Abstract
Although immunotherapy has revolutionized cancer treatment, many patients still experience limited benefit, highlighting the urgent need for improved biomarkers1. Although immunotherapy is founded on unleashing T cells2, most existing biomarkers remain tumour-centric and mainly overlook host immune competence. The thymus is a key immune organ that is crucial for T cell maturation, and we hypothesized that thymic functionality is associated with immunotherapy outcomes3. Here we show that thymic health, a radiographic measure of thymic functionality, is strongly associated with immunotherapy outcomes across several cancer types. Using a deep-learning framework applied to routine computed tomography images, we quantified thymic health in a pan-cancer cohort of 3,476 patients receiving immune checkpoint inhibitors. In patients with non-small cell lung cancer, higher thymic health was associated with reduced risks of progression and all-cause mortality. These associations remained significant across clinically relevant levels of programmed death ligand 1 (PD-L1) and tumour mutation burden. In the prospective TRACERx lung cancer study, thymic health was positively associated with T cell receptor diversity and T cell receptor excision circles, and correlated with immune-system signalling pathways, supporting radiographic thymic health as a proxy for thymic activity and adaptive immune competence. Analysis across patients with melanoma, breast cancer or renal cancer demonstrated pan-cancer relevance. Together, these findings identify thymic health as a previously unrecognized, tumour-agnostic determinant of immunotherapy efficacy, with potential implications for patient stratification, treatment timing and the development of immune-rejuvenating strategies in precision immuno-oncology.
Similar content being viewed by others
Main
In the past decade, immunotherapy has transformed cancer treatment, markedly improving patient outcomes across diverse tumour types2. Notably, beyond prolonging survival, immunotherapy can produce durable, complete responses, even in patients with advanced or metastatic disease4. Immunotherapy aims to unleash a host-derived T cell immune response against cancer cells, by blocking immune regulatory pathways and overcoming cancer immune evasion. With the introduction of immune checkpoint inhibitors (ICIs) that target programmed cell death protein 1 (PD-1), its ligand (PD-L1) or cytotoxic T-lymphocyte-associated protein 4 (CTLA-4), immunotherapy in the early or advanced setting has become standard of care across several tumour types, including melanoma5,6, non-small cell lung cancer (NSCLC)7, and renal cell carcinoma (RCC)8. However, only subsets of patients across tumour types have improved survival in response to ICIs9,10,11.
At present, biomarker research focuses on tumour-intrinsic factors, such as PD-L1 (ref. 12) or tumour mutation burden (TMB)13, but their utility as biomarkers is limited12,13,14, given that cancers across all biomarker levels may or may not have improved outcomes in relation to ICIs12,13. The poor performance of tumour-based biomarkers might be caused by a reliance on biopsies from tumours that are temporally and spatially heterogeneous15,16. However, it is also likely that the health of the patient’s immune system is important for immunotherapy response. No clinical test is at present performed to evaluate individual immune competence before immunotherapy.
Studies in recent years extend to tumour-extrinsic blood-based biomarkers, such as T lymphocytes, the T cell receptor repertoire, myeloid-derived suppressor cells or circulating tumour cells. This might pose a way forward, but research is still preliminary17,18. Indeed, the relevance of a patient’s immune health before treatment remains mostly unknown.
The thymus is an immune organ that is often overlooked in adults19. Because the thymus naturally decays through ageing, it is generally considered irrelevant in adults. However, the thymus is crucial for T cell maturation and for maintaining a diverse and specific adaptive immune response through life. Notably, previous studies suggest that the thymus is of pivotal relevance to health in adults20,21. However, the relevance of the thymus for cancer immunotherapy outcomes has never been investigated, and it remains unclear whether thymic health is associated with immuno-oncology outcomes and survival. The standard-of-care chest computed tomography (CT) scans obtained during routine cancer care capture the thymus in their field of view, which presents a unique opportunity to assess thymic health on the basis of radiographic characteristics. Especially given the breakthroughs in deep learning using self-supervised learning, this allows for detailed automated characterization of thymic health.
In this study, we investigate the associations of thymic health with outcomes to cancer immunotherapy in 3,476 real-world patients with various types of cancer (HARVARD-NSCLC and HARVARD-PAN; Fig. 1a). We further explore the immunological associations of thymic health in an independent deeply characterized NSCLC cohort from a prospectively enrolled lung cancer clinical trial (TRACERx, ClinicalTrials.gov: NCT01888601; Fig. 1a). We use a deep-learning system to automatically quantify thymic health on CT scans, and we demonstrate a significant positive association between thymic health and survival outcomes across cancer types. Notably, we show that thymic health is strongly associated with prolonged progression-free survival following immune checkpoint inhibitor therapy and prolonged overall survival (OS) in patients with NSCLC, performing similarly to TMB and PD-L1 for outcome prognostication. In the independent, prospectively recruited TRACERx study of treatment-naive patients with NSCLC, thymic health at diagnosis is associated with blood T cell receptor excision circle levels, T cell receptor diversity in blood and tumour and blood immune proteins, suggesting immunological relevance. Finally, in an analysis of real-world immunotherapy-treated patients with various cancer types, we observe a significant association between thymic health and survival outcomes, indicating the tumour-agnostic prognostic potential of thymic health. These results provide evidence of the crucial role of a patient’s thymic health for immunotherapy outcomes across cancer types, with key implications for precision medicine.
a, Overview of training data and the real-world Harvard-NSCLC and Harvard-PAN (melanoma, renal, breast, bladder, oesophagus, others) evaluation cohorts (total, n = 3,476). Harvard-PAN tumours with fewer than 100 patients per entity were pooled as ‘Others’. All patients were treated with immunotherapy at the Dana-Farber Harvard Cancer Center (DFHCC). External and biological validation was done in the prospectively collected TRACERx NSCLC cohort (n = 464) b, Overview of model development. A deep-learning system able to automatically predict thymic health, as a proxy for thymic functionality, based on standard-of-care chest CT scans was developed using 5,674 independent CT scans of the training data. We applied the model to the standard-of-care CT scans from the Harvard-NSCLC and Harvard-PAN cohorts as well as the external prospectively collected TRACERx cohort for statistical analysis. SSL, self-supervised learning. c, Overview of cohort descriptions. d, Representative images of high, average and low thymic health. The anatomical overview of cancer types in a was created in BioRender; Birkbak, N. https://BioRender.com/aa6hkul (2026).
Quantifying thymic health in NSCLC
We obtained imaging and clinical data from 1,218 real-world patients with NSCLC from the Harvard-NSCLC cohort (Fig. 1). All patients with NSCLC were treated with checkpoint inhibitor immunotherapy, either as immunotherapy (IO) monotherapy (n = 793; n = 245 (31%) as first-line treatment) or in combination with chemotherapy (chemo-IO; n = 425; n = 407 (96%) as first-line treatment). The demographic and clinical characteristics of the cohort are shown in Table 1. We applied a newly developed and independently validated deep-learning system to quantify imaging characteristics of the thymus on CT scans as a surrogate for thymic functionality. The system takes a CT scan as input and automatically outputs a continuous quantification of thymic health (Fig. 1b), in which higher thymic-health values are hypothesized to correspond to higher thymic functionality. For outcome analyses, patients were categorized into low (bottom 25%), average (middle 50%) and high (top 25%) thymic-health groups. These thresholds were supported by cut-point iterations (Extended Data Fig. 1).
Thymic health and outcomes in NSCLC
To investigate the associations of thymic health with progression-free survival (PFS) in patients treated for NSCLC, we used Kaplan–Meier and Cox proportional hazards analyses in the HARVARD-NSCLC cohort. Patients with average or high thymic health had significantly lower risks of progression or death following immune checkpoint inhibitor therapy than did those with low thymic health across both treatment groups (chemo-IO and IO monotherapy) (high versus low thymic health: hazard ratio, 0.65; 95% confidence interval (CI), 0.54–0.77; average versus low thymic health: hazard ratio, 0.67; 95% CI, 0.58–0.78; type III P < 0.001) (Fig. 2a), in patients treated with chemo-IO (high versus low thymic health: hazard ratio, 0.58; 95% CI, 0.41–0.82; average versus low thymic health, hazard ratio, 0.62; 95% CI, 0.47–0.82; type III P = 0.002) (Extended Data Fig. 2a) and in patients treated with IO monotherapy (high versus low thymic health: hazard ratio, 0.66; 95% CI, 0.54–0.80; average versus low thymic health: hazard ratio, 0.71; 95% CI, 0.59–0.85; type III P < 0.001) (Extended Data Fig. 2b). The effect sizes of these associations were stronger in first-line settings for patients treated with chemo-IO or IO monotherapy (Fig. 2b,c). All associations were preserved after adjustments that included—among others—sex, age, Eastern Cooperative Oncology Group (ECOG) performance status, histological subtype, PD-L1 and TMB (chemo-IO, first-line, type III P = 0.031; IO monotherapy, first-line, type III P = 0.016). Similar results were found after additional adjustments for body mass index (BMI) and smoking status (Supplementary Fig. 1), and also in subgroup analyses stratified by sex and age (Extended Data Fig. 3). Weaker associations of thymic health with prolonged PFS were found in patients who received immunotherapy as second-line or later treatment (Extended Data Fig. 2c,d).
