light green in 1% acetic acid for 5 min, washed with water, dehydrated and cover slips were mounted.
Human histology and OPAL analysis
Human breast tissue preparation
Immune cells were examined to visualize the expression of CD3, CD8, CD69, CD103 and pan-cytokeratin (AE1/AE3) in cancer-unaffected normal breast tissue, using the OPAL serial immunostaining protocol. In brief, FFPE tissue sections were cut at a thickness of 4 μm and melted at 60 °C for 45 min, followed by dewaxing in three changes of histolene for 11 min and 3 changes of 100% ethanol for 1 min each and 70% alcohol for 1 min. The Leica Bond auto-stainer protocol was performed according to the manufacturer’s instructions.
Antigen-retrieval process
Heat-induced antigen retrieval was achieved using a microwave. Tissue sections were placed into a plastic hellendahl jar (Trajan Scientific Australia) in EDTA (pH 8) antigen-retrieval buffer and citrate buffer (pH 6) antigen-retrieval buffer for CD3 staining and brought to the boil at a 100% power for 50 s and at 10% power for 15 min. The tissue sections were then left to cool for 30 min and washed in 0.02% TBST three times for 7 min each with gentle agitation.
Primary antibody staining
The sections were blocked in blocking buffer (Dako, X0909) for 10 min at room temperature before incubation with primary antibodies or isotype controls. For primary antibody staining, tissue sections were incubated for 30 min at room temperature with mouse anti-human pan-cytokeratin (AE1/AE3; 1:1,000, Opal 480; isotype: mouse IgG1), rabbit anti-human CD3e (SP7; 1:500, Opal 690; isotype: rabbit IgG), mouse anti-human CD8 (4B11; Leica; 1:800, 570–1:500, Opal 570; isotype: mouse IgG2b), rabbit anti-human CD69 (EPR21814; Abcam, Opal 520; isotype: rabbit IgG), rabbit anti-human CD103 (EPR416602; Abcam; 1:1,000, Opal 780; isotype: rabbit IgG) and cell-permeable DNA-binding dye (DAPI) obtained from Abcam. After primary incubation, the sections were washed five times in 0.02% TBST for 5 min each. The tissue sections were then incubated in 0.3% H2O2 for 10 min after the first primary antibody incubation and washed three times in 0.02% TBST for 5 min each. Respective isotype antibody control expression was examined under similar conditions and was tested on matched human normal breast and in human tonsil lymphatic tissues.
Secondary antibody detection
All sections were incubated with secondary-HRP conjugated antibodies (Leica Bond autostainer protocol) at a dilution of 1:1,000 for 10 min at room temperature. The sections were washed three times for 5 min each in 0.02% TBST followed by signal amplification using 100 μl of TSA Plus working solution per slide at a dilution of 1:50 in 1× amplification diluent, incubated at room temperature for 10 min as specified by the manufacturer (Opal 6-Plex Manual Detection Kit for Whole Slide Imaging, AKOYA Biosciences). Nuclei were counterstained with DAPI according to the manufacturer’s instructions diluted from 10× spectral DAPI (AKOYA Biosciences) for 2 min at room temperature. The sections were then washed twice in 0.02% TBST for 2 min each and mounted in Vectashield hard-set medium (Vector) and left to dry flat for 20 min in the absence of light.
Microscopy image analysis
OPAL m-IHC slides were imaged on the Pheno Imager HT (Polaris) (AKOYA Biosciences). The acquired images were spectrally unmixed using inform v.2 (AKOYA Biosciences) into individual component .tiff files. Whole-slide images were then stitched using the HALO Image Analysis Platform v.3.6.4134 (Indica Labs). For human tissue, quantification of immune cells, specifically the CD8+ T cell subpopulations in the mammary epithelial-dense regions after excluding adipose-dense regions across the entire tissue was performed using the HighPlex FL v.4.2.3 module. For mouse tissue, quantification of immune cell subsets was performed using the HighPlex FL v.4.2.3 module after excluding lymph node regions within the surrounding tissues of interest. For the distance between ECAD+ epithelial cells and CD3+CD8+CD103+ T cells, nearest neighbour was calculated on the HALO Image Analysis Platform v.3.6.4134. Masson’s-trichrome-positive staining was quantified using QuPath (v.0.6)54.
RNA extraction and qPCR with reverse transcription analysis
MFP tissue from C57BL/6 mice was manually minced and then homogenized in 500 µl TRIzol using metal bead lysing matrix (MP Biomedicals) on the FastPrep-24 bead-beating grinder and lysis system (MP Biomedicals). RNA was extracted from the homogenized tissue using the PureLink RNA Mini Kit (Invitrogen) according to the manufacturer’s instructions. Total RNA was then quantified using the NanoDrop 2000 Spectrophotometer (Thermo Fisher Scientific). First-strand complementary DNA (cDNA) was synthesized from extracted RNA using the SuperScript IV VILO Master Mix with ezDNase enzyme (Invitrogen) according to the manufacturer’s instructions. This was performed on a T100 Thermal Cycler (Bio-Rad).
Quantitative PCR (qPCR) was performed on the synthesized cDNA to assess relative gene expression using the TaqMan Gene Expression Assay with primers targeting Gapdh, Mm99999915_g1; Tgfb2, Mm00436955_m; Tnf, Mm00443258_m1; and Cxcl16, Mm00469712_m1. All qPCR reactions were performed on the StepOnePlus Real-Time PCR System (Thermo Fisher Scientific). Gapdh was used as the housekeeping gene. The \({2}^{-{\rm{\Delta \Delta }}{C}_{{t}}}\) method was used for calculating relative gene expression55.
