Abstract
Stem cell-derived embryo models could greatly facilitate our understanding of embryonic development. Although human and monkey embryo models have reached early gastrulation stage1,2,3,4,5,6,7, the development of robust models beyond this time remains to be accomplished8. Here, using an optimized 3D suspension culture system, we have successfully advanced the in vitro culture of a stem cell-derived monkey blastoid to day 25. Morphological and histological analyses showed that these monkey embryoids underwent gastrulation and largely recapitulated key developmental events of the late gastrulation stage observed in vivo, with the appearance of a neural plate, haematopoietic system, allantois, primitive gut, primordial germ cells, yolk sac structures and progenitors of other organs, excluding trophoblast derivatives. Single-cell transcriptomic analyses revealed that the lineage composition and differentiation trajectories of cells in these monkey embryoids were similar to those found in natural embryos during gastrulation. Thus, this primate stem cell-derived embryo model provides a valuable platform for dissecting the mechanisms of primate embryonic development from blastocyst to late gastrulation stage.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Customizable visualizations and analyses of our data can be executed at https://jiefrankli.shinyapps.io/monkey_embryoid_scexp/. The raw scRNA-seq data of monkey embryoids generated in this study have been deposited in the Genome Sequence Archive (https://ngdc.cncb.ac.cn/gsa) with the accession number CRA032070. The dataset of monkey blastoid IVC (D17, 2D) is available at the Gene Expression Omnibus (GEO) under accession no. GSE218375. The datasets of natural post-implantation cynomolgus monkey embryos and prolonged in vitro-cultured natural blastocysts are available at the GEO under accession nos. GSE74767 and GSE130114, respectively. The dataset of human CS7 embryos used as reference is available at Array Express under accession no. E-MTAB-9388. The dataset of natural monkey embryos (CS8–9) is available at the GEO under accession no. GSE193007. The dataset of in vitro-cultured natural monkey embryos (18–25 days post-fertilization) is available at the GEO under accession no. GSE207534, and the processed gene expression matrix was downloaded from Figshare (https://doi.org/10.6084/m9.figshare.22146638). The dataset of the human PCW3 embryo is available at the GEO under accession no. GSE155121. The dataset of the human endoderm is available at the Genome Sequence Archive under accession no. HRA000280. All raw image data and 3D morphological data generated by LiTone XL light-sheet microscope have been deposited in the BioImage Archive (https://www.ebi.ac.uk/bioimage-archive/, under the accession number: S-BIAD2201). All the monkey cell lines used in this study have been deposited to the National Stem Cell Resource Center of China with certificates of authentication (http://www.nscrc.cn/xbzy/zylb/frxb/dngxb/202309/t20230914_173915.html) and are available from the corresponding authors with a completed Materials Transfer Agreement. Source data are provided with this paper.
Code availability
This paper does not report original code. All custom codes to process the scRNA-seq data are available on GitHub (https://github.com/JieBest/MonkeyEmbryoidscRNA-seq) and they are not central to the conclusions of the paper.
References
Karvas, R. M. et al. 3D-cultured blastoids model human embryogenesis from pre-implantation to early gastrulation stages. Cell Stem Cell 30, 1148–1165.e7 (2023).
Oldak, B. et al. Complete human day 14 post-implantation embryo models from naïve ES cells. Nature 622, 562–573 (2023).
Liu, L. et al. Modeling post-implantation stages of human development into early organogenesis with stem-cell-derived peri-gastruloids. Cell 186, 3776–3792.e16 (2023).
Weatherbee, B. A. T. et al. A model of the post-implantation human embryo derived from pluripotent stem cells. Nature https://doi.org/10.1038/s41586-023-06368-y (2023).
Pedroza, M. et al. Self-patterning of human stem cells into post-implantation lineages. Nature 622, 574–583 (2023).
Ai, Z. et al. Dissecting peri-implantation development using cultured human embryos and embryo-like assembloids. Cell Res. 33, 661–678 (2023).
Li, J. et al. Cynomolgus monkey embryo model captures gastrulation and early pregnancy. Cell Stem Cell 30, 362–377.e7 (2023).
Ghimire, S., Mantziou, V., Moris, N. & Martinez Arias, A. Human gastrulation: the embryo and its models. Dev. Biol. 474, 100–108 (2021).
Gong, Y. et al. Ex utero monkey embryogenesis from blastocyst to early organogenesis. Cell 186, 2092–2110.e23 (2023).
Zhai, J. et al. Neurulation of the cynomolgus monkey embryo achieved from 3D blastocyst culture. Cell 186, 2078–2091.e18 (2023).
Zhai, J. et al. Primate gastrulation and early organogenesis at single-cell resolution. Nature 612, 732–738 (2022).
Bergmann, S. et al. Spatial profiling of early primate gastrulation in utero. Nature 609, 136–143 (2022).
Cui, G. et al. Spatial and molecular anatomy of germ layers in the gastrulating cynomolgus monkey embryo. Cell Rep. 40, 111285 (2022).
Zeng, B. et al. The single-cell and spatial transcriptional landscape of human gastrulation and early brain development. Cell Stem Cell 30, 851–866.e7 (2023).
Xu, Y. et al. A single-cell transcriptome atlas profiles early organogenesis in human embryos. Nat. Cell Biol. 25, 604–615 (2023).
Tyser, R. C. V. et al. Single-cell transcriptomic characterization of a gastrulating human embryo. Nature 600, 285–289 (2021).
Cui, L. et al. Spatial transcriptomic characterization of a Carnegie stage 7 human embryo. Nat. Cell Biol. 27, 360–369 (2025).
Xiao, Z. et al. 3D reconstruction of a gastrulating human embryo. Cell 187, 2855–2874.e19 (2024).
Yuan, Y. et al. 3D reconstruction of a human Carnegie stage 9 embryo provides a snapshot of early body plan formation. Cell Stem Cell 32, 1006–1024.e5 (2025).
Clark, A. T. et al. Human embryo research, stem cell-derived embryo models and in vitro gametogenesis: considerations leading to the revised ISSCR guidelines. Stem Cell Rep. 16, 1416–1424 (2021).
