Abstract
The transition to agriculture was a transformative process in human history with wide-ranging demographic and social consequences1. Across South America, agriculture was adopted at different times and through diverse pathways, resulting in a mosaic of regionally distinct farming histories2,3. The Uspallata Valley, at the southern frontier of Andean farming, offers a unique opportunity to examine a case of late adoption of agriculture. Here we show that agriculture in the Uspallata Valley was adopted by local hunter-gatherers, as evidenced by genetic continuity between pre-farming and farming populations inferred from 46 newly sequenced ancient human genomes. These groups carried a distinct genetic component in Indigenous American diversity, indicating a unique population history in the region. Palaeodietary isotopes (δ13C/δ15N) reveal fluctuating maize intake consistent with flexible farming. Strontium isotopes (87Sr/86Sr) indicate the arrival of migrants from nearby regions between around 810–700 cal years BP, shortly before the Inka expansion. Genomic and isotopic analyses show that these migrants belonged to the same regional metapopulation as local groups, relied heavily on maize, probably moved in matrilineally organized family groups, exhibited stress markers (including malnutrition and diseases, such as tuberculosis, as confirmed by pathogen genomics) and experienced a long-term demographic decline. Our results suggest that these groups used social organization and migration as resilience strategies in the face of a multidimensional crisis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The ancient human genomic data generated in this study are deposited in the Owey repository (https://doi.org/10.48802/owey.bdxssypL) under controlled access. We selected this repository to support long-term, community-responsive data governance: in recent years, Indigenous communities’ perspectives on the implications of genetic data and its re-use have evolved rapidly, and it is realistic to expect that expectations and access conditions may continue to evolve. Our engagement framework therefore aims to ensure that data access policies can be adapted over time, including the introduction of additional conditions or consultation requirements if requested by the communities. Owey allows us to retain direct oversight of access decisions and to update access terms without dataset migration or changes in repository governance. Access requests are managed by the data generators (N.R. and P.L.) in coordination with local partners and Indigenous communities. Requests should be submitted via the repository interface; a response is typically provided within two weeks. Approved uses are limited to non-commercial academic research and must not include redistribution of the data, upload to open-access databases, or uses that could stigmatize or harm Indigenous communities. Previously published genomic data used for comparative analyses are available from the original publications.
Code availability
R scripts generated to plot the genomic (P.L.) and isotopic (A. Tessone, M.L.C. and R.B.) analyses, as well as for Random Forest modelling of bioavailable strontium (M.L.C.) and OxCal (L.B.-V.) modelling are available at https://github.com/pierrespc/BarberenaLuisi_2025. All of the genomic data processing software used are contained in the singularity image provided by nf-core/eager/2.4.2 (https://github.com/nf-core/eager/releases/tag/2.4.2), Contamination software; Nuclear contamination, in the same singularity image, ContamLD (https://github.com/nathan-nakatsuka/ContamLD), ContamMix (https://github.com/DReichLab/ADNA-Tools), Haplocheck (https://github.com/genepi/haplocheck), Schmutzi (https://github.com/grenaud/schmutzi). Genomic analysis software: Haplogrep (https://haplogrep.i-med.ac.at/), Yleaf (https://github.com/genid/Yleaf), ADMIXTURE (https://dalexander.github.io/admixture/), ADMIXTOOLS (https://github.com/DReichLab/AdmixTools), POPSTATS (https://github.com/pontussk/popstats), BEAST and TreeAnnotator (https://beast.community/programs), TreeStat (http://tree.bio.ed.ac.uk/software/treestat/), hapROH (https://github.com/hringbauer/hapROH) and HapNe (https://github.com/PalamaraLab/HapNe).
References
The Emergence of Agriculture: A Global View (Routledge, 2007).
Pearsall, D. M. in The Handbook of South American Archaeology (eds Silverman, H. & Isbell, W. H.) 105–120 (Springer, 2008).
Lombardo, U. et al. Early Holocene crop cultivation and landscape modification in Amazonia. Nature 581, 190–193 (2020).
Fuller, D. Q., Denham, T. & Allaby, R. Plant domestication and agricultural ecologies. Curr. Biol. 33, R636–R649 (2023).
Lipson, M. et al. Parallel palaeogenomic transects reveal complex genetic history of early European farmers. Nature 551, 368–372 (2017).
Smith, B. D. Eastern North America as an independent center of plant domestication. Proc. Natl Acad. Sci. USA 103, 12223–12228 (2006).
Piperno, D. R., Ranere, A. J., Holst, I., Iriarte, J. & Dickau, R. Starch grain and phytolith evidence for early ninth millennium B.P. maize from the Central Balsas River Valley, Mexico. Proc. Natl Acad. Sci. USA 106, 5019–5024 (2009).
Yacobaccio, H. D. The domestication of South American camelids: a review. Animal Front. 11, 43–51 (2021).
