Abstract
Chronic inflammation is a well-established risk factor for cancer, but the underlying molecular mechanisms remain unclear1,2. Using a mouse model of colitis, we demonstrate that colonic stem cells retain an epigenetic memory of inflammation following disease resolution that persists for more than 100 days. Here we find that memory of colitis is characterized by a cumulative gain of activator protein 1 (AP-1) transcription factor activity, with durable changes to chromatin accessibility. Further, we develop SHARE-TRACE, a method that enables simultaneous profiling of gene expression, chromatin accessibility and clonal history in single cells, enabling high-resolution tracking of epigenomic memory. This approach reveals that memory of colitis is propagated cell-intrinsically and inherited through stem cell divisions, with some clones demonstrating stronger memory than others. Finally, we show that colitis primes stem cells for increased expression of an AP-1-regulated gene program following oncogenic mutation that accelerates tumour growth, a phenotype dependent on AP-1 activity. Together, our findings provide a mechanistic link between chronic inflammation and malignancy, revealing how long-lived epigenetic alterations in regenerative tissues may contribute to disease susceptibility and suggesting potential diagnostic and therapeutic strategies to mitigate cancer risk in patients with chronic inflammatory conditions.
Similar content being viewed by others
Main
Inflammation is a major risk factor for cancer, whether it is due to autoimmune disease, long-term infections or environmental exposures1,2, and risk often increases with the duration and severity of disease3,4. Whereas diverse mechanisms may explain this connection, including the acquisition of DNA mutations1,2, we propose that inflammation may cause lasting epigenetic alterations that lower the threshold for oncogenesis. In support, the role of the epigenome as a causal driver in cancer has become clear5,6,7. The epigenome is dynamically regulated as cells respond to environmental challenges by making new regions of their DNA accessible and active, directing transcription factor (TF) proteins to these sites and activating the expression of new genes and cellular functions. During regeneration and immunity, alterations to the epigenome can persist and accumulate following repeated exposure8,9,10, enhancing subsequent responses to secondary stimuli7,11,12,13,14,15,16,17. Although this epigenetic ‘memory’ is largely described as adaptive, evidence suggests that it may also carry maladaptive consequences and increase future risk of disease7,16,18. Here we look to study how epigenetic memory accumulates within cells, is inherited across clones and influences predisposition for cancer. These epigenetic mechanisms may prove to be of central importance to cancer biology, providing missing mechanisms connecting lifestyle and environmental exposures to malignancy.
The gastrointestinal tract is an intriguing system in which to study epigenetic memory given its immense exposure to the environment19. There are well-established clinical associations between inflammation in the gut and cancer, including between ulcerative colitis and colorectal carcinoma (CRC). Patients with ulcerative colitis are twofold to fivefold more likely to develop cancer, with those diagnosed during childhood or with pancolitis carrying a substantially higher risk3,4. The presumed cell-of-origin for CRC is the colonic stem cell20, a long-lived progenitor residing in the crypt base responsible for regenerating the colonic epithelium every few days21,22.
Stem cells retain memory of colitis
We proposed that exposure history of the intestine would be encoded in this cell population, representing a clear target for the study of how epigenetic memory may influence tissue health. To test this, we used a mouse model of chronic colitis in which the colon is repeatedly injured through low-dose dextran sodium sulfate (DSS) administration23 (Methods) and defined three states of disease progression: acute injury (one cycle of DSS), chronic injury (three cycles of DSS) and recovery (Fig. 1a,b). We found that within 21 days of DSS cessation, most animals recovered or exceeded their starting body weight, epithelial crypt structures reformed and immune infiltration subsided (Extended Data Fig. 1). Altogether, this indicated histological and morphological recovery at the cellular and organismal levels.
a, Model for studying colitis memory. b, Immunofluorescence for EPCAM at each stage. c, Uniform manifold approximation and projection (UMAP) embedding of single-cell assay for transposase-accessible chromatin using sequencing (scATAC-seq) and scRNA-seq data coloured by cell type. d, Mean gene expression changes through disease progression relative to control. n = 9 control, 4 acute, 5 chronic and 5 recovered mice. e, Top, number of genes significantly upregulated relative to controls at each stage. Bottom, fold change relative to controls in which each point represents 1 of the 246 differential genes identified. f, Top, relative expression of stem markers Lgr5 and Mki67. Bottom, colitis stage enrichment in stem cells, with colour representing the enrichment of each cell’s k-NN network for a given stage. g, Mean motif accessibility changes through disease progression relative to control. n = 9 control, 4 acute, 5 chronic and 5 recovered mice. FDR from two-sided t-test. h, AP-1 motif accessibility in stem cells. n = 500 cells across all animals at each stage. P value from Wilcoxon rank-sum test. i, Quantification of stem cells with high motif accessibility (score greater than 1.5). n = 9 control and 5 recovered mice. j, Immunofluorescence for FOS protein. Basal (B) and luminal (L) sides of crypts are marked. Arrows indicate crypt basal cells with high FOS levels. k, Quantification of CD44 and FOS colocalization. n = 3 mice per stage. l, Chromatin accessibility and FOS protein levels following extended recovery. For chromatin accessibility, control n = 14, acute n = 4, chronic n = 5, early recovery n = 5, 50-day recovery n = 2, 79-day recovery n = 2, 102-day recovery n = 4 mice. For FOS protein, control n = 3, acute n = 2, chronic n = 2, early recovery n = 3, 50-day recovery n = 3, 79-day recovery n = 2 and 102-day recovery n = 4 mice. The left y axis represents mean change in AP-1 motif accessibility in stem cells over control and the right y axis represents mean FOS+ percentage of CD44+ cells. All error bars are s.e.m. AE, absorptive enterocyte; EEC, enteroendocrine cell. Panels created in BioRender: a, Nagaraja, S. https://biorender.com/jqhj3wt (2026); a,l, Nagaraja, S. https://biorender.com/865k9jb (2026). Scale bars, 100 μm (b); 50 μm (j).
We jointly profiled chromatin accessibility and gene expression (SHARE-seq24) to identify memory signatures of chronic inflammation (Extended Data Fig. 2a–d). Measuring 52,540 single cells across 23 animals (n = 9 controls, 4 acute injury, 5 chronic injury, 5 recovered), we identified known populations of the colonic epithelium, including Lgr5+ intestinal stem and progenitor cells (Fig. 1c and Extended Data Fig. 2e–h). Consistent with the recovery of the tissue, we observed no significant difference in the proportions of stem cells or cells in the absorptive lineage between recovered and control animals (Extended Data Fig. 2i).
During acute injury, stem cells upregulated genes related to interferon signalling and immunomodulation, consistent with previous studies of epithelial inflammation25,26, and chronic injury further activated genes related to wound healing, cell junction reformation and extracellular matrix reconstruction (Fig. 1d and Extended Data Fig. 3a). These changes, in addition to more than 97% of the 246 genes transcriptionally activated in stem cells during either stage of disease, returned to baseline following recovery (Fig. 1e and Supplementary Table 1).
Given the transcriptome demonstrated minimal changes following recovery and DSS is not known to be mutagenic27, we proposed that molecular memory would be more apparent within the epigenomes of stem cells. To quantify whether cells were distinct in their overall epigenomic states, we constructed a low-dimensional embedding, found the k-nearest neighbours (k-NN) (k = 100) of each stem cell and, for each neighbourhood, quantified the proportion belonging to each stage of colitis (Extended Data Fig. 3b). Cells from recovered tissue were epigenomically distinct from cells derived from control tissues (Fig. 1f and Extended Data Fig. 3c). By contrast, analogous analysis with gene expression did not reveal discrete transcriptomic states in stem cells recovered from colitis (Extended Data Fig. 3d). Further analysis of heterogeneity within stem and progenitor cells revealed that cells acquire an intermediate epigenomic state between that of activation and quiescence (Supplementary Note 1 and Supplementary Fig. 1).
To more precisely define this epigenomic memory, we grouped TFs by binding motif sequence similarity (n = 299 families, Supplementary Table 2) and quantified changes in accessibility associated with these motifs28, revealing persistent alterations following recovery (Fig. 1g, Extended Data Fig. 3e and Supplementary Table 3). The most prominent of these was a cumulative gain in accessibility at activator protein 1 (AP-1) motif sites (false discovery rate (FDR) of 1.27 × 10−3), as well as further increases in motif accessibility in the ETS family (ETS, SPIB and SPI1), consistent with their known roles in oncogenesis and ulcerative colitis29,30,31,32. By contrast, CTCF sites showed a significant loss in accessibility during chronic colitis and recovery (FDR = 8.79 × 10−3). Examining differentiated cells, we find intermediate enterocytes also show memory of AP-1 motif accessibility (Extended Data Fig. 3f), although to a lesser extent, and no evidence of memory in fully differentiated enterocytes, altogether suggesting that stem cells lose their memory on differentiation.
Whereas AP-1 factors, such as FOS and JUN, are known to be activated by a diverse set of stimuli, including damage, growth and stress33,34, these TFs have been shown to be mediators of epigenetic memory13,14,15. However, these characterizations have largely been described across bulk populations of cells14,16. We quantified motif accessibility in single stem cells and identified substantial heterogeneity in TF memory, including a subpopulation with exceptionally high AP-1 motif accessibility following recovery from colitis (9.2% versus 1.6%, P = 1.44 × 10−15, Fig. 1h,i and Extended Data Fig. 3g). This subpopulation also demonstrated minor increases in Fos and Jun transcripts (Extended Data Fig. 3h–k). Therefore, we quantified protein levels and found FOS to be preferentially elevated in epithelial cells during chronic injury (Supplementary Fig. 2). Examining this longitudinally, we found a subset of stem and progenitor cells at the crypt base (CD44+) with elevated FOS protein in animals recovered from colitis but not controls (16.4% versus 0%, P = 0.011, Fig. 1j,k). These findings highlight that although the average stem cell shows moderate epigenomic memory of colitis, roughly 10% of cells carry a prominent AP-1 memory of injury following colitis recovery.
To assess the durability of this epigenetic memory, we extended recovery periods following colitis to more than 100 days (Fig. 1l and Extended Data Fig. 3l,m). We find that FOS protein levels peak during chronic injury and then sharply decline within 21 days of DSS withdrawal. By contrast, chromatin accessibility of AP-1 motif sites lag, attaining its highest value during early recovery and showing a slow attrition, with evidence of chromatin memory still found after 102 days, representing dozens of generations of colonic epithelial turnover21. Altogether, this suggests that cells cumulatively restructure their epigenomes in response to injury and subsequently maintain these changes following morphological and functional recovery, independent of TF protein levels.
Epigenetic states are clonally inherited
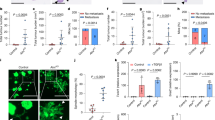
Immune and other non-epithelial cells are known to mediate the progression and resolution of colitis through local and systemic production of signalling factors35,36. Given this, we tested whether stem cell memory of colitis was cell-intrinsic by deriving organoids from colitis tissue during chronic injury. We found that, despite initially appearing similar to healthy controls, colitis organoids progressively obtained a regenerative and hyperplastic morphology37,38,39 over 34 days of culture (Fig. 2a). Consistently, colitis-derived organoids were more proliferative (Fig. 2b), representing a potential adaptation to repeated cycles of regenerating wounded tissue and demonstrating that cellular memory of colitis was maintained within stem cells following removal from the tissue microenvironment.
a, Organoid morphology of colitis and control organoids. Representative of n = 6 mice per group. b, Proliferation in colitis-derived and control organoids at 9 days of culture. n = 3 organoid lines per group. c, Schematic for lineage tracing. d, UMAP embedding of scATAC-seq data showing expression of enterocyte differentiation marker Car1 (left) and cell cluster (right). e, Examples of select clones. f, Model for distinguishing features that show clonal memory. µ, mean; σ, standard deviation. g, Left, AP-1 and CTCF motif accessibility among the 50 largest clones. Each violin plot represents an individual clone and points represent median values per clone. Right, AP-1 motif accessibility for top and bottom clones in which each point is a cell. h, Clonal memory of motif accessibility. The x axis represents the difference between observed clonal variance and random variance following clonal label permutation, and the y axis represents significance of this difference. Further details in Extended Data Fig. 6d. i, Median AP-1 motif accessibility across 80 control and 52 colitis clones. Crossbars indicate median by group across clones. j, Spearman correlation between gene program score and mean motif accessibility across clones. k, Program 20-related gene ontology. l, Proliferation following 24 h of AP-1 inhibition (T-5224, 10 μM) or matched vehicle control. n = 5 control vehicle, 5 control T-5224, 4 colitis vehicle, 3 colitis T-5224 organoid lines. NS, not significant. m, Organoid morphology following chronic AP-1 inhibition (T-5224, 10 μM) and washout. n, Quantification of organoid size following 20 days of washout. n = 86 control vehicle, 87 control T-5224, 67 colitis vehicle and 71 colitis T-5224 individual organoids. Representative of eight to nine wells per group. P values from two-sided Wilcoxon rank-sum test. o, Comparison of accessibility and methylation change per region between colitis and control organoids. FC, fold change. p, Genome tracks at P20 gene Thbs. Top, normalized ATAC-seq insertions in control and colitis organoids. Bottom, methylation in the selected region. n = 3 organoids lines per condition. All error bars are s.e.m. Exp., expression; Norm., normalized. For box and whisker plots, the centre line represents median, the box upper and lower quartiles and the whiskers 1.5× interquartile range (IQR). Panels created in BioRender: c, Nagaraja, S. https://biorender.com/f22bz54 (2026); m, Nagaraja, S. https://biorender.com/865k9jb (2026). Scale bars, 150 μm (a); 300 μm (m).
We next sought to determine whether cellular memory is clonally heritable and mediated by the epigenome. Recent advances in single-cell lineage tracing provide new opportunities to measure fate transitions through introduction of barcodes that are delivered to individual cells, propagated through cell division and transcriptionally expressed for detection through single-cell RNA sequencing (scRNA-seq)40,41. Inspired by this, we created SHARE-TRACE (SHARE-seq with clonal tracing) to simultaneously measure a cell’s clonal lineage history, gene expression, differentiation state and chromatin accessibility. We modified previous cell barcoding technology40 to improve nuclear retention and its compatibility with SHARE-seq (Extended Data Fig. 4a,b), thereby enabling the profiling of 52,564 cells across 6 organoid lines (n = 3 control, 3 colitis) and mapping of transcriptomic and epigenomic states in 172 clones (Fig. 2c–e).
The identification of hundreds of clones provided the opportunity to more sensitively quantify heritability of epigenomic states. We reasoned that for any clonally remembered state, cells within a clone would more closely resemble one another than a random selection of cells. We therefore developed a permutation-based statistical approach to characterize general principles of clonal heritability for each TF motif family, independent of past exposure to colitis (Fig. 2f,g, Extended Data Fig. 4c,d, Supplementary Table 4 and Supplementary Data 2). Whereas ETS and CTCF motifs demonstrated memory in vivo, we did not see clonal heritability of these motifs, suggesting that memory of these factors may be propagated by cell extrinsic mechanisms (Fig. 2h). However, we did find clonal memory of AP-1 with this approach, further highlighting the central role of this TF complex in regulating memory of colitis. Our analysis also uncovered clonal memory of several other TF families that do not demonstrate memory in vivo following colitis, potentially related to the position within the colon from which the organoid originated and the inherent differentiation capacity of a given clone42 (Extended Data Fig. 4e–g and Supplementary Note 2). However, AP-1 was the only TF family found to hold memory both in vivo following colitis and ex vivo through clonal lineages. Of note, we find our detection power to be reliant on the total number of clones (Extended Data Fig. 4h), suggesting that larger studies may identify further TF families with weaker clonal memory.
