Abstract
Cell competition and fitness comparison between cancer and tumor microenvironment (TME) cells determine oncogenic fate. Our previous study established a role for human Flower isoforms as fitness fingerprints, where the expression of Flower Win isoforms in tumor cells leads to growth advantage over TME cells expressing Lose isoforms. Here we demonstrate that the expression of Flower Lose and reduced microenvironment fitness is not a pre-existing condition but, rather, a cancer-induced phenomenon. Cancer cells actively reduce TME fitness by the exosome-mediated release of a cancer-specific long non-coding RNA, Tu-Stroma, which controls the splicing of the Flower gene in the TME cells and expression of Flower Lose isoform, which leads to reduced fitness status. This mechanism controls cancer growth, metastasis and host survival in ovarian cancer. Targeting Flower protein with humanized monoclonal antibody (mAb) in mice significantly reduces cancer growth and metastasis and improves survival. Pre-treatment with Flower mAb protects intraperitoneal organs from developing lesions despite the presence of aggressive tumor cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in the paper, its extended data and its supplementary information. All source data have been made available. The LNCipedia version 5.2 FASTA database is available at https://lncipedia.org/download. PhyloCSF scores were calculated using PhyloCSF score software (https://github.com/mlin/PhyloCSF/wiki). The Coding-Potential Assessment Tool (CPAT) was used based on code available at https://code.google.com/archive/p/cpat/. Tu-Stroma secondary structure was predicted using RNAfold server software, available at http://rna.tbi.univie.ac.at/. Alu sequence analysis was performed using RepeatMasker Web Server, available at http://www.repeatmasker.org/cgi-bin/WEBRepeatMasker. RBPsuite was used to identify putative RNA–protein interactions (http://www.csbio.sjtu.edu.cn/bioinf/RBPsuite/). DNA conservation score was calculated using Phastcon score, downloaded from the UCSC Genome Browser (ftp://hgdownload.cse.ucsc.edu/goldenPath/mm9/phastCons30way/vertebrate). Putative TFBS analysis was performed using MatInspector (Genomatix) with MatBase matrix library 8.0. MatInspector is available online at https://pmc.ncbi.nlm.nih.gov/articles/PMC9584888/. Source data are provided with this paper.
References
Merino, M. M. et al. Elimination of unfit cells maintains tissue health and prolongs lifespan. Cell 160, 461–476 (2015).
Merino, M. M., Rhiner, C., Portela, M. & Moreno, E. ‘Fitness fingerprints’ mediate physiological culling of unwanted neurons in Drosophila. Curr. Biol. 23, 1300–1309 (2013).
Rhiner, C. et al. Flower forms an extracellular code that reveals the fitness of a cell to its neighbors in Drosophila. Dev. Cell 18, 985–998 (2010).
Madan, E. et al. Flower isoforms promote competitive growth in cancer. Nature 572, 260–264 (2019).
Yekelchyk, M. et al. Flower lose, a cell fitness marker, predicts COVID-19 prognosis. EMBO Mol. Med. 13, e13714 (2021).
Merino, M. M., Levayer, R. & Moreno, E. Survival of the fittest: essential roles of cell competition in development, aging, and cancer. Trends Cell Biol. 26, 776–788 (2016).
Zhang, R. et al. Winner-takes-all resource competition redirects cascading cell fate transitions. Nat. Commun. 12, 853 (2021).
Roulis, M. et al. Paracrine orchestration of intestinal tumorigenesis by a mesenchymal niche. Nature 580, 524–529 (2020).
Flanagan, D. J. et al. NOTUM from Apc-mutant cells biases clonal competition to initiate cancer. Nature 594, 430–435 (2021).
van Neerven, S. M. et al. Apc-mutant cells act as supercompetitors in intestinal tumour initiation. Nature 594, 436–441 (2021).
Moya, I. M. et al. Peritumoral activation of the Hippo pathway effectors YAP and TAZ suppresses liver cancer in mice. Science 366, 1029–1034 (2019).
Hill, W. et al. EPHA2-dependent outcompetition of KRASG12D mutant cells by wild-type neighbors in the adult pancreas. Curr. Biol. 31, 2550–2560 (2021).
Kohashi, K. et al. Sequential oncogenic mutations influence cell competition. Curr. Biol. 31, 3984–3995 (2021).
Colom, B. et al. Mutant clones in normal epithelium outcompete and eliminate emerging tumours. Nature 598, 510–514 (2021).
Ayukawa, S. et al. Epithelial cells remove precancerous cells by cell competition via MHC class I–LILRB3 interaction. Nat. Immunol. 22, 1391–1402 (2021).
Fernandez-Antoran, D. et al. Outcompeting p53-mutant cells in the normal esophagus by redox manipulation. Cell Stem Cell 25, 329–341 (2019).
Liu, Z. et al. Differential YAP expression in glioma cells induces cell competition and promotes tumorigenesis. J. Cell Sci. 132, jcs225714 (2019).
Krotenberg Garcia, A. et al. Active elimination of intestinal cells drives oncogenic growth in organoids. Cell Rep. 36, 109307 (2021).
Yum, M. K. et al. Tracing oncogene-driven remodelling of the intestinal stem cell niche. Nature 594, 442–447 (2021).
Ball, M. P. et al. Targeted and genome-scale strategies reveal gene-body methylation signatures in human cells. Nat. Biotechnol. 27, 361–368 (2009).
Laurent, L. et al. Dynamic changes in the human methylome during differentiation. Genome Res. 20, 320–331 (2010).
Lev Maor, G., Yearim, A. & Ast, G. The alternative role of DNA methylation in splicing regulation. Trends Genet. 31, 274–280 (2015).
Rauch, T. A., Wu, X., Zhong, X., Riggs, A. D. & Pfeifer, G. P. A human B cell methylome at 100-base pair resolution. Proc. Natl Acad. Sci. USA 106, 671–678 (2009).
Shayevitch, R., Askayo, D., Keydar, I. & Ast, G. The importance of DNA methylation of exons on alternative splicing. RNA 24, 1351–1362 (2018).
Yearim, A. et al. HP1 is involved in regulating the global impact of DNA methylation on alternative splicing. Cell Rep. 10, 1122–1134 (2015).