Hastorf, C. A. Agriculture and the Onset of Political Inequality Before the Inka (Cambridge Univ. Press, 1993).
de Valdivia, L. Doctrina cristiana y catecismo en la lengua allentiac, que corre en la ciudad de San Juan de la frontera, con un confesionario, arte y vocabulario breves. An. Inst. Etnogr. Am. 1, 19–94 (1940).
Yáñez, B. et al. Pace and space in the practice of aDNA research: concerns from the periphery. Am. J. Biol. Anthropol. 180, 417–422 (2023).
de la Fuente Castro, C. & Figueiro, G. Human ancient DNA analysis in Latin America: current state and challenges. Archaeometry 67, S4–S26 (2025).
Novellino, P. S. et al. Sitios de entierro de la localidad arqueológica Barrancas, Maipú (Mendoza, Argentina). Intersec. Antropol. 14, 271–277 (2013).
Da Peña Aldao, G., Novellino, P. & Frigolé, C. Caracterización del Túmulo II (Uspallata, Mendoza): actualización del análisis del contexto funerario. Comechingonia 20, 21–49 (2016).
Guevara, D. et al. Estructura demográfica, dieta y migración en los Andes del sur: nuevos análisis del sitio Osario Potrero Las Colonias, Uspallata (Mendoza, Argentina). Intersec. Antropol. 23, 67–82 (2022).
Durán, V. et al. Barrio Ramos I. Funebria y modos de vida en el inicio del período de dominación inca del valle de Uspallata (Mendoza, Argentina). Relac. Soc. Arg. Antropol. XLIII, 55–86 (2018).
Barberena, R. et al. Scale of human mobility in the southern Andes (Argentina and Chile): a new framework based on strontium isotopes. Am. J. Phys. Anthropol. 164, 305–320 (2017).
Barberena, R. et al. Multi-isotopic and morphometric evidence for the migration of farmers leading up to the Inka conquest of the southern Andes. Sci. Rep. 10, 21171 (2020).
Jørkov, M. L. S., Heinemeier, J. & Lynnerup, N. The petrous bone—a new sampling site for identifying early dietary patterns in stable isotopic studies. Am. J. Phys. Anthropol. 138, 199–209 (2009).
Willerslev, E. & Meltzer, D. J. Peopling of the Americas as inferred from ancient genomics. Nature 594, 356–364 (2021).
Rohland, N. et al. Three assays for in-solution enrichment of ancient human DNA at more than a million SNPs. Genome Res 32, 2068–2078 (2022).
Nakatsuka, N. et al. ContamLD: estimation of ancient nuclear DNA contamination using breakdown of linkage disequilibrium. Genome Biol 21, 199 (2020).
García, A. et al. Ancient and modern mitogenomes from Central Argentina: new insights into population continuity, temporal depth and migration in South America. Human Mol. Genet. 30, 1200–1217 (2021).
Luisi, P. et al. Fine-scale genomic analyses of admixed individuals reveal unrecognized genetic ancestry components in Argentina. PLoS ONE 15, e0233808 (2020).
Arango-Isaza, E. et al. The genetic history of the Southern Andes from present-day Mapuche ancestry. Curr. Biol. 33, 2602–2615.e5 (2023).
Posth, C. et al. Reconstructing the deep population history of Central and South America. Cell https://doi.org/10.1016/j.cell.2018.10.027 (2018).
Pedersen, M. W. et al. Ancient Human genomes and environmental DNA from the cement attaching 2,000-year-old head lice nits. Mol. Biol. Evol. 39, 1–16 (2022).
De La Fuente Castro, C. et al. The genomic and cultural diversity of the Inka Qhapaq Hucha ceremony in Chile and Argentina. Genome Biol. Evol. 16, 1–15 (2024).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Excoffier, L., Smouse, P. E. & Quattro, J. M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131, 479–491 (1992).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Peralta, E. A. et al. Past maize consumption correlates with population change in Central Western Argentina. J. Anthropol. Archaeol. 68, 101457 (2022).
Falabella, F., Planella, M. T. & Tykot, R. H. El Maíz (Zea mays) en el mundo prehispánico de Chile Central. Latin Am. Antiq. 19, 25–46 (2008).
Finucane, B. C. Maize and sociopolitical complexity in the Ayacucho Valley, Peru. Curr. Anthropol. 50, 535–545 (2009).
Pezo-Lanfranco, L. & Colonese, A. C. The role of farming and fishing in the rise of social complexity in the Central Andes: a stable isotope perspective. Sci. Rep. 14, 4582 (2024).
Knüsel, C. J. & Schotsmans, E. M. J. The Routledge Handbook of Archaeothanatology: Bioarchaeology of Mortuary Behaviour (Routledge, 2022).
Sedig, J. W., Olalde, I., Patterson, N., Harney, É & Reich, D. Combining ancient DNA and radiocarbon dating data to increase chronological accuracy. J. Archaeolog. Sci. 133, 105452 (2021).
Marsh, E. J., Kidd, R., Ogburn, D. & Durán, V. Dating the expansion of the Inca empire: Bayesian models from Ecuador and Argentina. Radiocarbon 59, 117–140 (2017).