Consistent with in vivo stem cell memory, colitis-derived clones demonstrated an increase in mean AP-1 motif accessibility (P = 0.038) and a subpopulation of clones with exceptionally high accessibility (12.2% versus 2.7%; Fig. 2i). The resemblance of this distribution to that seen in vivo (Fig. 1h) suggests that heterogeneity in memory is maintained through clonal lineages and the range of these clonal states within the same microenvironment indicates that these states are maintained intrinsically rather than by paracrine signalling. Expanding on this, we assessed how previous exposure to colitis may influence clonal heritability of TF motif accessibility by using a mixed linear model, revealing that clonal identity to explain substantially more of the observed variance in most cases (Extended Data Fig. 4i–m and Supplementary Note 3). Together, these findings demonstrate that variability between individual stem cells could be propagated through clonal lineages for select motif families and this epigenetic memory can be further modulated by environmental stimuli.
We next wanted to understand how the memory of AP-1 relates to biological functions. To this end, we derived transcriptional programs composed of coregulated genes and, measuring clonality analogously to TFs, we observed that a subset of gene expression programs shows heritability across clones (Extended Data Fig. 5a,b). To identify the target processes of AP-1, we correlated gene programs with the motif accessibility of AP-1 and another strongly clonal TF, HNF4/PPAR, across clones. We found program 20 (P20) was strongly correlated with AP-1 (ρ = 0.52; Fig. 2j and Extended Data Fig. 5c) and consisted of genes related to wound healing43 (Clu), cytoskeletal remodelling (Flnb and Macf1), cell junction reformation (Cdh1 and Itgb6) and stem cell proliferation (Cd44 and Sox4; Fig. 2k, Extended Data Fig. 5d,e and Supplementary Table 5). By contrast, HNF4/PPAR-related programs were negatively correlated with AP-1 (P9 ρ = −0.36, P30 ρ = −0.14) and associated with colonic enterocyte maturation42 (Extended Data Fig. 5e–g and Supplementary Note 4). Examining the P20 gene program in tissues, we found that AP-1-related gene expression was reduced below baseline by 50 days and remained suppressed for 102 days, with only 1 statistically significant gene remembered after 50 days (Extended Data Fig. 5h–l). Overall, this demonstrated memory of AP-1 to be held specifically in the epigenome.
Given the P20 gene program peaked in expression during chronic injury and was enriched for repair-associated genes, we proposed that AP-1 was facilitating a proliferative and regenerative phenotype. To test this, we treated organoids with T-5224 (ref. 44), a chemical inhibitor that interferes with binding of AP-1 to DNA, and found that acute inhibition of AP-1 complex activity preferentially blocked proliferation in colitis-derived organoids (Fig. 2l). We then tested whether blockade of AP-1 activity was sufficient to permanently erase the colitis memory state. We found that following prolonged inhibition of AP-1 activity, colitis-derived organoids restored their hyperplastic morphology37,38,39 and demonstrated no difference in size compared with uninhibited colitis organoids (Fig. 2m,n).
We next proposed that memory of colitis was durably encoded through DNA methylation6. We thus performed whole-genome DNA methylation sequencing and found that colitis-induced changes in chromatin accessibility were highly negatively correlated with changes in DNA methylation (ρ = −0.51; Fig. 2o,p and Extended Data Fig. 6a,b). This included 4,397 regions that showed concordant changes in accessibility and methylation, exemplified by the P20 genes Thsb1 and Mecom. Extending this finding in vivo, we find AP-1 sites to preferentially demonstrate loss of DNA methylation following colitis and AP-1 inhibition insufficient to restore these marks (Extended Data Fig. 6c–e), demonstrating a role for other factors in maintenance of memory.
We demonstrate that cellular memory can be maintained within clonal lineages and result in cell populations with exceptionally altered epigenetic states. This molecular memory is facilitated by durable changes to chromatin accessibility and DNA methylation. The retrieval of these memories coincides with altered morphology and function and are reliant on the binding of AP-1 factors. Considering our finding that colitis memory promotes proliferation, the heritability of these states raises the intriguing prospect that clonal memory of chronic inflammation may give individual stem cells and their progeny a fitness advantage. Such a process may create fields within tissues with altered epigenetic states that affect future responses to stimuli and the development of disease.
FOX factors stabilize AP-1 binding
Given the broad expression of AP-1 across tissues, we sought to understand how AP-1 may be directed to have stem cell specific functions. Previous studies have reported that AP-1 interacts with tissue-specific factors to establish memory at specific genomic regions13,14,15,45. To uncover binding partners of AP-1 in colonic stem cells in vivo, we used seq2PRINT46, an approach that combines TF footprinting with deep learning to de novo discover DNA sequence motifs and localize binding events of TFs to regulatory elements (Fig. 3a). Training seq2PRINT on chromatin accessibility footprints of in vivo colitis progression yielded a total of 1,838 motifs, representing 890 known motifs and 948 unknown motifs (Supplementary Data 1) and uncovering DNA sequences related to dimerization, cobinding and stability not annotated in existing motif databases (Extended Data Fig. 7a).
a, Schematic for derivation of de novo motifs from footprinting data. b, Left, mean accessibility change over control for all de novo and known motifs. Right, sequence information content of de novo derived AP-1 composite motifs. c, Top, FOX/AP-1 composite motif accessibility change over control through colitis progression. Mean across n = 9 control, 4 acute, 5 chronic and 5 recovered mice. Bottom, predicted effect of motif presence on footprint scores. The x axis represents distance from motif, the y axis represents size of footprint and colour indicates predicted change of footprint score when motif is added. d, Disruption of cobinding with AP-1 for select TF families following 24 h of AP-1 inhibition. e, Cobinding of select TF families at IBD-specific AP-1 footprints in human organoids. f, Comparison of cobinding scores between mouse and human organoids. g, Schematic for quantification of in vitro binding capability. h, Example locus assayed by the in vitro binding assay. Top, seq2PRINT footprint scores from control and colitis-recovered mice. Bottom, in vitro TF binding score at the same sequence. i, In vitro binding scores for AP-1 and FOX TFs alone and in combination. The y axis represents the binding score at the AP-1 motif normalized to the FOS–JUN heterodimer alone. Error bars represent s.e.m. across all tested loci (n = 29 sequences for AP-1 alone, n = 34 sequences for FOX/AP-1). j, AlphaFold3 predicted structure for FOXP1, composite motif DNA and FOS–JUN dimer (left) or JUN alone (right). All error bars represent s.e.m. Significance values are from two-sided t-tests unless otherwise indicated. For box and whisker plots, the centre line represents median, the box upper and lower quartiles and the whiskers 1.5× IQR. Panels d and e created in BioRender; Nagaraja, S. https://biorender.com/or0ceke (2026).
Although this unsupervised approach once again uncovered AP-1 as the most prominent regulator of memory in vivo, it also identified composite motifs of AP-1 with other TF families (Fig. 3b and Extended Data Fig. 7b,c). Most notable among these was an AP-1/forkhead box (FOX) composite motif that increased in accessibility with memory of colitis and was predicted to increase accessibility by displacing nucleosomes (Fig. 3c). In addition, inhibition of AP-1 led to loss of both AP-1 and FOX footprints preferentially in colitis organoids over controls (Extended Data Fig. 7d), suggesting a disease-specific interaction between these TFs.
To determine whether disruption in FOX binding following AP-1 inhibition was through primary interactions or a consequence of secondary effects, we developed a computational approach to predict direct or indirect binding (Extended Data Fig. 7e–i and Supplementary Note 5). Applying this to footprint changes following AP-1 inhibition, we found that loss in FOX footprints strongly correlated with AP-1 loss in the same regulatory element, further indicating that FOX binding at these sites depended on AP-1 (Fig. 3d). Additionally, AP-1 footprints gained from colitis memory in vivo were associated most strongly with a gain in proximal FOX binding (Extended Data Fig. 7j–l). To determine the relevance of this association to human disease, we performed SHARE-seq in organoids derived from patients with inflammatory bowel disease (IBD) or healthy control participants (Supplementary Fig. 3). Our analysis predicted that novel AP-1 binding sites formed in IBD were strongly associated with proximal FOX binding as well, revealing a unique role for FOX TFs in memory of intestinal inflammation across many model systems and species (Fig. 3e,f).
To biochemically validate the association between FOX and AP-1, we used an in vitro binding assay that uses footprinting to quantify cobinding46 (Methods). By incubating DNA from genomic regions predicted to gain AP-1 binding following colitis (n = 63 regions) with purified AP-1 proteins in combination with FOXA1 and FOXP1, both of which were found to be highly expressed in colonic stem cells (Extended Data Fig. 8a), we were able to directly measure TF cobinding relationships (Fig. 3g,h, Extended Data Fig. 8b,c and Supplementary Table 6). Supporting our computational predictions, FOXA1 enhanced the binding of AP-1 TFs to DNA containing the composite AP-1/FOX motif (mean fold change 3.2, P = 7.14 × 10−11; Fig. 3i, Extended Data Fig. 8d,e and Supplementary Table 6). We found FOXP1 notably increased AP-1 binding both in the presence of a composite motif (mean fold change 8.3, P = 6.60 × 10−11) as well as the AP-1 motif alone (mean fold change 9.0, P = 3.11 × 10−6). This suggested FOXP1 and AP-1 cooperativity is facilitated by protein interactions independent of DNA sequence, a finding further supported by protein structure predictions (Fig. 3j and Extended Data Fig. 8f,g).
Finally, the cobinding assay also revealed that FOXP1 substantially increases the binding of JUN to DNA without FOS (AP-1 only mean fold change 5.2, P = 2.10 × 10−4; AP-1:FOX mean fold change 4.8, P = 4.30 × 10−7), further supported by several predicted interaction sites (Extended Data Fig. 8h). The cobinding of FOXP1 and JUN, without FOS, is intriguing and provides a biochemical mechanism supporting previous work describing epigenetic memory in the skin through the cobinding of JUN to tissue-specific TFs without FOS15. With FOX TFs having described roles in intestinal wound healing, CRC oncogenesis and cooperative AP-1 binding47,48,49, we build on previous studies to show that FOX stabilizes AP-1 binding at regions implicated in colitis memory.
Memory of colitis promotes tumour growth
We next proposed that memory of chronic colitis may prime the colonic epithelium for malignant transformation. To test this hypothesis, we induced adenomas through APC loss (Cdx2:CreERT2;APCfl/fl)50 in mice recovered from colitis and naive controls (Fig. 4a). We found that colitis-associated adenomas were grossly larger in size compared with controls, with a greater fraction of tumours larger than 1 mm in diameter (P = 0.042, Fig. 4b,c and Extended Data Fig. 9a).
a, Model for studying effects of colitis memory on tumour formation. i.p., intraperitoneal. b, Gross images of adenomas in recovered and control animals. Arrowheads indicate adenomas greater than 1 mm in diameter. c, Quantification of large adenomas (more than 1 mm diameter). n = 4 control and 5 colitis-recovered mice. d, Model for studying early tumour growth. e, Quantification of microscopic tumour size following low dose initiation. n = 88 tumours across 7 control mice and 256 tumours across 10 colitis-recovered mice. f, Schematic for performing SHARE-seq. g, Mean motif accessibility in adenoma cells. n = 3 control and 2 colitis-recovered mice. h, Gene expression changes in adenoma cells relative to non-neoplastic stem cells. i, Left, spatial Axin2 expression. Right, microscopic tumour identification with each tumour coloured by a distinct colour. j, Expression scores of AP-1/P20 across n = 117 control and n = 137 colitis adenomas. k, Spatial AP-1/P20 expression score in tumour cells. l, Left, schematic for in vivo AP-1 inhibition and tumour formation. Right, quantification of microscopic tumour size. n = 48 tumours across 6 control vehicle-treated mice, n = 43 tumours across 5 control T-5224-treated mice, n = 95 tumours across 7 vehicle-treated colitis-recovered mice and n = 163 tumours across 7 T-5224 treated colitis-recovered mice. p.o., per os (oral administration). m, Model of colitis memory priming the epithelium for increased tumour growth. Error bars are s.e.m. ECM, extracellular matrix. Panels a, d and l created in BioRender; Nagaraja, S. https://biorender.com/tra3y6g (2026). Panel m created in BioRender; Nagaraja, S. https://BioRender.com/tefh2xg (2026). Scale bars, 1 cm (b); 400 μm (i,k).
Colitis-recovered animals did not carry higher numbers of macroscopic tumours and adenomas did not demonstrate higher proliferation or clone-forming potential (Extended Data Fig. 9b–d). Thus, we proposed that epigenetic memory following colitis promoted initial tumour outgrowth and larger tumours occurred due to the close proximity and clustering of many microscopic tumours. To test this, we sparsely induced tumour formation and harvested tissue before gross adenoma formation, allowing the assessment of early microscopic growth51 (Fig. 4d and Extended Data Fig. 9e,f). Consistently, we found that these individual microscopic tumours were larger in mice that had recovered from colitis (P = 1.79 × 10−5, Fig. 4e and Extended Data Fig. 9g,h). Similar to the heterogeneity observed in clonal memory of AP-1 accessibility (Fig. 2i), we also found a greater proportion of these microscopic lesions were excessively large in colitis-recovered tissue (8.7% versus 2.5%), raising the intriguing possibility that these represent clonal fields expanded from stem cells with strong memory of inflammation. Altogether, these findings indicate that the protumorigenic effect of chronic colitis was due to a proliferative advantage at early stages of tumorigenesis, potentially reflecting the acquisition of more dominant oncogenic drivers through tumour progression. Whereas previous studies have found that colitis injury after inducing oncogenic mutations increases tumour formation52,53, we show here that the pro-oncogenic effect of colitis is maintained even after recovery and resolution.