Eads, C. A. et al. MethyLight: a high-throughput assay to measure DNA methylation. Nucleic Acids Res. 28, E32 (2000).
Pan, X. et al. Massively targeted evaluation of therapeutic CRISPR off-targets in cells. Nat. Commun. 13, 4049 (2022).
Lin, L. et al. Genome-wide determination of on-target and off-target characteristics for RNA-guided DNA methylation by dCas9 methyltransferases. Gigascience 7, 1–19 (2018).
Misumi, Y. et al. Novel blockade by brefeldin A of intracellular transport of secretory proteins in cultured rat hepatocytes. J. Biol. Chem. 261, 11398–11403 (1986).
Mondal, T. et al. MEG3 long noncoding RNA regulates the TGF-β pathway genes through formation of RNA–DNA triplex structures. Nat. Commun. 6, 7743 (2015).
Pandey, R. R. et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 32, 232–246 (2008).
Ostrowski, M. et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat. Cell Biol. 12, 19–30 (2010).
Coughlan, C. et al. Exosome isolation by ultracentrifugation and precipitation and techniques for downstream analyses. Curr. Protoc. Cell Biol. 88, e110 (2020).
Parisse, P. et al. Atomic force microscopy analysis of extracellular vesicles. Eur. Biophys. J. 46, 813–820 (2017).
Statello, L. et al. Identification of RNA-binding proteins in exosomes capable of interacting with different types of RNA: RBP-facilitated transport of RNAs into exosomes. PLoS ONE 13, e0195969 (2018).
Li, J., Tian, T. & Zhou, X. The role of exosomal shuttle RNA (esRNA) in lymphoma. Crit. Rev. Oncol. Hematol. 137, 27–34 (2019).
Calviello, L. et al. DDX3 depletion represses translation of mRNAs with complex 5′ UTRs. Nucleic Acids Res. 49, 5336–5350 (2021).
Welch, J. D., Hartemink, A. J. & Prins, J. F. MATCHER: manifold alignment reveals correspondence between single cell transcriptome and epigenome dynamics. Genome Biol. 18, 138 (2017).
Cartharius, K. et al. MatInspector and beyond: promoter analysis based on transcription factor binding sites. Bioinformatics 21, 2933–2942 (2005).
Armstrong, D. K. et al. NCCN Guidelines® Insights: Ovarian Cancer, Version 3.2022. J. Natl Compr. Canc. Netw. 20, 972–980 (2022).
Moisan, F. et al. Enhancement of paclitaxel and carboplatin therapies by CCL2 blockade in ovarian cancers. Mol. Oncol. 8, 1231–1239 (2014).
Burger, R. A. et al. Incorporation of bevacizumab in the primary treatment of ovarian cancer. N. Engl. J. Med. 365, 2473–2483 (2011).
Chung, Y. S. et al. Outcomes of non-high grade serous carcinoma after neoadjuvant chemotherapy for advanced-stage ovarian cancer: a Korean gynecologic oncology group study (OV 1708). BMC Cancer 19, 341 (2019).
Lisio, M. A., Fu, L., Goyeneche, A., Gao, Z. H. & Telleria, C. High-grade serous ovarian cancer: basic sciences, clinical and therapeutic standpoints. Int. J. Mol. Sci. 20, 952 (2019).
Millstein, J. et al. Prognostic gene expression signature for high-grade serous ovarian cancer. Ann. Oncol. 31, 1240–1250 (2020).
Kamelgarn, M. et al. Proteomic analysis of FUS interacting proteins provides insights into FUS function and its role in ALS. Biochim. Biophys. Acta 1862, 2004–2014 (2016).
Peinado, H. et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 18, 883–891 (2012).
Wang, P. et al. The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science 344, 310–313 (2014).
Zubeldia-Plazaola, A. et al. Comparison of methods for the isolation of human breast epithelial and myoepithelial cells. Front. Cell Dev. Biol. 3, 32 (2015).
Broner, E. C., Onallah, H., Tavor Re’em, T., Davidson, B. & Reich, R. Role of the exosome secretion machinery in ovarian carcinoma: in vitro and in vivo models. J. Oncol. 2020, 4291827 (2020).
Sun, Z., Nyberg, R., Wu, Y., Bernard, B. & Redmond, W. L. Developing an enhanced 7-color multiplex IHC protocol to dissect immune infiltration in human cancers. PLoS ONE 16, e0247238 (2021).
Mayer, A. & Churchman, L. S. A detailed protocol for subcellular RNA sequencing (subRNA-seq). Curr. Protoc. Mol. Biol. 120, 4.29.1–4.29.18 (2017).
Reimer, K. A. & Neugebauer, K. M. Preparation of mammalian nascent RNA for long read sequencing. Curr. Protoc. Mol. Biol. 133, e128 (2020).
Gogna, R., Madan, E., Kuppusamy, P. & Pati, U. Reactive oxygen species-mediated p53 core-domain modifications determine apoptotic or necrotic death in cancer cells. Antioxid. Redox Signal. 16, 400–412 (2012).
Gogna, R., Madan, E., Khan, M., Pati, U. & Kuppusamy, P. p53’s choice of myocardial death or survival: oxygen protects infarct myocardium by recruiting p53 on NOS3 promoter through regulation of p53-Lys118 acetylation. EMBO Mol. Med. 5, 1662–1683 (2013).
Madan, E. et al. The curcumin analog HO-3867 selectively kills cancer cells by converting mutant p53 protein to transcriptionally active wildtype p53. J. Biol. Chem. 293, 4262–4276 (2018).
Lin, M. F., Jungreis, I. & Kellis, M. PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics 27, i275–i282 (2011).
Wang, L. et al. CPAT: Coding-Potential Assessment Tool using an alignment-free logistic regression model. Nucleic Acids Res. 41, e74 (2013).
Kerpedjiev, P., Hammer, S. & Hofacker, I. L. Forna (force-directed RNA): simple and effective online RNA secondary structure diagrams. Bioinformatics 31, 3377–3379 (2015).