Human and mouse reagents and antibodies
Flow cytometry antibodies used for normal breast immunophenotyping include CD45-BV510, CD3-APCH7, CD103-BV421, CD8-BV605, PD-1-BV785, CCR7-BV711, CD3-BV711, CD4-BV650, CD69-PerCP-Cy5-5, Zombie red-PE-Tex-Red (BioLegend), HLA-ABC-PECy5, CD45-FITC, CD8-PE and CD45RA-FITC (BD Biosciences).
Flow cytometry antibodies used for mouse healthy MFP analysis and mammary tumour immunophenotyping include CD45-APCCY7, TCRβ-BB700, TCRβ-BV786, CD8-BV711, CD8-BUV395, CD4-BUV805, CD44-BV605, CD62L-PECY7, CD62L-BUV737, CD69-BUV395, CD69-PE, CD103-FITC, CD103-PE, CD11B-BV711, CD11C-BV786, F4/80-EF450, MHC-II APC and Fixable dyes Fix-Yellow-BV510, Zombie NIR or Zombie red PE-Tex-Red (BioLegend).
Antibodies used for spectral flow cytometry immunophenotyping were as follows: CD45-PerCP, CD19-AF488, TCRγδ-PE-Cy7, NK1.1-BV570, CD3-Spark NIR685, TCRβ-BB700, CD8α-BUV395, CD4-BUV805, CD44-APCR700, CD62L-BUV737, CD69-PE, CD103-BV421, CD11b-BV711, CD11c-BV785, F480-APC, Ly6G-BV650, Ly6C-BV605, MHC-II-Spark Blue 550, SIGLEC-F-PerCP-Fire 806, XCR1-Spark UV387 an CD64-PE-Fire 744, and H-2Kb-OVA-tetramer-PE or H-2Kb-OVA-tetramer-APC were used to detect and track CD8+ T cells specific to SIINFEKL (OVA257–264) peptide from OVA-expressing tumour cells. Antibodies used to generate the CD8α+ T cell UMAP were as follows: CD49a-BUV395, NK1.1-BUV563, CD8α-RB545, CD8b-BUV661, CD4-BUV805, SLAMF6-BV421, CD103-BV480, CD44-BV510, Ly6C-BV570, KLRG1-BV605, CXCR3-BV650, CD39-BV711, PD1-BV750, CD244-BV785, TCRγδ-BB700, CD1D-tetramer-PE, CXCR6-PeDazzle594, CD69-PE-Cy7, CD45.2-SparkNIR685, CD62L-APCR700, TCRβ-APC-Cy7, CD38-APC-Fire810, granzyme-A-e450, TCF1-AF488, granzyme-B-APC, TBET-PE/Fire810. Antibodies used to generate the AT3-OVA CD45+ UMAP were as follows: CD45-PerCP, CD19-AF488, TCRγδ-PE-Cy7, NK1.1-BV570, CD3-Spark NIR685, TCRβ-BB700, CD8α-BUV395, CD8β-BUV615, CD4-BUV805, CD44-APCR700, CD62L-BUV737, CD69-PE, CD103-BV421, CD11b-BV711, CD11c-BV785, F480-BUV661, Ly6G-BV650, Ly6C-BV605, MHC-II-Spark Blue 550, SIGLEC-F-PerCP-Fire 806, XCR1-Spark UV387, CD64-PE-Fire 744, H-2Kb-OVA-tetramer-APC, CD39-PE-Dazzle594, PD1-RY586, PD-L1-RB780, LAG3-RB744, KLRG1-BV510, CD38-APCFire810, FOXP3-e450, TBET-PE/Fire810, TCF1-RB705 and CTLA-4-PE-Fire-640. A representative flow cytometry gating strategy of immune cells in murine tissue is shown in Extended Data Fig. 10b.
Human MyBrCa clinical dataset
The full cohort described previously28 was filtered to cases with available RNA-seq data, complete parity information and stage 1–3 diseases. Where patients had multiple synchronous tumours that underwent RNA-seq, one sample per patient was selected giving preference to more complete clinical metadata and basal-like subtyping. Parity was defined as at least one live birth, and pregnancy-associated cancers were not excluded. The ESTIMATE immune infiltration scores calculated in the MyBrCa publication were used unmodified. An exploratory analysis of the associations of sequencing technical factors and known clinicopathological predictors influencing immune infiltration was conducted. An effect of sequencing batch on the ESTIMATE score was observed. There was otherwise variable impact of age at diagnosis, grade and stage on the ESTIMATE score that was not consistent across subtypes. Linear models were fit for each PAM50 subtype with the ESTIMATE score as the response (non-standardized) and parity (yes or no), age at diagnosis (continuous), grade (grade 1/2 or grade 3) and stage (I, II or III) as covariates, using the lm function in R. Next, stepwise model selection using the akaike information criterion (AIC) was performed using the step function. Three cases without PAM50 and 20 cases that contained normal-like classification were excluded from the analysis. Selected covariates differed across subtypes, with parity being the only selected covariate for the basal-like subtype. The most parsimonious model was therefore considered to include parity (yes or no) and sequencing batch (1 or 2) as covariates, and this was used for the final analysis for all subtypes. The 95% CIs for the estimated co-efficient were calculated using the confint function in R (v.1.11.0). Use of the AIC selected models with more covariates specific to each subtype did not alter the conclusion about the influence of parity on immune infiltration in basal-like cases.