To characterize the molecular differences associated with this phenomenon, we performed SHARE-seq on control and colitis-associated tumours, identifying both adenoma cells and adjacent non-neoplastic epithelium (n = 38,382 cells; Fig. 4f and Extended Data Fig. 9i–k). We found colitis-associated adenomas to demonstrate similar patterns of motif accessibility and TF binding cooperativity as non-neoplastic epithelium with memory of colitis (Fig. 4g and Supplementary Fig. 4). Notably, we found many P20 genes to be upregulated in adenomas relative to normal stem cells (Fig. 4h), suggesting an oncogenic role. To relate these molecular differences to individual tumours, we performed spatial transcriptomics (VisiumHD, Methods) following adenoma formation and identified 254 distinct tumours based on Axin2 expression and image segmentation (Fig. 4i and Extended Data Fig. 10a). An unbiased analysis across 117 control and 137 colitis-recovered tumours found acquisition of distinct transcriptional states, with colitis tumours showing higher expression of genes related to cell adhesion and epithelial repair (Extended Data Fig. 10b). Consistent with this, we found that colitis-associated adenomas upregulated the P20 program (P = 1.01 × 10−9, Fig. 4j,k), driven by the tumour-specific activation of specific subsets of genes (Extended Data Fig. 10c,d). In line with the heterogeneity of memory observed in stem cells and organoid clones, we found that a subset of colitis-associated tumours showed particularly strong expression of AP-1-associated genes (8.8% versus 1.7%), as well as a fraction resembling control tumours that probably represents colitis memory-independent tumour development. The tumours with exceptional P20 gene expression also upregulated programs related to platelet-derived growth factor signalling, proliferation and vasculogenesis (Extended Data Fig. 10e,f). Together, these findings demonstrate that recovery from colitis primes tumours for increased AP-1-associated gene expression and an exceptionally high AP-1 subpopulation robustly activates extra pro-oncogenic programs.
Finally, we sought to determine whether AP-1 activity drives the memory-associated increase in tumour growth by selectively inhibiting AP-1 during tumour initiation. We found AP-1 blockade during APC loss reduced median tumour size in the colitis-recovered tumours by nearly 40% (relative size 95% CI: 0.11–0.64; Fig. 4l and Extended Data Fig. 9l) and this growth restriction was not present in control mice. This reinforced that activity of the AP-1 complex was required to enact the maladaptive phenotype associated with colitis memory. As we observed that AP-1 inhibition is insufficient to erase molecular memory of colitis (Extended Data Fig. 6d), these findings together suggest that memory of colitis promotes carcinogenesis during the initial tumour growth following oncogene mutation.
Overall, these findings suggest a model whereby chronic inflammation encodes an epigenetic memory of repair in colonic stem cells that promotes tumour growth through progressive gain of AP-1 and tissue-specific TF accessibility at proproliferative genes (Fig. 4m). This memory promotes tumorigenesis by increasing malignant outgrowth once a stem cell acquires an oncogenic mutation, thereby contributing to the raised incidence of cancer associated with chronic inflammation.
Discussion
Overall, we reveal that chronic inflammatory disease creates cellular memory through the accumulation of epigenetic changes. Following repeated cycles of inflammation, damage and healing, colonic stem cells heterogeneously encode a memory of regeneration in their epigenome that persists for more than 100 days and lowers the threshold for tumour formation. We find that this high AP-1 state is propagated clonally and maintained following tumour initiation, promoting the expression of regenerative programs and enhancing proliferation.
The central role of AP-1 in memory of intestinal damage builds on growing evidence that this TF complex may serve as a shared regulator of cellular memory across a variety of tissues, including the skin, pancreas and immune system, as well as diverse stimuli, such as inflammation, repair and therapeutic resistance10,13,14,15,16,31,54. By combining single-cell epigenomics with lineage tracing, we further provide a method and framework to enable quantitative assessment of how such cellular states propagate through stem cell lineages. This approach reveals clonal heritability of TFs related to a variety of biological programs besides exposure to inflammation, including positional memory and differentiation capacity. Altogether, deciphering the environment–TF–gene encoding of cell type specific memories represents an exciting opportunity to understand the long-term impact of diverse environmental exposures and experiences.
These findings provide an epigenetic mechanism that connects inflammatory diseases with malignancy. Our observation of macroscopically larger tumours suggests that clonal fields of epigenetically primed cells emerge in close proximity to one another, consistent with a model of ‘field cancerization’55. Whereas this framework has typically been used to describe fields of somatic mutations, this work suggests a similar model in which chronic colitis creates clonal fields of cells carrying fitness-conferring ‘epi-mutations’. In line with this, recent work demonstrated that colons of patients with ulcerative colitis consist of a ‘patchwork’ of millimetre-sized clonal fields56, representing massive expansions of single stem cells when compared with healthy colonic epithelium. However, these fields did not show positive selection for typical CRC driver mutations or mutations enriched in colitis-associated tumours. Our findings indicate this may instead be explained by heritable epigenetic alterations, largely maintained by subpopulations of stem clones bearing increased AP-1 accessibility following chronic inflammation and the proproliferative effect of this memory.
With the incidence of early-onset colorectal cancer rising globally57, our findings carry both diagnostic and therapeutic implications for patients. Memory-related epigenetic signatures linked to cancer could allow for tracking of oncogenic risk in patients before the formation of visible neoplastic lesions. Similarly, therapeutic strategies for erasing pathologic cellular memory and mitigating its maladaptive consequences could offer a promising avenue for disease prevention in patients with chronic diseases.
Methods
Animals and cell lines
Animal work
Mouse (Mus musculus) strains C57BL/6J (strain no. 000664) and Cdx2:CreERT2;APCfl/fl;KrasWT (strain no. 035169)50 were obtained from The Jackson Laboratory. Mice were housed at room temperature and ambient humidity in individually ventilated cages at a maximum density of five mice per cage with ad libitum access to food and water in a specific-pathogen-free facility accredited by the Association and Accreditation of Laboratory Animal Committee. Cages contained Anderson’s Bed-o’Cob bedding (The Anderson Inc.), one nestlet (Ancare, 2 × 2-inch2 compressed cotton squares) and a red mouse hut (Bioserv). The colony room was kept on a 12 h–12 h light–dark cycle. All animal handling and experiments were conducted in accordance with procedures approved by the Institutional Animal Care and Use Committee at Harvard University (protocol no. 19-10-362). For tumour formation experiments, euthanasia criteria were weight loss of more than 20%, persistent grossly bloody stool for greater than or equal to 3 days and/or excessively lethargic or moribund state, as determined by veterinary care. These criteria were not exceeded in any experiments.
Cell culture
Human embryonic kidney 293T (HEK293T) cells (American Type Culture Collection (ATCC), CRL-3216; authenticated by short tandem repeat profiling and tested for mycoplasma by ATCC) were grown in DMEM (Thermo, 11965-092) with 10% fetal bovine serum (FBS) and 1% penicillin–streptomycin. Cells were incubated at 37 °C in 5% CO2 and maintained in exponential phase.
Mouse organoid derivation and culture
Colitis organoids were derived from whole colonic tissue 11 days following cessation of the third cycle of DSS. Animals were anaesthetized with 2,2,2-tribromoethanol (Sigma, T48402-25G) and cardiac perfusion was performed with PBS to remove peripheral immune cells. Epithelium was removed by incubating colonic tissue in EDTA solution (section ‘Colon tissue processing and cell sorting’ below) supplemented with 100 μg ml−1 primocin (Invitrogen, ant-pm-05) for 20–30 min and scraping the luminal surface with a glass slide. Epithelial fragments were washed once with Advanced DMEM/F12 (ADMEM) and resuspended in Crypt Basal (ADMEM, 10 mM HEPES, 1× GlutaMax (Thermo, 35050061), 1× Pen-Strep (Thermo, 15140122), 1× N2 Supplement (Thermo, 17502048), 1× B27 Supplement (Thermo, 17504044), 1 mM N-acetylcysteine (Sigma, A9165-5G)) before mixing with an equal volume of Matrigel (Corning, 47743-722). Crypts were plated as roughly 30-l domes in a six-well plate and allowed 10–15 min to polymerize. Colon organoids were grown in WENR media: 50% ENR (Crypt Basal with 50 ng ml−1 epidermal growth factor (EGF) (Thermo, PMG8041), 100 ng ml−1 Noggin (Peprotech, 250-38), 1:100 Rspondin conditioned media) and 50% Wnt conditioned media. Conditioned media was generated in-house from L WNT3A cells (ATCC, CRL 2647) or HA-R-Spondin1-Fc 293T Cells (R&D, 3710-001-01). Organoids were passaged every 7–10 days by mild dissociation in TrypLE for 8–10 min, triturating every 4–5 min and quenching with 10% FBS in ADMEM. When collected for SHARE-seq, organoids were dissociated to near single cell for 15 min in TrypLE and quenched before treating with 1:100 recombinant DNase (Roche, 04716728001) in ADMEM at room temperature for 5 min to reduce dead cell DNA contamination. Cells were then washed and frozen in CryoStor at −80 °C before SHARE-seq.
Human organoid derivation and culture
Human organoid lines were derived from de-identified biopsies from grossly unaffected tissue in patients undergoing endoscopy at Boston Children’s Hospital. Informed consent and developmentally appropriate assent were obtained at Boston Children’s Hospital from the donors’ guardian and the donor, respectively. All methods were approved and carried out in accordance with the Institutional Review Board of Boston Children’s Hospital (Protocol number IRB-P00000529).
Organoids were derived from biopsies as previously described in ref. 58. Briefly, intestinal crypts were isolated from frozen tissue and then resuspended and plated in 40-μl Matrigel domes. Once established, human rectal organoids were sustained in specialized growth media that has been previously described58. Media changes occurred every 2 days during expansion, with organoids being passaged once every 6–8 days as necessary. To induce differentiation, organoids were grown in growth media for 2 days postpassage to allow for stem cell expansion; after which, the organoids were transitioned to differentiation media. Media was changed every 2 days for the length of the experiment, with organoids being collected for analysis after a total of 10 days.
Experimental procedures
Colitis induction
Male mice aged between 8 weeks and 15 weeks were administered dextran sulfate sodium (VWR, IC16011080) in drinking water at 1–1.5% final concentration to induce chronic colitis. Animals were weighed every day during DSS administration and every 2–3 days during rest periods. On the fourth day of each DSS administration, stool was tested for occult blood (VWR, 10012-002) to ensure successful induction of colitis. DSS concentrations were reduced if excess disease severity was observed through any of the following metrics: frank blood in the stool at any point, weight of loss of more than 10% before the ninth day of any cycle, failure to recover back to 90% of starting weight before the next cycle or poor body condition.
Acute injury timepoints were collected 3 days after the end of the first DSS cycle (day 11), chronic injury 9–11 days after the third cycle (days 51–53) and recovery 21–22 days after the third cycle (days 63–64).
Colon tissue processing and cell sorting
For in vivo colitis memory SHARE-seq experiments, animals were anaesthetized and perfused as previously described. Entire colons were dissected, lumens were exposed and tissue was transferred to EDTA Dissociation Solution (10% FBS, 4 mM EDTA, 10 mM HEPES in PBS). Following rotation for 20–30 min at room temperature, epithelium was coarsely removed by scraping the luminal surface with a glass slide and remaining muscle and submucosal were crudely chopped with scissors. Both epithelial and tissue fragments were then dissociated to single cells in ADMEM (Fisher, 12-634-028) with 10 mM HEPES, 0.4 mg ml−1 collagenase (Millipore, C9263-25MG), 1.25 U ml−1 dispase (Millipore, D4693-1G), 1 U ml−1 DNase (Worthington Biochemical, LS002004) and 5 μM Y-27632 (R&D, 1254). Cells were washed with 0.1% BSA/PBS, stained with Calcein Red-AM (BioLegend, 425205) then with antibodies for EPCAM 1:100 (Fisher, 501129753), CD45 1:100 (BioLegend, 103116), Ly6g 1:100 (BioLegend, 127605) and SiglecF 1:100 (BioLegend, 155503). 4,6-diamidino-2-phenylindole (DAPI) (Fisher, 62248) dead cell staining was performed before sorting.
Stained cells were sorted on a BD FACSAria for epithelial (EPCAM+CD45−) and non-granulocyte (EPCAM−CD45+LY6G−SiglecF−) populations into ADMEM with 0.2% BSA, 0.1 U μl−1 Enzymatics RNase inhibitor (Qiagen, Y9240L) and 15 μM Y-27632. For extended recovery timepoints (50 days and longer), only EPCAM+ cells were sorted. Cells were pelleted, resuspended in CryoStor CS10 (StemCell Technologies, 07959) and stored at −80 °C.
Histology and colitis scoring
Animals were anaesthetized and perfused as described above. After colonic tissue was dissected and the luminal surface was exposed, Swiss rolls or tissue fragments were fixed in 4% PFA and PBS overnight at 4 °C and then placed in 70% ethanol previous dehydration and paraffin embedding. Haematoxylin–eosin (H&E) staining was performed for general histology evaluation.
Colitis scoring was performed as described in ref. 59 with researchers blinded to sample identity. Immune infiltration was scored as follows: mucosa, 0, normal; 1, mildly increased immune infiltrate; 2, modest infiltration; and 3, severe infiltration; submucosa, 0, normal; 1, mild to modest immune infiltration; and 2, severe infiltration; and muscularis, 0 normal and 1, modest to severe.
Immunohistochemistry was performed for CD45 (anti-CD45 1:500, Abcam, ab10558; anti-rabbit secondary 1:1,000, Vectastain Elite ABC, PK-6101) and the total CD45+ cells were counted in the mucosa and submucosa in all images before normalizing to total tissue area assessed. Researchers were blinded to conditions during imaging and quantification.
Immunofluorescence
Tissue was extracted, fixed with 4% PFA and PBS and subsequently cryogenically protected in 30% sucrose and PBS before optimal cutting temperature (OCT) compound embedding. Following sectioning, tissue was washed with PBS then permeabilized and blocked for 1 h in PBS with 3% normal donkey serum (Jackson Immuno, 017-000-121) and 0.5% Triton X-100. Sections were incubated overnight at 4 °C in Antibody Diluent (PBS with 1% NDS, 0.3% Triton X-100) with primary antibodies for EPCAM 1:500 (Abcam, ab213500), FOS 1:5,000 (Synaptic System, 226308), FOSB 1:800 (Fisher Scientific, PIMA515056), FOSL1 1:500 (Fisher Scientific, PIPA5115252), FOSL2 1:200 (Sigma, HPA004817) and/or CD44 1:100 (BioLegend, 103001). Excess antibody was removed with three PBS washes and secondary antibodies (Jackson Immuno, 712-605-15, 706-586-148, 711-545-152, 715-546-150) were added 1:500 in Antibody Diluent. Following 1–4 h of incubation at room temperature, excess antibody was washed away with two PBS washes. Nuclei were counterstained with DAPI and slides were mounted with Prolong Gold (Thermo, P36934). Imaging was performed on a Andor CR-DFLY-201-40 confocal spinning disc coupled to a Nikon Ti-E microscope. For FOS+CD44+ costaining analysis, FOS positivity in crypt epithelial CD44+ cells was measured and researchers were blinded during both imaging and quantification.
