Abstract
The mouse PIWI-interacting RNA (piRNA) pathway provides sustained anti-transposon immunity to the developing male germline by directing transposon DNA methylation1,2,3. The first step in this process is the recruitment of SPOCD1 to young LINE1 loci4. Thereafter, piRNA-mediated tethering of the PIWI protein MIWI2 (also known as PIWIL4) to the nascent transposon transcript recruits the DNA methylation machinery5,6. The piRNA pathway needs to methylate all active transposon copies but how this is achieved remains unknown. Here we show that nuclear piRNA and de novo methylation factors are all euchromatic, exposing constitutive heterochromatin as a genomic blind spot for the piRNA pathway. We discover a ‘nowhere-to-hide’ mechanism that enables piRNA pathway-mediated LINE1 surveillance of the entire genome. We find that SPOCD1 directly interacts with the nuclear pore component TPR, which forms heterochromatin exclusion zones adjacent to nuclear pores7. In fetal gonocytes undergoing piRNA-directed DNA methylation, TPR is found both at the nuclear periphery and throughout the nucleoplasm. We find that the SPOCD1–TPR interaction is required for complete non-stochastic piRNA-directed LINE1 methylation. The loss of the SPOCD1–TPR interaction results in a fraction of SPOCD1 and other chromatin-bound piRNA factors relocalizing to constitutive heterochromatin where they are no longer accessible to MIWI2 and the de novo methylation machinery. In summary, the piRNA pathway has co-opted TPR to guarantee that LINE1s are accessible to the piRNA and de novo methylation machineries.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The EM-seq data generated in this study have been deposited on ArrayExpress under the accession number E-MTAB-14862. Data for the IP–MS experiments have been deposited at ProteomeXchange under the accession number PXD060850. The CL–MS data have been deposited under PXD060851. EM-seq data for Spocd1−/− and Miwi2−/− P14 spermatogonia were retrieved from E-MTAB-7997, whereas that for C19orf84−/− spermatogonia were retrieved from E-MTAB-11612. The following Uniprot accession IDs for SPOCD1 proteins were used for conservation analyses: mouse (M. musculus, B1ASB6), golden hamster (M. auratus, A0A3Q0D6B7), Ord’s kangaroo rat (D. ordii, A0A1S3FIT4), western European hedgehog (E. europaeus, A0A1S3WPZ3), rabbit (O. cuniculus, G1SPR0), aardvark (O. afer afer, A0A8B7AXN8), bison (B. bison bison, A0A6P3HA20), bovine (B. taurus, F1MG39), goat (C. hircus, A0A452FMH8), sheep (O.aries, W5NRM3), pig (S. scrofa, F1SV96), horse (E. caballus, F6YBJ1), alpaca (V. pacos, A0A6J3AYV9), California sealion (Z. californianus, A0A6J2C2W2), northern fur seal (C. ursinus, A0A3Q7MZA7), Atlantic bottle-nosed dolphin (T. truncatus, A0A6J3PXS9), sperm whale (P. macrocephalus, A0A455B8T1), blue whale (B. musculus, A0A8B8W162), great Himalayan leaf-nosed bat (H. armiger, A0A8B7SLV8), large flying fox (P. vampyrus, A0A6P3Q928), leopard (P. pardus, A0A6P4UE11), cat (F. catus, A0A5F5XDK8), red fox (V. vulpes, A0A3Q7T0D7), dog (C. lupus familiaris, A0A8P0P5S7), polar bear (U. maritimus, A0A8M1EZU5), greater bamboo lemur (P. simus, A0A8C8YEI0), giant panda (A. melanoleuca, G1MHH0), rhesus macaque (M. mulatta, F7G2T4), northern white-cheeked gibbon (N. leucogenys, G1QNN7), Sumatran orangutan (P. abelii, H2N866), gorilla (G. gorilla gorilla, G3RKR7), chimpanzee (P. troglodytes, H2R1B9), human (H. spaiens, Q6ZMY3), American alligator (A. mississippiensis, A0A151MMW3), Chinese alligator (A. sinensis, A0A1U8CWC7), snapping turtle (C. serpentina, A0A8T1SPX3), central bearded dragon (P. vitticeps, A0A6J0UYI0), mainland tiger snake (N. scutatus, A0A6J1UEZ9), Indian cobra (N. naja, A0A8C6YBU2), American chameleon (A. carolinensis, H9GI50), western clawed frog (X. tropicalis, A0A8J0T0T7) and African clawed frog (X. laevis, A0A1L8HFK1). Source data are provided with this paper.