Wu, H., Pan, X., Yang, Y. & Shen, H. B. Recognizing binding sites of poorly characterized RNA-binding proteins on circular RNAs using attention Siamese network. Brief. Bioinform. 22, bbab279 (2021).
Martianov, I., Ramadass, A., Serra Barros, A., Chow, N. & Akoulitchev, A. Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature 445, 666–670 (2007).
Chu, C., Quinn, J. & Chang, H. Y. Chromatin isolation by RNA purification (ChIRP). J. Vis. Exp. https://doi.org/10.3791/3912 (2012).
Postepska-Igielska, A. et al. LncRNA Khps1 regulates expression of the proto-oncogene SPHK1 via triplex-mediated changes in chromatin structure. Mol. Cell 60, 626–636 (2015).
Madan, E. et al. HIF-transcribed p53 chaperones HIF-1α. Nucleic Acids Res. 47, 10212–10234 (2019).
Lagarde, J. et al. Extension of human lncRNA transcripts by RACE coupled with long-read high-throughput sequencing (RACE-Seq). Nat. Commun. 7, 12339 (2016).
Rosenberg, M. et al. Denaturing cross-linking immunoprecipitation to identify footprints for RNA-binding proteins. STAR Protoc. 2, 100819 (2021).
Davidson-Pilon, C. lifelines: survival analysis in Python. J. Open Source Softw. 4, 1317 (2019).
Ferber, S. et al. Co-targeting the tumor endothelium and P-selectin-expressing glioblastoma cells leads to a remarkable therapeutic outcome. eLife 6, e25281 (2017).
Landberg, G. et al. Patient-derived scaffolds uncover breast cancer promoting properties of the microenvironment. Biomaterials 235, 119705 (2020).
Koya, Y. et al. Murine experimental model of original tumor development and peritoneal metastasis via orthotopic inoculation with ovarian carcinoma cells. J. Vis. Exp. https://doi.org/10.3791/54353 (2016).
Nair, A. B. & Jacob, S. A simple practice guide for dose conversion between animals and human. J. Basic Clin. Pharm. 7, 27–31 (2016).
Hunter, M., Yuan, P., Vavilala, D. & Fox, M. Optimization of protein expression in mammalian cells. Curr. Protoc. Protein Sci. 95, e77 (2019).
R Core Team. R: A Language and Environment for Statistical Computing. https://www.r-project.org/ (R Foundation for Statistical Computing, 2020).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Angerer, P. et al. destiny: diffusion maps for large-scale single-cell data in R. Bioinformatics 32, 1241–1243 (2016).
Haghverdi, L., Buttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Grambsch, P. M. & Therneau, T. Modeling Survival Data: Extending the Cox Model (Springer, 2010).
Acknowledgements
We thank C. Beroud and P. Beroud for their expert input in writing and in generation of Supplementary Fig. 4b,c. This study was supported by VCU Massey Cancer Center and School of Medicine start-up funds, the Adenoid Cystic Carcinoma Research Foundation, the Swiss Cancer League, Seeds of Science, UAMS, SNSF, Fundação para a Ciência e a Tecnologia and Fundamental Mandates (Stichting tegen Kanker—Fondation contre le Cancer) to R.G.; VCU Massey Cancer Center and School of Medicine start-up funds and Tina’s Wish Foundation Award to E. Madan; FIS (Ministry of Health), Madrid, Spain, Grant PI17/01981 to A.L.-B.; ERC, SNSF, Josef Steiner Cancer Research Foundation, Swiss Cancer League and Champalimaud Foundation to E. Moreno; Novo Nordisk Foundation (NNF17CC0027852) and Lundbeck Foundation (R313–2019–421) to K.J.W.; support from the Thelma Newmeyer Corman Chair in Cancer Research, the VCU Commercialization Fund and NIH/NCI R01 CA259599 to P.B.F.; Fundação para a Ciência e a Tecnologia Grant 2020.05319.BD to A.M.P.; NIH Core Grant CA06927, the Dubrow Fund and the Bucks County Board of Associates to D. Connolly; and Fundação para a Ciência e a Tecnologia (FCT/MEC - PIDDAC) through projects UIDB/00100/2020 (CQE/IST), UIDP/00100/2020 and LA/P/0056/2020 to I.C. We would also like to acknowledge support from the University of Kansas Medical Center’s Biospecimen Repository Core Facility (BRCF) staff, including A. Webster, E. Johnson, L. DiMartino and S. Baker, for helping to identify, collect and process the human specimens; M. Kluthe for histology services; and R. Madan for pathology review. The BRCF was supported, in part, by the KU Cancer Center’s Support Grant (P30 CA168524) and the Kansas Institute for Precision Medicine (P20 GM130423 to A.K.G.). This study was supported, in part, by a grant from NIH National Cancer Institute (R01 CA260132 to A.K.G.) and the Kansas Institute for Precision Medicine (GM130423 to A.K.G.). Services and products in support of the research project were generated by the VCU Massey Comprehensive Cancer Center Proteomics Shared Resources, supported, in part, by funding from NIH-NCI Cancer Center Support Grant P30 CA016059.