a–f, Kaplan–Meier estimates and Cox proportional hazards models of PFS (a–c) and OS (d–f) according to thymic health (an imaging-based proxy for thymic functionality). Analyses show the comparison of high or average thymic health to the reference of low thymic health. The forest plots show the same data after multivariate adjustments. a,d, All patients with NSCLC who were treated with immunotherapy. Adjustments included sex, age, ECOG performance status, histological analysis, PD-L1, TMB, treatment line and stratification by treatment type (IO monotherapy or chemo-IO combination therapy). b,c,e,f, Subgroups of patients with NSCLC after excluding second-line-plus patients who were treated with chemo-IO combination therapy (b,e) or IO monotherapy (c,f). Adjustments included sex, age, ECOG performance status, histological analysis, PD-L1 and TMB. a–f, Age, PD-L1 and TMB were used as continuous covariates. Cox proportional hazards regression was used to estimate hazard ratios (HRs). In the forest plots, the centre of each box represents the estimated HR, and the whiskers denote the corresponding 95% CI; arrowheads indicate that the 95% CI extends beyond the visualized limits; shaded box size is for visualization only and does not encode statistical weight. The overall contribution of thymic health to uni- or multivariable models was evaluated using likelihood ratio tests (χ² tests) comparing full models with nested models excluding thymic health (type III test, two-sided), with no adjustments for multiple comparisons.
Next, we investigated the associations of thymic health with OS in the same patient cohort treated for NSCLC. In line with the associations with PFS, we found significantly lower risks of death among patients with average or high thymic health than among those with low thymic health, in all patients (high versus low thymic health: hazard ratio, 0.56; 95% CI, 0.46–0.68; average versus low thymic health: hazard ratio, 0.64; 95% CI, 0.55–0.76; type III P < 0.001) (Fig. 2d), in patients treated with chemo-IO (high versus low thymic health: hazard ratio, 0.54; 95% CI, 0.36–0.81; average versus low thymic health: hazard ratio, 0.54; 95% CI, 0.40–0.75; type III P < 0.001) (Extended Data Fig. 4a) and in patients treated with IO monotherapy (high versus low thymic health: hazard ratio, 0.56; 95% CI, 0.45–0.70; average versus low thymic health: hazard ratio, 0.70; 95% CI, 0.58–0.85; type III P < 0.001) (Extended Data Fig. 4b). Also here, the effect sizes of these associations were stronger after excluding patients who received immunotherapy as second-line or later treatment among patients treated with IO monotherapy, and similar for those who received chemo-IO as first-line treatment (Fig. 2e,f). All associations were preserved after adjustments that included—among others—sex, age, ECOG performance status, histological subtype, PD-L1 and TMB (type III P < 0.005 for both). Again, the associations were similar after additional adjustments for BMI and smoking status (Supplementary Fig. 1), and also in subgroup analyses stratified by sex and age (Extended Data Fig. 3). Weaker associations of thymic health with OS were found in patients who received second-line or later treatment (Extended Data Fig. 4c,d). We observed a consistent—although non-significant—trend towards an interaction between thymic health and immunotherapy treatment line, in which the association of thymic health with outcomes seemed to be stronger in first-line patients (Supplementary Fig. 2).
Thymic health versus PD-L1 and TMB
To compare thymic health with established clinical biomarkers, we first investigated thymic health across the clinically established PD-L1 and TMB subgroups. We found that average or high thymic health was associated with improved PFS and OS in each of the PD-L1 and TMB subgroups, as compared with low thymic health (Table 2 and Extended Data Fig. 5). Notably, among patients with the lowest PD-L1 expression (less than 1%), those who had high thymic health had a 44% lower risk of progression following immune checkpoint inhibitor therapy, compared with those who had low thymic health. Next, to benchmark the performance of thymic health against PD-L1 and TMB for their associations with clinical outcomes, we compared each biomarker’s relationship with PFS and OS, individually. We found that the average effect sizes of thymic health were similar to those of PD-L1 and TMB, for both PFS and OS in Cox models extensively adjusted for clinical variables (Fig. 3a,b and Supplementary Fig. 3). Patients who received first-line therapy for tumours with PD-L1 expression of 50% or higher had a risk of progression of 0.77 (95% CI, 0.62–0.96; P = 0.02) and a risk of death of 0.64 (95% CI, 0.49–0.83; P < 0.001), as compared with those with PD-L1 expression of less than 50%. By contrast, patients with high thymic health had risks of progression or death of 0.57 (95% CI, 0.42–0.78; type III P < 0.001) or 0.50 (95% CI, 0.35–0.72; type III P < 0.001), respectively, as compared with those with low thymic health (Extended Data Fig. 6a,b). There was no significant interaction between thymic health and PD-L1 (Supplementary Fig. 4) and no significant correlation with PD-L1 or TMB (Extended Data Fig. 6c,d). In addition, excluding thymic health from multivariate Cox models did not markedly alter the hazard ratio estimates of the remaining covariates (Supplementary Fig. 3). Together, these data indicate that thymic health provides independent and potentially complementary prognostic information beyond clinical variables and tumour-centric biomarkers.
Cox proportional hazards models of PFS and OS according to thymic health, PD-L1 and TMB, with adjustments for sex and continuous age. a,b, PFS (a) and OS (b) analyses for all patients with NSCLC who were treated with immunotherapy. Analyses according to PD-L1 and TMB were done at clinically relevant cut-points of 1% or 50% for PD-L1 and 10 mutations per Mb (mut per Mb) for TMB. a,b, Cox proportional hazards regression was used to estimate HRs. In the forest plots, the centre of each box represents the estimated HR, and the whiskers denote the corresponding 95% CI; shaded box size is for visualization only and does not encode statistical weight. The overall contribution of thymic health to multivariable models was evaluated using likelihood ratio tests (χ² tests) comparing full models with nested models excluding thymic health (type III test, two-sided). Statistical significance of individual covariate coefficients was assessed using two-sided Wald z-tests without adjustments for multiple comparisons.
Thymic health reflects thymic output
To determine the biological correlates of our image-based thymic-health quantification, we investigated thymic health in 464 prospectively enrolled, treatment-naive and operable stage I–IIIB patients with NSCLC from the TRACERx study22. Details of TRACERx patient selection and the number of patients with available data per analysis are shown in Supplementary Fig. 5.
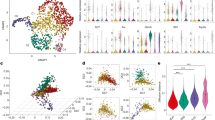
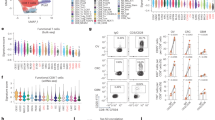
Signal joint T cell receptor excision circles (sjTRECs), a by-product of VDJ recombination during T cell maturation in the thymus23, are an established marker of thymic output20,23,24. To investigate the link between thymic output and thymic health, we quantified the levels of sjTRECs in blood from 45 patients with either low or high thymic health and available blood. Patients with high thymic health had significantly higher levels of sjTRECs (Fig. 4a), linking high thymic health with increased thymic output. These findings indicate that, in many individuals, thymic production of naive T cells persists into advanced age, potentially sustaining the T cell receptor (TCR) repertoire diversity24. To further investigate how thymic health may relate to T cell dynamics, we gathered tumour and blood-based immunological data for patients with available thymic health scores (Supplementary Fig. 5). We analysed peripheral blood TCR diversity in 50 TRACERx patients for whom data were available25. Our analysis revealed that patients with higher thymic health had greater peripheral blood α- and β-chain TCR diversity (Fig. 4b,c). Finally, we estimated the fraction of T cells in blood using T cell ExTRECT26. We found that patients with higher thymic health exhibited increased circulating T cell abundance (Fig. 4d), providing further evidence of higher thymic output in these patients.
a, Comparison of sjTREC to ribonuclease P protein subunit P30 (RPP30) ratio for patients with high or low thymic health. Split based on the full cohort. b,c, Comparison of TCR β-chain (b) and α-chain (c) normalized Shannon diversity for patients with above-median and below-median thymic health. Split based on median thymic health among patients for whom TCR data were available. d, Comparison of blood T cell fraction for patients with above-median and below-median thymic health. Split based on median thymic health among patients for whom data were available. e, Correlation between thymic health and blood protein levels, measured by Olink (n = 108). Associations were quantified using Spearman’s rank correlation, using algorithm AS 89 for P value calculation (two-sided). Associations were considered significant at a false discovery rate (FDR) < 0.25 (Benjamini–Hochberg method). f,g, REACTOME pathway enrichment analysis showing the top ten enriched pathways based on proteins associated with higher thymic health (f; n = 8 proteins, 6 in pathways) or proteins associated with lower thymic health (g; n = 91 proteins, 60 in pathways). P values were calculated using a hypergeometric test (one-sided) and adjusted using FDR (Benjamini–Hochberg method). GPCR, G-protein-coupled receptor; TNF, tumour necrosis factor. a–d, Box plots show the median (centre line), interquartile range (25th–75th percentiles; box), and whiskers extending to the minimum and maximum values within 1.5 × the interquartile range; statistical comparisons between groups were performed using two-sided Wilcoxon rank-sum tests; no adjustment for multiple comparisons was applied.