Skoglund, P. et al. Genetic evidence for two founding populations of the Americas. Nature 525, 104–108 (2015).
Fournier, R., Tsangalidou, Z., Reich, D. & Palamara, P. F. Haplotype-based inference of recent effective population size in modern and ancient DNA samples. Nat. Commun. 14, 7945 (2023).
Suchard, M. A. et al. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. 4, vey016 (2018).
O’Donnell, L., Hill, E. C., Anderson, A. S. A. & Edgar, H. J. H. Cribra orbitalia and porotic hyperostosis are associated with respiratory infections in a contemporary mortality sample from New Mexico. Am. J. Phys. Anthropol. 173, 721–733 (2020).
Bos, K. I. et al. Pre-Columbian mycobacterial genomes reveal seals as a source of New World human tuberculosis. Nature 514, 494–497 (2014).
Vågene, ÅJ. et al. Geographically dispersed zoonotic tuberculosis in pre-contact South American human populations. Nat. Commun. 13, 1195 (2022).
Washburn, E. et al. A multi-isotope approach to reconstructing human residential mobility and diet during the Late Intermediate Period (1000–1450 CE) in highland Ancash, Peru. J. Archaeol. Sci. 41, 103291 (2022).
McCool, W. C. et al. Climate change–induced population pressure drives high rates of lethal violence in the Prehispanic central Andes. Proc. Natl Acad. Sci. USA 119, e2117556119 (2022).
Fort, J. Tendencies in the tempo of prehistoric agricultural expansions. J. Archaeol. Res. https://doi.org/10.1007/s10814-025-09212-1 (2025).
Fehren-Schmitz, L. et al. Climate change underlies global demographic, genetic, and cultural transitions in pre-Columbian southern Peru. Proc. Natl Acad. Sci. USA 111, 9443–9448 (2014).
Maravall-López, J. et al. Eight millennia of continuity of a previously unknown lineage in Argentina. Nature https://doi.org/10.1038/s41586-025-09731-3 (2025).
Mallick, S. et al. The Allen Ancient DNA Resource (AADR): a curated compendium of ancient human genomes. Sci. Data 11, 182 (2024).
Gil, A. F., Giardina, M. A., Neme, G. A. & Ugan, A. Demografía humana e incorporación de cultígenos en el centro occidente argentino: explorando tendencias en las fechas radiocarbónicas. Revis. Española Antropol. Am. 44, 523–553 (2014).
Dawson, N. qgis/QGIS: 3.44.7. Zenodo https://doi.org/10.5281/zenodo.6139224 (2026).
De La Fuente, C. et al. Genomic insights into the origin and diversification of late maritime hunter-gatherers from the Chilean Patagonia. Proc. Natl Acad. Sci. USA 115, E4006–E4012 (2018).
Homburger, J. R. et al. Genomic insights into the ancestry and demographic history of South America. PLoS Genet. 11, e1005602 (2015).
R Core Team. R: A Language and Environment For Statistical Computing (R Foundation for Statistical Computing, 2023).
South, A., Michael, S. & Massicotte, P. rnaturalearthdata: World Vector Map Data from Natural Earth Used in ‘rnaturalearth’. R v.1.0.0.9000 https://docs.ropensci.org/rnaturalearthdata/ (R project, 2026).
Rusconi, C. Poblaciones Pre y Posthispanicas de Mendoza Vol. 3, Arqueología (Mendoza, 1962).
Rusconi, C. Poblaciones Pre y Posthispanicas de Mendoza Vol. 1, Etnografía (Mendoza, 1961).
Suby, J. A., Novellino, P., Da Peña, G. & Pandiani, C. D. Elongated odontoid process in late Holocene skeletal remains from B6 archaeological site, Mendoza, Argentina. Intl J. Paleopathol. 22, 86–91 (2018).
Hansen, H. B. et al. Comparing ancient DNA preservation in petrous bone and tooth cementum. PLoS ONE 12, e0170940 (2017).
Damgaard, P. B. et al. Improving access to endogenous DNA in ancient bones and teeth. Sci. Rep. 5, 11184 (2015).
Pinhasi, R. et al. Optimal ancient DNA yields from the inner ear part of the human petrous bone. PLoS ONE 10, e0129102 (2015).
Dabney, J. & Meyer, M. Extraction of highly degraded DNA from ancient bones and teeth. Methods Mol. Biol. 1963, 25–29 (2019).
Rohland, N. & Hofreiter, M. Ancient DNA extraction from bones and teeth. Nat. Protoc. 2, 1756–1762 (2007).
Allentoft, M. E. et al. Population genomics of post-glacial western Eurasia. Nature 625, 301–311 (2024).
Rohland, N., Harney, E., Mallick, S., Nordenfelt, S. & Reich, D. Partial uracil-DNA-glycosylase treatment for screening of ancient DNA. Philos. Trans. R. Soc. B 370, 20130624 (2015).