SHARE-seq
SHARE-seq was performed with minor modifications to the protocol described in ref. 24 (https://www.protocols.io/view/share-seq-v1-6qpvrdexpgmk/v1). For sorted cell and organoid experiments, frozen cells in CryoStor were briefly thawed (roughly 2 min) at room temperature before diluting with ice-cold PBS supplemented with 0.04% BSA, 0.1 U μl−1 Enzymatics RNase Inhibitor and 0.05 U μl−1 SUPERase RNase inhibitor (Thermo, AM2696). Cells were pelleted and supernatant was discarded before lysis with hypotonic lysis buffer (HLB), which is H-RSB (10 mM HEPES, 10 mM NaCl, 3 mM MgCl2) with 0.1% NP-40 (Thermo, 28324), 0.04% BSA, 0.1 U μl−1 Enzymatics RNase Inhibitor and 0.05 U μl−1 SUPERase RNase inhibitor. Following 5 min of incubation on ice, buffer was diluted with HDT-2RI (H-RSB, 0.04% BSA, 0.1% Tween-20, 0.01% digitonin (Thermo, 300410), 0.1 U μl−1 Enzymatics RNase Inhibitor and 0.05 U μl−1 SUPERase RNase inhibitor) and nuclei were pelleted. Supernatant was discarded and nuclei were resuspended in HDT-2RI at a density of 1 M ml−1 and fixed with 0.2% formaldehyde for 5 min at room temperature. Fixation was quenched with 140 mM glycine, 50 mM Tris pH 8.0 and 0.1% BSA on ice for 5 min. Fixed nuclei were washed once with HDT-2RI, once without SUPERase RNase inhibitor and stored at −80 °C until SHARE-seq was performed.
For adenoma tissue, nuclei were isolated from OCT-embedded tissue. Two to four 40-μm sections were collected from each tissue block, then excess peripheral OCT was removed and sections were placed into 1.5-ml tubes on dry ice, not allowing the tissue to thaw. Tubes were allowed to briefly warm before resuspending in 200 μl of H-RSB with 0.1% NP-40, 0.04% BSA, Enzymatics RNase inhibitor and SUPERase RNase inhibitor. Tissue was dissociated by triturating with a P1000 for 20 strokes then a P200 for 80 strokes. Nuclei were diluted with HDT-2RI, pelleted, resuspended in 500 μl of HDT-RI and filtered through a 40-μm filter (Millipore, BAH136800040-50EA) to remove large fragments of undissociated tissue. Nuclei were then fixed as described above.
Fixed nuclei were transposed as previously described in ref. 24 with Protease Inhibitor Cocktail (Sigma, P8340) and 0.1% NP-40. Reverse transcription was performed as described in ref. 24 except here we used 1× Smart-seq3 Buffer (40 mM DTT, 125 mM Tris pH 8.0, 5 mM GTP, 150 mM NaCl, 12.5 mM MgCl2) in place of Maxima RT Buffer. Washes and split-pool barcoding were performed with 0.1% Tween-20 and 0.01% digitonin instead of NP-40. Sublibrary generation, reverse crosslinking, complementary DNA (cDNA) pulldown and assay for transposase-accessible chromatin (ATAC) library preparation were all performed as previously described. Template switching was performed with 1× Smart-seq3 Buffer in place of Maxima RT Buffer. cDNA amplification and tagmentation were performed as previously described.
Organoid proliferation and AP-1 inhibition
Organoids were passaged once before AP-1 inhibition to remove dead or dying cells from primary plating. For acute AP-1 inhibition, after 3 days following passaging, the media was supplemented with 10 μM T-5224 (MedChemExpress, HY-12270) or an equal volume of dimethylsulfoxide (DMSO). After 24 h, 10 μM 5-ethynyl-2′-deoxyuridine (EdU) was added for 3 h before cell dissociation as described above. A portion of cells were banked for ATAC-seq and footprinting while the rest were fixed with 4% PFA/PBS for 10 min at room temperature, washed with PBS then permeabilized with 0.5% Triton X-100 in PBS. Following two washes with 3% BSA and PBS, EdU staining was performed with Click-iT EdU Assay kit (Life Tech, C10340) and cells were counterstained with DAPI. Percentage EdU was measured on an Attune CytPix cytometer as EdU+ cells over total DAPI+ cells. For baseline EdU differences between colitis and control organoids, EdU assays were performed at 9 days of culture.
For washout experiments, organoids were expanded for one passage to purify cultures before treatment with 10 μM T-5224 for 5 days, refreshing drug after the first 2 days. Cultures were routinely maintained for an extra 20 days as described above. Organoids were imaged using an EVOS M5000 at ×10 magnification. Organoid size was quantified using CellProfiler in which individual organoids were manually selected in the ‘MeasureObjectSizeShape’ module and the ‘Estimated Diameter Size’ was used. A two-sided Wilcoxon rank-sum test was used across all organoids quantified to compare treatment conditions.
Barcode vector cloning and library construction
The pLARRY empty vector (Addgene no. 140025) was first modified to insert a TruSeq sequencing adaptor (ACACTCTTTCCCTACACGACGCTCTTCCGATCT) upstream of the barcode insertion site to allow for direct amplification. A further sequence, including a mouse U1 hairpin, was introduced downstream of the barcode site to promote nuclear translocation of RNA transcripts and more efficient SHARE-seq capture. For nuclear localization validation, lentivirus was generated (using the method below), HEK293T cells were infected and sorted for a pure green fluorescent protein positive (GFP+) culture. Plasmids will be deposited in Addgene on publication.
Fluorescence in situ hybridization was performed using anti-GFP probes (LGC Biosearch, VSMF-1014-5) and imaging was performed on an Andor CR-DFLY-201-40 confocal spinning disc coupled to a Nikon Ti-E microscope.
For constructing barcode libraries, the following oligonucleotides were ordered from IDT:
Forward oligo: CCTATAGTGAGTCGTATTAGAGACATNNNNCTNNNNACNNNNTCNNNNGTNNNNTGNNNNCANNNNATNNNNGCATCATCAAGATCGGAAGAGCGTCGTG
Reverse oligo: CACGACGCTCTTCCGATCTTGATG
The two oligos were annealed with the following program: 95 °C for 5 min; 58 cycles of: 95 °C for 1 min, then −1 °C per cycle; 37 °C hold.
Double-stranded barcode inserts were then generated. Extension was performed by adding 1 U μl−1 Exo-Klenow (NEB, M0212L) and 1 mM dNTPs and incubating at 37 °C for 2 h, followed by enzyme inactivation at 75 °C for 20 min. The resulting annealed barcodes were purified into 20 μl of 10 mM Tris pH 8.0. The empty barcode vector (2 μg) was digested and dephosphorylated with BamHI (NEB, R3136T), XbaI (NEB, R0145S) and FastAP (Thermo, EF0652) for 4–12 h at 37 °C. Following purification, 1 μg of digested plasmid and 60 ng of annealed barcodes were assembled using NEBuilder 2× master mix (NEB, E2621S) for 1–4 h at 50 °C. The resulting product was purified into 10 μl of water, electroporated into Stbl4 ElectroMAX cells (Thermo, 11635018) and plated onto bioassay plates (Sigma, CLS431111-16EA) with carbenicillin. After growth at 30 °C for 20–24 h, colonies were scraped into media and grown for an extra 1–4 h before plasmid purification.
Organoid lentiviral infection
Lentivirus was generated by transfecting LentiX cells with second generation packaging constructs. Viral supernatant was concentrated by overnight incubation with 300 mM NaCl and 8% PEG-6000 (Millipore, 8074911000) and centrifugation at 3,000g for 30 min. Concentrated virus was resuspended in PBS before cell treatment.
Murine organoids were pretreated for 24 h with 5 μM Y-27632 before dissociation to near single cells. Cells were pre-incubated in infection media (WENR, 10 μM Y-27632, 10 μg ml−1 polybrene) for 15 min before addition of concentrated virus at less than or equal to 50% the volume of Infection Media. A rough ratio of 750 μl of concentrated virus to 25,000 cells was used. Spinoculation was performed by centrifuging at 600g for 1 h at 32 °C and the cell–viral mixture was incubated at 37 °C for 3–4 h before organoids were replated. A multiplicity of infection of less than 0.3 was used and verified by GFP expression.
SHARE-TRACE
Nuclear isolation, fixation and transposition were performed as described above. For clonal barcode capture from SHARE-seq, barcode-specific RT primer was spiked into the reverse transcription reaction at 10% general RT primer concentration:
(/5Phos/GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGNNNNNNNNNN/iBiodT/CTCATTCAGCCACGGTGG)
Split-pool barcoding, ligation and reverse crosslinking were all performed without modification. ATAC-seq libraries were generated as above. During cDNA PCR amplification, a barcode-specific forward primer mix was spiked into the reaction at 2 μM final concentration. The primer mix consisted of an equimolar mixture of:
ACACTCTTTCCCTACACGACGCTCTTCCGATC
ACACTCTTTCCCTACACGACGCTCTTCCGATCTNTAGACAT
ACACTCTTTCCCTACACGACGCTCTTCCGATCTNNTAGACAT
ACACTCTTTCCCTACACGACGCTCTTCCGATCTNNNTAGACAT
where N represent a random base mixture, introducing a frameshift in the amplification products and increased sequencing diversity of the barcode library.
Following total cDNA amplification and purification, 0.5 μl of product was removed and further amplified with the above primer mix and P7 primer for 10 cycles and Ampure purified into 10 μl. A barcode-enriched library was then generated by amplifying 5 μl of these products with P7 and a barcode-specific index primer:
AATGATACGGCGACCACCGAGATCTACAC(index5)ACACTCTTTCCCTACACGACGCTCTTCCGATCT
Whole-genome methylation profiling
Genomic DNA was extracted using the Qiagen DNeasy Blood & Tissue Kit (Qiagen, 69504). For organoid profiling, cultures were dissociated per standard protocol described above at 25 days of culture and processed per kit protocol. For whole tissue profiling, animals were perfused as described and whole colons were dissected and freshly embedded in blocks of OCT before freezing. Tissue was cryogenically sectioned and excess OCT was removed before processing per kit protocol.
For each sample, 200 ng of DNA was resuspended in water to 48 μl, mixed with control DNA from the NEBNext Enzymatic Methyl-seq V2 Kit (NEB, E8015S) total and sonicated on a Covaris S220 with settings: 60 s, 140 W, 10% duty and 200 cycles per burst. The resulting distribution of fragments was almost entirely between 200 base pairs (bp) and 600 bp. Sonicated DNA was then processed per kit protocol and libraries were sequenced on a NovaSeq X at roughly 20 times coverage.
In vivo AP-1 inhibition
The compound T-5224 was resuspended in DMSO at 200 mg ml−1 before being mixed at 10:90 with prewarmed corn oil, resulting in a 20 mg ml−1 final suspension. Mice were given 100 mg kg−1 of drug by oral gavage daily for 5 days (19 days post-DSS withdrawal to 23 days postwithdrawal). For concurrent tumorigenesis experiments, tamoxifen was administered intraperitoneally at the third day of T-5224 treatment.
Regional colon motif accessibility
The entirety of the colon (between caecum and rectum) from a healthy control mouse was extracted as previously described and then cut into six equal length fragments (each roughly 1 cm). These fragments were then individually treated with EDTA solution, crudely scraped and dissociated to single cells as previously described. Cells were resuspended in ADMEM, counted and 10,000 cells per segment were collected. Bulk ATAC-seq and library preparation were performed in the same manner as SHARE-seq without crosslinking.
Bulk ATAC-seq
Following acute AP-1 inhibition in organoids, bulk ATAC-seq was performed analogously to that of SHARE-seq without crosslinking. Briefly, cultures were dissociated to single cells and 10,000 cells were used per transposition reaction. Following 30 min of transposition, DNA was purified using a Qiagen MinElute kit and libraries were prepared following the same protocol as SHARE-seq.
FOS CUT&Tag
Epithelium was crudely scraped from control or recovered animal colons as described above and nuclei were isolated by resuspension in HLB supplemented with Protease Inhibitor Cocktail. Following 5 min of incubation on ice, nuclei were diluted with Working NE Buffer—Nuclear Extraction Buffer (20 mM HEPES pH 8, 10 mM KCl, 0.1% Triton X-100, 20% glycerol) supplemented with 0.5 mM spermidine and PIC. Following another 5 min of incubation, nuclei were centrifuged and 500,000 nuclei were resuspended in 100 μl of Working NE Buffer. Concanavalin A beads (Bangs Laboratories, BP531) were activated by washing twice in Bead Activation Buffer (20 mM HEPES pH 8, 10 mM KCl, 1 mM CaCl2, 1 mM MnCl2) and 10 μl of beads were added per sample. Nuclei were bound to beads for 10 min at room temperature, magnetized and supernatant was removed. Bound nuclei were resuspended in 50 μl of Digitonin150 Buffer (20 mM HEPES, 150 mM NaCl, 0.5 mM spermidine, 0.01% digitonin, PIC) supplemented with 2 mM EDTA, 1 μg (1:100) of anti-FOS antibody (AE-1059 from the laboratory of M. Greenberg) was added and samples were rotated overnight at 4 °C.
Beads were magnetized and resuspended in 50 μl of Digitonin150 Buffer before being added to anti-rabbit secondary (Novus, NBP1-72763). Samples were rotated for 30 min at room temperature and washed twice with Digitonin150 Buffer. Nuclei were then resuspended in 50 μl of Digitonin300 Buffer (20 mM HEPES pH 8, 300 mM NaCl, 0.5 mM spermidine, PIC, 0.01% digitonin) and 2.5 μl of CUTANA pAG-Tn5 was added (EpiCypher, 15-1017). Samples were rotated for 1 h at room temperature and washed twice with 200 μl of Digitonin300 Buffer. Nuclei were resuspended in tagmentation buffer (20 mM HEPES pH 8, 300 mM NaCl, 0.5 mM spermidine, PIC, 10 mM MgCl2) and placed on a thermocycler at 37 °C for 1 h. Tagmented nuclei were magnetized, washed with 50 μl of TAPS Buffer (10 mM TAPS, 0.2 mM EDTA) and resuspended in 5 μl of SDS Release Buffer (10 mM TAPS, 0.1% SDS). Following an incubation at 58 °C for 1 h, 15 μl of SDS Quench Buffer (0.67% Triton X-100) was added and libraries were prepared using NEBNext HiFi 2× Master Mix.
In vitro TF binding assay
Genomic regions to investigate were selected by filtering for peaks with a difference in normalized footprint score of at least 0.2 at a FOS or JUN motif between colitis-recovered and control tissue (described below in ‘Footprinting and seq2PRINT’). Regions were then partitioned into those with only an AP-1 or AP-1/FOX composite motif based on motif matching (also described below in the section ‘Footprinting and seq2PRINT’) and confirmed by visual inspection of DNA sequence. The top regions by change in footprinting score were then selected and a roughly 1,000-bp region from each was amplified from mouse genomic DNA. Following purification, amplicons were pooled in equimolar concentrations.
In vitro footprinting was performed as described in ref. 46 with the following modifications: briefly, selected sequences (25 ng per reaction) were incubated with various combinations of recombinant JUN (Active Motif, 31116), FOS (OriGene, TP760257), FOXP1 (OriGene, TP313862) and FOXA1 protein (OriGene, TP306045), along with tagmentation buffer (20 mM Tris, 10 mM MgCl2 and 20% dimethylformamide) and water in a 22.5-μl total volume at room temperature for 1 h. Then, 0.15 μl of preassembled Tn5 (seqWell, Tagify) was combined with 2.35 μl of dilution buffer (50 mM Tris, 100 mM NaCl, 0.1 mM EDTA, 1 mM DTT, 0.1% NP-40 and 50% glycerol), and subsequently added to samples (resulting in final TF concentrations of 300 nM each). Tagmentation was performed for 30 min at 37 °C. A Qiagen MinElute PCR clean-up kit was used to purify tagmented DNA, and samples were then PCR amplified for seven cycles. After pooling, sample libraries were sequenced on a Next-seq 500/550.