Next, we investigated the association of thymic health with tumour T cell infiltration. Notably, higher thymic health was correlated with a higher intratumoral β-chain TCR diversity, whereas this was not observed in the adjacent normal lung tissue (Extended Data Fig. 7a–d). In addition, we observed a higher fraction of T cells—estimated using T cell ExTRECT26—in tumours from patients with higher thymic health (Extended Data Fig. 7e). Splitting the data on the basis of the three thymic-health groups showed similar trends (Extended Data Fig. 7f–m). Together, these data suggest that thymic health is associated with the adaptive immune system, including peripheral blood abundance and diversity and the tumour-specific immune response.
Next, we analysed blood protein levels in 108 TRACERx patients with available Olink data27, to examine the associations between the plasma proteomic landscape and thymic health (Fig. 4e). Notably, pathway enrichment analysis revealed signalling pathways of the adaptive immune system that were positively correlated with thymic health (Fig. 4f,g). Furthermore, we investigated protein–protein interactions using the STRING database28. We found that proteins associated with higher thymic health were significantly enriched for protein–protein interactions (P = 0.017), and after expanding the network to include closely connected nodes, we identified interactions with several molecules involved in T cell regulation (Extended Data Fig. 8a). This indicates that there is a positive correlation between thymic health and proteomic mediators of adaptive immunity.
Finally, we investigated the association between thymic health and clinical factors, and found that thymic health was higher in female individuals and decreased with age (Extended Data Fig. 8b,c). As observed in the Harvard-NSCLC cohort for progression-free and overall survival, in the TRACERx cohort, patients with average to high thymic health had longer disease-free survival, as compared with those with low thymic health (high versus low thymic health: hazard ratio, 0.62; 95% CI, 0.43–0.89; average versus low thymic health: hazard ratio, 0.69; 95% CI, 0.51–0.93; type III P = 0.019; Extended Data Fig. 8d). The estimated effect size was preserved after multivariable adjustments (sex and age adjusted: high versus low thymic health: hazard ratio, 0.71; 95% CI, 0.49–1.05; average versus low thymic health: hazard ratio, 0.74; 95% CI, 0.54–1.00; type III P = 0.12; sex, age, stage and adjuvant treatment: high versus low thymic health: hazard ratio, 0.84; 95% CI, 0.57–1.24; average versus low thymic health: hazard ratio, 0.78; 95% CI, 0.49–1.25; type III P = 0.56; Extended Data Fig. 8d).
Together, these biological data provide evidence that radiographic quantification of thymic health is a proxy of thymic functionality and immune competence, and support the clinically relevant role of the thymus in patients with cancer—particularly in the context of immunotherapy.
Thymic health and pan-cancer survival
To investigate tumour-agnostic associations of thymic health with immunotherapy outcomes, we obtained imaging and clinical data from the Harvard-PAN cohort, consisting of an additional 2,258 real-world patients who were treated with ICIs for cancers of types other than NSCLC, including melanoma (n = 309), renal (n = 247), breast (n = 136), bladder (n = 133) and oesophageal (n = 120) cancers (Fig. 1). Smaller cohorts with fewer than 100 patients each were pooled as ‘others’ (n = 1,313). The demographic and clinical characteristics of the cohort are shown in Supplementary Tables 1–6, and associations between thymic health and age, sex and smoking in the pooled Harvard cohorts are shown in Extended Data Fig. 9. To investigate the association of thymic health with OS, we used Kaplan–Meier and Cox proportional hazards analyses across the different cancer types. We found lower risks of death in patients with average or high thymic health than in those with low thymic health across all examined cancer types, and significance was reached for patients with melanoma, breast cancer, renal cancer and pooled smaller types (Fig. 5). All associations were preserved with similar effect sizes after adjustments for sex and age. Together, these pan-cancer investigations suggest that thymic health has a clinically relevant role in determining outcomes to immunotherapy, and that this role is tumour-agnostic.
a–f, Kaplan–Meier estimates and Cox proportional hazards models of OS according to thymic health (an imaging-based proxy for thymic functionality). Analyses show the comparison of high or average thymic health to the reference of low thymic health. The forest plots show the same data after multivariate adjustments for sex and continuous age. Patients with melanoma (a), breast (b), renal (c), bladder (d), oesophageal (e) and pooled other cancer types with fewer than 100 cases per entity (f). a–f, Cox proportional hazards regression was used to estimate HRs. In the forest plots, the centre of each box represents the estimated HR, and the whiskers denote the corresponding 95% CI; arrowheads indicate that the 95% CI extends beyond the visualized limits; shaded box size is for visualization only and does not encode statistical weight. The overall contribution of thymic health to uni- or multivariable models was evaluated using likelihood ratio tests (χ² tests) comparing full models with nested models excluding thymic health (type III test, two-sided), with no adjustments for multiple comparisons.
Discussion
In this study we provide evidence for the clinical relevance of thymic health in immuno-oncology. We found that high thymic health was consistently associated with improved outcomes across a real-world cohort of 3,476 patients with various cancer types who were treated with ICIs. In our analysis, thymic health was a particularly strong indicator of outcome in lung cancer and melanoma—cancer types that are commonly treated with immunotherapy. Associations of thymic health and outcomes were also found in patients with breast and renal cancer, indicating that the prognostic relevance of thymic health is tumour-agnostic. Analyses in the independent and prospectively collected TRACERx cohort provide evidence that radiographic quantification of thymic health is a proxy of thymic functionality and related to immune competence. These findings suggest that an individual’s immune health relates to immunotherapy outcomes, highlighting the potential importance of considering host-specific factors as biomarkers, rather than relying solely on the tumour-centric biomarkers that at present dominate the field.
Although immunotherapy is increasingly being applied in oncology, only a subset of patients respond9,10,11. To optimize treatments and expand the population of patients who benefit from immunotherapy, improved biomarkers are needed to better estimate individual prognosis in this context. Despite the fact that ICIs derive their anti-tumour properties from T cells and their capability to invade and potentially destroy neoplasms, precision medicine in immuno-oncology has so far focused on biomarkers that predominantly capture cancer-specific biology12,13,14. We define thymic health as a universal prognostic biomarker on the basis of the radiographic characteristics of the thymus itself—an immune organ that is crucial for T cell maturation and generating a diverse T cell repertoire. These characteristics are essential for detecting foreign antigens and required for immunotherapy response.
Our results show that thymic health enables robust immuno-oncology prognostication using routine, standard-of-care thoracic CT scans without additional imaging, patient burden or delay. Although further studies are required to define biomarker-specific thresholds and assess negative and positive predictive values, our results nevertheless provide evidence that thymic health could serve as an instantly available, integral biomarker for immuno-oncology, either as a stand-alone measure or in combination with measures of PD-L1 and TMB. PD-L1 remains a well-established predictor of response, and, as expected, patients with low PD-L1 expression had worse survival following immunotherapy in NSCLC. However, our results suggest that thymic health adds independent prognostic value beyond PD-L1 by capturing the host’s immune competence. Together, these markers reflect distinct biological axes, tumour immune evasion and immune competence, and their combined assessment might offer a more complete framework for precision immuno-oncology. Our findings thus suggest that host-based biomarkers that quantify the state of adaptive immunity should be incorporated into the precision-medicine-based stratification of patients.
We show that we can identify patients with different cancer types who are likely to benefit from immunotherapy. It is conceivable that our approach could further accelerate the adoption of immunotherapy in new indications in which only a small subset of patients might benefit, and for which no biomarkers currently exist. Although higher thymic health was consistently associated with improved outcomes, the threshold for benefit seemed to vary across cancer types. In some cancers, both high and average levels of thymic health were associated with better outcomes; in others, only high levels were. This suggests that thymic health acts along a gradient, with tumour-type-specific factors, immune responsiveness and specific treatment protocols also being important factors18,29. A similar pattern has been reported for TMB, which showed stronger predictive value in some cancers, such as melanoma or NSCLC, than in others, such as breast or oesophageal cancer29. Similarly, our results suggest that thymic health assessment could facilitate the identification of patients at risk of poor outcomes following immunotherapy. Indeed, preventing non-beneficial immunotherapy is essential. It increases quality of life by reducing the risk of unnecessary adverse events, and facilitates and speeds up the selection of more favourable alternative treatments. Furthermore, it reduces population healthcare costs by avoiding unbeneficial, toxic21 and expensive therapy among these individuals.