Gamba, C. et al. Comparing the performance of three ancient DNA extraction methods for high-throughput sequencing. Mol. Ecol. Resour. 16, 459–469 (2016).
Kapp, J. D., Green, R. E. & Shapiro, B. A Fast and efficient single-stranded genomic library preparation method optimized for ancient DNA. J. Hered. 112, 241–249 (2021).
Fages, A. et al. Tracking five millennia of horse management with extensive ancient genome time series. Cell 177, 1419–1435.e31 (2019).
Yates, J. A. F. et al. Reproducible, portable, and efficient ancient genome reconstruction with nf-core/eager. PeerJ 9, e10947 (2021).
Schubert, M., Lindgreen, S. & Orlando, L. AdapterRemoval v2: rapid adapter trimming identification and read merging. BMC Res. Notes 9, 88 (2016).
Genovese, G., Handsaker, R. E., Li, H., Kenny, E. E. & McCarroll, S. A. Mapping the human reference genome’s missing sequence by three-way admixture in Latino genomes. Am. J. Hum. Genet. 93, 411–421 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Peltzer, A. et al. EAGER: efficient ancient genome reconstruction. Genome Biol. 17, 60 (2016).
Andrews, R. M. et al. Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat. Genet. 23, 147–147 (1999).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Neukamm, J., Peltzer, A. & Nieselt, K. DamageProfiler: fast damage pattern calculation for ancient DNA. Bioinformatics 37, 3652–3653 (2021).
Jun, G., Wing, M. K., Abecasis, G. R. & Kang, H. M. An efficient and scalable analysis framework for variant extraction and refinement from population-scale DNA sequence data. Genome Res. 25, 918–925 (2015).
Renaud, G., Slon, V., Duggan, A. T. & Kelso, J. Schmutzi: estimation of contamination and endogenous mitochondrial consensus calling for ancient DNA. Genome Biol. 16, 224 (2015).
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013).
Weissensteiner, H. et al. Contamination detection in sequencing studies using the mitochondrial phylogeny. Genome Res 31, 309–316 (2021).
Korneliussen, T. S., Albrechtsen, A. & Nielsen, R. ANGSD: analysis of next generation sequencing data. BMC Bioinformatics 15, 356 (2014).
Alaçamlı, E. et al. READv2: advanced and user-friendly detection of biological relatedness in archaeogenomics. Genome Biol. 25, 216 (2024).
Popli, D., Peyrégne, S. & Peter, B. M. KIN: a method to infer relatedness from low-coverage ancient DNA. Genome Biol. 24, 10 (2023).
van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum. Mutat. 30, E386–E394 (2009).
Motti, J. M. B. et al. Ancient mitogenomes from the Southern Pampas of Argentina reflect local differentiation and limited extra-regional linkages after rapid initial colonization. Am. J. Biol. Anthropol. 181, 216–230 (2023).
Tamburrini, C. Diversidad genética de muestras arqueológicas humanas del nordeste de la provincia del Chubut (Patagonia Argentina) durante el Holoceno (6000–200AP). Arqueología 30, 14390–14390 (2024).
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 214 (2007).
Ralf, A., Montiel González, D., Zhong, K. & Kayser, M. Yleaf: software for human y-chromosomal haplogroup inference from next-generation sequencing data. Mol. Biol. Evol. 35, 1291–1294 (2018).
Moreno-Mayar, J. V. et al. Early human dispersals within the Americas. Science 362, eaav2621 (2018).
Campelo dos Santos, A. L. et al. Genomic evidence for ancient human migration routes along South America’s Atlantic coast. Proc. Biol. Sci. 289, 20221078 (2022).
Bongers, J. L. et al. Integration of ancient DNA with transdisciplinary dataset finds strong support for Inca resettlement in the south Peruvian coast. Proc. Natl Acad. Sci. USA 117, 18359–18368 (2020).
Capodiferro, M. R. et al. Archaeogenomic distinctiveness of the Isthmo-Colombian area. Cell 184, 1706–1723 (2021).
Ferraz, T. et al. Genomic history of coastal societies from eastern South America. Nat. Ecol. Evol. 7, 1315–1330 (2023).
Kennett, D. J. et al. South-to-north migration preceded the advent of intensive farming in the Maya region. Nat. Commun. 13, 1530 (2022).
Kennett, D. J. et al. Archaeogenomic evidence reveals prehistoric matrilineal dynasty. Nat. Commun. 8, 14115 (2017).
Lindo, J. et al. The genetic prehistory of the Andean highlands 7000 years BP though European contact. Sci. Adv. 4, eaau4921 (2018).
Lindo, J. et al. The genomic prehistory of the Indigenous peoples of Uruguay. PNAS Nexus 1, pgac047 (2022).
Nakatsuka, N. et al. Ancient genomes in South Patagonia reveal population movements associated with technological shifts and geography. Nat. Commun. 11, 3868 (2020).