Adenoma induction, macroscopic quantification and proliferation
Mice were ordered from The Jackson Laboratory (035169) with genotype wild type for Kras<tm4Tyj>, homozygous for Apc<tm1Tno> and homozygous for Tg(CDX2–cre/ERT2)752Erf. Adenomas were induced at 21–23 days following DSS cessation. For macroscopic tumours, tamoxifen dissolved in corn oil was administered at 50 mg kg−1 by intraperitoneal injection on 3 consecutive days and animals were euthanized 25–28 days following the first injection. The entire length of the colon was removed and imaged. Owing to higher expression of the CDX2–cre driver of this mouse model in the proximal colon50, more tumours form in the 1 cm of colon adjacent to the caecum and adenomas were quantified in the distal 5 cm of colon for more accurate counting. Adenoma diameter was measured in ImageJ along the longest axis of each tumour and scaled to millimetres using a ruler placed in the same image. Researchers were blinded during quantification.
To measure proliferation, immunohistochemistry for Ki67 (anti-Ki67 1:100, Abcam, ab15580; secondary anti-rabbit 1:500, Vectastain Elite ABC, PK-6101) was performed and the proliferating fraction was quantified in only adenoma areas, as identified by H&E morphology on adjacent sections. Researchers were blinded during quantification.
Ex vivo adenoma organoid clonogenicity
Organoids were derived from adenoma tissue 23 days following the first tamoxifen administration. Tissue was crudely scraped following EDTA treatment and dissociated to near single cell as previously described. Cells were plated in Crypt Basal with 50 ng ml−1 EGF and 100 ng ml−1 Noggin and allowed to grow for at least 1 passage to purify cultures. Clonogenicity (colony-forming efficiency) was calculated on the secondary organoids by plating 1,000 cells passaged from the primary organoids and assessing organoid formation 7 days after initiation of cultures.
Microscopic tumour initiation and quantification
Adenomas were induced 21 days following DSS cessation with a single dose of tamoxifen at 10 mg kg−1 and euthanized 13–14 days following. For AP-1 inhibition assays, T-5224 was administered 2 days before tamoxifen induction, the day of induction and 2 days following, as described above in the section ‘In vivo AP-1 inhibition’. The entire length of colon was removed, fixed and paraffin embedded as a Swiss roll as previously described. Immunohistochemistry for β-catenin (anti-β-catenin 1:200, BD, 610153; secondary anti-mouse 1:500 Vectastain Elite ABC, PK-7200) was performed. The entire section was first imaged at low magnification (×2) to quantify total tissue area, then microscopic tumours were identified by high nuclear β-catenin staining and imaged under high magnification (×20). Both total tissue and individual tumour areas were quantified using ImageJ, using scale bars as reference, and tumour area summed across all tumours and reported as a percentage of total tissue. Researchers were blinded during imaging and quantification.
Spatial transcriptomics on adenoma tissue
Spatial RNA-seq was performed on fresh frozen Swiss rolls of colons with adenomas induced at 50 mg kg−1 × 3. The Visium HD Kit was used according to standard protocol with the following modifications: the OCT-embedded Swiss rolls were sectioned on a cryostat at 10-μm thickness and mounted on a Fisherbrand Superfrost Plus glass slide. The slides were fixed using 4% PFA and stained using H&E method. During that process the wash buffers were supplemented with either ribonucleoside vanadyl complex or a commercial RNase inhibitor. After imaging of the H&E staining, the samples were destained and permeabilized using 1% SDS and then prechilled 70% methanol. Tissue processed this way was then analysed using a standard 10X Visium HD method described in the GenBank Nuccore User Guide CG000685 (Rev. A).
Data processing and analysis
SHARE-seq raw data processing
Raw SHARE-seq data were processed as previously described24 with minor modifications (code available at https://github.com/masai1116/SHARE-seq-alignmentV2/). Briefly, raw fastq files were demultiplexed using custom Python scripts. ATAC-seq reads were aligned to the mm10 or hg38 genome using bowtie2 (v.2.3.3.1)60, removing fragments with length longer than 2 kb. RNA-seq reads were aligned using STAR (v.2.5.3a)61, removing reads with greater than 20 alignments or score less than 0.3. Both library types were further filtered to remove mitochondrial reads and reads with a mapping quality less than 30, and chrY reads were removed from ATAC libraries. Filtered ATAC reads were then deduplicated and further filtered for cell barcodes with at least 100 raw reads. Filtered RNA reads were assigned to genes using featureCounts62 using only primary mapping coordinates and unique molecular identifiers (UMIs) were counted using umi_tools (v.1.0.1)63, removing those consisting of only ‘G’s. Libraries were then filtered for RNA cell barcodes with at least 300 UMIs or ATAC cell barcodes with a library size of 500 reads before further processing and filtering.
scRNA-seq processing
All filtered cell barcodes were normalized by the total number of transcripts detected. The top 5,000 most variable genes were selected and principal components analysis was performed on the log2 + 1 transformed values of these genes. Library sizes were smoothened over the 20 k-NN in this space and clumps of cells were manually identified as those barcodes with extremely high smoothened library sizes. Once clumps were removed, the remaining barcodes were processed with scrublet64 to identify doublets and barcode collisions, in which doublet score thresholds were manually selected. The remaining singlet filtered barcodes were again normalized before training scVI65 models (20-dimensional latent space; negative-binomial likelihood) with batch as a covariate to learn a shared low-dimensional representation of cells. The resulting latent features were used to build a k-NN graph and to compute uniform manifold approximation and projection (UMAP) embeddings for visualization. Gene expression was visualized after using k = 5 k-NN for smoothening normalized values and capping maximum z score values at 3.
scATAC-seq processing
Peaks were called on a merged set of fragments from all sublibraries for each dataset (in vivo tissue memory, ex vivo organoid culture or adenoma tissue) using MACS2 (v.2.2.9.1)66. These peaks were then filtered using a previously described approach67 in which summits were padded with 400 bp on either end, overlapping windows were filtered for those with higher significance and finally resized to a set of non-overlapping 301-bp peaks. For adenoma tissue samples, this process was done in conjunction with peaks identified from the in vivo tissue memory dataset. Fragments were counted within peaks for each cell barcode and barcodes with low library size (in vivo tissue memory and ex vivo organoids, 1,000 fragments and adenoma tissue, 2,000 fragments) or fraction of reads in peaks (in vivo tissue memory and adenoma tissue fraction of reads in peaks less than 0.2, ex vivo organoids fraction of reads in peaks less than 0.25) were removed. cisTopic (v.0.3.0)68 models were generated on all filtered cell barcodes for 10 × n topics for n = 1 to 9, with 150 iterations and burnin of 120. Library sizes were smoothened over the 20 k-NN in this space and clumps of cells were manually identified as those barcodes with extremely high smoothened library sizes. These clump barcodes, as well as clump and doublet barcodes identified in the matched RNA cell barcodes, were removed to identify singlet cell barcodes. Using the value of n selected on all cell barcodes, cisTopic models were generated again on singlet barcodes using 10 × (n − 1), 10 × n, 10 × (n + 1) topics. UMAP, k = 5 k-NN and Louvain graph-based clustering analysis were performed in the resulting topic space. Gene expression visualization was performed by matching ATAC cell barcodes to corresponding RNA cell barcodes to get smoothened normalized expression values. For genome track visualization, fragments were separated by colitis stage, normalized to 10 million total reads, Tn5 insertion sites were counted and images were made using UCSC Genome Browser.
Cell type identification
For each single-cell ATAC using sequencing (scATAC-seq) Louvain cluster, the fraction of cells expressing each marker gene and the average normalized expression within those cells was computed and plotted. Clusters with high expression of Ptprc or Acta2 were designated as non-epithelial cells. Clusters with high expression of Muc1, Chga/b or Dclk1 were designated as secretory cells and subdivided as goblet, enteroendocrine or tuft cells when possible. In non-neoplastic tissue, the clusters with high expression of Lgr5 and Lrig1 were assigned as stem and progenitor cells and those with high expression of Car4, Car1, Lypd8 or Aqp8 were assigned as differentiated absorptive enterocytes. The remaining clusters with moderate expression of those genes were assigned as intermediate absorptive enterocytes. Analogous assignment was performed for ex vivo organoid clusters. In extended timepoints analysis (50 days, 79 days and 102 days), data from each timepoint was processed as described above to identify singlets. Each individual timepoint was then merged with the larger main dataset to rederive topics (up to 150) and cocluster. Cell type labels from the main dataset were used to identify new clusters and cell types in the new timepoint datasets. Labels from the original identification were retained.
For adenoma experiments, the clusters with high expression of Axin2 and high motif scores for LEF1, in conjunction with high expression of Lgr5, Lrig1 and Mki67, were assigned as adenoma cells. The remaining epithelial non-secretory cells were then partitioned as described for non-neoplastic tissue.
Gene expression change analysis
For differential analysis, raw UMI counts were first pseudobulked by cell type in each animal then tested using DESeq2 (v.1.42.1)69. For in vivo memory analysis, genes were filtered for those with a minimum reads per million (RPM) of 10 in at least one pseudobulk, then each disease stage was compared to controls and an FDR of 0.05 was used for significance. Genes with suspected multimapping artefacts were identified as those with excessively large expression values across all cells and blacklisted (14 genes). The log2[fold change] values were taken from DESeq2 output. For adenoma gene activation, all adenoma pseudobulks were tested against all stem and progenitor pseudobulks, regardless of colitis condition. When plotting individual genes, raw UMI counts were pseudobulked across stem and progenitor cells within each animal and RPM values were calculated using total assigned UMIs. Z scores were calculated across all animals for each gene and change was calculated by subtracting average value across control pseudobulks. Gene ontology analysis70 was done with all differential genes.
k-NN enrichment analysis
For the in vivo memory dataset, cells were subsetted to stem and progenitors only and 100 k-NN were determined for each cell using cells from all other biological replicates. The cisTopic space defined above was used for scATAC-seq and scVI was used for dimensional reduction for scRNA-seq. To obtain the batch-corrected latent representation of scRNA-seq data, count matrices are normalized by total counts per single cell and log-transformed. In total 10,000 highly variable genes were selected while accounting for batch effects using scanpy (highly_variable_genes with ‘flavour’ set as ‘seurat_v3’ and ‘batch_key’ as batch id.) We then used k-NN from scVI trained models, as described above. For each condition (control, acute injury, chronic injury, recovery), the expected value of k-NN was calculated for random assignment to the 100 k-NN and therefore was the fraction of cells in each condition. Enrichment for each condition was then calculated as the (observed percentage of k-NN) − (expected percentage of k-NN). The analogous procedure was used for human organoids except using all cells.
Motif accessibility analysis
Peaks were annotated as containing a motif using motifmatchr (v.1.12.0) for known cisBP71 motifs. For de novo derived motifs, peak annotation was performed as described in the ‘Footprinting and seq2PRINT’ method section. Single-cell motif deviations and accessibility scores were calculated with chromVAR (v.1.24.0)28 using 250 background peaks across all single cells within each dataset (in vivo memory, ex vivo organoids or adenoma-induced tissue). A motif similarity matrix was calculated on all known and de novo motifs using Tomtom (memesuite v.5.5.7)72 and a q value cut-off of 0.05 was used to group motifs into families based on sequence similarity. This step ensures reliable motif–motif comparisons for downstream analysis. During the bagging process, motifs are sorted based on their variation across cells and those with highest variation were retained as ‘leaders’, whereas other motifs with high similarity scores to these representatives are merged into their respective ‘families’, effectively consolidating similar motifs into unified groups. These motif families were created using the in vivo tissue memory dataset and held constant across all other murine tissue and organoid experiments. Motif families were derived analogously in the human organoid dataset.
For motif accessibility testing, the mean value was computed across single-cell scores of all cells of a given type within each animal or all cells in an organoid line. For in vivo memory, P values were computed for the 50 most variable families by t-test and adjusted using the Benjamini–Hochberg method. Motif accessibility change was defined as the difference between the score for each replicate and the mean across control animals or organoid lines. When visualizing the in vivo change per mouse as a heatmap, samples with fewer than 200 stem cells were excluded. For extended recovery analysis (50 days, 79 days and 102 days), all cells were rescored together with the new data and AP-1 motif accessibility change was recalculated using controls from all timepoints.
Heterogeneity in single-cell motif accessibility was evaluated by randomly downsampling each condition to 500 stem and progenitor cells and computing P values for selected motifs with a two-sided Wilcoxon rank-sum test. Activated cells were defined as stem and progenitor cells with a score greater than 1.5 and the fraction of activated cells was calculated for each animal using all stem and progenitor cells.
Stem and progenitor subcluster analysis
For reclustering to stem cells, all cells were subsetted to those identified as stem and progenitor (section ‘Cell type identification’) and cisTopics scores generated on the overall dataset were used. UMAP embeddings and clusters (k = 5) were generated as was done on the all cell dataset. Groups were assigned manually. In assessing stemness genes, expression was averaged across all cells within a group from a given animal and the mean across animals was then plotted.
The AP-1 high subpopulation was identified as stem cells with AP-1 motif score greater than 1.5. Differential testing was performed using a two-sided Wilcoxon rank-sum test between these AP-1 high cells and all other stem cells. For motifs, this was done across the top 50 most variable motif families, as identified in the overall dataset previously. For genes, the top 1,000 most variable genes across stem cells were tested using expression values normalized to reads per cell.
SHARE-TRACE clone assignment
Demultiplexed reads were processed with custom Python scripts to search for common barcode vector sequence TAGACAT, allowing at most one mismatch. This sequence and all preceding base pairs were trimmed, and any reads with UMIs consisting of only Gs were removed. The remaining 48 bp of barcode sequences were validated by checking for staggered invariant sequences every 4 bases (CT,AC,TC,GT,TG,CA,AT,GC), removing off-target PCR products. To account for sequencing base call errors, the number of reads for each cell–UMI–barcode triple were counted and those with fewer than five reads were removed.
To identify clonal barcode sequences, a Levenshtein distance matrix was calculated across all remaining barcode sequences. For each barcode sequence, all other sequences within a distance of four were found and the barcode was assigned to the most abundant of those sequences. Distance between members of this set of most abundant sequences was computed once more and any sequences within a distance of two were collapsed to the more abundant sequence, generating a consensus set of clonal barcodes. The remaining reads were used to assign each cell–UMI pair to a clone.
To account for transcript mixing that occurs during SHARE-seq split-pooling, we leveraged the fact that each clone should be unique to organoids generated from a single mouse. Cell-clone assignments were matched to validated ATAC or RNA singlets and each clone was assigned to the animal from which the most cells were present (typically more than 95% of barcode reads). Only clones with at least five cells were used for subsequent analysis.
SHARE-TRACE clonal variance calculation
For each feature (motif families or gene programs), the standard deviation of the single-cell scores was calculated across all cells in a clone and the observed clonal variance was then defined as the median of these values across clones to the second power. Cell-clone assignments were then randomly permuted and the shuffled clonal variance was computed analogously. This process was repeated 1,000 times and the mean and standard deviation of the resulting distribution of randomized clonal variance was used to compute a P value:
Z = (observed clonal variance − mean shuffled clonal variance)/(s.d. shuffled clonal variance)
P = 2 × pnorm(−abs(Z))
The FDR was then calculated using the Benjamini–Hochberg method of P value adjustment.