The biological mechanism behind the observed poorer clinical outcomes seen in individuals with low thymic health is most likely to be a reduction in the output of naive T cells. This is supported by our findings in the TRACERx cohort, in which we found that relative to patients with low thymic health, those with high thymic health had significantly higher levels of sjTRECs, a by-product of thymic T cell production that is considered to be a direct readout of thymic activity23. Moreover, we showed a correlation between higher thymic health and increased T cell diversity. A similar link between the state of the thymus and T cell diversity has been reported previously20,24. Our findings thus suggest that the state of the adaptive immune system is crucial for outcomes and that adaptive immune capacity is reflected in thymic characteristics.
Notably, thymic health was prognostic in patients with advanced NSCLC receiving immunotherapy, including those who had previously been treated with cytotoxic chemotherapy. Effect sizes were strongest in the first-line immunotherapy setting, with a consistent—although non-significant—trend towards thymic health and pretreatment interaction, suggesting that the association of thymic health with outcomes is particularly relevant before chemotherapy-induced immune suppression. Given previous evidence that chemotherapy can impair thymic function19,30, our results raise the possibility that maintaining thymic health could influence the long-term benefit derived from systemic cancer therapies. Of note, the persistence of prognostic value in pretreated patients receiving immunotherapy underscores the robustness of thymic health as a host immune-competence marker even in this subset of patients. Our results might have broader implications for personalized cancer treatment31. It has previously been shown that the balance between myeloid and lymphoid cells changes with age, favouring an innate immune response. Work in mice has shown that specific depletion of myeloid precursor stem cells can restore the balance between lymphoid and myeloid compartments and reinvigorate the immune system32. Similar results could potentially be obtained in humans using similar approaches, or through specific treatments aimed at restoring thymic function33. However, any such approach to manipulate the basic elements of the immune system will require rigorous preclinical trials.
Previous studies found that thymic involution—that is, a reduction in thymic health—was a driver of immunosenescence and increased disease risk in adults34,35,36. Attempts have been made to visually score thymic involution on CT scans by estimating the degree of fatty degeneration in the thymic bed, and although no outcome implications were reported, these studies found basic associative results that are consistent with our study, such as differences in thymic involution depending on sex, age and smoking status24,37,38. Conversely to our results, these studies estimated that most adults have a fully fatty degenerated thymus24,37, whereas we found significantly improved health outcomes in association with high and average thymic health, representing 75% of the examined pan-cancer population. This indicates that using an automated deep-learning approach allows for a more detailed thymic health assessment. Indeed, our findings of maintained thymic function in most adults are supported by our findings of positive associations with T cell output in the independent TRACERx cohort; by previously reported sustained T cell output in presumed fully fatty degenerated thymic glands24; and by mathematical modelling35. Consistent with our findings and our accompanying article39, lifestyle might be directly associated with thymic health, emphasizing the need for studies that address potential preventive or regenerative strategies34,40,41.
In our study, we investigated various cancer types across independent cohorts, comprising the real-world Harvard-NSCLC and Harvard-PAN cohorts for outcome analyis and the prospective observational TRACERx study for biological correlations. We provide robust evidence of a positive association between thymic health and outcomes in patients with various cancer types who were treated with immunotherapy, strongly suggesting that thymic health should be assessed for the stratification of patients for immunotherapy treatment. This has crucial implications beyond cancer and for current clinical practice; for example, in cardiothoracic surgery and radiation oncology, the thymus is currently not considered an essential organ, and could potentially be removed20 or included in high-dose irradiation fields. However, in both instances, the thymus might be preserved with minimal adjustments to current clinical practice. Our results, supported by those of others20, suggest that there may be benefits to preserving the thymus. However, further studies are required to define the specific clinical context in which this applies. We emphasize the importance of the thymus for adult health in our accompanying article39, in which we demonstrate the relevance of the thymus for long-term health and lifespan in more than 25,000 presumably healthy individuals.
One limitation of our study is that although the included patients encompass a wide age range for both sexes, they are predominantly white, and further testing in diverse ethnic populations is required. Before the thymic-health model can be applied in clinical settings, it is essential to prove generalizability across scanners, institutions, countries and populations. Given these limitations, the presented thymic-health analyses were done using population-specific thresholding, and no universal cut-offs can be assumed at this time. However, because the development and application of the thymic-health model were performed in fully independent datasets, with high robustness as demonstrated by test–retest stability, successful external validation is likely. The next steps will include international and external validation in diverse populations of people with cancer.
Although our study design, which did not include a matched non-immunotherapy comparator, prevents the direct assessment of the predictive implications of thymic health, our analysis is consistent across multiple cohorts and cancer types. However, further work in properly powered cohorts will be required to determine whether a patient’s thymic health could serve as an independent biomarker of immunotherapy response that is equal to established tumour-intrinsic biomarkers such as PD-L1 or TMB, and that extends to cancer types other than NSCLC. This argues strongly that thymic health is a functionally important and independent component of tumour-agnostic immunotherapy efficacy.
In summary, this study provides evidence of the previously unknown importance of the thymus for immuno-oncology. Together, our work, supported by that of others19,20,24,35,37, indicates that the state of the immune system is highly individualized, varies by age and sex and associates with survival following immunotherapy.
Most current biomarkers are cancer-type-specific and are commonly derived from tumour tissue. Conversely, thymic health is host-specific and measured independently of the tumour. At present, the thymus is not examined in routine clinical care, and no established clinical standards exist. Our approach is based on CT scans and provides fast, non-invasive and personalized assessments that can be applied widely across oncology. Our observations could provide a starting point for additional studies investigating the relevance of the thymus and the immune system in maintaining health in the face of cancer and other diseases. This might lead to a more holistic approach to personalized medicine in cancer, in which the overall health of the patient and the immune system are considered together with relevant tumour biomarkers to improve treatment outcomes.
Methods
Patient cohorts
Patients from the Harvard-NSCLC and Harvard-PAN cohorts were included for the application of thymic-health quantification (see Supplementary Fig. 6 for a flow chart of patient inclusion). Patients were eligible for inclusion if they were 18 years of age or older and had baseline chest CT imaging within three months before the start of immunotherapy. This resulted in a total of 3,476 eligible patients (Harvard-NSCLC, n = 1,218; Harvard-PAN, n = 2,258). Patients had NSCLC or cancers of different entities and were treated with immunotherapy at the Dana-Farber Harvard Cancer Center (DFHCC). Clinical and imaging data were obtained using the Mass General Brigham Research Patient Data Registry (RPDR), the Oncology Data Retrieval System (OncDRS) and Medical Imaging Data As A Service (MIDAS), and, for Harvard-NSCLC, additional manual chart reviews. Patient inclusion was retrospective, and all patients with cancer who consented and were treated with the US Food and Drug Administration (FDA)-approved ICIs atezolizumab, pembrolizumab, nivolumab, ipilimumab, durvalumab, avelumab, cemiplimab, relatlimab or dostarlimab were screened for study inclusion.
For external and biological validation, we analysed 464 treatment-naive and operable patients with stage I–IIIB NSCLC from the TRACERx study, where scans and complementary data were made available (Supplementary Fig. 5). The TRACERx22 study (‘Tracking non-small cell lung cancer evolution through therapy (Rx)’; ClinicalTrials.gov: NCT01888601) is an ongoing observational trial in NSCLC that began in 2014. Data collection takes place at several hospitals across the UK, and is overseen by the sponsors of the study (Cancer Research UK and UCL Cancer Trials Center). All participants provide written informed consent. Participants are followed for up to five years from the point of primary diagnosis, through surgical resection to cure, cancer progression(s) and death. In this paper, we use disease-free survival as an outcome measure—measured from the time of study registration to the date of first lung recurrence or death from any cause. Patients who do not have these events are censored at the date they were last known to be alive or after five years. The study protocol with inclusion and exclusion criteria has been published previously42. Further details about settings and participants are provided elsewhere27,43.
Patients from all datasets were imaged with thoracic CT with or without intravenous contrast and with hard or soft kernel reconstruction according to clinical or trial protocols, generating a diverse and heterogeneous dataset ideally suited for generalizable deep-learning model application.
Inclusion and ethics
This study was approved by the respective institutional review boards at the DFHCC (DFHCC-approved protocols; 13-055, 02-180, 17-000, 20-000 and 11-117). All patients provided written informed consent at study enrolment. The TRACERx study was approved by an independent research ethics committee (13/LO/1546).