Author information
Authors and Affiliations
Contributions
E. Madan performed experiments, analyzed the data, wrote the manuscript and provided funding and supervision. A.M.P., V.V., A.K. and P.B. performed experiments, prepared the figures, analyzed the data and wrote the manuscript. J.W. performed biostatistical analysis, prepared the figures, analyzed the data and wrote the manuscript. T.B. and D.A. performed microscopy experiments, analyzed and interpreted the data, prepared figures and wrote the manuscript. P.C.M., G.B., M.D.B., I.D., H.K. and D.O. performed biostatistical analysis, helped with interpretation of the data and prepared the figures. A. Gogna and A.B. helped with preparation of figures and live-cell and animation videos. M.L.P., M.Y., D.Z., S.P., S.K., K.B., P.M., I.C., E.A.F., A.K.P., S.M., C.J.P., A.L., M.N., S.C., K.G., K.C.G. and K.S.S. helped with collection of pathology tissue material and interpretation and analysis of pathology data, performed experiments and prepared the figures. T.F.A., E.G., A. Gustafsson, G.L., L.N., E.Y. and R.S.-F. helped with design and performance of 3D scaffold and organoid experiments, data analysis and interpretation. K.D.P.D. helped with PDX experiments and interpretation and analysis of data. N.M., M.J.P., K.J.-J. and S.H. helped with design and performance of exosome biology experiments, writing of the manuscript and data analysis and interpretation. Z.S. and W.L.R. helped with design and performance of multiplexing experiments and with data analysis and interpretation. M.B. and P.J. helped with design and with performance, interpretation and analysis of the FISH experiment. H.O.R., F.A.T. and R.B. helped with MR imaging and with interpretation and analysis of data. K.G.A., J.A., G.X., G.S. and S. Singh helped with design and production of the mAbs. S.D., C.L.d.S., A.M.F., D.T., S.R.G., K.P., G.G., G.L.W.G.R. and H.W. helped with editing of the manuscript. A.M.H., A.R. and C.L. helped with design and performance of mass spec analysis. A.S. and P.B.F. helped with writing of the manuscript and by providing laboratory resources, availability of postdoctoral researchers to collaborate and additional laboratory space. G.M.C., Y.L., A.L.O., L.L. and A.A.T. helped with design, performance, data analysis and interpretation of CRISPR-off experiments. A.S.P., U.M., R.D., A.K.G., K.S. and D. Chelmow helped with collection of surgery and pathology tissue material, helped with PDX models, provided expertise on ovarian cancer, helped with design of biomarker potential of the study and helped with design of therapy protocol for ovarian cancer. R.P., J.K., S.R., H.R., S. Sayeed, K.P.K., D. Connolly, K.P.N., M.O.I., M.P.K., C.M., M.A., A.L.-B., D.J., K.D.M., A. Tzankov, S.B., E.A.R. and R.C.M. helped with collection of surgery and pathology tissue material and helped with interpretation of pathology data. A. Tsung and D.S. helped with collection of surgery material, fostered clinical collaborations and helped with data analysis and interpretation. J.T. helped with mouse tumor models and data interpretation and analysis and provided funding and supervision. A.P.S., R.W., K.J.W. and E. Moreno helped with manuscript writing and data interpretation and analysis and provided funding and supervision. R.G. designed the study experiments, organized all collaborations, analyzed and compiled all the data, wrote the manuscript and provided funding and supervision.
Corresponding authors
Ethics declarations
Competing interests
All authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks George Calin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Flower exon 3 demonstrates increased enrichment of alternative splicing factors and epigenetic modifications.
A&B) Box plots show increased DNA methylation of the hFwe exon 3 (unmatched p = 3.2x10−3 and matched p = 1.1x10−3 normal vs. HGSC; unmatched and matched p < 1x10−10 HGSC vs. stroma), as well as an increased DNMT3A binding to the hFwe exon 3 (unmatched p = 5.8x10−7 and matched p = 1.9x10−4 normal vs. HGSC; p < 1x10−10 HGSC vs. stroma) in the stroma vs normal and HGSC tissue of patient derived samples. Scatter plots show a significant correlation between DNA Methylation of the hFwe exon 3 and hFwe-Lose expression (p = 4.9x10−4), as well as between DNMT3A binding to the hFwe exon 3 and hFwe- Lose expression (p = 2.5x10−6) in the stroma. In the box plots, the color scale represents log2 fold change of hFwe-Lose expression, while in the correlation analysis, the color delineates normal, tumor and stromal samples. (Biological replicates for normal tissue n=15, biological replicates for tumor and stroma tissue n=30). C) The control for qChIP experiment demonstrated in Extended Data Fig. 1A &B are presented. D) Heatmap of qChIP in stromal tissue from patient derived HGSC samples on Flower exon 3 shows association of 39 proteins which participate in DNA methylation induced RNA splicing via DNA methylation changes. H3K9Me3 (p = 1x10−5), HP1 (p = 1x10−4), SRSF3 (p = 5x10−5), GLP (p = 4x10−5) and G9a (p = 1x10−4) are the most abundant, (Biological replicates, n=5).
Extended Data Fig. 2 The enrichment of splicing machinery at Flower exon 3 is specific to HGSC stromal tissue.
A) A heatmap depicting relative enrichment from ChIP influenced gene walking experiment showing association of DNA methylation, G9a, GLP, H3K9Me3, HP1, SRSF3, and RNA Pol II, on each exon (1-6) and 200bp flanking regions upstream (5’) and downstream (3’) from each Exon of the hFwe gene locus. The bar representing the value ranges 0.3 and 4 (color yellow-green-blue) illustrates the relative binding/association of the epigenetic markers (left column) on the respective exons 1-6 and 200bp 5’ & 3’ flanking regions (Biological replicates, n=8). B) Methylight assay performed in all hFwe exon bodies in normal, HGSC tumor, and stromal samples showing specific and exclusive methylation of hFwe exon 3 in HGSC stromal tissue. Data are presented as mean values +/- SEM, (Biological replicates, n=15).
Extended Data Fig. 3 Biological assay using CRISPR-off at Flower exon 3 impacts Flower isoform expression.
A) A model depicts CRISPR-off strategy for hFwe exon 3. B) Agarose gel electrophoresis of Bi-sulfite-treated and untreated samples derived from WT stromal cells and two clones of CRISPR-off engineered stromal cells. Stromal cells of Clone 1 were co-transfected with CRISPR-off plasmids dCas9-KRAB, dCAS9-D3A-3L, sgRNA targeting hFwe exon 3. Stromal cells of Clone 2 were co- transfected with CRISPR-off plasmids dCas9-DNMT3A, dCAS9- DNMT3B, sgRNA targeting hFwe exon 3. To confirm DNA methylation, we performed Methylation- specific PCR in samples treated or untreated with bi- sulfite and run the products in agarose gels. C) Co-culture of WT tumor and stromal cells shows binding of LINC01914 to hFwe exon 3 DNA, high hFwe exon 3 DNA methylation, enrichment of splicing machinery and high expression of hFwe-Lose isoforms, in stromal cells. Co-culture of LINC01914 KO tumor cells and hFwe exon 3 CRISPR-off stromal cells show no binding of LINC01914 to hFwe exon 3, high DNA methylation, enrichment of splicing machinery at Flower exon 3, and high expression of Flower Lose.