Nakatsuka, N. et al. Genetic continuity and change among the Indigenous peoples of California. Nature 624, 122–129 (2023).
Nakatsuka, N. et al. A paleogenomic reconstruction of the deep population history of the Andes. Cell 181, 1131–1145 (2020).
Raghavan, M. et al. Genomic evidence for the Pleistocene and recent population history of Native Americans. Science 349, aab3884 (2015).
Rasmussen, M. et al. The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature 506, 225–229 (2014).
Salazar, L. et al. Insights into the genetic histories and lifeways of Machu Picchu’s occupants. Sci. Adv. 9, eadg3377 (2023).
Villa-Islas, V. et al. Demographic history and genetic structure in pre-Hispanic Central Mexico. Science 380, eadd6142 (2023).
Harris, D. N. et al. Evolutionary genomic dynamics of Peruvians before, during, and after the Inca Empire. Proc. Natl Acad. Sci. USA 115, E6526–E6535 (2018).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
Bergström, A. et al. Insights into human genetic variation and population history from 929 diverse genomes. Science 367, eaay5012 (2020).
Reich, D. et al. Reconstructing Native American population history. Nature 488, 370–374 (2012).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Maier, R. et al. On the limits of fitting complex models of population history to f-statistics. eLife 12, e85492 (2023).
Kamvar, Z. N., Tabima, J. F. & Grünwald, N. J. Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2, e281 (2014).
pontussk/popstats https://github.com/pontussk/popstats (GitHub, 2025).
Ringbauer, H., Novembre, J. & Steinrücken, M. Parental relatedness through time revealed by runs of homozygosity in ancient DNA. Nat. Commun. 12, 5425 (2021).
Buikstra, J. & Ubelaker, D. Standards for Data Collection from Human Skeletal Remains Vol. 44 (Archaeological Survey Research Series, 1994).
Chamberlain, A. T. Demography in Archaeology (Cambridge Univ. Press, 2006).
Gifford-Gonzalez, D. An Introduction to Zooarchaeology (Springer, 2018).
Adams, B. J. & Konigsberg, L. W. Estimation of the most likely number of individuals from commingled human skeletal remains. Am. J. Phys. Anthropol. 125, 138–151 (2004).
Gregoricka, L. A. et al. The impact of secondary mortuary practices on representation and distribution of commingled elements from Umm an-Nar human skeletons in communal tombs. Adv. archaeol. https://doi.org/10.1017/aap.2024.47 (2025).
Copeland, S. R. et al. Using strontium isotopes to study site accumulation processes. J. Taphon. 8, 115–127 (2010).
Bataille, C. P., Crowley, B. E., Wooller, M. J. & Bowen, G. J. Advances in global bioavailable strontium isoscapes. Palaeogeogr. Palaeoclimatol. Palaeoecol. 555, 109849 (2020).
Le Corre, M. et al. An ensemble machine learning bioavailable strontium isoscape for Eastern Canada. FACETS 10, 1–17 (2025).
Spies, M. J. et al. Strontium isoscapes for provenance, mobility and migration: the way forward. R. Soc. Open Sci. 12, 250283 (2025).
Barberena, R. et al. Bioavailable strontium, human paleogeography, and migrations in the Southern Andes: a machine learning and GIS approach. Front. Ecol. Evol. 9, 584325 (2021).
Breiman, L. Random Forests. Mach. Learn. 45, 5–32 (2001).
Scaffidi, B. K. & Knudson, K. J. An archaeological strontium isoscape for the prehistoric andes: Understanding population mobility through a geostatistical meta-analysis of archaeological 87Sr/86Sr values from humans, animals, and artifacts. J. Archaeolog. Sci. 117, 105121 (2020).
Barberena, R. et al. Bioavailable strontium in the Southern Andes (Argentina and Chile): a tool for tracking human and animal movement. Environ. Archaeol. 26, 323–335 (2021).
Wunder, M. B. Determining geographic patterns of migration and dispersal using stable isotopes in keratins. J. Mammal. 93, 360–367 (2012).
Ma, C., Vander Zanden, H. B., Wunder, M. B. & Bowen, G. J. assignR: an R package for isotope-based geographic assignment. Methods Ecol. Evol. 11, 996–1001 (2020).
Sealy, J. C. et al. Comparison of two methods of extracting bone collagen for stable carbon and nitrogen isotope analysis: Comparing whole bone demineralization with gelatinization and ultrafiltration. J. Archaeolog. Sci. 47, 64–69 (2014).
Hogg, A. G. et al. SHCal20 Southern Hemisphere calibration, 0–55,000 years cal BP. Radiocarbon 62, 759–778 (2020).
Bronk Ramsey, C. Bayesian analysis of radiocarbon dates. Radiocarbon 51, 337–360 (2009).
Martel-Cea, A. et al. A multiproxy approach to reconstruct the Late Holocene environmental dynamics of the semiarid Andes of central Chile (29°S). Front. Ecol. Evol. 11, 1227020 (2023).