Shahbazi, M. N., Siggia, E. D. & Zernicka-Goetz, M. Self-organization of stem cells into embryos: a window on early mammalian development. Science 364, 948–951 (2019).
Lewis, J. et al. Developmental and stem cell biology’s bright future. Cell 187, 3224–3228 (2024).
Wu, J. & Fu, J. Toward developing human organs via embryo models and chimeras. Cell 187, 3194–3219 (2024).
Tarazi, S. et al. Post-gastrulation synthetic embryos generated ex utero from mouse naïve ESCs. Cell 185, 3290–3306.e25 (2022).
Amadei, G. et al. Embryo model completes gastrulation to neurulation and organogenesis. Nature 610, 143–153 (2022).
Lau, K. Y. C. et al. Mouse embryo model derived exclusively from embryonic stem cells undergoes neurulation and heart development. Cell Stem Cell 29, 1445–1458.e8 (2022).
Li, H. et al. A complete model of mouse embryogenesis through organogenesis enabled by chemically induced embryo founder cells. Cell 188, 5912–5930.e20 (2025).
O'Rahilly, R. & Müller, F. Developmental Stages in Human Embryos (Carnegie Institution Of Washington, 1987).
Nakamura, T. et al. A developmental coordinate of pluripotency among mice, monkeys and humans. Nature 537, 57–62 (2016).
Sasaki, K. et al. The germ cell fate of cynomolgus monkeys is specified in the nascent amnion. Dev. Cell 39, 169–185 (2016).
Ma, H. et al. In vitro culture of cynomolgus monkey embryos beyond early gastrulation. Science 366, eaax7890 (2019).
Aguilera-Castrejon, A. et al. Ex utero mouse embryogenesis from pre-gastrulation to late organogenesis. Nature 593, 119–124 (2021).
Bedzhov, I., Leung, C. Y., Bialecka, M. & Zernicka-Goetz, M. In vitro culture of mouse blastocysts beyond the implantation stages. Nat. Protoc. 9, 2732–2739 (2014).
Ee, L. S. et al. Enhancer remodeling by OTX2 directs specification and patterning of mammalian definitive endoderm. Dev. Cell https://doi.org/10.1016/j.devcel.2025.07.020 (2025).
O'Rahilly, R. & Muller, F. Neurulation in the normal human embryo. Ciba Found. Symp. 181, 70–82 (1994).
Zhang, X. et al. Pax6 is a human neuroectoderm cell fate determinant. Cell Stem Cell 7, 90–100 (2010).
Sheng, G., Martinez Arias, A. & Sutherland, A. The primitive streak and cellular principles of building an amniote body through gastrulation. Science 374, abg1727 (2021).
Thomas P. Q., Brown, A. & Beddington, R. S. Hex: a homeobox gene revealing peri-implantation asymmetry in the mouse embryo and an early transient marker of endothelial cell precursors. Development 125, 85–94 (1998).
Silberg, D. G., Swain, G. P., Suh, E. R. & Traber, P. G. Cdx1 and cdx2 expression during intestinal development. Gastroenterology 119, 961–971 (2000).
Li, L. C. et al. Single-cell patterning and axis characterization in the murine and human definitive endoderm. Cell Res. 31, 326–344 (2021).
Han, L. et al. Single cell transcriptomics identifies a signaling network coordinating endoderm and mesoderm diversification during foregut organogenesis. Nat. Commun. 11, 4158 (2020).
Lawson, K. A. et al. Bmp4 is required for the generation of primordial germ cells in the mouse embryo. Genes Dev. 13, 424–436 (1999).
Kobayashi, T. et al. Principles of early human development and germ cell program from conserved model systems. Nature 546, 416–420 (2017).
Hancock, G. V., Wamaitha, S. E., Peretz, L. & Clark, A. T. Mammalian primordial germ cell specification. Development 148, dev189217 (2021).
Schule, K. M. et al. Eomes restricts Brachyury functions at the onset of mouse gastrulation. Dev. Cell 58, 1627–1642.e7 (2023).
Dias, A. & Martinez Arias, A. Laying out the mammalian body plan to a T. Dev. Cell 58, 1625–1626 (2023).
Amin, S. et al. Cdx and T Brachyury Co-activate growth signaling in the embryonic axial progenitor niche. Cell Rep. 17, 3165–3177 (2016).
Yamaguchi, T. P., Takada, S., Yoshikawa, Y., Wu, N. & McMahon, A. P. T (Brachyury) is a direct target of Wnt3a during paraxial mesoderm specification. Genes Dev. 13, 3185–3190 (1999).
Mallo, M. Reassessing the role of Hox genes during vertebrate development and evolution. Trends Genet. 34, 209–217 (2018).
Russ A. P. et al. Eomesoderminis required for mouse trophoblast development and mesoderm formation. Nature 404, 95–99 (2000).
Eakin, G. S. & Behringer, R. R. Diversity of germ layer and axis formation among mammals. Semin. Cell Dev. Biol. 15, 619–629 (2004).
Linneberg-Agerholm, M. et al. The primitive endoderm supports lineage plasticity to enable regulative development. Cell 187, 4010–4029.e16 (2024).
Girgin, M. U. et al. Bioengineered embryoids mimic post-implantation development in vitro. Nat. Commun. 12, 5140 (2021).
Masamsetti, V. P. et al. Characterization of the mesendoderm progenitors in the gastrulating mouse embryo. Dev. Cell https://doi.org/10.1101/2024.04.28.591221 (2024).
de Bree, K., de Bakker, B. S. & Oostra, R.-J. The development of the human notochord. PLoS ONE https://doi.org/10.1126/science.aag0053 (2018).
Cao, J. et al. Live birth of chimeric monkey with high contribution from embryonic stem cells. Cell 186, 4996–5014.e24 (2023).
Hamazaki, N. et al. Retinoic acid induces human gastruloids with posterior embryo-like structures. Nat. Cell Biol. 26, 1790–1803 (2024).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e4 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Phipson, B. et al. propeller: Testing for differences in cell type proportions in single cell data. Bioinformatics 38, 4720–4726 (2022).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Wickham, H. stringr: Simple, consistent wrappers for common string operations. R package version 1.5.1 (2023).