Extended Data Fig. 4 Silencing of LINC01914 expression prevents Flower exon 3 methylation in stromal cells.
A) Model depicting the insertion of the Poly-A sequence within LINC01914 locus and respective homology arms. B) Model depicting the experiments performed in stromal cells after co-culture with OVCAR8 cells. Experiments related to LINC01914 deletion and rescue are performed in co-culture. For the control experiment, OVCAR8 LINC01914 WT and GFP-labeled fibroblasts were cultured for 96h. For the deletion of hFwe exon 3 binding site, we co-cultured GFP-labeled fibroblasts either with OVCAR8 LINC01914 KO or with OVCAR8 LINC01914 with Poly-A insertion. For the rescue experiments, we co-cultured GFP-labeled fibroblasts either with OVCAR8 LINC01914 KO overexpressing a WT LINC01914, a MT (mutated hFwe exon 3 binding site) LINC01914, or a LINC01914 deletion construct (missing exon 3 binding site). C) Methylation-specific PCR (MSP) of hFwe exon 3 in stromal cells co-cultured with OVCAR8 cells is shown. Each panel represents a specific co-culture experiment with and without bisulfite treatment (indicated at bottom of gel with +/- signs). Bisulfite treatment of stromal DNA shows that stromal cells have hFwe exon 3 methylation after co-culture with WT OVCAR8 cells (panel 1, lane 6). The methylation of hFwe exon 3 cannot be observed in stromal cells co-cultured with LINC01914 KO OVCAR8 cells since only un-methylated DNA specific primers show amplification of hFwe exon 3 after bisulfite treatment (panel 2, lane 6). The methylation of hFwe exon 3 in stromal cells is rescued when these cells are co-cultured with LINC01914 KO OVCAR8 cells overexpressing the WT LINC01914 (panel 3, lane 5). Stromal cells co-cultured with LINC01914 KO OVCAR8 cells overexpressing the MT LINC01914 are unable to rescue the WT methylation pattern. D) Analysis of hFwe isoform expression pattern upon treatment with the cytokine inhibitor Brefeldin A. OVCAR8 WT and GFP-labeled fibroblasts were co- cultured in media containing Brefeldin A for 96h (left). GFP-labeled fibroblasts were also cultured alone and treated with conditional media containing Brefeldin A (right). The gels show that treatment with Brefeldin A does not affect exon 3 skipping and hFwe-Lose expression in stromal cells. E) Model depicting ChOP to observe the DNA-RNA adduct between LINC01914 and hFwe exon 3.
Extended Data Fig. 5 LINC01914 binds to Flower at exon 3 through Hoogsteen base-pairing.
A) ChOP shows an aggregate association of LINC01914 to the exon 3 body of the hFwe gene loci. The experiment is conducted in primary cancer (labeled with GFP) and stromal cells extracted and co-cultured (for 8 days) from 8 HGSC patient samples. Sorted primary stromal cell fraction upon incubation with LINC01914 anti-sense oligos show association exclusively on the exon 3 body (blue peak) and not on any other exon body or the 5’ & 3’ flanking regions. The ChOP pull-down analyzed by qPCR is calculated as the percentage of input (p < 1x10-10). B) Model depicting the experimental confirmation of the formation of a DNA-RNA adduct between hFwe and Tu-Stroma mediated by Hoogsteen base-pairing. C&D) Validation of the physical association of LINC01914 and exon 3 DNA body (containing complementary GA-rich sequences) via Hoogsteen base-pairing is performed by in-vitro capture assay using biotinylated LINC01914. Data shows a peak (red) generated specifically on the exon 3 body, while incubation with 7-deaza-purine nucleotides (blue) abolishes the Hoogsteen base-pairing (p < 1x10−10). The interaction is further validated in the nuclei isolated from HGSC stromal cells (8 HGSC patients). A consistent and exclusive enrichment of peak (red) is primarily observed on the exon 3 body upon incubation of the stromal nuclei with the ectopic biotinylated LINC01914 by qPCR analysis (p = 6.2x10−7). E&F) The in-vivo capture assay shows the physical association and DNA-RNA adduct formation between LINC01914 and exon 3 DNA body in the primary stromal cells extracted from the HGSC patients. Results show an exclusive enrichment of peak (red) on the exon 3 DNA body and not on other Exons or the 5’ & 3’ flanking regions of hFwe gene, upon transfection of stromal cells with biotinylated LINC01914 (p < 1x10−10). Results show a significant adduct enrichment on exon 3 body (red) in the photo-activated nuclear extracts of the stromal transfected cells in comparison to the non-photo- activated nuclear extracts (blue) (p < 1x10−10). G) exon 3 dsDNA and LINC01914 ssRNA shown in blue and orange spectra respectively correspond as the controls. Theoretical mean spectra [(LINC01914+ exon 3)/2] shown in red is the sum of the individual CD spectra for exon 3 dsDNA and LINC01914 ssRNA. H) Data shows the location of the exon 3 DNA body (lane 1), and the location of single-stranded LINC01914 (lane 2). Lane 3 shows the formation of a DNA-RNA complex upon mixing; the bottom shows DNA, the middle shows RNA, and a DNA-RNA complex is observed as a supershift (top band marked with red arrow). In lane 4, the unbound single-stranded RNA is destroyed upon the addition of RNase H and only the DNA-RNA adduct band is visible at the top. I) Data shows the location of the exon 3 DNA body (lane 1) and the location of single-stranded LINC01914 (lane 2). Lanes 3 to 7 show the formation of a DNA-RNA complex (top band marked with red arrow) upon mixing with decreasing concentrations of LINC01914 (middle band), which leads to a decrease in the intensity of the supershift (top band marked with red arrow). J)Data shows the location of the exon 3 DNA body (lane 1), and the location of single-stranded LINC01914 (lane 2). Lanes 3 to 7 show the formation of a DNA-RNA complex (top band marked with red arrow) upon mixing with labeled LINC01914 (middle band) and increasing concentrations of unlabeled LINC01914, which leads to a decrease in the intensity of the supershift (top band marked with red arrow). K) Data in shows the location of the exon 3 DNA body (lane 1) and the location of single-stranded LINC01914 (lane 2). Lane 3 shows the formation of a DNA-RNA complex (top band marked with red arrow) upon mixing with the first half of the LINC01914 sequence (middle band). In lane 4, the unbound single-stranded RNA is destroyed upon the addition of RNase H and only the DNA-RNA adduct band is visible at the top. L) Data shows the location of the exon 3 DNA body (lane 1), and the location of single-stranded LINC01914 (lane 2). Lane 3 does not show the formation of a DNA-RNA complex upon mixing with the second half of the LINC01914 sequence (top band). In lane 4, the unbound single-stranded RNA is destroyed upon the addition of RNase H and only the DNA band is visible.