For the breastfeeding analysis, cases were divided into those with any history of breastfeeding regardless of duration (BF), no history of breastfeeding (no-BF) and nulliparous cases (N). Three-way ANOVA models were fit for each subtype with the ESTIMATE score as the response (non-standardized), breastfeeding status as the grouping variable (BF, no-BF or N) and sequencing batch (1 or 2) as a covariate as above using the aov function in R (v.1.11.0). Homogeneity of variances between groups in each subtype was confirmed with Bartlett’s test using the bartlett.test function in R. Departure from normality was tested using the Shapiro–Wilk test and examination of QQ plots. This showed a minor departure from normality for the basal-like group ESTIMATE scores, which was not considered detrimental as there are at least 30 cases in each group. Between group mean differences were tested with Tukey’s honest significant difference method using the TukeyHSD function in R, also reporting a 95% CI for the mean difference and P value adjusted for multiple comparisons with the Tukey–Kramer method.
For GSEA in the MyBrCa dataset, RNA-seq was conducted and processed to raw counts as described previously28. The RNA-seq counts were normalized using RUVIII56 to remove batch effects and other unwanted variations. We next performed differential expression analysis comparing parous and nulliparous basal samples as well as breastfeeding parous and non-breastfeeding parous samples using the R package limma47. GSEA was performed using the fgsea (v.1.30.0) library in R to detect gene set enrichment of the TRM gene signatures, outputting a normalized enrichment score (NES) and FDR.
The P values reported for subtype-specific analyses were not corrected for multiple testing. The rationale for this is that these subtypes display clearly distinct characteristics, and there is a large body of literature supporting the differential biology of these subtypes and the variable significance of immune infiltration between them. There is no reasonable expectation of equipoise regarding the influence of parity on immune infiltration across subtypes.
Human MyBrCa Immunohistochemistry
IHC-stained FFPE tissue sections were prepared according to a standard diagnostic workflow. Four slides per tumour sample were stained with primary antibodies targeting specific antigens: anti-CD3 (2GV6, predilute; Ventana Medical Systems), anti-CD8 (SP57, predilute; Ventana Medical Systems), with an additional slide stained with H&E. The antibody staining was conducted using a Roche Ventana Bench Mark XT Autostainer, and the stained slides were digitized using the Aperio AT2 whole-slide scanner (Leica Biosystems).
Semi-automated scoring of immune markers
For CD3+ and CD8+ T cell IHC scoring, Aperio Imagescope (Leica Biosystems) was used to view the scanned images and 50 annotation boxes, each of size 0.7 mm × 0.4 mm, were drawn within the borders of the invasive tumour region. The borders of the invasive tumour region were determined with reference to the H&E-stained slides. Next, the Pixel Positive v.9 algorithm implemented in Aperio Imagescope was used to calculate the number of positively stained pixels within the annotation boxes at a 0.16 colour saturation threshold. The number of positive pixels was then divided by the total number of pixels within the annotation boxes to obtain the proportion of the annotated area that was positively stained for CD3+ and CD8+ immune cell types.
Human MyBrCa data analysis
β regression was used to model the relationship between T cell density (represented as a proportion of stained to total pixels) and parity/breastfeeding, as implemented in the R package betareg (v.3.2-1). Exploratory analysis determined that age at diagnosis of cancer and tumour grade were associated with T cell density, and these were used as covariates in addition to parity/breastfeeding as predictor variable. The emmeans R package (v.1.11.0) was used to calculate the AME and CIs between groups.
Breastfeeding patient cohort
Patient data were acquired from kConFab57, including women diagnosed with breast cancer over the age of 18, with a known germline pathogenic variant who were diagnosed between the years of 1980 and 2022. Only patients who had hormone-receptor-negative primary breast cancers, had at least one full-term pregnancy and confirmed breastfeeding status (yes/no) were included for survival analysis. Patients who had subsequent pregnancies after breast cancer diagnosis were not included in the cohort. All relevant clinical data are shown in Supplementary Table 4. Data were not blinded. Breastfeeding duration was recorded based on total length across one or more children, where available.
Stromal TIL scoring
TILs were scored in accordance with the guidelines from TILs International Working Group as the percentage of infiltration of the tumoural stroma30 from digitized H&E slides. Slides were acquired where possible (n = 136 out of 270).
To account for potential confounding effects of patient age on stromal TIL percentages, we performed an age-adjusted analysis. Stromal TIL data were obtained from the updated dataset containing clinical annotations and pathological assessments of breast cancer patients. The remaining patients were stratified into defined breastfeeding duration groups (0, <6 months, 6–12 months and >12 months).
We adjusted the stromal TIL percentages for patient age using a linear regression model, fitting stromal TIL percentage as the dependent variable and age at diagnosis as the independent predictor. Adjusted stromal TIL values were derived from the residuals of this linear model, standardized to the mean intercept value, effectively removing linear age-related variation. A Kruskal–Wallis test was used to evaluate differences in age-adjusted stromal TIL percentages across breastfeeding groups.
Breastfeeding patient survival analysis
The analysis examined the association between breastfeeding and survival outcomes using both univariate and multivariate Cox proportional hazards models. Survival data were censored at 15 years, resulting in 45 observed events. Univariate analyses evaluated the direct effect of breastfeeding status on OS, while multivariate analyses controlled for potential confounders including patient age at breast cancer diagnosis, years from breast cancer diagnosis from last live birth (≤10 years versus >10 years), chemotherapy treatment (yes/no), and mastectomy (yes/no). Robust variance estimates were applied and the proportional hazards assumption was tested using the Schoenfeld residuals test.
PB-TRM signature and TILs correlation
We accessed the ht-seq gene counts from TCGA using the Broad Institute Firehose platform. The counts were normalized to remove library size effects using edgeR58. The PB-TRM signature scores were computed using the genefu package46 in R. Stromal TILs were scored for each TCGA case by a pathologist (R.S.) using whole-slide H&E images according to established methods30. PB-TRM signature scores and TILs were correlated using the Spearman’s correlation coefficient in R.