Model development and application
We developed a deep-learning system that automatically extracts a thymic-health quantification ranging from zero (complete thymic decay with fatty involution) to one (no thymic decay) from a given CT scan that covers the thoracic region. A detailed description of the model development, including training data and architecture, can be found in Supplementary Methods section S1 and Supplementary Fig. 7. In short, the deep-learning system (1) identifies the thymus centre-of-mass through seed-point annotation; (2) uses self-supervised learning-based encoding of the thymic region to capture intrinsic information in the imaging data; and (3) uses supervised prediction of thymic health from representations learned through self-supervised learning. Model development was done in a completely independent collection of 5,674 CT scans covering the thoracic region. All patients analysed and reported in this study were defined as test sets and remained unseen until the fully independent and externally developed model was locked. A technical evaluation of the performance of the end-to-end deep-learning system can be found in Supplementary Methods section S2, Supplementary Figs. 8–17 and Supplementary Tables 7–11.
Biological validation in TRACERx
We acquired preoperative CT scans for 464 patients from the TRACERx cohort with matching clinical or biological data (Supplementary Fig. 5). Immune health was estimated using our deep-learning system. For biological analysis, we excluded patients with non-fat non-thymic-soft-tissue attenuation (such as fat stranding, oedema or scarring) in the thymic bed, because we assumed potential alteration of biological mechanisms among these patients19,34,44 (Supplementary Methods section 1.3). The abundance of sjTRECs was assessed from buffy-coat samples in a subset of 45 patients (25 with high thymic health and 20 with low thymic health). Buffy coat (100 µl) was isolated and DNA extracted using a modified protocol of the QIAGEN DNeasy Blood & Tissue Kit (69506, QIAGEN). In brief, samples were lysed with proteinase K and buffer AL, followed by ethanol precipitation and purification on QIAGEN spin columns, with elution in 50 µl EB buffer. DNA concentration was measured spectrophotometrically, and 600 ng DNA per reaction was used. For droplet digital PCR (ddPCR), DNA was pre-digested with HindIII (BioNordika) to linearize sjTREC molecules (20 min at 37 °C, enzyme inactivation at 80 °C for 10 min). sjTREC levels were quantified using primers from a previously published assay24: 5′-CACATCCCTTTCAACCATGCT-3′ (forward), 5′-GCCAGCTGCAGGGTTTAGG-3′ (reverse) and probe 5′-ACACCTCTGTTTTTGTAAAGGTGCCCACT-3′ (5′6-FAM, Iowa Black quencher). As a reference, the housekeeping gene RPP30 was amplified in parallel using forward primer 5′-AGATTTGGACCTGCGAGCG-3′, reverse primer 5′-GAGCGGCTGTCTCCACAAGT-3′ and probe 5′-TTCTGACCTGAAGGCTCTGCGCG-3′ (5′6-HEX, Iowa Black quencher). Each 20-µl reaction contained 10 µl ddPCR Supermix for Probes (no dUTP, Bio-Rad, 1863024), 1 µl sjTREC assay, 1 µl RPP30 assay and 8 µl digested DNA. Droplets were generated on a QX200 Automated Droplet Generator (Bio-Rad), sealed and amplified on a C1000 Touch Thermal Cycler (Bio-Rad) with the following cycling protocol: 95 °C for 10 min; 40 cycles of 94 °C for 30 s and 55 °C for 1 min (ramp rate 2 °C per s); 98 °C for 10 min; and hold at 12 °C. Plates were read on a QX200 Droplet Reader (Bio-Rad), and data were analysed using QuantaSoft and QX Manager software (Bio-Rad). All samples were run in triplicate. sjTREC abundance was normalized to RPP30 copy number, and results presented as the sjTREC/RPP30 ratio. TCR diversity was calculated on the basis of previously published TCR-sequencing data25. In short, TCR α- and β-chain sequencing were performed on whole RNA extracted from each tissue type: tumour tissue, adjacent normal lung tissue and cryopreserved peripheral blood mononuclear cell (PBMC) samples. TCR data were analysed in a tissue-specific manner and were not combined across tissues. Raw reads were processed using Decombinator45 to assemble TCR clones. Normalized Shannon diversity was calculated from productive TCR sequences using the formula: −1/log(N) × ∑((N, i = 1) pi × log (pi)), where N is the total number of TCRs in a sample and pi is the frequency of TCR i. Median TCR diversity was used for patients with multiple tumour regions available. For TCR analysis, we included an additional patient (CRUK0291) with TCR-sequencing data and a CT scan available, who was excluded from clinical follow-up owing to incomplete resection. Blood and tumour T cell fractions were calculated from available whole-exome sequencing (WES) data using T cell ExTRECT, as described previously26. In brief, T cell ExTRECT estimates the fraction of T cells in a WES sample by quantifying the amount of TCR excision at the TCRA locus, on the basis of the drop in read depth at this site. For blood T cell fraction, samples with a fraction of 0 were considered to be technical artefacts and excluded from the analysis (n = 31). In tumour samples, the T cell fraction was adjusted based on TCRA copy number and purity. Median T cell fraction was used for patients with multiple tumour regions available. Olink Explore 3072 proteomics data, measuring the relative plasma concentration of a panel of proteins in blood plasma samples, were available for 108 patients as part of a previous publication27. Proteins associated with thymic health were found by calculating Spearman’s rank correlation coefficients (P values from algorithm AS 89, two-sided) between our thymic-health score and the relative plasma concentration of proteins across all patients, using the rstatix package (v.0.7.2). To ensure accurate correlation estimates, we excluded proteins with Normalized Protein eXpression (NPX) levels below the limit of detection in more than 20% of patients, resulting in 2,743 included proteins.
Pathway enrichment analysis was performed on the basis of REACTOME46 pathway gene sets (v.78, from MSigDB v7.5.1). UniProt IDs were mapped to ENTREZ IDs using org.Hs.eg.db (v.3.14.0) To use a panel-specific null hypothesis, pathways were filtered to only include proteins that were included in the filtered Olink Explore 3072 panel. Enrichment score was calculated using the clusterProfiler package (v.4.2.0), using pathways that included at least ten genes. Protein–protein interactions were determined using the STRING protein–protein association database (v.12.028), using the STRING web-based user interface (https://string-db.org/), with an interaction confidence threshold of 400. The five most closely related proteins were added by using the ‘add more proteins to your network’ functionality once. Enrichment analysis used the STRINGdb package (v.2.6.0), with proteins from the filtered Olink panel as background.
Statistical analysis
To facilitate thymic-health interpretation, the raw thymic-health values were calibrated across the study population by applying matched-percentile ranking. This transformed the raw thymic-health values to a range from 0 to 100, matching the individual distribution rank; that is, a patient with a calibrated thymic-health value of 50 would be found in the median of the patient population distribution. Furthermore, to facilitate potential clinical translation, we categorized thymic health into low, average and high, based on the first and third population quartiles; that is, patients with calibrated thymic health ≤25, 25–75 and >75 were categorized as low, average and high thymic health, respectively. Summary statistics for continuous variables used the mean, median or range; categorical variables were summarized as proportions or percentages. PFS was determined from the start date of immunotherapy until the date of disease progression or death. OS was determined from the start date of immunotherapy until the date of death. If patients were alive at the last contact, they were censored accordingly. The follow-up was truncated at five years. Associations were computed using Wilcoxon rank-sum tests and linear regression as appropriate. Time-to-event distributions used the Kaplan–Meier estimator, and Cox proportional hazards models were used to calculate estimates of hazard ratios in univariate and multivariate modelling. The Schoenfeld residuals were assessed, and the proportional hazards assumptions were met. P values are two-sided, the level of significance was pre-defined at <0.05 and CIs are at the level of 95%. No statistical methods were used to predetermine sample size. Statistical analyses were performed using R v.4.2.2 (R Project for Statistical Computing).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Both the Harvard-NSCLC and the Harvard-PAN dataset used in this project were curated with institutional permission through approval by the institutional review board for the current study, and thus cannot be made publicly available in compliance with patient privacy obligations. All requests for processed data, including the thymic-health scores, will be evaluated according to institutional and departmental policies to determine whether the data requested are subject to intellectual property or patient privacy obligations. Data that can be transferred will require a material or data transfer agreement between the institutions and will limit the utility of the data to non-commercial academic research purposes. The exact timeline will depend on the execution of such agreements. Please email all requests to the corresponding author. For the TRACERx dataset, the WES data used in this publication are available at the European Genome–phenome Archive (EGA), which is hosted by the European Bioinformatics Institute (EBI) and the Centre for Genomic Regulation (CRG), under the accession code EGAS00001006494 (controlled access due to the nature of the data and commercial licences). Specifically, data are available through the Cancer Research UK and UCL Cancer Trials Centre (ctc.tracerx@ucl.ac.uk) for academic non-commercial research purposes only, and this is subject to review of a project proposal by the TRACERx data access committee, entering into an appropriate data access agreement and any applicable ethical approvals. A response to the request for access is typically provided within ten working days of the committee receiving the relevant project proposal and all other required information. The TRACERx TCR-sequencing FASTQ data are available at the Sequence Read Archive (SRA) under accession code BioProject: PRJNA544699. TRACERx thymic-health scores and associated data used in this analysis are available through Zenodo: https://doi.org/10.5281/zenodo.18330021 (ref. 47). Protein interactions were determined using the STRING database (v.12; https://string-db.org). Pathway analysis used the REACTOME pathways (v.78; https://reactome.org). Public imaging data collections downloaded from the Imaging Data Commons (IDC; https://portal.imaging.datacommons.cancer.gov/) were used in the development of the deep-learning model. An overview of these datasets is provided in Supplementary Table 7.