Extended Data Fig. 6 Increased levels of mature LINC01914 in stroma is secondary to elevated expression in tumor tissue.
A) qChIP is performed on the promoter region of LINC01914 (exon1 and 700 bp upstream of exon 1) in normal ovarian (N - grey color), HGSC tumor (H - red color) and stromal (S - blue color) tissue samples (from 8 HGSC patients) using anti-H3K27Ac (p < 1x10−4), anti- H3K36Me3 (p < 1x10−4), and anti-H3K27Me3 antibodies (p = 1.7x10−4). The results show repressed LINC01914 promoter activity in the normal ovarian, and HGSC stromal tissue whereas an active LINC01914 promoter status in the HGSC tumor tissue. Data is expressed as the percent of pre-IP input for each sample ± SE and is representative of at least 3 independent IPs. B) A comprehensive set of internal controls such as scramble primer, IgG antibody, Actin antibody, and no antibody were included in the ChIP assay along with several antibodies H3K27ac, H3K36me3, H3K27me3. C) Model depicting the analysis of the expression of LINC01914 nascent RNA (left). Nascent RNA transcript analysis from normal, HGSC and stromal tissue shows that LINC01914 is expressed 12-fold in the HGSC tumor tissue compared to the normal ovarian and the stromal tissue. Mature RNA signal found in the stromal tissue is most likely a result of RNA transport from the tumor tissue (p = 1.5x10−5) (right). D) Analysis of RT-qPCR of LINC01914 using WT LINC01914 primers and 5’ & 3’ barcoded-specific primers. (Top)The plot shows the expression of endogenous LINC01914 in monocultures of OVCAR8 WT cells (lane 2), OVCAR8 LINC01914 KO cells (lane 3), OVCAR8 LINC01914 KO cells with overexpression of barcoded LINC01914 (lane 4), and stromal cells (lane 5, p = 4,7x10−3). The expression of endogenous LINC01914 in stromal cells is also shown upon co-culture with OVCAR8 WT cells (lane 6), OVCAR8 LINC01914 KO cells (lane 7, p = 4.4x10−6), OVCAR8 LINC01914 KO cells with overexpression of barcoded LINC01914 (lane 8, p = 6.4x10−6), and OVCAR8 LINC01914 KO cells with overexpression of barcoded LINC01914 with simultaneous overexpression of WT LINC01914 in stromal cells (lane 9). The expression of endogenous LINC01914 in stromal cells is also shown upon co-culture with OVCAR8 WT cells (lane 6), OVCAR8 Rab27a/b, LINC01914 triple KO cells (lane 7), OVCAR8 Rab27a/b, LINC01914 triple KO cells with overexpression of barcoded LINC01914 (lane 8, p = 1.5x10−4), and Rab27a/b, LINC01914 triple KO cells with overexpression of barcoded LINC01914 with simultaneous overexpression of WT LINC01914 in stromal cells (lane 9). The green arrows mark the lanes in which the barcoded LINC01914 was overexpressed by cancer cells. The * signals the overexpression of WT LINC01914 in stromal cells. A model of the location of WT LINC01914 primers and 5’ & 3’ barcoded-specific primers is shown at the bottom of each plot. LINC01914 enrichment was quantified with N=5 and presented on a logarithmic scale.
Extended Data Fig. 7 HGSC tumor cells pack Tu-Stroma in exosomes via DDX3X binding.
A) Representative nanoparticle tracking analysis showing a snapshot of the video recordings of the particles recorded and the graph showing the particle size and concentration in control and HGSC serum samples. B) Western blot analysis confirms the purity of the exosomes isolated from the blood from the HGSC patients and normal non-HGSC individuals, as indicated by the expression of calnexin, CD9 and TSG101 (markers of exosomes). C) Exosome particles confirmed by image stream flow cytometry (the bar surrounding color indicated exosome count) in control, & HGSOC serum samples. D) Relative expression of Rab27a and Rab27b are higher in WT OVCAR8 cells compared to OVCAR8 cells with Rab27a and Rab27b double knockout (Technical replicates, n=3). E) Data shows nanoparticle tracking (NTA) with exosomes extracted from KO cells having a larger size compared to exosomes from WT cells (189.1 ±12.4 nm vs. 141.8 ± 5.13 nm). F) Mean particle concentration with 95% confidence intervals, determined by NTA, demonstrated a significantly lower concentration of exosomes from Rab27 KO cells compared to WT cells. G) RT-qPCR experiment is performed to understand the cellular localization of LINC01914 (Tu-Stroma) in matched normal, tumor, and stromal tissue from distinct patient samples. In HGSC tissue it is highly expressed in exosomal fraction, followed by the cytoplasmic fraction (p<1X10−4). In the stromal tissue, maximum expression of LINC01914 is observed in the nuclear fraction (p<1X10−4), (Technical replicates, n=3). H) The visualization of LINC01914 transcripts expression in FFPE HGSC sections was achieved through ViewRNA™ in situ hybridization analysis. Large cut sections of selected high-grade ovarian tumor tissue samples were used for staining HOTAIR and LINC01914. Counterstaining was performed using DAPI. Positive LINC01914 staining was observed in HGSC (red spots). I) To determine the binding of LINC01914 exclusively to DDX3X, RIP experiment is performed in matched normal ovarian tissue, HGSC cancer, and HGSC stromal tissue. All the 30 RBPs are immunoprecipitated from each of the three samples and analyzed for the enrichment of LINC01914 by qPCR analysis. The results show that DDX3X is significantly and exclusively upregulated in the cancer tissue (p < 1x10−10), (Biological replicates, n=3). J) RT-qPCR analysis is conducted exosomal fractions isolated from matched normal ovarian tissue, HGSC cancer, and HGSC stromal tissue. RT-qPCR results show that DDX3X exclusively binds to the LINC01914 in exosomal fraction isolated from the HGSC tumor tissue by a 9-fold high expression compared to the stromal exosomal fraction (unmatched p = 3.6x10−6 and matched p = 3.9x10−6). The western blot results confirm that high expression of DDX3X exclusively in the exosomes from HGSC tumor tissue, (Biological replicates, n=5). K) Western blot analysis is conducted in exosomal fractions isolated from normal ovarian tissue, HGSC cancer, and HGSC stromal tissue (15 HGSC patients). The western blot results show high expression of DDX3X in the exosomes from HGSC tumor tissue. L) RT-qPCR measurement is conducted to study the fold change expression of DDX3X mRNA in pooled set of laser-captured. Results show a significant increase in the DDX3X mRNA expression in HGSC tumor tissue compared to the stroma (unmatched and matched p < 1x10−10), (Biological replicates for tumor and stroma tissue, n=30).