PB-TRM signature score and survival
We accessed the normalized microarrays from the METABRIC study31 using cBioportal59 and computed the PB-TRM signature scores using genefu (v.2.36.0)46. The samples were stratified into two groups around the median signature score (high versus low) and Kaplan–Meier survival estimates were compared between groups using the log-rank test. Hazard ratios were also calculated in R using the PB-TRM signature scores by fitting a Cox proportional-hazards regression model.
Statistics, codes and reproducibility
All statistical analyses and the associated information relating error bars, box plots and sample size used to determine statistical significance are provided in the figure legends, and the statistical tools used are given in respective methods section. For Fig. 1b, the proportions of CD45+, CD3+ and CD8+ T cells over total cells were provided, and CD69+CD103– and CD69+CD103+ T cells over total CD45RA−CCR7−CD8+ T cells (y axes) were determined by flow cytometry analysis of normal breast of women with high breast cancer risk. For box plots, the horizontal bar shows the median, the box limits show the IQR and the whiskers extend to the most extreme dot point within 1.5 × IQR, and points beyond show outliers. Multiplex OPAL microscopy data associated with representatives in Fig. 1c were also verified by flow cytometry and OPAL verification cohort were repeated n = 5 per group in independent (N versus P) human breast tissue samples with comparable results; single-colour-stained images with markers and the collated data are provided in Extended Data Fig. 2a,b. All experiments were reproduced twice, and the results represent two combined biologically independent experiments unless specified. Breastfeeding patient cohort statistical information and statistical analyses of human data were conducted using R v.4. For mouse data, we used GraphPad Prism v.9. Violin plots with error bars representing the IQR were used to compare immune cell proportions. No codes or software tools were specifically developed for this study. Plots were produced using the ggplot2 R package (v.3.5.1), and data input and processing were performed with dplyr v.1.1.2, tidyr v.1.3.0, readxl v.1.4.3, readr v.2.1.4 and magrittr v.2.0.3.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Processed and raw bulk RNA-seq counts resulting from FACS-sorted CD69+CD103+CD8+ T cells used to generate the PB-TRM signature can be accessed at the GEO (GSE271307). scRNA-seq data and all associated cell annotations are available from the original publication16. Code associated with analyses can be accessed at Zenodo60 (https://doi.org/10.5281/zenodo.17120517). Data associated with human normal breast and murine experimental analyses are available as source data. Clinical metadata and sequencing data for the human MyBrCa dataset are accessible with permission as per the original publication28. Human breastfeeding data with associated OS and TIL scores are provided by kConFab. As this is an ongoing prospective cohort, the ethics pertaining to this data requires that data access be approved via the process described at https://www.kconfab.org/Data%20Access/AppProcess.html. using the contact details provided at https://www.kconfab.org/contact_us.html. The corresponding author (S.L.) can assist requesting parties in facilitating applications to access the data. Responses are provided within a 2-month time frame. All other data supporting the findings of this study are provided in the Supplementary Information. Source data are provided with this paper.
References
Jung, A. Y. et al. Distinct reproductive risk profiles for intrinsic-like breast cancer subtypes: pooled analysis of population-based studies. J. Natl Cancer Inst. 114, 1706–1719 (2022).
Collaborative Group on Hormonal Factors in Breast Cancer. Breast cancer and breastfeeding: collaborative reanalysis of individual data from 47 epidemiological studies in 30 countries, including 50302 women with breast cancer and 96973 women without the disease. Lancet 360, 187–195 (2002).
Feigman, M. J. et al. Pregnancy reprograms the epigenome of mammary epithelial cells and blocks the development of premalignant lesions. Nat. Commun. 11, 2649 (2020).
D’Cruz, C. M. et al. Persistent parity-induced changes in growth factors, TGF-β3, and differentiation in the rodent mammary gland. Mol. Endocrinol. 16, 2034–2051 (2002).
Russo, J., Moral, R., Balogh, G. A., Mailo, D. & Russo, I. H. The protective role of pregnancy in breast cancer. Breast Cancer Res. 7, 131–142 (2005).
Islami, F. et al. Breastfeeding and breast cancer risk by receptor status-a systematic review and meta-analysis. Ann. Oncol. 26, 2398–2407 (2015).
Palmer, J. R. et al. Parity, lactation, and breast cancer subtypes in African American women: results from the AMBER Consortium. J. Natl Cancer Inst. 106, dju237 (2014).
Harris, M. A. et al. Towards targeting the breast cancer immune microenvironment. Nat. Rev. Cancer 24, 554–577 (2024).
Savas, P. et al. Clinical relevance of host immunity in breast cancer: from TILs to the clinic. Nat. Rev. Clin. Oncol. 13, 228–241 (2016).
Virassamy, B. et al. Intratumoral CD8+ T cells with a tissue-resident memory phenotype mediate local immunity and immune checkpoint responses in breast cancer. Cancer Cell 41, 585–601 (2023).
Savas, P. et al. Single-cell profiling of breast cancer T cells reveals a tissue-resident memory subset associated with improved prognosis. Nat. Med. 24, 986–993 (2018).
Hitchcock, J., Hughes, K., Pensa, S., Lloyd-Lewis, B. & Watson, C. J. The immune environment of the mammary gland fluctuates during post-lactational regression and correlates with tumour growth rate. Development 149, dev200162 (2022).
Dawson, C. A. et al. Tissue-resident ductal macrophages survey the mammary epithelium and facilitate tissue remodelling. Nat. Cell Biol. 22, 546–558 (2020).