Code availability
The scripts used for the EM-seq analysis are available on GitHub (https://github.com/tamchow/spocd1_pirna-directed-dna-met-variance/ (version of the record deposited on Zenodo54, https://doi.org/10.5281/zenodo.17162836) and https://github.com/rberrens/SPOCD1-piRNA_directed_DNA_met (version of the record deposited on Zenodo55, https://doi.org/10.5281/zenodo.10509247)). The Fiji/ImageJ2 plugin used to calculate Pearson’s correlation coefficients in germ cell nuclei is also available on GitHub (https://github.com/COIL-Edinburgh/ROI_NucleusColocalisation; a version of the record has been deposited on Zenodo41, https://doi.org/10.5281/zenodo.17200734).
References
Aravin, A. A., Sachidanandam, R., Girard, A., Fejes-Toth, K. & Hannon, G. J. Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316, 744–747 (2007).
Carmell, M. A. et al. MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev. Cell 12, 503–514 (2007).
Kuramochi-Miyagawa, S. et al. DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev. 22, 908–917 (2008).
Dias Mirandela, M. et al. Two-factor authentication underpins the precision of the piRNA pathway. Nature 634, 979–985 (2024).
De Fazio, S. et al. The endonuclease activity of Mili fuels piRNA amplification that silences LINE1 elements. Nature 480, 259–263 (2011).
Schopp, T. et al. TEX15 is an essential executor of MIWI2-directed transposon DNA methylation and silencing. Nat. Commun. 11, 3739 (2020).
Krull, S. et al. Protein Tpr is required for establishing nuclear pore-associated zones of heterochromatin exclusion. EMBO J. 29, 1659–1673 (2010).
Walsh, C. P., Chaillet, J. R. & Bestor, T. H. Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nat. Genet. 20, 116–117 (1998).
Greenberg, M. V. C. & Bourc’his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Biol. 20, 590–607 (2019).
Kafri, T. et al. Developmental pattern of gene-specific DNA methylation in the mouse embryo and germ line. Genes Dev. 6, 705–714 (1992).
Monk, M., Boubelik, M. & Lehnert, S. Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ cell lineages during mouse embryo development. Development 99, 371–382 (1987).
Seisenberger, S. et al. The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol. Cell 48, 849–862 (2012).
Bourc’his, D. & Bestor, T. H. Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature 431, 96–99 (2004).
Ozata, D. M., Gainetdinov, I., Zoch, A., O’Carroll, D. & Zamore, P. D. PIWI-interacting RNAs: small RNAs with big functions. Nat. Rev. Genet. 20, 89–108 (2019).
Aravin, A. A. et al. A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol. Cell 31, 785–799 (2008).
Molaro, A. et al. Two waves of de novo methylation during mouse germ cell development. Genes Dev. 28, 1544–1549 (2014).
Barau, J. et al. The DNA methyltransferase DNMT3C protects male germ cells from transposon activity. Science 354, 909–912 (2016).
Jain, D. et al. rahu Is a mutant allele of Dnmt3c, encoding a DNA methyltransferase homolog required for meiosis and transposon repression in the mouse male germline. PLoS Genet. 13, e1006964 (2017).
Zoch, A. et al. SPOCD1 is an essential executor of piRNA-directed de novo DNA methylation. Nature 584, 635–639 (2020).
Zoch, A. et al. C19ORF84 connects piRNA and DNA methylation machineries to defend the mammalian germ line. Mol. Cell https://doi.org/10.1016/j.molcel.2024.01.014 (2024).