Wickham, H. et al. dplyr: A grammar of data manipulation. R package version 1.1.4 (2025).
Wickham, H. Reshaping data with the reshape package. J. Stat. Softw. 21, 1–20 (2007).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Kolde, R. Pheatmap: pretty heatmaps. CRAN https://doi.org/10.32614/CRAN.package.pheatmap (2019).
Acknowledgements
We thank all members of Zhen Liu laboratory, M.-M. Poo (CEBSIT) and M. Zhu (Harvard University) for helpful comments and discussion; and the non-human primate research facility, optical imaging core facility and the molecular and cellular biology core facility of CEBSIT and Optical Imaging Core Facility of Shanghai Center for Brain Science and Brain-Inspired Technology for supporting this project. We also acknowledge M. Saitou from Kyoto University for granting permission to reproduce the images of natural monkey embryos at E13 and E16 from their published work and H. Wang from Institute of Zoology, Chinese Academy of Sciences for granting permission to reproduce the images of natural monkey embryos at E22 from their published work. This work was funded by grants from the National Natural Science Foundation of China (32525026, 82021001, 92168109 and 32400675), the National Key Research and Development Program of China (2021YFA1101804, 2022YFF0710901, 2021ZD0200900 and 2024YFA0916602), the Young Climbing Program Project of CAS Shanghai Branch, the 2025 SANS Exploration Scholars project, the Youth Innovation Promotion Association of Chinese Academy of Sciences, the Shanghai Municipal Science and Technology Major Project, the Shanghai Yangfan Project (23YF1453000 and 24YF2752000), the Shanghai Oriental Talents-youth program (QNKJ2024018) and the Postdoctoral Fellowship Program of China Postdoctoral Science Foundation (no. GZC20241791). A.M.-A. is funded by an ERC AdG (Mini Embryo Blueprint_ 834580) and the ‘Maria de Maeztu’ Programme for Units of Excellence in R&D (grant no. CEX2018-000792-M).
Author information
Authors and Affiliations
Contributions
Z.L. and J.L. (1st) designed the project. Z.L., Q.S., A.M.-A. and X.W. supervised the study. J.L. (1st), S.S. and J.C. performed all the experiments. J.L. (2nd) performed all the bioinformatic analysis. L.Z., F.G., J.F., H.C., G.C. and H.W. provided technical support in the experiments. J.L. (1st), J.L. (2nd), Z.L. and A.M.-A. prepared the manuscript with input from the other authors. All authors contributed to the interpretation of the data and read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.M.-A. is an inventor on two patents on human polarized three-dimensional cellular aggregates (patent applicant: Alfonso Martinez-Arias; name of inventor(s): Alfonso Martinez-Arias, Naomi Moris; application number: PCT/GB2019/052670; status of application: pending) and polarized three-dimensional cellular aggregates (patent applicant: Alfonso Martinez-Arias; name of inventor(s): Alfonso Martinez-Arias, Naomi Moris; application number: PCT/GB2019/052668; status of application: abandoned). The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Thorold Theunissen, Gioele La Manno and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Increased induction efficiency of monkey blastoids and characterization of monkey embryoids from D9 to D17.
a, Representative images of monkey blastoids derived from different passage 4CL naive ESCs on D3 and D7 in aggrewell. Scale bars, 100 μm. b, Monkey blastoid formation efficiency at indicated passages of monkey 4CL naïve ESCs (P4-P7, n = 13; P8-P12, n = 5 from 3 independent experiments). Each dot represents a sample. Two-sided unpaired t-test, *** = P ≤ 0.001 (P = 5.12E-10). c, Relative expression levels of naive marker genes in different passage 4CL naive ESCs (n = 3 biologically independent samples). Each dot represents one repeat. d, Representative brightfield image of monkey 4CL naive ESCs at different passages. Scale bars, 100 μm. e, Cell proliferation rate analysis of 4CL naive ESCs at different passages. P4-P7: n = 12 and P8-P12: n = 8 from 3 independent experiments. Each dot represents a sample. Two-sided unpaired t-test, ** = P ≤ 0.01 (P = 0.001467). f, Representative images of monkey embryoids from D0 to D9 in aggrewell. Scale bars, 100 μm. g, Representative images of IF staining of monkey blastoid on D7 with OCT4 (Green) for epiblast, GATA4 (Red) for hypoblast and GATA3 (Cyan) for trophoblast. Scale bars, 100 μm. h, Representative images of IF staining of monkey pre-gastruloid on D9 with OCT4 (Red) for epiblast, SOX17 (Cyan) for hypoblast and GATA3 (Green) for trophoblast. Scale bars, 100 μm. i, Representative images of monkey embryoids in each group during in vitro culture from D9 to D17. The blue arrow represents group of succeeded developed monkey embryoids with embryonic disc and properly formed amnion/yolk sac cavity; the green arrow represents group of failed monkey embryoids with embryonic disc but no amnion/yolk sac cavities; the red arrow represents another group of failed monkey embryoids without embryonic disc. Scale bars, 200 μm. j, Statistical analysis of formation efficiency of each group in i (n = 8 biologically independent experiments for D9; n = 10 biologically independent experiments for D11, n = 8 biologically independent experiments for D13; n = 11 biologically independent experiments for D17). k, Developmental efficiency of monkey embryoids from D9 to D17 (n = 8 for D9; n = 6 for D11; n = 11 for D13; n = 11 for D17). Optimal formed monkey embryoids with embryonic disc under standard range (Diameter <0.1 mm for D9; 0.1–0.2 mm for D11; 0.15–0.5 mm for D13; 0.15–0.5 mm for D17) were calculated for statistical analysis. Each dot represents one independent experiment. l, Measurement of long diameter of whole embryo and embryonic disc of monkey embryoids from D9 to D17 respectively (n = 35 for each group from 5 independent experiments). m, Measurement of embryonic disc diameter of natural human embryos from CS6 (n = 7) and CS7 (n = 12) (Data from Carnegie stage, O’Rahilly et al. 1987) and monkey embryoids from D13 and D17, respectively (n = 35 for each group from 5 independent experiments). Each dot represents a sample. Two-sided unpaired t-test was used for D13 versus CS6-H and D17 versus CS7-H groups respectively, ns = P > 0.05 (P = 0.8599 for D13 versus CS6-H group; P = 0.0900 for D17 versus CS7-H group). n, Ratio of length of PS to length of embryonic disc between groups of monkey embryoids at different stages, natural monkey embryos at E16-18 and natural human embryos at CS7 stage. N = 5 for D13; n = 8 for D15 and D17 groups from 3 independent experiments; n = 4 for E16-18 group; n = 12 for CS7 group. Each dot represents a sample. One-way ANOVA with Dunnett’s T3 multiple comparisons test, ns = P > 0.05; * = P ≤ 0.05. D13 versus D15: P = 0.0203; D15 versus D17: P = 0.0256; D17 versus E16-18-M: P > 0.9999; D17 versus CS7-H: P = 0.8756; E16-18-M versus CS7-H: P = 0.8321. b-d, j-n, Data are presented as mean ± SEM.