For comparisons between colitis and control organoids, the median value of the scores for all cells belonging to each clone was computed and a two-sided t-test was performed on these values. Clones with high AP-1 accessibility were defined as those with a median score greater than 1.25.
Linear mixed model of single-cell variance
A linear model was created to evaluate the contribution to variance for past exposure to colitis and clonal identity. A data table was formed in which each cell barcode had a designation as having originated from a mouse that experienced colitis (‘is_colitis’) and clone assigned (‘clone’). This was represented as:
form ~ (1|is_colitis) + (1|clone)
Variance contributions were then calculated using the variancePartition R package using fitExtractVarPartModel(), providing the matrix of single-cell motif scores across all motif families. This was compared to randomized clonal distribution by permuting clonal labels within colitis conditions. Cells were subsetted to those exposed to colitis and clone labels were permuted within this, before performing the same procedure on cells not exposed to colitis.
Gene expression program derivation and scoring
For gene expression program modelling, the input was a cell-by-gene count matrix. Unlike the cistopic approach for scATAC-seq data68, this matrix was not binarized but remains in raw count format. Latent Dirichlet allocation, implemented in the Mallet package, was used to infer: (1) the probability distribution of topics for each cell, and (2) the probability distribution of genes for each topic. Latent Dirichlet allocation models were trained with a range of topic numbers (30–90), and the model with the highest log-likelihood was selected, following a procedure similar to cisTopic.
To score single cells on these programs, we adapted the chromVAR algorithm for RNA topics. The input cell-by-peak accessibility matrix was replaced with the cell-by-gene transcription matrix, and the motif-by-peak matching matrix is substituted with the topic-by-gene probability distribution matrix inferred by latent Dirichlet allocation. The rest of the calculations remain identical to the original chromVAR workflow. Background genes were generated by grouping all genes in 20 bins of equivalent size based on average expression and 250 background genes were chosen for each gene in the annotated set. The resulting cell-by-topic scores represented the activity levels of RNA topics in each cell while controlling for sequencing depth, gene expression level and other biases.
To identify AP-1- and HNF4/PPAR-associated gene programs, the mean motif score and gene program score across all cells within each clone was calculated. These mean values were then correlated across clones to get motif–gene program correlation values. The top programs were selected for each motif family (gene P20 for AP-1, and gene programs 9 and 30 for HNF4/PPAR) and the top 150 genes by weight of contribution to the program were selected for Gene Ontology analysis and subsequent gene program scoring. For plotting scores on UMAP projections, single-cell score values were capped at −3 and 3.
For single gene analysis in organoids, scRNA-seq reads were pseudobulked by organoid clone and normalized to total reads per clone. Fold change was calculated as the ratio between the mean normalized expression value across all colitis clones and all control clones. Values were scaled by gene before plotting as heatmap. Plotting individual gene change in tissue across disease timepoints was done as described in the section ‘Gene expression change analysis’. Differential testing comparison between chronic injury and 50-day recovery was done with DESeq2 as previously described in ‘Gene expression change analysis’ and then one-sided values were computed as one_sided_P = two_sided_P/2 for genes with positive log2[fold change] and 1 − two_sided_P/2 for negative log2[fold change].
EM-seq methylation data processing
EM-seq data were processed and aligned to the mm10 reference genome using the nf-core/methylseq pipeline (v.4.0.0)73 with GPU-enabled bwa-meth, and methylation calls were obtained with MethylDackel (v.0.6.1). We derived two types of feature: (1) ATAC-seq peak-anchored methylation, defined as the fraction of methylated cytosines within ±500 bp of ATAC-seq peak summits, and (2) per-CpG methylation, calculated by combining strand-specific counts at each CpG site, and further derived the fraction of methylated cytosines.
Methylation fraction quantification
For testing change in methylation, peaks were first filtered for those with at least 10% methylation in at least one sample and a standard deviation in fraction methylation of at least 0.05. A two-sided t-test was then performed between colitis-derived organoids and controls across 58,454 regions resulting from this filtering and FDR was calculated with Benjamini–Hochberg adjustment. These methylation change values were compared to ATAC-seq signal by pseudobulking scATAC-seq counts by organoid line, normalizing to 1 million reads per sample and calculating fold change per peak as the average RPM for all colitis organoid lines over all control organoid lines. The methylation at individual CpGs was then visualized by creating a per-base heatmap in a given genomic interval coloured by percentage methylation, with the values for each CpG being extended half way to the next CpG to cover non-CpG bases.
For analysis of AP-1 sites, motif annotations from seq2PRINT (section ‘Footprinting and seq2PRINT’) were used to identify relevant peaks. Variance in methylation fraction across these peaks was calculated and the top 500 peaks were selected to visualize. Average fraction methylation was calculated across all animals in a given condition and the difference between these values was plotted. For colitis recovered versus control comparison, this was all DMSO-treated colitis-recovered animals and DMSO-treated control animals. For T-5224 comparison, this was all T-5224-treated colitis-recovered animals and DMSO-treated colitis-recovered animals. The matched background set of peaks (100 per AP-1 peak selected before) was generated with chromVAR across single cells, controlling for average accessibility and GC-content. P values were then computed using two-sided Wilcoxon rank-sum test between AP-1 peaks and matched background peaks.
Footprinting and seq2PRINT
Footprint scores were calculated using the scPrinter package (v.1.2.0)46. The data used were as follows: in vivo memory, all epithelial cells; AP-1 inhibition, all bulk ATAC-seq reads; human organoids, all cells and mouse adenomas and only adenoma cells. Briefly, the seq2PRINT model was trained to take DNA sequences spanning a candidate cis-regulatory element (cCRE) of interest and its surrounding regions (±920 bp) as input and predict multiscale footprints derived using the PRINT method. After training, DeepLIFT is used to extract sequence attribution scores, which represent the contribution of each base pair in the input sequence to the predicted footprints. These sequence attribution scores enable highly accurate TF binding predictions through a neural network model trained on chromatin immunoprecipitation sequencing data46 and are referred to as seq2PRINT footprint scores. In this study, raw seq2PRINT footprint scores were binned at a 10-bp resolution for all cCREs. Bins with maximum seq2PRINT footprint scores below 0.2 across all conditions are excluded from further analysis. To facilitate comparisons across different conditions, the scores were quantile-transformed for each condition using the quantile_transform function in scikit-learn (n_quantiles=100000, target_distribution=‘uniform’; v.1.5.2).
Sequence attribution scores from seq2PRINT were further analysed to identify de novo motifs. TF-MoDISco (v.2.2.1) is used to align and cluster seqlets (local subsequences with high sequence attribution scores) into groups of de novo motifs. To assign these motifs to cCREs, the software finemo (v.0.40) was used, which takes the output de novo motifs from TF-MoDISco and the sequence attribution scores as input for motif matching.
Motif accessibility change in de novo motifs was calculated as described above, in which single-cell scores were averaged across all stem and progenitor cells in a given animal, P values computed by t-test compared with control animals and change was defined as the difference from the control animal average.
To visualize how the seq2PRINT model learns the association between input DNA sequences and output multiscale footprints, we generated marginalized predictions (referred to as delta effects) for given motifs using the delta_effects_seq2print function from the scprinter package. Briefly, we randomly selected 10,000 CRE sequences, inserted the consensus sequence of a specific motif at the centre and averaged the difference in model predictions with and without the inserted motif across the 10,000 CREs.
In vitro binding score analysis
Sequencing data were processed using the bulk ATAC-seq pipeline described previously in ref. 46. All conditions were downsampled to an equivalent number of insertions in selected regions. Multiscale footprinting was performed using a one-sided binomial test, testing for depletion of insertions in centre versus flanking windows of varying radius lengths (2–100). To test for depletion and account for intrinsic Tn5 sequence preferences, expected insertions in centre and flanking windows were calculated using observed frequencies in a naked (no TF) control condition. As in PRINT46, binding scores represent −log[P] of depletion, relative binding score per locus was then calculated by dividing the binding score for each TF combination by the binding score for FOS and/or JUN at the same region. P values were calculated for each TF combination by a one-sample t-test against a μ value of 1, separately for all AP-1 only loci and all AP-1:FOX composite loci.
AlphaFold3-based structure prediction
To nominate FOX factors to investigate, scRNA-seq reads of stem cells from each mouse were pseudobulked and normalized to RPM. The full amino acid sequences and features of murine FOS (P01101), JUN (P05627), FOXP1 (P58462), FOXA1 (P35582), FOXN2 (E9Q7L6) and FOXJ2 (Q9ES18) were downloaded from UniProt. Structures were predicted using AlphaFold3 (r2024.05.23)74 by means of the AlphaFold Server by combining nucleotide sequences and amino acid sequences. Each structure included the motif nucleotide sequence, its reverse complement, and the amino acid sequences of FOS, JUN and FOX proteins. PyMOL (v.3.1.3) was used to select for interacting residues and structure visualization. To reduce the amount of disordered regions in each structure and to focus on protein-to-protein and protein-to-DNA interactions, the beginning and end of the protein sequences were truncated to only include regions containing residues within 3.5 Å of neighbouring protein and DNA structures. AlphaFold3 was once again used to predict the truncated structures containing the complete DNA sequence, truncated FOS chain, truncated JUN chain and truncated FOX chain. FOX residues within 3.5 Å of either FOS or JUN were highlighted in the truncated structures as an estimate of interaction between proteins. The location of the interaction residues were compared to known features for each FOX protein. For visualization purposes, structures were modified to only include regions with a B-factor greater than 50.
CUT&Tag processing and analysis
FOS CUT&Tag reads were aligned to the mm10 genome annotation and processed analogously to SHARE-seq (ignoring the single-cell barcode considerations). FOS binding signal analysis was performed by counting CUT&Tag reads within ATAC-seq peaks called from SHARE-seq data and summing reads across replicates for control or recovered samples. RPM was calculated at each peak and variable peaks were identified as the top 10,000 by standard deviation across RPM values after filtering for peaks with a minimum of 20 raw reads in at least 1 sample.
For footprinting prediction performance, peaks were first called on FOS CUT&Tag data using MACS2. Reads were counted within peaks, normalized to 10 million total reads and a t-test was performed between recovered and control samples. Differential peaks were defined as having a P value of less than 0.05. Normalized footprint scores were then condensed by motif, in which overlapping AP-1 motifs were collapsed to one site and change in footprint score was calculated between recovered and control. Performance was then evaluated by calculating the sensitivity and specificity of identifying differential peaks on the basis of varying thresholds of footprint score difference.
Cobinding analysis
Change in footprint score was calculated as the difference between acute, chronic or recovered scores and their batch matched controls. Memory footprints were defined as those showing a minimum change of 0.2. Using all pairs across the selected families, peaks were labelled as containing a memory motif for the first family only, second family only, both families or neither family, and an odds ratio of the resulting contingency table was calculated. Cobinding scores were then calculated by log2-transforming these odds ratios, performing 10th to 90th quantile normalization across all pairs on positive and negative ratios separately, and linearly scaling positive values between 0 and 1 and negative values between 0 and −1. For primary tissue memory, this normalization and scaling was performed on values across all timepoints. For human IBD organoids and mouse adenomas, footprint differences were calculated as (IBD − Control) and (Recovered adenoma − Control adenoma), respectively. For AP-1 cobinding loss, the analysis was performed analogously using a decrease in footprint score of 0.2 (T-5224 − DMSO). Distance between TFs was calculated by finding the midpoints of all memory sites and calculating all distances between sites within the same peak. Negative footprint scores were set to 0 before plotting.
Motif families were identified as above under ‘Motif accessibility analysis’ with the following modifications:
ETS: ETS family with SPI1, SPIB and SPIC added.
RUNX: RUNX1, RUNX2 and RUNX3.
SNAI/MESP: SNAI2, SNAI3, MESP1, MESP2, TCF3 and TCF4.
ESRR: ESRRA, ESRRB and ESRRG.
Retinoid: RXRA, RXRB, RXRG, RARA and RARB.
Visium HD data processing and visualization
Raw BCL files were converted to fastqs using spaceranger (v.3.0.1) mkfastq and gene expression values were computed using spaceranger count using the Visium_Mouse_Transcriptome_Probe_Set_v2.0 and mm10 genome reference data. Bins of a size of 16 μm were used. For plotting of individual genes, beads were first filtered for a minimum of 300 reads, and then normalized to average read depth across all remaining beads. Principal components analysis was performed on the log2 + 1 transformation of these values and the top 20 principal components were used to find k-NN (k = 20) for smoothening before plotting of Z-scored expression values (capped at −3 and 3).
Spatial transcriptomic adenoma identification and gene scoring
To identify tumour cells, the principal component smoothened Axin2 expression values were Z scored and adenoma cells were defined as beads with a minimum value of 1. Tumours were then called by finding k = 5 k-NN using spatial coordinates of adenomas cells only and performing Louvain graph-based clustering.
For AP-1-associated genes, the top 150 genes from P20 were overlapped with probes present in the Visium output and then this set of genes was scored analogously to single cells as above, treating bins as cells. In single bin spatial plots, scores were plotted without smoothening and capped at −2 and 2. For whole tumour analysis, raw expression values were pseudobulked by tumour call and these pseudobulks were scored analogously as cells. Tumours with high AP-1-associated gene expression were identified as those with scores greater than 1.5. For analysis of single AP-1 related genes, RPM values were computed on single bins. Visualization of these genes in adenomas only was performed by subsetting bins to only tumour cell calls, renormalizing and resmoothening expression values in only these cells, smoothening once more across the k = 20 x–y coordinate k-NN and capping Z scores at −3 and 3 before plotting.
Adenoma heterogeneity analysis
Expression values from scRNA-seq were pseudobulked by tumour and normalized as described. For unbiased analyses, the top 1000 most variable genes across tumour pseudobulks were selected. For P20, genes were filtered to be expressed in at least 20% of tumours. Fold change was calculated as the ratio between average expression across colitis tumours and across control tumours. Expression values were scaled by gene for heatmap visualization. Differential testing between high P20 tumours and others, as well as colitis tumours and controls, was performed using tumour pseudobulks and DESeq2 as described in ‘Gene expression change analysis’.
Statistical analysis
Except where noted in figures, legends or methods, all experiments in this study were repeated at least three times. All sample numbers (n) of biological replicates and definitions of centre and dispersion are defined in the figure legends. No statistical methods were used to predetermine sample size. Light microscopy, immunofluorescence, immunohistochemistry and gross morphology images each represent one of six or more biological replicates unless otherwise stated. All values are shown as mean ± standard error of the mean (s.e.m.) unless otherwise specified. No animals were excluded from statistical comparisons. Age-matched mice were randomly assigned to treatment groups. Blinding was not performed except for imaging experiments, as noted in the Methods.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All single-cell sequencing data have been provided to the Impact of Genomic Variation on Function consortium and may be accessed through https://tinyurl.com/nagaraja-buenrostro-2025. All bulk and spatial sequencing are available through the Gene Expression Omnibus as a part of series at GSE316619. Reference genomes for H. sapiens (hg38) and M. musculus (mm10) are available from the University of California at https://hgdownload.soe.ucsc.edu/goldenPath/. Source data are provided with this paper.