Tiner, R. J., Negrini, R. M., Antinao, J. L., McDonald, E. & Maldonado, A. Geophysical and geochemical constraints on the age and paleoclimate implications of Holocene lacustrine cores from the Andes of central Chile. J. Quaternary Sci. 33, 150–165 (2018).
Martel-Cea, A. et al. Late Holocene environmental changes as recorded in the sediments of high Andean Laguna Chepical, Central Chile (32°S; 3050 m a.s.l.). Palaeogeogr. Palaeoclimatol. Palaeoecol. 461, 44–54 (2016).
Rein, B. et al. El Niño variability off Peru during the last 20,000 years. Paleoceanography 20, 2004PA001099 (2005).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Pebesma, E. Simple features for R: standardized support for spatial vector data. R J. 10, 439 (2018).
Acknowledgements
We deeply thank the Huarpe communities from Uspallata (Guaytamari and Llahué Xumec) for their cooperation in the development of this multivocal project. We acknowledge the former and current authorities of Cultural Heritage of Mendoza Province, Argentina (H. Chiavazza, C. Sonego), and the Director of the Museo de Ciencias Naturales y Antropológicas Juan C. Moyano (G. Campos) for the authorization to study the remains from UV. M. Vázquez and J. Avalos, from the Registro Nacional de Yacimientos Colecciones y Objetos Arqueológicos (RENYCOA) of Argentina, for their assistance in the exportation of the studied samples. We thank the aDNA facility from the Institute of Genomics, University of Tartu, and especially C. L. Scheib, H. Kabral, K. Tambets and L. Saag for their support. We would also like to thank the HPC Core Facility of Institut Pasteur (IP) for their support for computational analyses and M. Monot, L. Motreff and F. Jagorel from the IP Biomics Platform (supported by France Génomique ANR-10-INBS-09-09 and IBISA) for their assistance in sample sequencing. We are thankful to E. Willerslev, O. Caceres Rey, T. O’Connor and V. Borda for facilitating access to sequencing data for some modern individuals included in this study. We acknowledge P. Rossetti and the CIRAM Lab for their advice on radiocarbon dating and the HPC Core Facility of Institut Pasteur for their support with computational work. Research was financed by European Research Council ERC-2020-STG - PaleoMetAmerica – 948800 (N.R.), Institut Pasteur and CNRS UMR funding (N.R.), INCEPTION program (Investissement d’Avenir Grant ANR-16-CONV-0005) (N.R.), National Geographic Society grant no. NGS-92679R-22 (R.B.), Wenner-Gren Foundation (grant no. 2368532037 to R.B. and ERG-60 to P.L.), Fondation pour la Recherche Médicale Postdoctoral fellowship (E.A.N.), EU-MSCA postdoctoral fellowship PATHOGEN (E.A.N.), Consejo Nacional Nacional de Investigaciones Científicas y Técnicas (CONICET) Extraordinary Postdoctoral fellowship (P.L.), ERC-2023-SyG-Horse Power–101071707 (L.O.) and PIP-CONICET (grant no. 11220210100098CO to P.N.).
Author information
Authors and Affiliations
Contributions
N.R., R.B. and P.L conceptualized the work. R.B., D.G., G.D.P., P.N., C.A.M., A. Tessone, A. Troncoso and V.D. conducted the sampling. M.L., G.T., M.I.O., E.A.N., L.O. and H.S. performed the aDNA laboratory work. P.L.R., J. Sealy, K.G., J.L. and A. Tessone performed the isotope laboratory work. P.N., D.G., G.D.P., C.A.M., J. Suby and L.M. performed the bioarchaeological analysis, whereas P.L., N.R., R.B., A. Tessone, L.B.-V., M.L.C., M. Cardillo, J.M.C., G.L., A.G., E.A.N., M.R., M.E.d.P. and E.P. performed the data analysis. R.B., P.N., D.G., G.D.P., C.A.M., M.F.Q., M.L.L., V.D., C.M. and F.S.-S. performed the archaeological work. N.R., R.B., L.O. and H.S. acquired funding. N.R., P.L. and R.B. wrote the original draft, whereas L.Q.-M., H.S., E.P., J. Sealy, A. Tessone, L.B.-V., L.M. and A.G reviewed and edited it. R.B., P.L., N.R., P.N., D.G., C.H., G.C., M. Candito and E.A.N. were responsible for community engagement.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks R. Alexander Bentley and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
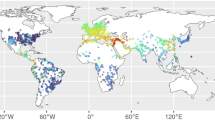
Extended Data Fig. 1 Reference modern groups for human population analyses.
Geographical location of modern individuals with genotype data included in population genetic analyses. Groups included in Unmasked categories include individuals whose sequencing data from the whole-genome was used (data from103,107,108,109 reprocessed by us). Other categories for groups including individuals whose genotyping data was reported in24,53,54,110, and for which we used masked genotype data for genetic ancestry Indigenous to the Americas. Groups whose individuals do not self-identify as Indigenous are represented in grey shades, while groups represented in other colours were reported as Indigenous in the original publications. Dark shade delimits the study area (Fig. 1). Map data from rnaturalearthdata plotted with ggplot2138 and sf139 packages in R55.