Extended Data Fig. 2 Quantitative assessment of amnion/yolk sac cavity and downstream subtypes differentiation of trophoblasts in monkey embryoids.
a, Representative 3D images of whole-mount IF staining showing the typical amniotic cavity and yolk sac cavity in monkey embryoids on D13, D15 and D17 with OCT4 (gray), TFAP2A (red), and SOX17 (green) (Supplementary Videos 4–6). OCT4, marker for epiblast; TFAP2A, marker for amnion; SOX17, marker for yolk sac. A, anterior; P, posterior. Scale bars, 200 μm. b, Representative a monolayer of whole-mount IF staining showing the typical amniotic cavity and yolk sac cavity in monkey embryoids on D13, D15 and D17 with OCT4 (gray), TFAP2A (red), and SOX17 (green). Scale bars, 200 μm. c, Statistical analysis of amnion cavity area of natural monkey embryos and monkey embryoids on D13 and D17, respectively. N = 3 for natural monkey embryos on E13 and E16-17; n = 20 for monkey embryoids on D13; n = 28 for monkey embryoids on D17 from 3 independent experiments. P = 0.4275 for D13 group; P = 0.7168 for D17 group. d, Statistical analysis of yolk sac cavity area of natural monkey embryos and monkey embryoids on D13 and D17, respectively. N = 3 for natural monkey embryos on E13 and E17; n = 20 for monkey embryoids on D13 and n = 28 for monkey embryoids on D17 from 3 independent experiments. P = 0.8229 for D13 group; P = 0.9976 for D17 group. e-h, Statistical analysis of the long axis and wide axis of amnion and yolk sac cavity in natural monkey embryos and monkey embryoids on D13 and D17, respectively. N = 3 for natural monkey embryos on E13 and E16-17; n = 18 for monkey embryoids on D13and n = 28 for monkey embryoids on D17 from 3 independent experiments. P = 0.9554 for long axis of amnion in D13 group; P = 0.4638 for wide axis of amnion in D13 group; P = 0.1971 for long axis of amnion in D17 group; P = 0.7381 for wide axis of amnion in D17 group; P = 0.4011 for long axis of yolk sac in D13 group; P = 0.1972 for wide axis of yolk sac in D13 group; P = 0.3608 for long axis of yolk sac in D17 group; P = 0.7493 for wide axis of yolk sac in D17 group. i, Representative images showing the typical lateral view of monkey embryoids on D13 and D17 by immunostaining with OCT4 (gray), KRT7 (red) and MMP2 (green) or SDC1 (green) or TP63 (green). OCT4, a marker for epiblast; KRT7, a marker for trophoblasts; MMP2, a marker for EVT; SDC1, a marker for STB; TP63, a marker for CTB. EVT, Extravillous trophoblast; STB, Syncytiotrophoblast; CTB, Cytotrophoblast. Scale bars, 100 μm. j, Real-time qPCR analysis of relative expression level of marker genes for EVTs, STBs and CTBs in monkey embryoids on D13 and D17 (n = 3 biologically independent samples). Two-sided unpaired t-test, ns = P > 0.05; * = P ≤ 0.05; ** = P ≤ 0.01. P = 0.8733 for MMP2; P = 0.1313 for ITGA5; P = 0.1352 for HLA-G; P = 0.2550 for CGA; P = 0.3474 for CGB; P = 0.2146 for SDC1; P = 0.1313 for TP63; P = 0.0123 for VGLL1; P = 0.0381 for KRT7; P = 0.0251 for NR2F2; P = 0.0081 for GATA3. c-h, Two-sided unpaired t-test was used for D13 and D17 groups respectively, ns = P > 0.05. c-j, Data are presented as mean ± SEM. Each dot represents a sample.
Extended Data Fig. 3 Optimization of 3D suspension in vitro culture condition and quality classification of monkey embryoids from D17 to D25.
a, Comparison of different prolonged culture conditions of monkey embryoids from D17. Representative brightfield images of monkey embryoids on D25 were shown in the right. Scale bars, 200 μm. b, Developmental efficiency of in vitro-cultured monkey embryoids from D17 to D25 under different conditions were statistically analyzed. c, Representative images of monkey embryoids divided from three groups, optimal, suboptimal and medium group according to clear standards during in vitro culture from D17 to D25. N = 3 for each group to present. A, anterior; P, posterior. Scale bars, 100 μm. d, Statistical analysis of formation efficiency of each group from D17 to D25. n = 79 for Optimal group, n = 21 for Suboptimal group and n = Medium group from 3 independent experiments. e, Measurement of embryonic disc diameters of natural human embryos from CS8 (n = 8) and CS9 (n = 8) (Data from Carnegie stage, O’Rahilly et al. 1987) and monkey embryoids from D22 and D25, respectively (n = 15 for each group from 3 independent experiments). Data are presented as mean ± SEM. Each dot represents a sample. Two-sided unpaired t-test was used for D22 versus CS8-H and D25 versus CS9-H groups respectively, ns = P > 0.05 (P = 0.0604); *** = P ≤ 0.001. P = 0.0604 for D22 versus CS8-H group; P = 4.25E-06 for D25 versus CS9-H group. f, Representative images of monkey embryoids cultured beyond D25. N = 3 for each group to present. Scale bars, 100 μm.