Code availability
The code for reproducing the analysis is available at GitHub (https://github.com/snagaraja13/colitis_memory). A static version of the code used for specific analyses is available at Zenodo (https://doi.org/10.5281/zenodo.18487777)75.
References
Coussens, L. M. & Werb, Z. Inflammation and cancer. Nature 420, 860–867 (2002).
Grivennikov, S. I., Greten, F. R. & Karin, M. Immunity, inflammation, and cancer. Cell 140, 883–899 (2010).
Beaugerie, L. & Itzkowitz, S. H. Cancers complicating inflammatory bowel disease. N. Engl. J. Med. 373, 195 (2015).
Ekbom, A., Helmick, C., Zack, M. & Adami, H. O. Ulcerative colitis and colorectal cancer. A population-based study. N. Engl. J. Med. 323, 1228–1233 (1990).
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Flavahan, W. A., Gaskell, E. & Bernstein, B. E. Epigenetic plasticity and the hallmarks of cancer. Science 357, eaal2380 (2017).
Naik, S. & Fuchs, E. Inflammatory memory and tissue adaptation in sickness and in health. Nature 607, 249–255 (2022).
Bell, C. C., Faulkner, G. J. & Gilan, O. Chromatin-based memory as a self-stabilizing influence on cell identity. Genome Biol. 25, 320 (2024).
Bintu, L. et al. Dynamics of epigenetic regulation at the single-cell level. Science 351, 720–724 (2016).
Ostuni, R. et al. Latent enhancers activated by stimulation in differentiated cells. Cell 152, 157–171 (2013).
Kaufmann, E. et al. BCG educates hematopoietic stem cells to generate protective innate immunity against tuberculosis. Cell 172, 176–190 (2018).
Netea, M. G. et al. Trained immunity: a program of innate immune memory in health and disease. Science 352, aaf1098 (2016).
Naik, S. et al. Inflammatory memory sensitizes skin epithelial stem cells to tissue damage. Nature 550, 475–480 (2017).
Gonzales, K. A. U. et al. Stem cells expand potency and alter tissue fitness by accumulating diverse epigenetic memories. Science 374, eabh2444 (2021).
Larsen, S. B. et al. Establishment, maintenance, and recall of inflammatory memory. Cell Stem Cell 28, 1758–1774 (2021).
Del Poggetto, E. et al. Epithelial memory of inflammation limits tissue damage while promoting pancreatic tumorigenesis. Science 373, eabj0486 (2021).
Foster, S. L., Hargreaves, D. C. & Medzhitov, R. Gene-specific control of inflammation by TLR-induced chromatin modifications. Nature 447, 972–978 (2007).
Falvo, D. J. et al. A reversible epigenetic memory of inflammatory injury controls lineage plasticity and tumor initiation in the mouse pancreas. Dev. Cell 58, 2959–2973 (2023).
Di Tommaso, N., Gasbarrini, A. & Ponziani, F. R. Intestinal barrier in human health and disease. Int. J. Environ. Res. Public Health 18, 12836 (2021).
de Sousa e Melo, F. et al. A distinct role for Lgr5 stem cells in primary and metastatic colon cancer. Nature 543, 676–680 (2017).
Sender, R. & Milo, R. The distribution of cellular turnover in the human body. Nat. Med. 27, 45–48 (2021).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Wirtz, S. et al. Chemically induced mouse models of acute and chronic intestinal inflammation. Nat. Protoc. 12, 1295–1309 (2017).
Ma, S. et al. Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell 183, 1103–1116 (2020).
Agudo, J. et al. Quiescent tissue stem cells evade immune surveillance. Immunity 48, 271–285 (2018).
Biton, M. et al. T helper cell cytokines modulate intestinal stem cell renewal and differentiation. Cell 175, 1307–1320 (2018).
Okayasu, I., Ohkusa, T., Kajiura, K., Kanno, J. & Sakamoto, S. Promotion of colorectal neoplasia in experimental murine ulcerative colitis. Gut 39, 87–92 (1996).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Li, L. et al. Epithelial-specific ETS-1 (ESE1/ELF3) regulates apoptosis of intestinal epithelial cells in ulcerative colitis via accelerating NF-κB activation. Immunol. Res. 62, 198–212 (2015).
Sizemore, G. M., Pitarresi, J. R., Balakrishnan, S. & Ostrowski, M. C. The ETS family of oncogenic transcription factors in solid tumours. Nat. Rev. Cancer 17, 337–351 (2017).
Alonso-Curbelo, D. et al. A gene-environment-induced epigenetic program initiates tumorigenesis. Nature 590, 642–648 (2021).
Vallejo, A. et al. An integrative approach unveils FOSL1 as an oncogene vulnerability in KRAS-driven lung and pancreatic cancer. Nat. Commun. 8, 14294 (2017).
Yap, E.-L. & Greenberg, M. E. Activity-regulated transcription: bridging the gap between neural activity and behavior. Neuron 100, 330–348 (2018).
Galbraith, M. D. & Espinosa, J. M. Lessons on transcriptional control from the serum response network. Curr. Opin. Genet. Dev. 21, 160–166 (2011).
Han, Y. et al. Cortical inflammation is increased in a DSS-induced colitis mouse model. Neurosci. Bull. 34, 1058–1066 (2018).
Bamias, G., Pizarro, T. T. & Cominelli, F. Immunological regulation of intestinal fibrosis in inflammatory bowel disease. Inflamm. Bowel Dis. 28, 337–349 (2022).
Qu, M. et al. Establishment of intestinal organoid cultures modeling injury-associated epithelial regeneration. Cell Res. 31, 259–271 (2021).
Nusse, Y. M. et al. Parasitic helminths induce fetal-like reversion in the intestinal stem cell niche. Nature 559, 109–113 (2018).
Yui, S. et al. YAP/TAZ-dependent reprogramming of colonic epithelium links ECM remodeling to tissue regeneration. Cell Stem Cell 22, 35–49 (2018).
Weinreb, C., Rodriguez-Fraticelli, A., Camargo, F. D. & Klein, A. M. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science 367, eaaw3381 (2020).
Wagner, D. E. & Klein, A. M. Lineage tracing meets single-cell omics: opportunities and challenges. Nat. Rev. Genet. 21, 410–427 (2020).
Chen, L. et al. A reinforcing HNF4-SMAD4 feed-forward module stabilizes enterocyte identity. Nat. Genet. 51, 777–785 (2019).
Ayyaz, A. et al. Single-cell transcriptomes of the regenerating intestine reveal a revival stem cell. Nature 569, 121–125 (2019).
Tsuchida, K. et al. Discovery of nonpeptidic small-molecule AP-1 inhibitors: lead hopping based on a three-dimensional pharmacophore model. J. Med. Chem. 49, 80–91 (2006).
Vierbuchen, T. et al. AP-1 transcription factors and the BAF complex mediate signal-dependent enhancer selection. Mol. Cell 68, 1067–1082 (2017).
Hu, Y. et al. Multiscale footprints reveal the organization of cis-regulatory elements. Nature 638, 779–786 (2025).
Zu, G. et al. The transcription factor FoxM1 activates Nurr1 to promote intestinal regeneration after ischemia/reperfusion injury. Exp. Mol. Med. 51, 1–12 (2019).
Laissue, P. The forkhead-box family of transcription factors: key molecular players in colorectal cancer pathogenesis. Mol. Cancer 18, 5 (2019).
Xu, C. et al. Systematic dissection of sequence features affecting binding specificity of a pioneer factor reveals binding synergy between FOXA1 and AP-1. Mol. Cell 84, 2838–2855 (2024).
Maitra, R. et al. Development and characterization of a genetic mouse model of KRAS mutated colorectal cancer. Int. J. Mol. Sci. 20, 5677 (2019).
Imada, S. et al. Short-term post-fast refeeding enhances intestinal stemness via polyamines. Nature 633, 895–904 (2024).
Tanaka, T. et al. A novel inflammation-related mouse colon carcinogenesis model induced by azoxymethane and dextran sodium sulfate. Cancer Sci. 94, 965–973 (2003).
Tanaka, T. et al. Dextran sodium sulfate strongly promotes colorectal carcinogenesis in Apc(Min/+) mice: inflammatory stimuli by dextran sodium sulfate results in development of multiple colonic neoplasms. Int. J. Cancer 118, 25–34 (2006).
Li, J. et al. AP-1 mediates cellular adaptation and memory formation. Nat. Commun. https://doi.org/10.1038/s41467-026-70862-w (2026).
Zhou, R. W., Harpaz, N., Itzkowitz, S. H. & Parsons, R. E. Molecular mechanisms in colitis-associated colorectal cancer. Oncogenesis 12, 48 (2023).
Olafsson, S. et al. Somatic evolution in non-neoplastic IBD-affected colon. Cell 182, 672–684 (2020).
Siegel, R. L. et al. Global patterns and trends in colorectal cancer incidence in young adults. Gut 68, 2179–2185 (2019).
Zeve, D. et al. Robust differentiation of human enteroendocrine cells from intestinal stem cells. Nat. Commun. 13, 261 (2022).
Remke, M. et al. Histomorphological scoring of murine colitis models: a practical guide for the evaluation of colitis and colitis-associated cancer. Exp. Mol. Pathol. 140, 104938 (2024).
Langmead, B., Wilks, C., Antonescu, V. & Charles, R. Scaling read aligners to hundreds of threads on general-purpose processors. Bioinformatics 35, 421–432 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in unique molecular identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Gayoso, A. et al. A Python library for probabilistic analysis of single-cell omics data. Nat. Biotechnol. 40, 163–166 (2022).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Lareau, C. A. et al. Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility. Nat. Biotechnol. 37, 916–924 (2019).
Bravo González-Blas, C. et al. cisTopic: cis-regulatory topic modeling on single-cell ATAC-seq data. Nat. Methods 16, 397–400 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Mi, H., Muruganujan, A., Casagrande, J. T. & Thomas, P. D. Large-scale gene function analysis with the PANTHER classification system. Nat. Protoc. 8, 1551–1566 (2013).
Weirauch, M. T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Nagaraja, S. Colitis memory release 2026-02-06 v2. Zenodo https://doi.org/10.5281/zenodo.18487777 (2026).
Acknowledgements
We are grateful to the Buenrostro Laboratory for helpful feedback throughout this body of work. We thank K. McKinley for advice and support throughout this project. We also thank the Harvard Stem Cell & Regenerative Biology Histology Core (C. MacGillivray and D. Faria) for tissue processing and staining and the Broad Spatial Technology Platform (M. Lipinski, T. Aleksanyan and S. L. Farhi) for performing Visium HD. Finally, we thank M. Greenberg for the generous donation of the FOS antibody for CUT&Tag experiments. We acknowledge support by grants from the NHGRI Impact of Genomic Variation on Function consortium (grant no. UM1 HG011986), Harvard Stem Cell Institute (grant no. BA-0002-21-00) and NIDDK (grant no. P30DK034854). This work was delivered as part of the PROSPECT team supported by the Cancer Grand Challenges partnership funded by Cancer Research UK (grant no. CGCATF-2023/100043), the National Cancer Institute (grant nos. 1OT2CA297577-01, 3OT2CA297577-01S1), the French National Cancer Institute and the Bowelbabe Fund for Cancer Research UK. Supplementary Fig. 4 was created in BioRender; Nagaraja, S. https://biorender.com/tra3y6g (2026).
Author information
Authors and Affiliations
Contributions
S.N. and J.D.B. conceived the project and designed experiments. S.N., L.O.-M. and E.O. performed in vivo animal work and ex vivo mouse organoid experiments. S.N., L.O.-M., R.Z. and Y.H. performed data analysis. C.H. performed in vitro TF binding assays and associated data analysis. Q.Z. performed clonogenicity experiments with support of O.H.Y. D.Z. and K.S. performed human organoid work with support of D.T.B. R.R.H. and A.C. assisted with animal care and experimentation. J.D.B. supervised all aspects of the study. S.N. and J.D.B. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
J.D.B. holds patents related to ATAC-seq, is on the scientific advisory board for Camp4 and seqWell, is a co-founder of Switchpoint Bio, and is a consultant at the Treehouse Family Foundation. O.H.Y. holds equity in Jumbl therapeutics and Ava Bioscience and consults for Nestle and Prescrypt therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature thanks Eduard Batlle and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Tissue and organismal recovery following colitis.
a, Mouse weights, normalized to day 1 weight, through chronic colitis and recovery. Each gray line represents an individual animal and the black line shows the average across all animals. b, Colon swiss rolls of mice 21 days following three cycles of 1.25% DSS (Recovery timepoint) and healthy controls. Scale bars 1 mm. c, H&E images of distal colon immune infiltration in healthy controls, 21 days following three cycles of 1.25% DSS, 20 days following one cycle of 2% DSS, and 20 days following one cycle of 2.5% DSS. Scale bars 200 μm. For b and c, representative of n = 3 control and 1.25% DSS mice, n = 2 2% and 2.5% mice. d, Colitis grading of distal colon immune infiltration per Remke et al.59 (n = 3 control, 2 2.5%-DSS, 2 2%-DSS, 3 1.25%-DSS mice). e, CD45 immunohistochemistry in the distal colon 21 days following three cycles of 1.25% DSS. f, Quantification of CD45 cell density in the mucosa and submucosa of the distal colon at 21 days of recovery. n = 3 mice per group. All error bars are s.e.m. Significance values are from two-sided t-tests.
Extended Data Fig. 2 Quality control and identification of cell types.
a, TSS enrichment of scATAC-seq data. The X-axis represents distance from the center of all transcription start sites and the Y-axis shows signal normalized to -2 kb. b, Density plot of library size vs FRIP for all scATAC-seq barcodes. Barcodes passing filter indicate number before matching with scRNA-seq data. c, Estimated library size per cell by sample for scRNA-seq data. d, Number of unique genes detected per cell for scRNA-seq data. For c and d, n = 50,723 control, 19,698 acute, 25,128 chronic and 16,328 recovered cells. e, UMAP embedding of scATAC-seq data colored by cluster. f, UMAP embedding of scATAC-seq and scRNA-seq data colored by disease stage. g, Key marker gene expression shown on scATAC-seq UMAP embedding. Relative expression was computed for each gene independently. h, Marker gene expression for each cell type identified within the absorptive lineage. For each gene, the size of the circle indicates the fraction of cells in which at least one transcript was detected and the color of the circle indicates average expression within the cell type. i, Absorptive epithelial cell type distribution between colitis recovered and control mice. Each circle represents one animal. All error bars are s.e.m. Significance values are from two-sided t-tests. For box-whisker plots, the center line represents median, the box upper and lower quartiles and the whiskers 1.5 x IQR.