Extended Data Fig. 2 Admixture analyses.
a. Admixture analyses performed on ancient American genomes and using the 1240 K SNP panel, filtering out individuals with less than 30000 positions with known genotypes. b. Admixture analyses including modern and ancient individuals with >30000 genotyped SNPs. In both cases, for each value of K (2–15), we ran 10 replicates and showed only the maximum-likelihood run. Cross-Validation scores are provided in parentheses.
Extended Data Fig. 3 Genetic affinity of UV ancient individuals to other Ancient American Groups.
a-b. Heatmaps summarizing Z-scores derived from f4(Mbuti, Uspallata; Ancient American Group 1, Ancient American Group 2), where Ancient American Groups 1 and 2 are represented by point type and colours on the x and y axes, respectively. Ancient groups whose individuals were sequenced through shotgun are marked with black dots. Ancient American groups are ordered by major geographical ranges and periods. Z-scores greater and lower than 0 are coloured in shades of yellow to red and light blue to dark blue, respectively. Results are shown when only including American groups from Early (EH) and Mid-Holocene (MH) in the comparisons (b) or also including Late Holocene (LH) groups (a). f4 were computed considering all autosomal positions on the 1240 K (see Fig. S24 for results on other SNP panels). c-d. Z-scores derived from f4(Mbuti, Ancient Central or North American Group; UV, Ancient South American Group) are shown by grouping the Ancient Central and North American groups along the x-axis by geographical range and period. Points are represented according to the Ancient South American Group. In one column (one violin plot), South American Groups appear N times when the grouping on the x-axis includes N Central or North American groups. Dashed lines for |Z| = 3 are used as thresholds for significance. Results are shown when only including South American groups from Early (EH) and Mid- Holocene (MH) in the comparisons (d) or also including Late Holocene groups (LH) (c). f4 were computed considering all autosomal positions on the 1240 K (see Fig. S25 for results on other SNP panels).
Extended Data Fig. 4 Genetic differentiation within UV and with major ancient American groups and phylogenetic trees from mitogenomes.
a. Results for Analysis of MOlecular VAriance. Black dots represent the index of differentiation, ϕ, within and among major American groups (categories above the plot) containing at least two population groups (each column of coloured data points) with at least 2 individuals (3 in the case of Central Andes). Standard deviations were obtained from permutations. Error bars show ±3 standard deviations estimated from permutations for each ϕ value. The yellow rectangle shows the range encompassing ϕ among Uspallata and Calingasta Valleys (UV&CV) groups ± 3 standard deviations. Within UV&CV, individuals were grouped as Migrant Farmers (yellow circle; n = 16), Local Farmers (brown circle; n = 2) and Hunter-Gatherers from UV (brown diamond; n = 3) and CV (red circle; n = 2). Population groups’ data points follow the colour code shown in Main Fig. 2a, and the individuals from each group included in this analysis are given in Supplementary Table S4. This figure is associated with Table S8. b. Z-scores derived from f4(Mbuti, Ancient American Group; UV1, UV2) are shown, with pairs of UV1 and UV2 along the x-axis. Points are represented according to the Ancient American Group. Larger points are when the Ancient American Group is one group from UV, following the code shown in legend. Dashed lines for |Z | = 3 are used as thresholds for significance. f4 were computed considering all autosomal positions on the 1240 K (see Fig. S3 for results on other SNP panels). This figure, associated with Table S8 and Supplementary Data S6e, shows that UV individuals can be assigned to one unique group to study the potential origin of the genetic ancestry component specific to UV, and its links with ancient American groups. c. The Maximum parsimony trees for 38 mitogenomes newly sequenced, along with already published mitogenomes from South American individuals. Filled colour corresponds to a geographical region, and border type to a time period. Mutations are shown on the branches; they are transitions unless the base change is explicitly indicated. The prefix @ indicates the reversion of a mutation occurring earlier in the phylogeny. Recurrent mutations in the phylogeny are underlined. Sub-haplogroups defined for the first time in this study (C1b37 and B2b16a) are highlighted with a red fill. Meta-data concerning the mitogenomes analysed are provided in Table S6. d. Consensus bayesian phylogenetic tree from mitogenomes in UV and BB6. Only unrelated up to the 2nd-degree individuals from the present study were considered. Light blue circles size represent branches’ posterior probabilities. The branch lengths are provided in years. Leaves represent mitogenomes from individuals encoded according to the group assigned. LH-HG: Hunter-Gatherer; LH-MF: Migrant Farmer; LH-Mout: Migrant Farmer outlier, LH-LF: Local Farmer; LH-LFplc: Local Farmer from Potrero Las Colonias.
Extended Data Fig. 5 qpWave with duplet left populations being UV and another ancient South American group.