Extended Data Fig. 4 3D analysis of monkey embryoids morphology.
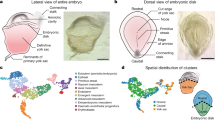
a, Representative 3D images showing the typical lateral view of monkey embryoids from D17 to D25 by light sheet microscope analysis after immunostaining with OCT4 (red), OTX2 (green) and FOXA2 (gray) (Supplementary Videos 7–10). OCT4, a marker for epiblast; OTX2, a marker for neuroectoderm and endoderm; FOXA2, a marker for endoderm and some mesoderm. A, anterior; P, posterior. Scale bars, 200 μm. b, Monolayer from 3D morphology in (a) showing the typical lateral view of monkey embryoids from D17 to D25 by light sheet microscope analysis after immunostaining with OCT4 (red), OTX2 (green) and FOXA2 (gray). Two domains of OTX2, one that overlaps FOXA2 (ventral) indicating endoderm and another that overlaps OCT4 (dorsal) indicating NP. NP, neural plate; Endo, endoderm. Scale bars, 200 μm. c, Representative 3D images showing the typical dorsal view of monkey embryoids from D17 to D25 by light sheet microscope analysis after immunostaining with OCT4 (red), OTX2 (green) and FOXA2 (gray) (Supplementary Videos 7–10). Scale bars, 200 μm. d, Monolayer from 3D morphology in (b) showing the typical dorsal view of monkey embryoids from D17 to D25 by light sheet microscope analysis after immunostaining with OCT4 (red), OTX2 (green) and FOXA2 (gray). Scale bars, 200 μm. e, Representative 3D images showing the typical transversal view of monkey embryoids from D17 to D25 by light sheet microscope analysis after immunostaining with OCT4 (red), OTX2 (green) and FOXA2 (gray) (Supplementary Videos 7–10). NG, neural groove. D, dorsal; V, ventral. Scale bars, 200 μm. f, Schematic diagram of monkey embryoids at different stages from lateral, dorsal, and transversal views. Schematic was created using BioRender (https://biorender.com).
Extended Data Fig. 5 Monkey embryoids induction in vitro from monkey ESC-2# and iPSCs.
a, Representative brightfield images of dynamic induction process of monkey blastoid from D0 to D7 and IVC1-treated monkey embryoids on D9 in aggrewell. Scale bars, 100 μm. b, Representative brightfield images of lateral viewed monkey embryoids from D13 to D25. A, anterior; P, posterior. Scale bars, 200 μm. c, Developmental efficiency of 3.5 cm dish-cultured monkey embryoids from D13 to D17 (n = 3 independent experiments for each group). Monkey embryoids with clear embryonic disc-like structure including epiblast, amnion cavity and yolk sac cavity were calculated for statistical analysis. Each dot represents one repeat. d, Development efficiency of single cultured monkey embryoids from D17 to D25. Monkey embryoids with clear embryonic disc-like structure were calculated for statistical analysis (53 monkey embryoids on D17 from 3 independent experiments were used to track developmental efficiency to D25). e, Measurement of long diameters of whole embryo and embryonic disc length of monkey embryoids from D13 to D25 respectively (n = 14 for D13 and D15 groups; n = 9 for D17 group; n = 13 for D20 group; n = 11 for D23 group; n = 12 for D25 group from 4 independent experiments). f, Representative 3D images (upper) and monolayer (lower) showing the typical lateral view of monkey embryoids on D17 by light sheet microscope analysis after whole-mount IF staining with OCT4 (gray), TFAP2A (red), and SOX17 (green) (Supplementary Video 12). OCT4, a marker for epiblast; TFAP2A, a marker for amnion cells; SOX17, a marker for yolk sac. Scale bars, 200 μm. g, Representative 3D images showing the typical lateral view and dorsal view of monkey embryoids on D20 by light sheet microscope analysis after whole-mount IF staining with TBXT (gray) and DAPI (Supplementary Video 13). TBXT, a marker for PS (primitive streak). Scale bars, 200 μm. h, Representative brightfield images of dynamic induction process of monkey blastoid from D0 to D7 and IVC1-treated monkey embryoids on D9 in aggrewell. Scale bars, 100 μm. i, Representative brightfield images of lateral viewed monkey embryoids from D13 to D25. A, anterior; P, posterior. Scale bars, 200 μm. j, Developmental efficiency of 3.5 cm dish-cultured monkey embryoids from D13 to D17 (n = 5 independent experiments for each group). Monkey embryoids with clear embryonic disc-like structure including epiblast, amnion cavity and yolk sac cavity were calculated for statistical analysis. Each dot represents one repeat. k, Development efficiency of single cultured monkey embryoids from D17 to D25. Monkey embryoids with clear embryonic disc-like structure were calculated for statistical analysis (62 monkey embryoids on D17 from 3 independent experiments were used to track developmental efficiency until to D25). l, Measurement of long diameters of whole embryo and embryonic disc length of monkey embryoids from D13 to D25 respectively (n = 20 for D13, D15, D17, D20 and D23 groups; n = 16 for D25 group from 5 independent experiments). m, Representative 3D images showing the typical lateral view and dorsal view of monkey embryoids on D17 by light sheet microscope analysis after whole-mount IF staining of with TBXT (gray) and DAPI (Supplementary Video 14). TBXT, a marker for PS (primitive streak). Scale bars, 200 μm. n, Representative 3D images showing the typical lateral view of monkey embryoids on D15 by light sheet microscope analysis after whole-mount IF staining with OCT4 (gray), TFAP2A (red), and SOX17 (green) (Supplementary Video 15). OCT4, a marker for epiblast; TFAP2A, a marker for amnion cells; SOX17, a marker for yolk sac. Scale bars, 200 μm. c, e, j, l, Data are presented as mean ± SEM.
Extended Data Fig. 6 Transcriptomic features of the cultured monkey embryoids.