Hanasoge Somasundara, A. V. et al. Parity-induced changes to mammary epithelial cells control NKT cell expansion and mammary oncogenesis. Cell Rep. 37, 110099 (2021).
Kumar, T. et al. A spatially resolved single-cell genomic atlas of the adult human breast. Nature 620, 181–191 (2023).
Reed, A. D. et al. A single-cell atlas enables mapping of homeostatic cellular shifts in the adult human breast. Nat. Genet. 56, 652–662 (2024).
Corral, D. et al. Mammary intraepithelial lymphocytes promote lactogenesis and offspring fitness. Cell 188, 1662–1680 (2025).
Watanabe, M. et al. Changes in T, B, and NK lymphocyte subsets during and after normal pregnancy. Am. J. Reprod. Immunol. 37, 368–377 (1997).
Mackay, L. K. et al. Long-lived epithelial immunity by tissue-resident memory T (TRM) cells in the absence of persisting local antigen presentation. Proc. Natl Acad. Sci. USA 109, 7037–7042 (2012).
Snyder, M. E. et al. Generation and persistence of human tissue-resident memory T cells in lung transplantation. Sci. Immunol. 4, eaav5581 (2019).
Santucci-Pereira, J. et al. Genomic signature of parity in the breast of premenopausal women. Breast Cancer Res. 21, 46 (2019).
Cansever, D. et al. Lactation-associated macrophages exist in murine mammary tissue and human milk. Nat. Immunol. 24, 1098–1109 (2023).
Mackay, L. K. et al. T-box transcription factors combine with the cytokines TGF-beta and IL-15 to control tissue-resident memory T cell fate. Immunity 43, 1101–1111 (2015).
Casey, K. A. et al. Antigen-independent differentiation and maintenance of effector-like resident memory T cells in tissues. J. Immunol. 188, 4866–4875 (2012).
Di Pilato, M. et al. CXCR6 positions cytotoxic T cells to receive critical survival signals in the tumor microenvironment. Cell 184, 4512–4530 (2021).
Martinson, H. A., Jindal, S., Durand-Rougely, C., Borges, V. F. & Schedin, P. Wound healing-like immune program facilitates postpartum mammary gland involution and tumor progression. Int. J. Cancer 136, 1803–1813 (2015).
Pan, J. W. et al. The molecular landscape of Asian breast cancers reveals clinically relevant population-specific differences. Nat. Commun. 11, 6433 (2020).
Pan, J. W. et al. Clustering of HR+/HER2− breast cancer in an Asian cohort is driven by immune phenotypes. Breast Cancer Res. 26, 67 (2024).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Salgado, R. et al. The evaluation of tumor-infiltrating lymphocytes (TILs) in breast cancer: recommendations by an International TILs Working Group 2014. Ann. Oncol. 26, 259–271 (2015).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Liu, J. et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400–416 (2018).
Plaks, V. et al. Adaptive immune regulation of mammary postnatal organogenesis. Dev. Cell 34, 493–504 (2015).
Galati, F. et al. Pregnancy-associated breast cancer: a diagnostic and therapeutic challenge. Diagnostics 13, 604 (2023).
Amant, F. et al. The definition of pregnancy-associated breast cancer is outdated and should no longer be used. Lancet Oncol. 22, 753–754 (2021).
Murphy, T. L. & Murphy, K. M. Dendritic cells in cancer immunology. Cell. Mol. Immunol. 19, 3–13 (2022).
Jaquish, A. et al. Mammary intraepithelial lymphocytes and intestinal inputs shape T cell dynamics in lactogenesis. Nat. Immunol. 26, 1411–1422 (2025).
Im, S. A. et al. Results from the randomized KEYNOTE-355 study of pembrolizumab plus chemotherapy for Asian patients with advanced TNBC. NPJ Breast Cancer 10, 79 (2024).
Wang, H. et al. The microbial metabolite trimethylamine N-oxide promotes antitumor immunity in triple-negative breast cancer. Cell Metab. 34, 581–594 (2022).
Cereser, B. et al. The mutational landscape of the adult healthy parous and nulliparous human breast. Nat. Commun. 14, 5136 (2023).
Lowenfeld, L. et al. Dendritic cell vaccination enhances immune responses and induces regression of HER2pos DCIS independent of route: results of randomized selection design trial. Clin. Cancer Res. 23, 2961–2971 (2017).
Strickley, J. D. et al. Immunity to commensal papillomaviruses protects against skin cancer. Nature 575, 519–522 (2019).
Datta, J. et al. Progressive loss of anti-HER2 CD4+ T-helper type 1 response in breast tumorigenesis and the potential for immune restoration. Oncoimmunology 4, e1022301 (2015).
Chakraborty, B. et al. Estrogen receptor signaling in the immune system. Endocr. Rev. 44, 117–141 (2023).
Coles, C. E. et al. The Lancet Breast Cancer Commission. Lancet 403, 1895–1950 (2024).
Gendoo, D. M. A. et al. Genefu: an R/Bioconductor package for computation of gene expression-based signatures in breast cancer. Bioinformatics 32, 1097–1099 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2021).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Alquicira-Hernandez, J. & Powell, J. E. Nebulosa recovers single-cell gene expression signals by kernel density estimation. Bioinformatics 37, 2485–2487 (2021).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Bouchard, G. et al. Stimulation of triple negative breast cancer cell migration and metastases formation is prevented by chloroquine in a pre-irradiated mouse model. BMC Cancer 16, 361 (2016).
Beavis, P. A. et al. Dual PD-1 and CTLA-4 checkpoint blockade promotes antitumor immune responses through CD4+Foxp3− cell-mediated modulation of CD103+ dendritic cells. Cancer Immunol. Res. 6, 1069–1081 (2018).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the \({2}^{-{\rm{\Delta \Delta }}{C}_{{\rm{T}}}}\) method. Methods 25, 402–408 (2001).