Yoshioka, H., McCarrey, J. R. & Yamazaki, Y. Dynamic nuclear organization of constitutive heterochromatin during fetal male germ cell development in mice. Biol. Reprod. 80, 804–812 (2009).
Santos-Rosa, H. et al. Active genes are tri-methylated at K4 of histone H3. Nature 419, 407–411 (2002).
Vasiliauskaite, L. et al. A MILI-independent piRNA biogenesis pathway empowers partial germline reprogramming. Nat. Struct. Mol. Biol. 24, 604–606 (2017).
Cordes, V. C., Hase, M. E. & Muller, L. Molecular segments of protein Tpr that confer nuclear targeting and association with the nuclear pore complex. Exp. Cell Res. 245, 43–56 (1998).
Cordes, V. C., Reidenbach, S., Rackwitz, H. R. & Franke, W. W. Identification of protein p270/Tpr as a constitutive component of the nuclear pore complex-attached intranuclear filaments. J. Cell Biol. 136, 515–529 (1997).
Mitchell, P. J. & Cooper, C. S. The human tpr gene encodes a protein of 2094 amino acids that has extensive coiled-coil regions and an acidic C-terminal domain. Oncogene 7, 2329–2333 (1992).
Singh, D. et al. The molecular architecture of the nuclear basket. Cell 187, 5267–5281.e13 (2024).
Nakano, H., Funasaka, T., Hashizume, C. & Wong, R. W. Nucleoporin translocated promoter region (Tpr) associates with dynein complex, preventing chromosome lagging formation during mitosis. J. Biol. Chem. 285, 10841–10849 (2010).
Bartlett, B. M. et al. TPR is required for cytoplasmic chromatin fragment formation during senescence. eLife https://doi.org/10.7554/eLife.101702 (2024).
Coyle, J. H., Bor, Y. C., Rekosh, D. & Hammarskjold, M. L. The Tpr protein regulates export of mRNAs with retained introns that traffic through the Nxf1 pathway. RNA 17, 1344–1356 (2011).
Pastor, W. A. et al. MORC1 represses transposable elements in the mouse male germline. Nat. Commun. 5, 5795 (2014).
Vasiliauskaite, L. et al. Defective germline reprogramming rewires the spermatogonial transcriptome. Nat. Struct. Mol. Biol. 25, 394–404 (2018).
Watanabe, T. et al. Role for piRNAs and noncoding RNA in de novo DNA methylation of the imprinted mouse Rasgrf1 locus. Science 332, 848–852 (2011).
Di Giacomo, M. et al. Multiple epigenetic mechanisms and the piRNA pathway enforce LINE1 silencing during adult spermatogenesis. Mol. Cell 50, 601–608 (2013).
Carrieri, C. et al. A transit-amplifying population underpins the efficient regenerative capacity of the testis. J. Exp. Med. 214, 1631–1641 (2017).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Di Giacomo, M., Comazzetto, S., Sampath, S. C., Sampath, S. C. & O’Carroll, D. G9a co-suppresses LINE1 elements in spermatogonia. Epigenetics Chromatin 7, 24 (2014).
Pandey, R. R. et al. Tudor domain containing 12 (TDRD12) is essential for secondary PIWI interacting RNA biogenesis in mice. Proc. Natl Acad. Sci. USA 110, 16492–16497 (2013).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Stringer, C. & Pachitariu, M. Cellpose3: one-click image restoration for improved cellular segmentation. Nat. Methods 22, 592–599 (2025).
Kelly, D. & Chowdhury, T. COIL-Edinburgh/ROI_NucleusColocalisation: nucleus colocalisation v2.01 (2.01). Zenodo https://doi.org/10.5281/zenodo.17200734 (2025).
Schopp, T. et al. The DUF3715 domain has a conserved role in RNA-directed transposon silencing. RNA 29, 1471–1480 (2023).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Chambers, M. C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30, 918–920 (2012).
Mendes, M. L. et al. An integrated workflow for crosslinking mass spectrometry. Mol. Syst. Biol. 15, e8994 (2019).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Jurrus, E. et al. Improvements to the APBS biomolecular solvation software suite. Protein Sci. 27, 112–128 (2018).