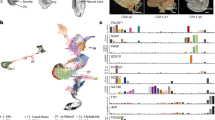
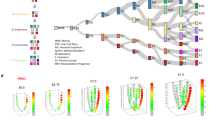
a, Bar graph showing the number of collected single cells and filtered cells of each monkey embryoid. b, Violin and box (median, 25th and 75th percentile; whiskers show 1.5 × the interquartile range) plots depicting the distributions of the number of UMI and detected genes in monkey embryoid cells (n = 101,160 cells from 7 monkey embryoids). c, Violin plot displaying the expression of representative marker genes for the indicated cell clusters of in vivo monkey embryos and embryoids. d, UMAP plot showing the integration of monkey embryoids (D17-D25) with in vitro-cultured natural monkey embryos (18 d.p.f. to 25 d.p.f.). Cells are colored by study (top panel) and annotated cluster (middle and bottom panel). The deficient cell clusters in monkey embryoids compared to in vitro cultured monkey embryos are marked in red in the legend. Progenitors included epiblast and ectoderm; PS/PM, primitive streak/paraxial mesoderm; PGCLC, primordial germ-cell-like cell; LPM, lateral plate mesoderm; EXM, extraembryonic mesoderm; Mes, mesenchymal cell; Al, allantois; Cardi., Cardiomyocyte; ys.Meso, yolk sac mesoderm; DE/VE, definitive endoderm/visceral endoderm; ys.Endo, yolk sac endoderm; STB, syncytiotrophoblast; EVT, extravillous trophoblast; CTB, cytotrophoblast; EC, endothelial cell.
Extended Data Fig. 7 Further characteristics of neuroectoderm lineages development.
a-b, UMAP plot showing neural related cell clusters in a and heatmap in b of marker gene expressions for each cluster. ECT, ectoderm; NPL, neural plate progenitors like cells; border, neural-plate-border like cells. c, UMAPs showing the expression of feature genes in neural-related cell clusters (SOX2 and OTX2 for ECT; SLC7A3 and CLDN10 for NPL; DLX5 and CXCL12 for border; PMP22 and DLK1 for epidermis). Color bars represent the normalized expression level of indicated genes from high (red) to low (grey). ECT, ectoderm; NPL, neural plate progenitors like cells. d, Bar plot shows the cell proportions of neural-related cell clusters in indicated developmental stage. e, Monocle pseudotime trajectory showing the progression of ectoderm (ECT), neural plate progenitors like cells (NPL), and neural-plate-border like cells (border). f, UMAP plot showing the sub-clusters of NPL and border (upper panel) and heatmap (lower panel) of marker gene expressions for the identified sub-clusters. g-h, Cell proportions and featured genes expression of sub-clusters in f among indicated developmental stages. i, Representative images showing the dorsal view and lateral view of whole-mount IF staining of monkey embryoids on D25 with SOX2 (cyan), SOX9 (red), and TWIST1 (green) to exhibit the structure of neural tube (SOX2+). Arrowheads indicate SOX9+TWIST1+ neural crest-like cells. A, anterior; P, posterior; D, dorsal; V, ventral. Scale bars, 100 μm. j, Representative images showing the typical transversal view of whole-mount IF staining of monkey embryoids on D25 by immunostaining with SOX2 (gray), SOX9 (green) and TWIST1 (red). SOX2, a marker for epiblast; SOX9 and TWIST1, markers for neural crest-like cells. Scale bars, 100 μm. k, Representative images showing the typical lateral view of monkey embryoids on D25 (1# and 2#) by immunostaining with SOX2 (gray), DLX6 (red) and DLX5 (green). SOX2, a marker for epiblast; DLX5 and DLX6, markers for placode-like cells. Scale bars, 100 μm. l-m, Lateral view and ventral view of 3D reconstruction of D25-2# monkey embryoids by Light sheet microscope analysis after immunostaining with SOX2 (gray), DLX6 (red) and DLX5 (green) (Supplementary Video 16). Scale bars, 100 μm.
Extended Data Fig. 8 PS derived lineages identification and gene expression analysis of mesoderm related clusters using scRNA-seq data.
a, Heatmap showing the expression of markers associated with 12 mesoderm clusters. b, Marker genes of PS cluster and their expression in CS8-9 staged natural monkey embryo, in vitro-cultured monkey embryos and monkey embryoids. EPI, epiblast; ECT, ectoderm; PS, primitive streak; PGCLC, primordial germ cell-like cell; APS, anterior primitive streak; DE/VE, definitive endoderm/visceral endoderm; ys.Endo, yolk sac endoderm; Nas.Meso, nascent mesoderm; LP.Meso, lateral plate mesoderm; Cardi., cardiomyocyte; Cardi.Meso, cardiac mesodermal cells; Exe.Meso, extraembryonic mesoderm; Mes, mesenchymal cell; Al, allantois; ys.Meso, yolk sac mesoderm; EC, endothelial cell. Caud.Meso, caudal mesoderm; Pharyg.Meso, pharyngeal mesoderm; BP, blood progenitor; Mac, macrophage; Ery1, erythocyte1; Ery2, erythocyte2; NC, neural crest; SE, surface ectoderm; Neural Plate, Neural plate-like cell; AM, axial mesoderm; PM, paraxial mesoderm; LPM, lateral plate mesoderm; EXM, extraembryonic mesoderm; ys.Meso, yolk sac mesoderm; ParE, parietal endoderm; Border, neural-plate-border-like cells. c, Schematic diagram of monkey embryoid on D20 and representative images of H&E staining and whole-mount IF staining of monkey embryoids on D20 with TBXT (green), TBX6 (magenta), and DAPI. TBXT, marker for PS and Nas.Meso; TBX6, marker for Nas.Meso, NMP and other mesoderm. Red arrows indicate the TBXT+TBX6+ Nas.Meso cells. Yellow dot lines indicate embryonic disc of D20 monkey embryoid. AM, amnion; NP, neural plate; PS, primitive streak; NMP, neuromesodermal progenitor; DE, definitive endoderm; Nas.Meso, nascent mesoderm; A, anterior; P, posterior. Scale bars, 100 μm. Schematic was created using BioRender (https://biorender.com). d, Bar plot shows the cell proportions of indicated developmental stage in each mesoderm cluster. e, Bar plot shows the cell proportions of cell cycle phases in each mesoderm cluster in Cell-cycle phases include G1, S, and G2/M.