Molania, R. et al. Removing unwanted variation from large-scale RNA sequencing data with PRPS. Nat. Biotechnol. 41, 82–95 (2023).
Thorne, H., Mitchell, G. & Fox, S. kConFab: a familial breast cancer consortium facilitating research and translational oncology. J. Natl Cancer Inst. Monogr. 2011, 79–81 (2011).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2009).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Science Signal. 6, pl1 (2013).
Virassamy, B. Code for ‘Parity and lactation induce T-cell-mediated breast cancer protection’. Zenodo https://doi.org/10.5281/zenodo.17120517 (2025).
Acknowledgements
We thank the staff at the Peter MacCallum Cancer Centre Flow Cytometry Facility, A. Lee, S. Bencraig and T. Semple from the Molecular Genomics Core, the Animal Core Facility and N. Anthony from the Centre for Advanced Histology and Microscopy at the Peter MacCallum Cancer Centre; members of the Australian genome research facility (AGRF) for their assistance with RNA-seq; R. Young and D. Byrne for the OPAL microscopy reagents and histology sectioning support; C. Mintoff, M. Neeson, T. Tan and V. Setiono for assistance with human breast tissue processing; H. Makmur and M. Khairi from Cancer Research Malaysia for their assistance with compiling the clinical and digital pathology data from the human MyBrCa cohort; M. Pereira for providing D2A1 cells; the Kconfab team members G. Glavich, A. Smith, L. Mokdsi and E. Niedermayr for their ongoing support; and J. Parrodi for her administrative support. This work was supported by the Mark Foundation ASPIRE II Award (5820526561), Australia, S.L. is supported by the National Breast Cancer Foundation of Australia (NBCF) (EC-17-001), the Breast Cancer Research Foundation, New York (BCRF-21-102) and a National Health and Medical Council of Australia (NHMRC) Investigator Grant (1162318), and the Peter MacCallum Cancer Centre Foundation; P.S. by a National Health and Medical Council of Australia (NHMRC) Investigator Grant (2017477) and the Peter MacCallum Cancer Centre Foundation; B.V. by a Postdoctoral Research Fellowship from Cancer Council Victoria and the Department of Health, Victoria, Australia; and P.K.D. by a National Health and Medical Research Council Principal Research Fellowship (1136680). L.K.M. is a Senior Medical Research Fellow supported by the Sylvia and Charles Viertel Charitable Foundation.
Author information
Authors and Affiliations
Consortia
Contributions
B.V., F.C., P.S. and M.A.H. performed primary human and mouse experiments, analysed results, coordinated the project and wrote the research manuscript. F.C., P.S., J.W. and J.-W.P. generated programming scripts and analysed the bioinformatics results. E.B. and D.J.C. performed mouse experiments, dosing, tumour cell injections, associated procedures and maintenance of mouse models. M.M.R.O., C.T.v.G., M.H., T.N.B. and S.S. supported with interpretation of results and performed experiments. J.K., J.D.B., L.E.L.G., K.C., S.T., H.X.A.Y., R.I. and M.J. supported with experiments. kConFab Investigators contributed to the recruitment of participants and coordinated biospecimen acquisition, clinical data curation and governance. H.T. from KConFab contributedto recruitment of participants, biospecimen acquisition and clinical data curation. R.S., H.T., C.P., S.N., T.P.S., J.V., P.J.N., P.K.D. and L.K.M. provided scientific expertise. All of the authors approved the final research manuscript. S.L. as senior author conceived, led and coordinated the project, and provided the project funding.
Corresponding authors
Ethics declarations
Competing interests
P.S. has acted as a consultant (not compensated) to Roche-Genentech and received research funding to his institution from Roche-Genentech. S.L. receives research funding to her institution from Novartis, Bristol Meyers Squibb, Merck, Puma Biotechnology, Eli Lilly, Nektar Therapeutics, Astra Zeneca, Roche-Genentech and Seattle Genetics; has acted as a consultant to Seattle Genetics, Novartis, Bristol Meyers Squibb, Merck, AstraZeneca, Eli Lilly, Roche-Genentech, Aduro Biotech, GlaxoSmithKline, Silverback Therapeutics, G1 Therapeutics, PUMA Biotechnologies, Pfizer, Gilead Therapeutics, Daiichi Sankyo, Amuni and Tallac Therapeutics. P.K.D. receives research funding from Myeloid Therapeutics, Prescient Therapeutics and Bristol-Myers Squibb. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Sandra Demaria, Matteo Lambertini and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cell types by parity status in cancer unaffected human breast tissue.
a-e, Abundance of immune cell types quantified from scRNAseq by parity status in human normal breast tissue shown as ratio of cell type in women of average breast cancer risk. Cell types are annotated as per ref. 16. Y-axes indicate cell type frequency as calculated by cell subpopulations as indicated over total epithelial cells. a, Macrophages, n = 170 women (N, n = 53) vs (P, n = 117), M1 and M2 macrophage cell sub-populations, n = 105 women (N, n = 30) vs (P, n = 75). b, Monocytes, n = 105 women (N, n = 30) vs (P, n = 75). c, Dendritic cells n = 124 women (N, n = 37) vs (P, n = 87). d, B cells n = 123 women (N, n = 37) vs (P, n = 86). e, Fibroblasts, n = 170 women (N, n = 53) vs (P, n = 117), Fibroblast matrix associated cells, n = 102 women, (N, n = 27) vs (P, n = 75) and Fibroblast SFRP4+ subpopulations, n = 90 women (N, n = 25) vs (P, n = 65). Violin plots show data distribution with boxplots indicate IQR extension from Q1 to Q3 and median as a white bar, whiskers show minimum and maximum within 1.5 times the IQR and datapoints show outliers. Two-sided Wilcoxon rank sum test, unadjusted exact p-values are shown.