Extended Data Fig. 8 Uptake of tumor- derived exosomes changes the expression of Flower isoforms in stromal cells.
A) qPCR experiment is conducted to study the expression of hFwe-Win and hFwe-Lose isoforms in the primary HGSC stromal cells co-cultured with GFP-infected primary HGSC cancer cells on day 0, day 2, day 4, day 6, and day 8. Exogenous addition of exosome inhibitor (GW4869) in the co-cultured cells abrogates the increase in total hFwe mRNA and hFwe-Lose expression (p < 1x10−10). Knockdown of DDX3X by lentivirus-mediated shRNA in the co-culture mix results in a comparatively higher expression of total hFwe mRNA with similar expression levels of hFwe-Lose isoform as observed in exosome inhibitor-treated (p < 1x10−10). B) qPCR and Immunoprecipitation experiments are conducted to validate the role of DDX3X in the exosomal packaging of Tu-Stroma, in the exosomal fractions of HGSC cells on day 0, day 2, day 4, day, and day 8. IP results show a steady and an increasing DDX3X expression in the exosomal fractions from day 0 to day 8. A coherent and significant increase in the Tu-Stroma expression is observed in the same exosomal fractions, as analyzed by qPCR analysis (p < 1x10−10). C) RT-qPCR measurement is performed to analyze the expression of Tu-Stroma mRNA in the exosomal fractions of primary HGSC cancer cells post DDX3X shRNA transfection (from day 0 to day 8), (Technical replicates, n=3). D) qPCR is conducted to study the effect of exosome inhibitors and DDX3X knockdown on Tu-Stroma localization and expression in the nuclear and cytoplasmic fractions of primary HGSC cancer cells. Results show no change in the nuclear to the cytoplasmic distribution of Tu-Stroma expression in the exosome inhibitor treated HGSC cancer cells compared to untreated cells. However, knockdown of DDX3X show an increase in the Tu-Stroma expression in nucleus compared to cytoplasm. Tu-Stroma expression is studied in the nuclear and cytoplasmic fraction of the HGSC stromal cells post co-culture with HGSC cancer cells from day 0-day 8 by qPCR analysis. P-values are indicated in the figure. E) Live-cell imaging of co-cultures experiments for 24h. The top panel shows the co-culture of OVCAR8 DDX3X KO cells expressing RFP-blank with WT stromal cells. The images show that both RFP-tagged OVCAR8 DDX3X KO cells and stromal cells proliferate and survive over time. The middle panel shows the co-culture of OVCAR8 DDX3X KO cells expressing RFP-fused DDX3X with WT stromal cells. The images show that WT stromal cells start expressing RFP within 24 hours. The bottom panel shows the co-culture of OVCAR8 DDX3X KO cells expressing RFP-fused DDX3X with WT stromal cells in the presence of exosome inhibitors.
Extended Data Fig. 9 Epistatic pathway of Tu-Stroma mediated exon 3 methylation and Flower Lose generation in stromal cells.
A) A functional genetic assay is performed on HGSC and Stromal cells cultured individually (row 1) or in co- culture using a series of loss-of-function experiments using Tu-Stroma (row 2) and DDX3X (row 3) shRNA (HGSC), double KO of Rab27a and Rab27b (row 4), Exosome (row 5) and DNMT3A (row 6) inhibitors (co- culture) or shRNA against Tu-Stroma (row 7), DDX3X (row 8), DNMT3A (row 9), G9a (row 10), GLP (row 11), HP1 (row 12) and SRSF3 (row 13). We used five sets of molecular experiments to determine the steps in this molecular mechanism. These experiments were a) ChOP on exon 3 region of HGSC stromal cells to identify the association of Tu-Stroma at this gene locus, b) MethyLight Assay to evaluate level of DNA methylation on this region, c) a series of ChIP experiments (with input, no Ab, Actin Ab, IgG Ab, and Scramble primers as controls) to identify DNA methylation status, and association of DNMT3A, G9a, GLP, H3K9Me3, HP1, and SRSF3 on hFwe exon 3 gene locus in HGSC stromal cells, d) qPCR-based analysis of hFwe-Lose isoform, hFwe-Win isoform, and exon 3 expression in HGSC stromal cells, and e) RT-PCR based validation of Exon 3 cassette deletion. Using the results from this assay, a series of hypotheses are generated to construct the epistatic relationship between individual components in the pathway and depicted as a model. The p-values shown in d) compare the expression of hFwe-Lose in each group with the expression in the control. Microscopy images of each experiment are shown in the first column (scale 10 µm). B) Pseudotime algorithm model to determine the sequence of epigenetic events at the exon 3 hFwe locus. The current model represents the most plausible series of steps (significance of the model (p) is 0.043, which is calculated as a sum of the type I and type II errors across all steps). This pseudo time analysis combined with the known biological function of individual components reveals the mechanism of Tu-Stroma mediated exclusion of exon 3 in stromal cells.
Extended Data Fig. 10 Knockout of Tu- Stroma and DDX3X expression leads to reduced tumor growth and improves overall survival.