Yariv, B. et al. Using evolutionary data to make sense of macromolecules with a “face-lifted” ConSurf. Protein Sci. 32, e4582 (2023).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2 — a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
UniProt, C. UniProt: the Universal Protein Knowledgebase in 2025. Nucleic Acids Res. 53, D609–D617 (2025).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Chowdhury, T. tamchow/spocd1_pirna-directed-dna-met-variance: Version of record for manuscript submission (1.0). Zenodo https://doi.org/10.5281/zenodo.17162836 (2025).
Berrens, R. rberrens/SPOCD1-piRNA_directed_DNA_met: 20240114_release (v1.0.2). Zenodo https://doi.org/10.5281/zenodo.10509247 (2024).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Acknowledgements
This research was supported by the Welcome Trust funding to D.O. (225237), A.G.C (200898), the Wellcome Centre for Cell Biology (203149) and the multi-user equipment grants (108504 and 092076). This work was supported by funding for the Wellcome Discovery Research Platform for Hidden Cell Biology (226791), and we acknowledge support from the Microscopy, Proteomics and Bioinformatics cores. T.C. is funded by the Darwin Trust of Edinburgh. A.Z. was funded by a German Research Foundation fellowship (DFG; award ZO 376/1-1). S.B. and W.A.B. are funded by MRC University Unit grant MC_UU_00035/7 and by a Wellcome Trust Investigator Award 217120/Z/19/Z. This work utilized the University of Edinburgh Protein Production Facility (EPPF), the Wellcome Centre for Cell Biology Centre Optical Instrumentation Laboratory/Light Microscopy Core, Proteomics and Bioinformatics Core Platforms as well as the Centre for Regenerative Medicine’s FACS facility. We also acknowledge the EMBL’s GeneCore facility for preparing the methyl-seq libraries and sequencing all libraries.
Author information
Authors and Affiliations
Contributions
T.C. contributed to the design, execution and analysis of most experiments. S.B. performed the FISH experiments. A.Z. performed the IP–MS experiments. X.X. performed the bioinformatic analysis of the EM-seq data. M.D.M. and H.F. performed the initial biochemical experiments and prepared the mESCs expressing SPOCD1–HA. J.Z. performed the analysis of the CL–MS data. C.S. analysed the IP–MS data. D.K. developed the plugin to calculate the colocalization coefficients. D.O., A.G.C. and W.A.B. supervised the study. D.O. conceived the study. D.O. and T.C. wrote the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Zhao Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Nuclear piRNA factors and the de novo methylation machinery are euchromatic.
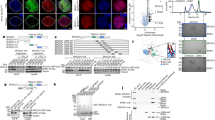
a, Representative E16.5 gonocyte, somatic cell and corresponding seminiferous cord stained for DAPI, HP1B (red) and H3K9me3 (green, top row) or HP1A (green, bottom row) in foetal testis sections from wild-type mice. Scale bars, 2 μm (top row), 10 μm (bottom row). White rectangles on images of seminiferous cords highlight cells shown zoomed-in. b-e, Representative E16.5 seminiferous cord stained for HA (green), HP1B (red, top row) or H3K4me3 (red, bottom row) and DAPI (blue) in foetal testis sections from Spocd1HA/+ (b), Miwi2HA/+ (c), Dnmt3lHA/HA (d) and Dnmt3cHA/HA (e) mice. f-g, Representative E16.5 seminiferous cord stained for C19ORF84 (f) or SPIN1 (g) (both green), HP1B (red, top row) or H3K4me3 (red, bottom row) and DAPI (blue) in foetal testis sections from wild-type mice. Also shown in b-g are the channel intensity profiles (RFU, relative fluorescence units) along the indicated line passing through major heterochromatin domains (for HP1B) or avoiding nucleoli (for H3K4me3) for the representative gonocytes presented in Fig. 1. Images shown in both rows of f are from the same quadruple stain (DAPI, C19ORF84, H3K4me3 and HP1B) of a seminiferous cord. Images shown in both rows of g are from another quadruple stain (DAPI, SPIN1, H3K4me3 and HP1B) of a seminiferous cord. Images in a through g are representative of n = 4 (b, c, e) or n = 3 (a, d, f, g) biological replicates; scale bars, 10 μm for seminiferous cords and 2 μm for single gonocytes. White rectangles highlight cells shown in Fig. 1 and in the profile plots.