Code availability
The code used to perform the statistical analysis and create figures is available through Zenodo: https://doi.org/10.5281/zenodo.18330021 (ref. 47). The software used in the publication is available on GitHub for public, non-commercial use in our GitHub (https://github.com/AIM-Harvard/thymus_health_deeplearning_system.git). Additional technical details about the development and evaluation of our deep-learning framework are provided in the Supplementary Information. The models’ weights are subject to intellectual property obligations and cannot be shared publicly, but may be made available through academic collaboration. For more details, please contact the corresponding author.
References
Das, S. & Johnson, D. B. Immune-related adverse events and anti-tumor efficacy of immune checkpoint inhibitors. J. Immunother. Cancer 7, 306 (2019).
Waldman, A. D., Fritz, J. M. & Lenardo, M. J. A guide to cancer immunotherapy: from T cell basic science to clinical practice. Nat. Rev. Immunol. 20, 651–668 (2020).
Goldrath, A. W. & Bevan, M. J. Selecting and maintaining a diverse T-cell repertoire. Nature 402, 6–13 (1999).
Robert, C. et al. Durable complete response after discontinuation of pembrolizumab in patients with metastatic melanoma. J. Clin. Oncol. 36, 1668–1674 (2018).
Luke, J. J. et al. Pembrolizumab versus placebo as adjuvant therapy in completely resected stage IIB or IIC melanoma (KEYNOTE-716): a randomised, double-blind, phase 3 trial. Lancet 399, 1718–1729 (2022).
Rozeman, E. A. et al. Identification of the optimal combination dosing schedule of neoadjuvant ipilimumab plus nivolumab in macroscopic stage III melanoma (OpACIN-neo): a multicentre, phase 2, randomised, controlled trial. Lancet Oncol. 20, 948–960 (2019).
Singh, N. et al. Therapy for stage IV non-small-cell lung cancer without driver alterations: ASCO living guideline. J. Clin. Oncol. 40, 3323–3343 (2022).
Xu, W., Atkins, M. B. & McDermott, D. F. Checkpoint inhibitor immunotherapy in kidney cancer. Nat. Rev. Urol. 17, 137–150 (2020).
Eggermont, A. M. M. et al. Adjuvant pembrolizumab versus placebo in resected stage III melanoma. N. Engl. J. Med. 378, 1789–1801 (2018).
Larkin, J. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Hellmann, M. D. et al. Nivolumab plus ipilimumab in advanced non-small-cell lung cancer. N. Engl. J. Med. 381, 2020–2031 (2019).
Davis, A. A. & Patel, V. G. The role of PD-L1 expression as a predictive biomarker: an analysis of all US Food and Drug Administration (FDA) approvals of immune checkpoint inhibitors. J. Immunother. Cancer 7, 278 (2019).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Wang, D.-R., Wu, X.-L. & Sun, Y.-L. Therapeutic targets and biomarkers of tumor immunotherapy: response versus non-response. Signal Transduct. Target. Ther. 7, 331 (2022).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Haragan, A. et al. Heterogeneity of PD-L1 expression in non-small cell lung cancer: Implications for specimen sampling in predicting treatment response. Lung Cancer 134, 79–84 (2019).
Li, S., Zhang, C., Pang, G. & Wang, P. Emerging blood-based biomarkers for predicting response to checkpoint immunotherapy in non-small-cell lung cancer. Front. Immunol. 11, 603157 (2020).
Usset, J. et al. Five latent factors underlie response to immunotherapy. Nat. Genet. 56, 2112–2120 (2024).
Wang, W., Thomas, R., Sizova, O. & Su, D. M. Thymic function associated with cancer development, relapse, and antitumor immunity – a mini-review. Front. Immunol. 11, 773 (2020).
Kooshesh, K. A., Foy, B. H., Sykes, D. B., Gustafsson, K. & Scadden, D. T. Health consequences of thymus removal in adults. N. Engl. J. Med. 389, 406–417 (2023).
Fenioux, C. et al. Thymus alterations and susceptibility to immune checkpoint inhibitor myocarditis. Nat. Med. 29, 3100–3110 (2023).
Bailey, C. et al. Tracking cancer evolution through the disease course. Cancer Discov. 11, 916–932 (2021).
Douek, D. C. et al. Changes in thymic function with age and during the treatment of HIV infection. Nature 396, 690–695 (1998).
Sandstedt, M. et al. Complete fatty degeneration of thymus associates with male sex, obesity and loss of circulating naïve CD8+ T cells in a Swedish middle-aged population. Immun. Ageing 20, 45 (2023).
Joshi, K. et al. Spatial heterogeneity of the T cell receptor repertoire reflects the mutational landscape in lung cancer. Nat. Med. 25, 1549–1559 (2019).
Bentham, R. et al. Using DNA sequencing data to quantify T cell fraction and therapy response. Nature 597, 555–560 (2021).
Al-Sawaf, O. et al. Body composition and lung cancer-associated cachexia in TRACERx. Nat. Med. 29, 846–858 (2023).
Szklarczyk, D. et al. The STRING database in 2023: protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Sinha, N. et al. Immune determinants of the association between tumor mutational burden and immunotherapy response across cancer types. Cancer Res. 82, 2076–2083 (2022).
Kinsella, S. & Dudakov, J. A. When the damage is done: injury and repair in thymus function. Front. Immunol. 11, 1745 (2020).
Rohaan, M. W. et al. Tumor-infiltrating lymphocyte therapy or ipilimumab in advanced melanoma. N. Engl. J. Med. 387, 2113–2125 (2022).
Ross, J. B. et al. Depleting myeloid-biased haematopoietic stem cells rejuvenates aged immunity. Nature 628, 162–170 (2024).
Napolitano, L. A. et al. Growth hormone enhances thymic function in HIV-1-infected adults. J. Clin. Invest. 118, 1085–1098 (2008).
Duah, M. et al. Thymus degeneration and regeneration. Front. Immunol. 12, 706244 (2021).
Palmer, S., Albergante, L., Blackburn, C. C. & Newman, T. J. Thymic involution and rising disease incidence with age. Proc. Natl Acad. Sci. USA 115, 1883–1888 (2018).
Thomas, R., Wang, W. & Su, D.-M. Contributions of age-related thymic involution to immunosenescence and inflammaging. Immun. Ageing 17, 2 (2020).
Araki, T. et al. Normal thymus in adults: appearance on CT and associations with age, sex, BMI and smoking. Eur. Radiol. 26, 15–24 (2016).
Ackman, J. B. et al. Sex difference in normal thymic appearance in adults 20-30 years of age. Radiology 268, 245–253 (2013).
Bernatz, S. et al. Thymic health consequences in adults. Nature https://doi.org/10.1038/s41586-026-10242-y (2026).
Duggal, N. A., Pollock, R. D., Lazarus, N. R., Harridge, S. & Lord, J. M. Major features of immunesenescence, including reduced thymic output, are ameliorated by high levels of physical activity in adulthood. Aging Cell 17, e12750 (2018).
Chaudhry, M. S., Velardi, E., Dudakov, J. A. & van den Brink, M. R. M. Thymus: the next (re)generation. Immunol. Rev. 271, 56–71 (2016).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Frankell, A. M. et al. The evolution of lung cancer and impact of subclonal selection in TRACERx. Nature 616, 525–533 (2023).
Nunes-Alves, C., Nobrega, C., Behar, S. M. & Correia-Neves, M. Tolerance has its limits: how the thymus copes with infection. Trends Immunol. 34, 502–510 (2013).
Thomas, N., Heather, J., Ndifon, W., Shawe-Taylor, J. & Chain, B. Decombinator: a tool for fast, efficient gene assignment in T-cell receptor sequences using a finite state machine. Bioinformatics 29, 542–550 (2013).
Milacic, M. et al. The Reactome Pathway Knowledgebase 2024. Nucleic Acids Res. 52, D672–D678 (2024).
Prudente, V. C. G. et al. Resources “Thymic health and immunotherapy outcomes in patients with cancer”. Zenodo https://doi.org/10.5281/zenodo.18330021 (2026).