Extended Data Fig. 3 Cellular memory following chronic colitis.
a, Gene ontology for genes upregulated in stem cells at acute or chronic injury. b, Schematic for computing neighborhood enrichment by sample. c, scATAC-seq UMAP embedding showing enrichment for each colitis stage in k-NN networks, with each point representing a single cell colored by the enrichment of its k-NN network for a given stage. d, scRNA-seq UMAP embedding showing enrichment for each colitis stage in k-NN networks, with each point representing a single cell colored by the enrichment of its k-NN network for a given stage. e, Change in motif accessibility in stem cells at each stage of colitis. The size of the circle represents the -log10(FDR) and the color represents the magnitude of change in motif score. f, Left, change in mean motif accessibility relative to controls across colitis progression in differentiated epithelial cells. Right, barplot demonstrating change relative to control in motif accessibility, where each point represents mean change across all cells of that type within each animal. n = 9 control, 4 acute, 5 chronic and 5 recovered mice. g, Single cell motif accessibility for select motif families. n = 500 cells across all animals at each stage. P-value from two-sided Wilcoxon rank-sum test. h, Top, AP-1 motif accessibility scores plotted on stem/progenitor UMAP re-embedding. Bottom, identification of high AP-1 cells (motif score > 1.5) identified in Fig. 1. i, Differential gene expression between high AP-1 cells and all others. The X-axis represents log2(fold change) between AP-1 high cells and all others, and the Y-axis represents FDR following two-sided Wilcoxon rank sum test. j, Differential motif accessibility between high AP-1 cells and all others. The X-axis represents change in motif score between AP-1 high cells and all others, and the Y-axis represents FDR following two-sided Wilcoxon rank sum test. k, Violin plots of single cell motif accessibility values for AP-1 high and all other stem/progenitor cells. l, Change in Fos+ percentage across extended recovery, where each point represents one animal. control n = 3, acute n = 2, chronic n = 2, early recovery n = 3, 50-day recovery n = 3, 79-day recovery n = 2, and 102-day recovery n = 4 mice. m, Change in AP-1 motif accessibility across extended recovery, where each point represents average across all cells belonging to one animal. control n = 14, acute n = 4, chronic n = 5, early recovery n = 5, 50-day recovery n = 2, 79-day recovery n = 2, 102-day recovery n = 4 mice. All error bars are s.e.m.
Extended Data Fig. 4 Clonal inheritance of epigenetic states.
a, Fluorescent in situ hybridization against GFP for clonal barcode transcripts in HEK293T cells. Top, cells infected with original LARRY barcode from Weinreb et al.40 Bottom, cells infected with LARRY barcode modified by addition of mU1 hairpin sequence. Scale bar 50 μm. Representative of n = 100 cells across 2 independent experiments. b, Schematic detailing molecular process of capturing clonal barcodes and computational workflow for assigning cells to clones. c, HNF4/PPAR motif accessibility amongst the 50 largest clones by cell number. Each violin plot represents distribution of motif scores for an individual clone and point represents median values per clone. d, Framework for computing clonal variance and identifying clonal features. e, Motif accessibility relative to position in the colon. The X-axis represents segment of the colon, with segment 1 being most proximal and segment 6 being most distal. The Y-axis shows change in motif accessibility of a given segment relative to segment 1. f, Accessibility of all TF motif families relative to position in the colon. Each row represents a TF motif family and color indicates change in motif accessibility relative to segment 1. g, Expression stemness (Lgr5, Lrig1) and differentiation (Car1) markers in relation to HNF4/PPAR motif score. h, Downsampling analysis of permutation-based clonal heritability. Clones were randomly sampled to the number indicated on the x-axis and permutation-based testing was performed identically to the complete dataset. The total number of motif families identified to have lower clonal variance as compared to random (FDR < 0.05) for each clone number is shown on the y-axis. i, Linear mixed model of random effects from exposure to colitis and clonal identity. j, Variance explained by clonal identity from the linear mixed model. k, Comparison of variance explained by clonal identity and exposure to colitis for all TF motif families. l, Left, variance explained by clonal identity and exposure to colitis for top TF motif families. Right, variance attributed to each factor following randomization of clonal identities within colitis or control conditions. m, Comparison of clonality identified by permutation and linear mixed model. The X-axis shows -log10(FDR) from the permutation method (Fig. 2h) and the Y-axis shows percent variance attributed to clonal identity in the linear mixed model (panel j). Panels created in BioRender: e, Nagaraja, S. https://biorender.com/jqhj3wt (2026); i, Nagaraja, S. https://biorender.com/4mut8sd (2026).
Extended Data Fig. 5 Gene expression programs in colitis memory.
a, Schematic for deriving gene expression programs. b, Clonal memory of gene expression programs. The X-axis represents the difference between observed clonal variance and random variance following clonal label permutation and the Y-axis represents significance of this difference described in Extended Data Fig. 6d. c, Spearman correlation of mean AP-1 motif score per clone and mean P20 gene program score by clone. d, Individual gene contributions to program 20. The Y-axis represents the weight assigned by modified cisTopic model and the top 150 genes colored in red were used for rescoring other datasets. e, Relative gene expression values for the top 150 P20 genes across all organoid clones. The side bar chart shows log2(fold change) of colitis organoid clones over controls for each gene. f, Gene ontology biological programs for HNF4/PPAR related genes (programs 9 and 30). g, UMAP embeddings showing motif accessibility scores (top) and gene program expression scores (bottom). h, P20 gene expression scores through extended recovery. The Y-axis shows mean change in gene expression program score over control in stem cells. i, Change in mean P20 gene expression program score per animal. For h and i, control n = 14, acute n = 4, chronic n = 5, early recovery n = 5, 50-day recovery n = 2, 79-day recovery n = 2, 102-day recovery n = 4 mice. j, Gene expression (left) and chromatin accessibility loci (right) at P20 genes Adgrl3 and Pvt1 (n = 9 control, 4 acute, 5 chronic and 5 recovered mice). k, Change in mean expression of individual P20 genes over control across disease progression. n = 9 control, 4 acute, 5 chronic and 5 recovered mice. l, Comparison of P20 gene expression change between chronic injury and 50 day recovery. Each point represents a P20 gene and the axis value represents a signed log10(p-value) from a one-sided test between chronic injury (Y-axis) or 50-day recovery (X-axis), with negative values representing reduction in expression. All error bars are s.e.m. Significance values are from two-sided t-tests unless otherwise indicated. Panels created in BioRender: a, Nagaraja, S. https://biorender.com/2ju935u (2026); h, Nagaraja, S. https://biorender.com/865k9jb (2026).
Extended Data Fig. 6 Methylation changes in epigenetic memory of colitis.
a, Differential methylation between colitis and control organoids. The X-axis represents change in methylation percent and the Y-axis represents -log10(FDR) following two-sided t-test. b, Genome tracks at P20 gene Mecom, where the x-axis shows genomic position. Top, ATAC-seq chromatin accessibility data in control and colitis organoids, where the y-axis shows normalized insertions. Bottom, methylation in the selected region, with 3 organoids lines per condition as rows and color indicating percent methylation at CpGs. c, Schematic for AP-1 inhibition in vivo. d, Change in methylation at top 500 most variable AP-1 motif sites and matched background set of peaks. Left, change in methylation following recovery from colitis relative to controls, both DMSO treated. Right, change in methylation following AP-1 inhibition relative to DMSO, both following recovery from colitis. n = 500 top variable AP-1 peaks with AP-1 motifs and n = 36,367 matched background peaks. P-values from two-sided Wilcoxon rank-sum tests. e, Genome tracks at genes Dcaf5 and Rbm27, where the x-axis shows genomic position. Top, ATAC-seq chromatin accessibility data in control and recovered cells (from Fig. 1), where the y-axis shows normalized insertions. Bottom, whole tissue methylation in the selected region, with 3 mice per condition as rows and color indicating percent methylation at CpGs. For box-whisker plots, the center line represents median, the box upper and lower quartiles and the whiskers 1.5 x IQR. Panel c created in BioRender; Nagaraja, S. https://biorender.com/tra3y6g (2026).
Extended Data Fig. 7 Transcription factor networks in colitis memory.
a, Examples of de novo derived motifs. The y-axis on motif sequences shows information content. Line plots showing mean change in motif accessibility in stem cells through colitis progression as compared to control. b, Examples of de novo derived AP-1 composite motifs. The y-axis on motif sequences show information content. Line plots show mean change in motif accessibility in stem cells through colitis progression over controls. c, Left, mean change in chromatin accessibility at HNF4/PPAR composite motif sites through colitis progression over control. Right, predicted effect of motif presence on footprint scores. The X-axis represents distance from motif, the Y-axis represents size of footprint and color indicates predicted change of footprint score when motif is added. For a-c, n = 9 control, 4 acute, 5 chronic and 5 recovered mice. d, Comparison of change footprint score between colitis and control organoids following 24 h of AP-1 inhibition (T-5224, 10 μM). Each axis represents the fraction of footprints showing a reduction in footprint score of at least 0.2. e, Genome tracks showing high resolution footprinting in epithelial cells. Horizontal position represents genomic location and vertical height represents either ATAC-seq accessibility (top) or footprint score (bottom). f, Genome tracks of a Fos memory site showing footprint score (top) and respective Fos CUT&Tag (bottom). g, Fos CUT&Tag signal in control and recovered tissue at the top 10,000 variable sites following normalization to read depth. h, Differential footprint score performance in predicting CUT&Tag signal. The x-axis represents varying thresholds for difference in AP-1 footprint score and the y-axis shows sensitivity and specificity for detecting Fos CUT&Tag signal difference between control and recovered tissue. i, Comparison of AP-1 footprint score difference and Fos CUT&Tag signal difference at all identified Fos binding sites. Red points represent sites called as different by footprint score difference of 0.2. j, Co-binding scores with AP-1 factors in primary epithelial tissue following colitis recovery. Co-binding score quantifies the enrichment of concurrent footprint gain for a given TF family with AP-1 over chance. k, TF family co-binding change across colitis progression. l, Distance from each TF family memory footprint (gain of at least 0.2 following recovery) to the nearest AP-1 memory footprint. All error bars are s.e.m. Significance values are from two-sided t-tests unless otherwise indicated. Panel j created in BioRender; Nagaraja, S. https://biorender.com/jqhj3wt (2026).
Extended Data Fig. 8 In vitro binding relationships between AP-1 and FOX transcription factors.
a, Gene expression of FOX family transcription factors in all stem cells in primary tissue (n = 23 mice). b, Multi-scale footprinting at sample genomic loci. Top, expanded copy of multiscale plot below. The X-axis represents distance in base pairs from the AP-1 motif center and the Y-axis represents radius of footprint being evaluated. The color represents -log10(p-value) from a one-sided binomial test of the predicted footprint at given radius and genomic position. For all in vitro binding scores, a 14 bp footprint radius was used. c, Genomic tracks of in vivo tissue footprint score (top), Tn5 insertions relative to naked DNA alone (middle), and in vitro binding score for given TF combinations. d, All loci and combinations of TFs tested by in vitro binding assay. Columns are positions relative to AP-1 motif site center and rows are each individual genomic locus. Color represents in vitro binding score calculation of -log10(p-value) at 14 bp radius. e, Average in vitro binding scores across all loci in panel (d). f, AlphaFold predicted structures for Fos-Jun dimer, composite motif and either Foxa1, Foxn2 or Foxj2. g, UniProt domain annotation for FOX TFs with black boxes indicating regions predicted to interact with Fos/Jun heterodimer. Numbers indicate amino acid positions. DBD = DNA binding domain, ZF = Zinc finger, LZ = Leucine zipper. h, Predicted interactions with Foxp1 and Jun alone. All error bars are s.e.m.
Extended Data Fig. 9 Features of adenomas following colitis.
a, Distribution of macroscopic adenoma size between control and colitis recovered animals. Each point represents a single tumor. P-value from K-S test. b, Proliferation in adenomas following high dose tamoxifen induction (50 mg/kg x 3). n = 3 mice per condition. c, Clonogenicity in adenoma-derived organoid lines. n = 3 control and 4 colitis recovered organoid lines. d, Visible adenoma number following high dose tamoxifen induction. n = 4 control and 5 colitis recovered mice. e, Representative example of microscopic tumor following low dose tamoxifen induction (10 mg/kg x 1). Scale bar 100 μm. f, Method for computing tumor area. Left, low magnification image used to compute total tissue area. Middle, high magnification image where individual tumors are individually traced. Right, equation for computing percent tumor area. g, Quantification of microscopic tumor area following low dose initiation. Each point represents the total area per animal as a percentage of total tissue assessed. n = 7 control mice and 10 colitis recovered mice. h, Microscopic tumor number normalized to total tissue area. n = 7 control mice and 10 colitis recovered mice. i, UMAP embedding of scATAC-seq data of primary tissue following adenoma induction with cells colored by cluster. j, UMAP embeddings showing Lef1 motif accessibility (top), Axin2 relative gene expression (middle), and sample condition (bottom). k, Marker gene expression for each epithelial cell type identified. For each gene, the size of the circle indicates the fraction of cells in which at least one transcript was detected and the color indicates average expression within the cell type. l, Beta-catenin immunohistochemistry images of microscopic tumors following in vivo AP-1 inhibition. Circles indicate examples of lesions. Scale bar 100 μm. Representative of n = 6 control vehicle-treated, 7 colitis recovered vehicle-treated and 7 colitis recovered T-5224-treated mice. All error bars are s.e.m.
Extended Data Fig. 10 Spatial transcriptomics of adenomas.
a, Axin2 based tumor cell calls. Top left, H&E of adjacent section. Scale bar 2 mm. Top right, spatial expression of Axin2. Bottom, boxed regions showing alignment of H&E morphology, Axin2 expression and individual tumor cell calls. Scale bar 50 μm. b, Left, gene expression heatmap of top 1000 most variable genes across all 254 tumors. Each row is a gene and each column is an individual tumor. Right, average expression in adenomas of select genes (n = 117 control tumors, n = 137 colitis recovered tumors). P-values from a two-sided test from DESeq2. The Y-axis represents the average reads per million expression across all tumors. c, Gene expression heatmap of P20 genes across all 254 tumors. Barplot represents log2(fold change) in colitis recovered tumors relative to controls. d, Spatial expression of AP-1/P20 associated genes in identified tumor cells. Scale bars 400 μm. e, Differential gene expression between tumors with high AP-1/P20 associated gene expression (score > 3) and tumors with low expression. The x-axis represents the log2(fold change) of high AP-1 tumors over all others and the y-axis represents significance following DESeq2 testing. f, Biological processes associated with genes upregulated in high AP-1 tumors.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–5, Figs. 1–4 and References. This file contains notes, relevant references and legends for data.
Supplementary Data 1 (download PDF )
De novo identified motifs by seq2PRINT All motifs identified de novo by seq2PRINT trained on in vivo SHARE-seq data (control, acute injury, chronic injury and early recovery), with predicted partial or full matches of known motifs.
Supplementary Data 2 (download TXT )
Single-cell motif scores and clone calls. Motif scores, clone calls and colitis exposure conditions by cell.
Supplementary Tables (download ZIP )
Supplementary Tables 1–6.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nagaraja, S., Ojeda-Miron, L., Zhang, R. et al. Epigenetic memory of colitis promotes tumour growth. Nature (2026). https://doi.org/10.1038/s41586-026-10258-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10258-4