Extended Data Fig. 2 piRNA pathway factors are euchromatic from initiation of expression.
a-c, Representative seminiferous cords stained for HA (SPOCD1-HA) (a), MIWI2 (b) and DNMT3L (c), H3K4me3 and DAPI from Spocd1HA/+ (a) or wild-type mice (b-c) at the indicated developmental stages. Shown to the right are boxplots of Pearson’s correlation coefficients of SPOCD1 (a), MIWI2 (b) and DNMT3L (c) with H3K4me3 in gonocyte nuclei at the indicated developmental stages. MIWI2 and DNMT3L are not expressed at E14.5, so co-localization was not calculated for this developmental stage. Scale bars, 10 μm. Images in a through c are representative of n = 3 biological replicates. Box indicates interquartile range, central line represents the mean, and whiskers extend to median ± 1.5× the interquartile range or data limits, whichever is smaller.
Extended Data Fig. 3 TPR is highly expressed in male foetal germ cells undergoing de novo genome methylation.
a, Volcano plot showing enrichment (log2(mean LFQ ratio of anti-HA immunoprecipitates from Miwi2HA/HA/wild-type E16.5 foetal testis lysates) and statistical confidence of proteins co-purifying with HA-MIWI2 (n = 4 with 50 testes per replicate, from previously published data19). b, Volcano plot showing enrichment (log2(mean LFQ ratio of anti-C19ORF84 immunoprecipitated/anti-rabbit serum immunoprecipitated from wild-type E16.5 foetal testis lysates) and statistical confidence of proteins co-purifying with C19ORF84 (n = 3 with 25 testes per replicate, from previously published data20). c, Single representative E16.5 germ and somatic cells stained for TPR (red) and DAPI (blue) in Spocd1HA/+ foetal testis sections. Scale bars, 2 μm. d, Representative E16.5 seminiferous cord stained for HA (green), TPR (red) and DAPI in foetal testis sections from Spocd1HA/+ mice. White rectangles highlight cells shown in Fig. 2f and Extended Data Fig. 3c. Scale bars, 10 μm. e, Representative gonocytes stained for TPR (red) and DAPI (blue) from wild-type mice at the indicated developmental stages. Scale bars, 2 μm. f, Representative seminiferous cords stained for TPR (red) and DAPI (blue) from wild-type mice at the indicated developmental stages. White rectangles highlight cells shown in Extended Data Fig. 3e. Scale bars, 10 μm. Images in c through f are representative of n = 3 biological replicates.
Extended Data Fig. 4 SPOCD1-K464A does not interact with TPR.
a, Multiple sequence alignment of the SPOCD1 TFIIS-M domains from the indicated species. Red rectangles highlight lysine residues making direct crosslinks with TPR-M in the CL-MS data. Sequence identity conservation is shown by depth of colour. b, Representative Coomassie-stained gel image of n = 3 MBP pull-down experiments with the indicated recombinant mouse SPOCD1 and human TPR fragments. For uncropped source gel image, see Supplementary Fig. 1d.
Extended Data Fig. 5 The Spocd1K464A mouse allele.