Acknowledgements
We acknowledge financial support from the National Institutes of Health (NIH) (H.J.W.L.A.: NIH-USA U24CA194354, NIH-USA U01CA190234, NIH-USA U01CA209414 and NIH-USA R35CA22052; D.B.: NIH-USA R01CA294033), and the European Union–European Research Council (H.J.W.L.A.: 866504). S.B. acknowledges funding from the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) - 502050303, and N.J.B. acknowledges funding from the Lundbeck Foundation (R272-2017-4040), the Novo Nordisk Foundation (NNF21OC0071483, NNF23OC0085954) and a Savvaerksejer Jeppe Juhl og Hustru Ovita Juhl research stipend. M.J.-H. has received funding from Cancer Research UK, the NIH National Cancer Institute, the IASLC International Lung Cancer Foundation, the Lung Cancer Research Foundation, Rosetrees Trust, UKI NETs and NIHR. We thank the National Cancer Institute for collecting and making the data from the National Lung Screening Trial (NLST) accessible, and the Cancer Imaging Archive (TCIA) and the Imaging Data Commons (IDC) for making this and other imaging collections used for developing our deep-learning model available on their platforms. We acknowledge the DFCI Oncology Data Retrieval System (OncDRS) for the aggregation, management and delivery of the clinical and operational research data used in this project. The content is solely the responsibility of the authors.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.B., S.P., V.P., A.K.A., N.J.B. and H.J.W.L.A. Methodology: S.B., S.P., V.P., A.K.A., L.N., C.A., N.J.B. and H.J.W.L.A. Software: S.P., V.P. and S.B. Validation: S.B., V.P., S.P., A.K.A., N.M., A.H., R.H.M., M.J.-H., C.S., N.J.B. and H.J.W.L.A. Formal analysis: S.B., V.P., S.P., A.K.A., A.R. and S.V. Investigation: S.B., V.P., S.P. and A.K.A. Resources: H.J.W.L.A., M.J.-H., C.S., N.J.B. and D.B. Data curation: S.B., V.P., S.P., A.D.F., L.D., J.V.A., P.A.O., E.S., D.B., M.A. and B.R. Writing (original draft): S.B., V.P., S.P., A.K.A., H.J.W.L.A. and N.J.B. Writing (review and editing): all authors. Visualization: S.B., A.K.A., S.P., V.P., L.N., H.J.W.L.A. and N.J.B. Supervision: H.J.W.L.A. and N.J.B. Project administration: H.J.W.L.A. and N.J.B. Funding acquisition: H.J.W.L.A., S.B. and N.J.B.
Corresponding author
Ethics declarations
Competing interests
M.J.-H. has consulted for Astex Pharmaceutical and Achilles Therapeutics, is a member of the Achilles Therapeutics scientific advisory board (SAB) and steering committee and has received speaker honoraria from Pfizer, Astex Pharmaceuticals, Oslo Cancer Cluster, Bristol Myers Squibb and Genentech. M.J.-H. is listed as a co-inventor on a European patent application relating to methods to detect lung cancer (PCT/US2017/028013); this patent has been licensed to commercial entities and, under terms of employment, M.J.-H. is due a share of any revenue generated from such licence(s). M.J.-H. is also listed as a co-inventor on a GB priority patent application (GB2400424.4: ‘Treatment and Prevention of Lung Cancer’). C.S. has received support from the Francis Crick Institute and the Royal Society; has received grants or contracts from AstraZeneca, Boehringer Ingelheim, Bristol Myers Squibb, Invitae (formerly Archer Dx), Ono Pharmaceuticals, Pfizer and Roche-Ventana; has received consulting fees from Bicycle Therapeutics, Genentech, Medicxi, Metabomed and Novartis; is a member of the GRAIL SAB, the Relay Therapeutics SAB, China Innovation Centre of Roche (CICoR), the SAGA Diagnostics SAB and the Sarah Cannon Research Institute; has received honoraria from Amgen, AstraZeneca, Bristol Myers Squibb, Illumina, GlaxoSmithKline, MSD, Roche-Ventana and Pfizer; holds patents, planned, issued or pending, including PCT/GB2017/053289, PCT/EP2016/059401, PCT/EP2016/071471, PCT/GB2018/052004, PCT/GB2020/050221, PCT/GB2018/051912, PCT/US2017/28013 and PCT/GB2018/051892; has leadership or fiduciary roles with Cancer Research UK and the American Association for Cancer Research (AACR); holds stock or stock options in ApoGen Biotech, Epic Biosciences, GRAIL and Achilles Therapeutics; and has other financial or non-financial interests with AstraZeneca and GRAIL Bio UK. N.M. holds patents related to determining human leukocyte antigen (HLA) loss of heterozygosity (LOH) (PCT/GB2018/052004), determining the B cell fraction in mixed samples (PCT/EP2024/062999), determining lymphocyte abundance in mixed samples (PCT/EP2022/070694), identifying responders to cancer treatment (PCT/GB2018/051912), targeting neoantigens (PCT/EP2016/059401), identifying patient response to immune checkpoint blockade (PCT/EP2016/071471) and predicting survival rates of patients with cancer (PCT/GB2020/050221), and has a patent pending in determining HLA disruption (PCT/EP2023/059039). S.V. is a co-inventor on a patent of methods for detecting molecules in a sample (US 10,578,620). A.H. has received fees for being a member of independent data monitoring committees for Roche-sponsored clinical trials and academic projects coordinated by Roche. S.B. reports consulting fees from Ambient. N.J.B. reports consulting fees from Ambient and is listed as a co-inventor on a patent application (PCT/GB2020/050221) on methods for cancer prognostication and a patent on methods for predicting anti-cancer response (US14/466,208). H.J.W.L.A. reports consulting fees and/or stock from Onc.AI, Love Health, Sphera, HealthAI, Ambient and AstraZeneca. R.H.M.: advisory board (ViewRay and AstraZeneca), consulting (AstraZeneca, Varian Medical Systems, Sio Capital Management and Pfizer), honoraria (Novartis, Springer Nature), research funding (NIH, ViewRay, AstraZeneca, Siemens Medical Solutions USA and Varian Medical Systems). D.B. is a member of the MercurialAI SAB. L.D. has sponsored research agreements with C2i Genomics, Natera, AstraZeneca, Photocure and Ferring; an advisory or consulting role at Ferring, MSD, Cystotech and UroGen; and has received speaker honoraria from AstraZeneca, Pfizer and Roche and travel support from MSD. A.D.F. reports fees and advisory fees from Novartis, IQVIA and Hanson-Wade. C.A. is an employee of Astrazeneca and owns stocks in Astrazeneca; and holds patents/patent applications in minimal residual disease detection (PCT/GB2017/053289, PCT/US2017/028013 and PCT/EP2022/077987). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature thanks Bharat Thyagarajan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Association of increasing thymic health cut-point thresholds with long-term risk of mortality.
a, Risk of death according to increasing thymic-health thresholds in the pooled pan-cancer cohort (Harvard-NSCLC and Harvard-PAN; total, n = 3,476) revealed no clear binarization threshold. This analysis compares the entire population divided into two groups at various percentile thresholds (e.g., above vs. below a given percentile). b, Risk of death in each thymic-health decile versus the lowest decile, as reference, in the pooled pan-cancer cohort (n = 3,476) indicated low, average, and high-performance pan-cancer decile subsets. This analysis provides a more granular analysis by comparing individual deciles (i.e., specific 10% slices) against the lowest decile (bottom 10% thymic health). Panel b demonstrates that no single decile disproportionately drives the association, indicating the absence of an outlier effect and supporting the stability and robustness of the associations. This explains why the binary splits shown in subpanel a consistently produce HRs within a similar range, i.e., the underlying risk reduction is broadly distributed. a,b, Cox proportional hazards regression was used to estimate HRs. In the forest plots, the centre of each box represents the estimated hazard ratio, and the whiskers denote the corresponding 95% CI; arrowheads indicate that the 95% CI extends beyond the visualized limits; shaded box size is for visualization only and does not encode statistical weight. Statistical significance of the binarized thymic-health covariate coefficients at the different cut-points was assessed using two-sided Wald z-tests without adjustments for multiple comparisons.
Extended Data Fig. 2 Association of thymic health with PFS in patients with NSCLC.