Extended Data Fig. 9 GO term functional analysis of mesoderm related clusters using scRNA-seq data.
a, Heatmap displays the expression of top 5 differentially expressed genes (DEGs) for 9 subclusters related to blood. b, Bar plots show the represented GO terms of HAB, EC, aEC, Mac, monocyte, Mk, HPC and Ery (lower right panel). The x axis represents the adjusted p value. Meso.progenitor, mesoderm progenitor; HAB, hemogenic angioblast; EC, endothelial cell; aEC, arterial endothelial cell; Mac, macrophage; Mk, megakaryocyte. c, UMAP plot showing the integration of EC and blood cells between monkey embryoids and in vitro-cultured embryos (left 3 panels) and bar plot shows the cell proportion of indicated clusters between monkey embryoid and in vitro cultured embryo (right panel). d, Representative images showing IF staining of sectioned slides of monkey embryoid on D25 with CD34 (red), MPO (green), and PECAM1 (gray). Red arrows indicate CD34+MPO+PECAM1+ HPC cells located in SYS cavity. AM, amnion; SYS, secondary yolk sac. Scale bars, 100 μm.
Extended Data Fig. 10 The identification of endoderm related cells in monkey embryoids.
a, UMAP plot shows the integration of cells related to endoderm lineage between monkey embryoids and in vivo monkey embryos (left 3 panels) and bar plot shows the cell proportion of indicated clusters between monkey embryoid and in vivo monkey embryo (right panel). b, Representative images of whole-mount IF staining of monkey embryoids on D20 with OCT4 (gray), OTX2 (red), and FOXA2 (green). Arrowheads indicate the OTX2+FOXA2+OCT4− DE-like cells. Yellow dot lines indicate embryonic disc of D20 monkey embryoid. DE, definitive endoderm; A, anterior; P, posterior. Scale bars, 100 μm. c, Representative images of whole-mount IF staining of monkey embryoids on D20 with OCT4 (gray), GATA4 (red), and SOX17 (green). Arrowheads indicate the GATA4+SOX17+OCT4low VE-like cells. Yellow dot lines indicate embryonic disc of D20 monkey embryoid. VE, visceral endoderm. Scale bars, 100 μm. d, Representative images of IF staining of in vivo-derived monkey embryo on E22 with DAPI (gray), OTX2 (magenta), and FOXA2 (green). Arrowheads indicate the OTX2+FOXA2+ DE-like cells. Scale bars, 100 μm. Panel reproduced from ref. 11, Springer Nature, Ltd. e, Representative images of IF staining of monkey embryoids on D25 with SOX2 (grey), FOXA2 (red), and ISL1 (green). Yellow dot lines indicate embryonic disc of D25 monkey embryoid. Red arrows indicate the FOXA2+ISL1+ primitive foregut-like cells. Schematic diagram of monkey embryoid on D25 and H&E staining of D25 monkey embryoids (Right). NF, neural folding; PS, primitive streak; AM, amnion; SYS, secondary yolk sac; Meso, mesoderm. Scale bars, 100 μm. Schematic was created using BioRender (https://biorender.com). f, Representative images of IF staining of monkey embryoids on D25 with SOX2 (grey), FOXA2 (red), and CDX2 (green). Yellow dot lines indicate embryonic disc of D25 monkey embryoid. Red arrows indicate the FOXA2+CDX2+ primitive hindgut-like cells. Scale bars, 100 μm. g, UMAP showing the annotated clusters of DE and gut cells. The overlaid line represents the trajectory lineage based on Slingshot pseudotime analysis. h, UMAP showing the continuous CytoTRACE score, from 1 (highest) to 0 (lowest) level of plasticity.
Extended Data Fig. 11 PGCLCs differentiation in monkey embryoids and GO term functional analysis of PGCLCs in monkey embryoids.
a, Schematic diagram showing the PGC specification in natural monkey embryo at indicated time. EPI, epiblast; AM, amnion; EXMC, extra-embryonic mesenchyme cells; PGC, primordial germ cell; SYS, secondary yolk sac. Schematic was created using BioRender (https://biorender.com). b, UMAPs showing expression of marker gene for each PGCLC subcluster. Colors from gray (low) to red (high) indicating gene expression level. c, UMAP plot displays the expression of MKI67 in PGCLCs. d, Heatmap of DEGs of each sub-cluster of PGCLCs (left panel). The representative GO terms (right panel) are shown. e, UMAP plot showing the integration of PGCLCs in monkey embryoids and in vitro monkey embryos. The cells are colored by studies. f, UMAP plot showing the subclusters of PGCLCs in monkey embryoids. g, UMAP plot showing the subclusters of PGCLCs in in vitro monkey embryos. h, Heatmap illustrates the similarity between subclusters of PGCLCs in monkey embryoids and in vitro monkey embryos.
Extended Data Fig. 12 Genetic perturbation of TBXT in monkey embryoids.