a, Schematic representations of the mouse Spocd1 locus and encoded 1015 amino acid protein are shown, along with a schematic of the CRISPR targeting strategy showing the location of single-stranded oligo DNA donor (ssODN) and homology arms (HA) used. The sgRNA used for generation of the Spocd1K464A allele (red), adjacent PAM sites (red) and the primers used for genotyping (blue) are indicated, along with a representative sequencing trace of Spocd1K464A exon 7 harbouring the 3 bp mutation encoding the K464A mutation (highlighted in red). Sequencing was performed on n = 3 F1 animals. b, Representative image of PCR genotyping result for Spocd1+/+, Spocd1+/K464A and Spocd1K464A mice. PCR genotyping was performed for over 600 mice with similar results. c, Volcano plot showing enrichment (LFQ ratio of anti-SPOCD1 immunoprecipitates from wild-type/Spocd1K464A E16.5 foetal testis lysates) and statistical confidence (−log10(P-value of two-sided Student’s t-test)) of proteins co-purifying with wild-type SPOCD1 (right quadrant) or SPOCD1-K464A (left quadrant) (n = 3 with 24 foetal testes per replicate per genotype). All identified proteins meeting the enrichment cut-off are listed in Supplementary Table 2. d, Representative adult testis sections of n = 3 wild type, Spocd1K464A and Spocd1−/− stained in blue for DAPI and red for the DNA damage marker γH2AX (top) or apoptotic cells by terminal deoxynucleotidyl transferase dUTP nick end labelling (TUNEL) assay (bottom). Scale bars, 50 μm.
Extended Data Fig. 6 The SPOCD1-TPR interaction ensures that SPOCD1-SPIN1-C19ORF84 avoids heterochromatin.
a-c, e-g, Representative E16.5 seminiferous cord stained for DAPI, HP1B (red) and SPOCD1 (a), SPIN1 (b), C19ORF84 (c), MIWI2 (e), DNMT3L (f), or HA (HA-DNMT3C) (g) (all green) stains in wild-type (top row) or Spocd1K464A (bottom row) foetal testis sections from littermates. Images are representative of n = 8 (a), n = 4 (c) or n = 3 (b, d-f) biological replicates of each genotype. White rectangles highlight cells shown zoomed-in in Fig. 4a–f. d, Representative E16.5 gonocyte (top two rows) and seminiferous cords (bottom two rows) stained for DAPI (blue), H3K4me3 (red or green), SPOCD1 (green) and HP1B (red) stains in wild-type (first and third row) or Spocd1K464A (second and last row) foetal testis sections from littermates. Images are representative of n = 4 biological replicates of each genotype. White rectangles in the last two rows highlight cells shown in the first two rows. Also shown below are channel intensity profiles (RFU, relative fluorescence units) along the indicated line passing through major heterochromatic domains in the representative gonocyte. Shown to the right are boxplots of Pearson’s correlation coefficients of SPOCD1 with H3K4me3 in gonocyte nuclei of the indicated genotypes. Box indicates interquartile range, central line represents the mean, and whiskers extend to median ± 1.5× the interquartile range or data limits, whichever is smaller. P-values are from unpaired, two-tailed Student’s t-tests. h, Representative E16.5 seminiferous cord stained for DAPI, SPOCD1 and LINE1 DNA in foetal testis sections of the indicated genotypes. Images are representative of n = 6 biological replicates of each genotype. White rectangles highlight cells shown in Fig. 4g. a-h, Scale bars, 10 μm for seminiferous cords and 2 μm for single gonocytes.
Extended Data Fig. 7 The SPOCD1-TPR interaction is necessary for Rasgrf1 methylation.
a, Heatmap presentation of mean CpG methylation of imprinted loci in P14 undifferentiated spermatogonia of the indicated genotypes. The imprint control region (ICR) of Rasgrf1 is shown in detail on the bottom. b, Cartoon representation of piRNA-directed epigenetic silencing of LINE1 elements in mice. TPR ensures SPOCD1-targeted LINE1 loci are present in euchromatin and accessible to the piRNA-directed DNA methylation machinery.
Supplementary information
Supplementary Information (download PDF )
This file contains Supplementary information summary, Supplementary Figures 1–2, Supplementary Tables 1–2, and legend for Supplementary Table 3.
Supplementary Table 3 (download XLSX )
Identified cross-links between SPOCD1-TFIISM and TPR-M. Table listing all Xi-validated cross-linked peptides detected in and between SPOCD1-TFIISM and TPR-M for n=2 cross-linking reactions (analysed together), relevant to Fig. 2e.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chowdhury, T., Boyle, S., Zoch, A. et al. A nowhere-to-hide mechanism ensures complete piRNA-directed DNA methylation. Nature 650, 779–785 (2026). https://doi.org/10.1038/s41586-025-09940-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09940-w