Kaplan–Meier estimates and Cox proportional hazards models of PFS according to thymic health, i.e., an imaging-based proxy for thymic functionality. Analyses show the comparison of high or average thymic health to the reference of low thymic health. The forest plots show the same data after multivariate adjustments. Patients were treated with a, immunotherapy in combination with chemotherapy (Chemo-IO), or b, immunotherapy monotherapy (IO mono) - with adjustments for sex, age, ECOG performance status, histologic analysis, PD-L1, TMB, and treatment line. c,d, Subgroup of second-line plus patients, i.e., after exclusion of first-line patients, with NSCLC who were treated with c, Chemo-IO combination therapy or d, IO monotherapy - adjustments in d, included sex, age, ECOG performance status, histologic analysis, PD-L1, and TMB, while in c, adjustments were not performed due to limited sample size. a–d, Cox proportional hazards regression was used to estimate HRs. In the forest plots, the centre of each box represents the estimated hazard ratio, and the whiskers denote the corresponding 95% CI; arrowheads indicate that the 95% CI extends beyond the visualized limits; shaded box size is for visualization only and does not encode statistical weight. The overall contribution of thymic health to multivariable models was evaluated using likelihood ratio tests (χ² tests) comparing full models with nested models excluding thymic health (type III test, two-sided) with no adjustments for multiple comparisons.
Extended Data Fig. 3 Association of thymic health with progression-free and overall survival in patients with NSCLC in sex and age subgroups.
Cox proportional hazards models of PFS and OS of the Harvard-NSCLC cohort in subgroups according to a, sex with adjustments for age, ECOG performance status, and histologic subtype, b, age split at 65 years with adjustments for sex, ECOG performance status, and histologic subtype. a,b, Cox proportional hazards regression was used to estimate HRs. In the forest plots, the centre of each box represents the estimated hazard ratio, and the whiskers denote the corresponding 95% CI; shaded box size is for visualization only and does not encode statistical weight. The overall contribution of thymic health to multivariable models was evaluated using likelihood ratio tests (χ² tests) comparing full models with nested models excluding thymic health (type III test, two-sided) with no adjustments for multiple comparisons. GTE, greater than or equal to; LT, less than.
Extended Data Fig. 4 Association of thymic health with overall survival in patients with NSCLC.
Kaplan–Meier estimates and Cox proportional hazards models of OS according to thymic health, i.e., an imaging-based proxy for thymic functionality. Analyses show the comparison of high or average thymic health to the reference of low thymic health. The forest plots show the same data after multivariate adjustments. Patients were treated with a, immunotherapy in combination with chemotherapy (Chemo-IO), or b, immunotherapy monotherapy (IO mono) - with adjustments for sex, age, ECOG performance status, histologic analysis, PD-L1, TMB, and treatment line. c,d, Subgroup of second-line plus patients, i.e., after exclusion of first-line patients, with NSCLC who were treated with c, Chemo-IO combination therapy or d, IO monotherapy - adjustments in d, included sex, age, ECOG performance status, histologic analysis, PD-L1, and TMB, while in c, adjustments were not performed due to limited sample size. a–d, Cox proportional hazards regression was used to estimate HRs. In the forest plots, the centre of each box represents the estimated hazard ratio, and the whiskers denote the corresponding 95% CI; arrowheads indicate that the 95% CI extends beyond the visualized limits; shaded box size is for visualization only and does not encode statistical weight. The overall contribution of thymic health to multivariable models was evaluated using likelihood ratio tests (χ² tests) comparing full models with nested models excluding thymic health (type III test, two-sided) with no adjustments for multiple comparisons.
Extended Data Fig. 5 Association of thymic health with overall survival in patients treated with immunotherapy for NSCLC across clinically relevant levels of PD-L1.
Survival of patients with NSCLC according to programmed death-ligand 1 (PD-L1) expression levels of a, less than 50%, or b, at least 50%. c,d, Hazard ratio adjusted by sex- and age Cox proportional hazards models according to the same PD-L1 expression levels of c, less than 50%, or d, at least 50%. a–d, Cox proportional hazards regression was used to estimate HRs. In the forest plots, the centre of each box represents the estimated hazard ratio, and the whiskers denote the corresponding 95% CI; shaded box size is for visualization only and does not encode statistical weight. The overall contribution of thymic health to multivariable models was evaluated using likelihood ratio tests (χ² tests) comparing full models with nested models excluding thymic health (type III test, two-sided) with no adjustments for multiple comparisons.
Extended Data Fig. 6 Association of thymic health, PD-L, and TMB with progression-free and overall survival in patients who had first-line treatment for NSCLC, and associations of thymic health with PD-L1 and TMB across the whole NSCLC cohort.
a,b, Cox proportional hazards models of PFS and OS according to thymic health, PD-L1, and TMB with adjustments for sex and age. a, PFS and b, OS analyses for first-line patients with NSCLC who were treated with immunotherapy. Analyses according to PD-L1 and TMB were done at clinically relevant cut-points of 1% or 50% for PD-L1 and 10 mutations per Megabase (mut/Mb) for TMB. a,b, Cox proportional hazards regression was used to estimate HRs. In the forest plots, the centre of each box represents the estimated hazard ratio, and the whiskers denote the corresponding 95% CI; shaded box size is for visualization only and does not encode statistical weight. The overall contribution of thymic health to multivariable models was evaluated using likelihood ratio tests (χ² tests) comparing full models with nested models excluding thymic health (type III test, two-sided). Statistical significance of individual covariate coefficients was assessed using two-sided Wald z-tests without adjustments for multiple comparisons. c,d, Scatter plots of female (green) and male (purple) patients from the Harvard-NSCLC cohort with no missing data for c, PD-L1 and d, TMB, as analysed in main Fig. 3 (n = 750), are shown. Linear regression line and Spearman rank correlation analyses are shown independently for male and female patients. Spearman rank correlation coefficients (r) and two-sided P values were computed separately for each group using asymptotic t approximation. No adjustment for multiple comparisons was applied.
Extended Data Fig. 7 Associations of thymic health with T cell biology in TRACERx.
a–d, Association of thymic health with TCR α- and β-chain normalized Shannon diversity in a,b, tumour or c,d, adjacent lung tissue. Split by median thymic health among patients with the corresponding TCR data available. e, Comparison of tumour T cell fraction for patients with over and under median thymic health. Split by median thymic health among patients with data available. f–k, Association of thymic health with TCR α- and β-chain normalized Shannon diversity in f,g, blood, h,i, tumour tissue, or j,k, adjacent lung tissue. l,m, Comparison of thymic health and T cell fraction in l, blood, or m, tumour tissue. Low, average, and high correspond to bottom 25%, middle 50%, and top 25% thymic health in the full TRACERx cohort. Tumour diversity and T cell fraction were summarized to median for patients with multiple tumour regions available. Box plots show the median (centre line), interquartile range (25th–75th percentiles; box), and whiskers extending to the minimum and maximum values within 1.5 x the interquartile range. Statistical comparisons between groups were performed using two-sided Wilcoxon rank-sum tests. f–m, Low thymic health was used as the reference group. No adjustment for multiple comparisons was applied.
Extended Data Fig. 8 Biological and clinical associations with thymic health in TRACERx.
a, STRING protein–protein interaction network of proteins associated with higher thymic health (n = 8 proteins; from Olink proteomics analysis in Fig. 4e). The network is expanded to include the five most closely connected nodes. b,c, Association of thymic health with b, sex and c, age groups. Box plots show the median (centre line), interquartile range (25th–75th percentiles; box), and whiskers extending to the minimum and maximum values within 1.5 x the interquartile range. Statistical comparisons between groups were performed using two-sided Wilcoxon rank-sum tests. <65 was used as the reference group in c. d, Kaplan–Meier estimate and Cox proportional hazards model of disease-free survival in the TRACERx cohort according to thymic health. Analyses show the comparison of high or average thymic health to the reference of low thymic health. The top forest plot shows the same data after multivariate adjustments for sex and age. The bottom forest plot is additionally adjusted for disease stage and adjuvant treatment in the subset in whom this data is available (n=337). The overall contribution of thymic health to uni- or multivariable models was evaluated using likelihood ratio tests (χ² tests) comparing full models with nested models excluding thymic health (type III test, two-sided).
Extended Data Fig. 9 Associations of thymic health with age, sex and smoking history.
Analyses include the pooled Harvard-NSCLC and Harvard-PAN cohorts. Beta values are the effect estimate of the linear regression models using age, sex, or smoking history as predictors to model thymic health as outcome. In the forest plots, the centre of each box represents the estimated regression coefficient, and the whiskers denote the corresponding 95% CI. Statistical significance of individual coefficients was evaluated using two-sided t-tests. No adjustment for multiple comparisons was applied.
Supplementary information
Supplementary Information (download PDF )
This file contains two sections of Supplementary Methods that describe the technical details of the development and validation of the deep-learning approach. In addition, 17 Supplementary Figures and 11 Supplementary Tables present results related to the above-mentioned technical methods, or supplementary exploratory analysis related to the analyses presented in the manuscript.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bernatz, S., Prudente, V., Pai, S. et al. Thymic health and immunotherapy outcomes in patients with cancer. Nature (2026). https://doi.org/10.1038/s41586-026-10243-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10243-x