Extended Data Fig. 2 T cells in cancer unaffected human normal breast.
a, Representative OPAL multiplex immunofluorescence single-colour and merged images of cancer unaffected normal breast tissue from nulliparous and parous women stained for DAPI, pan-cytokeratin (AE1/AE3) CD3, CD8, CD69, and CD103. Scale bar is defined in the panels. b, Quantification of OPAL immunofluorescence in cancer unaffected human normal breast (n = 5 per group) for T cells subpopulations in mammary regions indicating density of CD3+, CD8+ and CD69+CD103+CD8+CD3+ T cells per mm2 from nulliparous (N) and parous(P) women. Data in graphs represent mean ± s.e.m, two-sided Wilcoxon rank-sum test and dots points represent indicated immune cell quantities from independent normal breast tissue samples. c, Immune cell abundance (right) and enrichment of PB-TRM gene signature (left) in the TEM and TRM-like cell clusters as annotated by originating publication, ref. 16. Effector CD8+ T cells (n = 26,333 cells) from nulliparous vs parous women. Wilcoxon rank-sum test (two-sided) and unadjusted p-value is shown.
Extended Data Fig. 3 Immune cell subsets in healthy murine mammary tissue.
a, Frequency of indicated immune cell subsets among all CD45+ cells from the MFP of virgin (n = 9) and d28-inv (n = 12) C57BL/6 mice. b, Number of indicated immune cell subsets per gram of MFP from virgin (n = 9) and d28-inv (n = 12) C57BL/6 mice. c, Number of indicated immune cell subpopulations per gram of MFP from virgin (n = 8) and d10-FW (n = 8) BALB/c mice. Data in graphs represent mean ± s.e.m, results represent two combined independent experiments. Statistical significance determined by two-sided Mann-Whitney test and exact p values are shown.
Extended Data Fig. 4 CD8+ T cell sub-populations in healthy murine MFP.
a, Top: UMAP of merged CD8α+ T cells generated by flow cytometry data from the MFP of virgin (n = 8) and d28-inv (n = 9) C57BL/6 mice with three enriched clusters (C1, C2, C3) as indicated. Each dot represents an individual cell. Feature plots showing expression of indicated markers projected on UMAP of CD8α+ T cells are presented in the bottom layout.
Extended Data Fig. 5 CD8+TRM-like cells in healthy murine MFP.
a, Relative expression of indicated expressed gene transcripts relative to GAPDH control isolated from MFP from virgin (n = 10) vs d28-inv (n = 9) timepoints from C57BL/6 mice. b, Representative OPAL multiplex images indicating the presence of DAPI+, CD3+, CD8+, CD103+ cells (white dots, depicts the co-expression of CD3, CD8 and CD103 surface markers on T cells) in the MFP of virgin, d10-FW and D28-inv C57BL/6 mice. Blue regions indicate local mammary draining lymph node and yellow represent mammary fat pad tissue boundary. c, Total number of CD3+CD8+CD103+ T cells in the 4th MFP of virgin (n = 7), d10-FW (n = 4) and d28-inv (n = 6) C57BL/6 mice measured by OPAL microscopy per mammary fat pad section. d, Representative OPAL images indicate the single-colour staining of DAPI, E-cadherin (ECAD), CD3, CD8 and CD103 on T cells in the MFP of virgin, d10-FW and D28-inv C57BL/6 mice. e, Mean distance between ECAD+ cells and the closest CD3+ CD8+ CD103+ T cells of virgin (n = 7), d10-FW (n = 4) and d28-inv (n = 6) C57BL/6 mice. Data in graphs represent mean ± s.e.m, results represent two combined independent experiments. Statistical significance determined by two sided Mann-Whitney test (a) or Kruskal-Wallis test (c, e). Exact p values are shown.
Extended Data Fig. 6 Immune cell infiltration in TNBC murine mammary tumours.
a, Numbers of indicated immune cell subpopulations per gram of AT3-OVA tumour from virgin (n = 9) and d28-inv (n = 10) C57BL/6 mice. b, Numbers of indicated immune cell subpopulations per gram of AT3-OVA tumour from virgin (n = 8) and d10-FW (n = 8) C57BL/6 mice. c, Numbers of indicated immune cell subpopulations per gram of D2A1 tumour from virgin (n = 8) and d10-FW (n = 8) BALB/c mice. d, Numbers of indicated immune cell subpopulations per gram of D2A1 tumour from virgin (n = 6) and d28-inv (n = 7) BALB/c mice. Data in graphs represent mean ± s.e.m, results represent two combined independent experiments. Statistical significance determined by two-sided Mann-Whitney test. Exact p values are shown.
Extended Data Fig. 7 TNBC tumour growth and immune infiltration in murine models.
a, Tumour growth (left) and endpoint tumour volume (right) of AT3-OVA cells in the 4th MFP of RAG2−/−γc−/− mice pre-inoculated with naive OT-I cells seven days prior to mating in d10-FW (n = 4) and age-matched virgin (n = 4) control mice. b, Numbers of indicated immune cell populations in RAG2−/−γc−/− mice pre-inoculated with effector OT-I cells seven days prior to mating in d28-inv (n = 10) and age-matched virgin (n = 10) control mice. c, Numbers of indicated immune cell subpopulations per gram of AT3-OVA tumour from virgin (n = 6) and d10-FW (n = 6) RAG2−/−γc−/− mice at four weeks post tumour cell injection. d, Tumour growth (left) and endpoint tumour burden (right) of AT3-OVA cells in the 4th MFP of RAG−/−1−/− mice. Mice were injected with 20 × 106 effector gBT-I cells or PBS control eight weeks prior to injection of AT3-OVA cells (n = 5 per group for tumour growth and 10 for endpoint weight). Data in graphs represent mean ± s.e.m. Results are representative of n = 2 independent experiments (a, d left) or two combined independent experiments (b-d right). Statistical significance determined by two-sided Mann-Whitney test. Exact p values are shown.