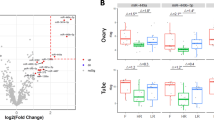
A) Tu-Stroma WT or KO OVCAR8 were orthotopically implanted in female NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) mice and tumor spread in ovaries was determined using MRI. B) RT-qPCR data shows high expression of hFwe-Win in tumor tissue and hFwe-Lose in stromal tissue in WT OVCAR8 xenografts. In Tu-Stroma KO OVCAR8 xenografts, hFwe-Win expression is high in both tumor and stromal tissue and hFwe-Lose expression in stromal tissue was diminished, (Biological replicates, n=5). C) Kaplan-Meier curves are colored as per the label coding key shown in the figure. The p-values show the comparison in survival between Tu-Stroma KD in combination with chemotherapy (PDX1: p = 2.1x10−2; PDX2: 1.8x10−3) and DDX3X KD in combination with chemotherapy (PDX1: p = 1.8x10−3; PDX2: 1.3x10−3) with the WT. The number of mice use in each group was equal to 5. D) After orthotopic implantation, mice with each knockout line (DDX3X KO, a Tu-Stroma KO, and a combined DDX3X/Tu-Stroma KO) were either untreated or received Docetaxel/Carboplatin therapy. Representative tumors from each group (n = 5) are shown with OVCAR8 tumors in the top half and OVCAR4 tumors in the bottom half. Tumors treated with Docetaxel/Carboplatin are indicated with an open box.
Supplementary information
41587_2024_2453_MOESM1_ESM.pdf (download PDF )
Supplementary Figs. 1–16, Table 1, Figure Legends 1–16, Table 1 Legend, Video Legends 1–5, unprocessed western blots and gels of supplementary figures.
Supplementary Video 1 (download AVI )
Live-cell imaging of co-culture with OVCAR8 DDX3X-RFP cells and primary stromal cells for 24 h. In Supplementary Video 1, the data show OVCAR8 cells marked with red resulting from the expression of an RFP fused to DDX3X and untagged primary stromal cells. The 24-h live-cell imaging shows untagged primary stromal cells starting to express RFP. These data show that primary stromal cells uptake DDX3X exported through exosomes by OVCAR8 cells.
Supplementary Video 2 (download MOV )
A 360° visualization of light-sheet microscopy imaging data showing GFP-labeled WT OVCAR8 cells in patient-derived 3D scaffolds carrying primary stromal cells extracted from patients. In Supplementary Video 2, the data are presented for a 3D scaffold where the cancer cells are marked with green, and the 3D scaffold structure and stromal cell populations are marked in purple. The scaffolds are scanned with light-sheet microscopy, and videos were rendered to highlight the structure of the 3D scaffold.
Supplementary Video 3 (download MOV )
A 360° visualization of light-sheet microscopy imaging data showing GFP-labeled Tu-Stroma KO OVCAR8 cells in patient-derived 3D scaffolds carrying primary stromal cells extracted from patients. The data are presented for a 3D scaffold where the cancer cells are marked with green, and the 3D scaffold structure and stromal cell populations are marked in purple. The 3D scaffold set carrying the Tu-Stroma KO OVCAR8 cells is scanned with light-sheet microscopy, and videos were rendered to highlight the structure of the 3D scaffold. The comparison of data in Supplementary Videos 1 and 2 clearly shows considerably reduced ability of Tu-Stroma KO OVCAR8 cells to spread in the 3D scaffold and replace stromal cells, when compared to the WT OVCAR8 cells.
Supplementary Video 4 (download MP4 )
Live-cell imaging of Flower Win expressing (RFP) cells in presence of Flower Lose expressing (GFP) cells in a cell competition assay as described previously4. Data show that Flower Lose expressing GFP cells do not undergo apoptosis in the presence of Flower mAb. Thus, the mAb successfully blocks Flower Win–Lose competitive interaction.
Supplementary Video 5 (download MOV )
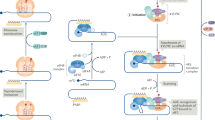
The main findings from our study are summarized as an animated presentation in Supplementary Video 5. HGSC tumor cells export STAT3 to fibroblasts via exosomes. Once inside these cells, STAT3 enters the nucleus and binds to the hFwe promoter, which induces the expression of Flower isoforms. The difference between hFwe-Win and hFwe-Lose isoforms is the presence of exon 3 in hFwe-Win isoforms. Tumor cells produce a lncRNA containing a 21-bp region with 90% complementarity to hFwe exon 3, named Tu-Stroma. The binding of DDX3X with Tu-Stroma packs this lncRNA into exosomes, which then are exported to fibroblasts. Once there, Tu-Stroma translocates to the nucleus and binds to hFwe DNA at exon 3, leading to exon 3 trimethylation by recruiting the DNA methyltransferases G9a, GLP and DNMT3A. Consequently, HP1 binds to the methylated DNA and recruits SRSF3, which excludes hFwe exon 3 from the final mRNA transcript of hFwe. This chain of events results in the upregulation of hFwe-Lose isoforms in fibroblasts and culminates with their elimination by cell competition with tumor cells.
Supplementary Data 1 (download XLSX )
Source data for supplementary figures.
Supplementary Data 2 (download PDF )
Unprocessed western blots and gels of supplementary figures.
Source data
Source Data Figs. 1–5 (download PDF )
Unprocessed western blots and gels of main figures.
Source Data Figs. 1–6 (download XLSX )
Statistical source data of main figures.
Source Data Extended Data Figs. 3–5 and 7–9 (download PDF )
Unprocessed western blots and gels of extended data figures.
Source Data Extended Data Figs. 1–3 and 5–10 (download XLSX )
Statistical source data of extended data figures.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Madan, E., Palma, A.M., Vudatha, V. et al. Ovarian tumor cells gain competitive advantage by actively reducing the cellular fitness of microenvironment cells. Nat Biotechnol 43, 1833–1847 (2025). https://doi.org/10.1038/s41587-024-02453-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41587-024-02453-3
This article is cited by
-
Multi-scale and multi-context interpretable mapping of cell states across heterogeneous spatial samples
Nature Communications (2025)