The lower panel summarizes the results, with each line for each ancient South American group included as left population along with UV. Each column represents a qpWave comparison according to the SNP set (shown at the bottom) and the right populations (groups included as right populations are highlighted with a skyblue rectangle in the upper panel). Comparisons that were not performed are shown with grey cells, otherwise the cell colours indicate the one-sided significance for rejection of Rank 0, without correction for multiple testing (white: p-value > 0.05; yellow: 0.01 <p-value < 0.05; dark yellow: 0.005 <p-value < 0.01; orange: 0.005 <p-value < 0.001; and red: p-value < 0.001). This figure is associated with Supplementary Data S7.
Extended Data Fig. 6 Distribution of the main genetic ancestry components observed in Southern Cone modern groups.
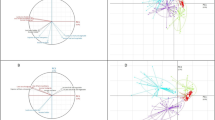
Geographic distribution of the six main ancestry genetic components identified in the Southern Cone (shown in Extended Data Fig. 2b) in modern groups across the Americas: a. blue/South Patagonia component; b. green/Brazil component; c. dark brown/Main Central Andes component; d. cardinal red/secondary Central Andes component; e. light brown/Central Chile component, and f. yellow/Uspallata component. For each panel, the right and left plots show the mean proportion ancestry estimates for each South American Indigenous group and Southern Cone non-Indigenous group, respectively, as a pie-chart located at the coordinates reported for that group. Map data from rnaturalearthdata plotted with ggplot2138 and sf139 packages in R55.
Extended Data Fig. 7 Strontium analyses for assessing mobility.
a-b. Bioavailable 87Sr/86Sr isoscape for the southern Andes of Argentina and Chile (a) with its associated spatial error (b). (7a and b were produced in R55, see Methods). c. Variable importance plot depicting the influence of the different covariates on the predicted bioavailable 87Sr/86Sr in the random forest regression. d. Partial dependence plots depicting the relationship between the predictors retained for the final random forest regression model and the predicted bioavailable 87Sr/86Sr. Hash marks on the x-axes correspond to the deciles of the predictor distribution. Description of the predictors is provided in Table S3. e. Bayesian geographical assignment of most likely provenience of individual AR0510 (PLC) based on the random forest strontium isoscape for the southern Andes (updated from125).
Extended Data Fig. 8 Radiocarbon chronology and Bayesian modelling.
Bars underneath each distribution denote 95.4% Confidence Interval (CI). The start of Inka occupation in the region38 is included at the bottom in red. The start and end estimates are 700-670 cal BP (median 685 cal BP) and 625-585 cal BP (median 610 cal BP), respectively, with a likely duration of between 55 to 100 (median 75 years). a. Bayesian model for the migrant phase at Potrero Las Colonias. The analyses were performed including kinship data (DOD values) as prior information. The radiocarbon dates are in purple following colouring in Fig. 4b. b. Bayesian model for the migrant phase at Túmulo III. The radiocarbon dates are in green following colouring in Fig. 4b. c. Consistency of the three Bayesian models. Probability density functions for the difference (‘D’) between the start and end boundaries for the three models created (Fig. 4b). Results show that apart from the start of the migrant phase at UV and the start of the Inka occupation, there is no significant difference between the modelled outputs as the distributions include zero at 95.4% CI.
Extended Data Fig. 9 Effective Population Size.
a. Conditional Heterozygosity (CH) for migrant and local farmers and a set of ancient American groups. Bars indicate ±2 standard errors estimated through block jackknife across the genome. Results are ordered by region according to CH means across groups within each region, and then by CH within each group. Analysis performed on the SG SNP panel and including only ancient individuals with shotgun sequencing data. b. Effective population size (Ne) estimated by maximum likelihood from the number and length distributions of runs of homozygosity under the hapROH model. Bars indicate ±2 likelihood-based standard errors. Ne estimates were obtained for local and migrant farmers, pre-farming hunter-gatherers, and a set of ancient American groups, ordered by region according to Ne mean across groups within region and then by Ne within group. The individuals from each group included in these analyses are given in Supplementary Table S4.
Extended Data Fig. 10 Paleoclimate reconstruction.
Macro-regional palaeocological archives in the southern Andes for the last 2400 years, including the following records: Laguna El Calvario, Laguna Cerritos Blancos, Laguna Chepical, and a proxy of ENSO activity in a sediment core located offshore from Peru134,135,136,137. The orange horizontal bar highlights the period of migrations and the grey bar a broader 400 year period.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–26 and Tables 1–8.
Supplementary Note (download PDF )
Translation of the paper to Spanish. This article was peer-reviewed in the English language. The translation to Spanish provided as Supplementary Note 1 was not peer-reviewed or checked for correctness by Springer Nature.
Supplementary Data (download ZIP )
Supplementary Data 1–8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Barberena, R., Luisi, P., Novellino, P. et al. Local agricultural transition, crisis and migration in the Southern Andes. Nature (2026). https://doi.org/10.1038/s41586-026-10233-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10233-z