a, Representative images of 4CL naïve monkey ESCs of NTC and TBXT KO converted from primed ones. Scale bars, 200 μm. b, Representative brightfield images of dynamic induction process of monkey embryoids derived from NTC and TBXT KO naïve monkey ESCs. Red arrows indicated typical monkey embryoids on D9. Scale bars, 100 μm. c, Representative brightfield images of lateral viewed monkey embryoids from D13 to D25 derived NTC and TBXT KO naïve monkey ESCs. Red dot lines indicate embryonic disc of monkey embryoids at different times. A, anterior; P, posterior. Scale bars, 100 μm. d, Representative images of whole-mount IF staining of monkey embryoid on D13 and D17 from NTC and TBXT KO with OCT4 (Red) and TBXT (Green). OCT4, a marker for epiblast; TBXT, a marker for gastrulating/PS cells. Scale bars, 100 μm. e, Proportion of TBXT+ cells in NTC controls or TBXT KO monkey embryoids. n = 10 for D13 NTC group, n = 10 for D17 NTC, n = 6 for D13 TBXT KO and n = 10 for D17 TBXT KO from 3 independent experiments. P = 2.09E-05 for D13 group and P = 8.81E-06 for D17 group. f, Developmental efficiency of monkey embryoids on D13 and D17 derived from derived NTC and TBXT KO naïve monkey ESCs (n = 8 independent experiments for D13 and D17 in NTC group; n = 3 independent experiments for D13 and D17 in TBXT KO group). The efficiency was calculated by the ratio of monkey embryoids with clear embryonic disc-like structure including epiblast, amnion cavity and yolk sac cavity and all aggregates. P = 0.2345 for D13 group; P = 0.0275 for D17 group. e, f, Data are presented as mean ± SEM. Each dot represents a sample. Two-sided unpaired t-test was used for D13 and D17 groups respectively, ns = P > 0.05 (P = 0.6266); * = P ≤ 0.05; *** = P ≤ 0.001. g, Measurement of embryonic disc of monkey embryoids from NTC and TBXT KO, respectively (In NTC group, n = 22 for D13; n = 23 for D17; n = 22 for D20; n = 21 for D25 from 3 independent experiments. In TBXT KO group, n = 22 for D13; n = 23 for D17; n = 18 for D20; n = 15 for D25 from 3 independent experiments). Data are presented as mean ± SEM. Two-way ANOVA with Šídák’s multiple comparisons test, ns = P > 0.05; *** = P ≤ 0.01. P = 0.4976 for D13 group; P = 0.6008 for D17 group; P = 0.0006 for D20 group; P < 0.0001 for D25 group. h, Representative 3D images of NTC and TBXT KO monkey embryoids on D25 by light sheet microscope (Supplementary Videos 17, 18). Scale bars, 200 μm.
Supplementary information
Supplementary Figures (download PDF )
This file contains Supplementary Figures 1–14.
Supplementary Table 1 (download XLSX )
DEGs of each cluster (D17-D18 3D cultured monkey embryoids) and Data for integration analysis with in vivo and in vitro monkey embryos.
Supplementary Table 2 (download XLSX )
scRNA seq of monkey embryoids and annotated cell clusters.
Supplementary Table 3 (download XLSX )
Top 100 DEGs of ECT, neural plate, border and epidermis.
Supplementary Table 4 (download XLSX )
Top 100 DEGs of neural plate, placode and neural crest.
Supplementary Table 5 (download XLSX )
DEGs of mesoderm related cell clusters.
Supplementary Table 6 (download XLSX )
DEGs of 9 endothelial and hematopoietic related subclusters.
Supplementary Table 7 (download XLSX )
DEGs of endoderm related cell clusters.
Supplementary Table 8 (download XLSX )
DEGs between Foregut1 and Foregut2.
Supplementary Table 9 (download XLSX )
DEGs between Hindgut1 and Hindgut2.
Supplementary Table 10 (download XLSX )
DEGs of PGCLC clusters.
Supplementary Table 11 (download XLSX )
Antibody used in this study.
Supplementary Table 12 (download XLSX )
Oligonucleotide used in this study.
Supplementary Data (download ZIP )
Source Data for Supplementary Figures 2, 7 and 14.
Supplementary Video 1 (download MP4 )
3D morphology reconstruction of monkey embryoid on D13 by light sheet microscopy imaging showing PS (primitive streak).
Supplementary Video 2 (download MP4 )
3D morphology reconstruction of monkey embryoid on D15 by light sheet microscopy imaging showing primitive streak.
Supplementary Video 3 (download MP4 )
3D morphology reconstruction of monkey embryoid on D17 by light sheet microscopy imaging showing primitive streak.
Supplementary Video 4 (download MP4 )
3D morphology reconstruction of monkey embryoid on D13 by light sheet microscopy imaging showing amnion and yolk sac.
Supplementary Video 5 (download MP4 )
3D morphology reconstruction of monkey embryoid on D15 by light sheet microscopy imaging showing amnion and yolk sac.
Supplementary Video 6 (download MP4 )
3D morphology reconstruction of monkey embryoid on D17 by light sheet microscopy imaging showing amnion and yolk sac.
Supplementary Video 7 (download MP4 )
3D morphology reconstruction of monkey embryoid on D17 by light sheet microscopy imaging showing ECT and endoderm.
Supplementary Video 8 (download MP4 )
3D morphology reconstruction of monkey embryoid on D20 by light sheet microscopy imaging showing neural plate and endoderm.
Supplementary Video 9 (download MP4 )
3D morphology reconstruction of monkey embryoid on D23 by light sheet microscopy imaging showing NF (neural folding).
Supplementary Video 10 (download MP4 )
3D morphology reconstruction of monkey embryoid on D25 by light sheet microscopy imaging showing NG (neural groove).
Supplementary Video 11 (download MP4 )
3D morphology reconstruction of monkey embryoid on D25 by light sheet microscopy imaging showing the three germ layers.
Supplementary Video 12 (download MP4 )
3D morphology reconstruction of monkey embryoid on D17 generated by monkey ESCs-2# by light sheet microscopy imaging showing the embryonic disc, amnion and yolk sac.
Supplementary Video 13 (download MP4 )
3D morphology reconstruction of monkey embryoid on D20 derived from monkey ESCs-2# by light sheet microscopy imaging showing the PS.
Supplementary Video 14 (download MP4 )
3D morphology reconstruction of monkey embryoid on D20 derived from monkey iPSCs by light sheet microscopy imaging showing the PS.
Supplementary Video 15 (download MP4 )
3D morphology reconstruction of monkey embryoid on D17 derived from monkey iPSCs by light sheet microscopy imaging showing the embryonic disc, amnion and yolk sac.
Supplementary Video 16 (download MP4 )
3D morphology reconstruction of monkey embryoid on D25 by light sheet microscopy imaging showing the placode-like cells.
Supplementary Video 17 (download MP4 )
3D morphology reconstruction of NTC monkey embryoid on D25 by light sheet microscopy imaging.
Supplementary Video 18 (download MP4 )
3D morphology reconstruction of TBXT KO monkey embryoid on D25 by light sheet microscopy imaging.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Li, J., Cao, J. et al. Modelling late gastrulation in stem cell-derived monkey embryo models. Nature 649, 161–172 (2026). https://doi.org/10.1038/s41586-025-09831-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09831-0