Extended Data Fig. 8 T cell depletions and TIL differences in TNBC murine models.
a, Number of indicated immune cells per gram of AT3-OVA tumour from virgin mice treated with isotype control(n = 16), anti-CD4 (n = 12), anti-CD8α (n = 9), anti-CD8β(n = 12) or anti-CD4/CD8a (n = 14). b, Number of indicated immune cells per gram of AT3-OVA tumour from d28-inv mice treated with isotype control (n = 16), anti-CD4 (n = 13), anti-CD8α (n = 9), anti-CD8β (n = 14) or combined anti-CD4/CD8α (n = 14). c, AT3-OVA endpoint tumour weight in C57BL/6 mice in virgin (n = 16) and d28-inv (n = 16) isotype treated controls. d, Number of indicated immune cells per gram of AT3-OVA tumour from d28-inv virgin (n = 16) and d28-inv (n = 16) mice treated with isotype control. e, Representative UMAP of 20,000 CD45+ immune cells by flow cytometry from AT3-OVA tumours of C57BL/6 d28-inv isotype control and CD4 depleted mice with two enriched clusters (C1,C2) indicated and CD4+ T cells highlighted. Each dot represents an individual cell. f, UMAP plots of indicated CD45+ cells from AT3-OVA tumours in d28-inv treated with isotype, anti-CD4, anti-CD8α, anti-CD8β or anti-CD4/CD8α. g, Frequency of cluster 1 among all CD45+ cells in AT3-OVA tumours from d28-inv isotype (n = 8) and CD4 depleted (n = 7) C57BL/6 mice. h, Relative expression of indicated markers in cluster 1 relative to all CD45+ cells from d28-inv isotype (n = 8). i, Frequency of cluster 2 among all CD45+ in AT3-OVA tumours from d28-inv isotype (n = 8) and CD4 depleted(n = 7) C57BL/6 mice. j, Relative expression of indicated markers in cluster 2 relative to all CD45+ from d28-inv isotype control mice (n = 8). k, Number of indicated immune cells per gram of AT3-OVA tumour from d28-inv mice treated with isotype (n = 16) or anti-CD4 (n = 13). l, Representative images of AT3-OVA tumours stained with Masaon’s trichrome. Arrows indicate positive green staining of collagen in virgin (left) and d28-inv (right) tissue. m, Proportion of Masson’s trichrome staining in AT3-OVA tumours from virgin (n = 6) and d28-inv (n = 5) C57BL/6 measured by Masson’s trichrome staining. Two combined independent experiments (a-c,k-m) representative of two independent experiments (e-j). mean ± s.e.m, two-sided Mann-Whitney test. Exact p values are shown.
Extended Data Fig. 9 Parity, breastfeeding and TIL associations in human breast cancer subtypes.
a, Breastfeeding cohort multivariate analysis with clinical prognostic factors shown (n = 270). OS events were censored at 15 years follow up and adjusted for breastfeeding status (BF), age at diagnosis (Age.dx), years since last live birth to BC diagnosis (time since last birth), and treatment with chemotherapy (treatment_chemo) or mastectomy (treatment_mastectomy). Hazard ratios are presented with 95% confidence intervals, shown as squares and horizontal lines, respectively. b, PB-TRM signature correlation with tumour infiltrating lymphocyte (TIL) counts in TNBC (Basal subtype, n = 136) from the TCGA dataset. R value indicate Spearman’s correlation coefficient (two-sided p value from correlation test). c, Enrichment of the PB-TRM signature in the indicated BC subtypes from the Metabric dataset, Basal (n = 329), HER2 (n = 240), LumA (n = 718) and LumB (n = 488). Kruskal Wallis test, p-value shown is unadjusted, d, Kaplan–Meier survival analysis indicates disease-free survival and e, overall survival from (n = 329) primary basal-like/TNBCs with prognostic separation according to PB-TRM signature from parous normal breast tissue. Log-rank, p-values and hazard ratios are shown. f, Intratumoural T cell density in HER2 and Luminal BC subtypes determined by immunohistochemistry for CD8+ and CD3+ in MyBrCa dataset, comparing nulliparous (N) and parous (P) women with differing breastfeeding histories prior to cancer diagnosis (bf: any recorded breastfeeding activity). CD8+ and CD3+ T cell density quantified as ratio of stain-positive pixels to all pixels within tumour margins, represented as a percentage. Modelled with beta regression adjusting for covariates of age at diagnosis and tumour grade. Unadjusted p values with the AME for each group compared to N are presented. Box plots: horizontal bar shows median, hinges represent IQR, and whiskers extend to the most extreme dot point within 1.5xIQR, and points beyond show outliers. Individual points show data from each case, coloured according to the status shown on the x-axis. Exact p values are shown.
Supplementary information
Supplementary Information (download PDF )
Guide to Supplementary Tables 1–4.
Supplementary Tables (download XLSX )
Supplementary Tables 1–4.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Virassamy, B., Caramia, F., Savas, P. et al. Parity and lactation induce T-cell-mediated breast cancer protection. Nature 649, 449–459 (2026). https://doi.org/10.1038/s41586-025-09713-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09713-5