Abstract
Yersinia pestis, the bacterium that causes the plague, has a dynamic genome with highly conserved fragments prone to rearrangement, influencing gene function and evolution. However, understanding these patterns is limited by few complete genomes and analytical methods. We developed a dual-validation strategy to analyze 242 complete genomes of Y. pestis natural isolates from diverse phylogroups. We detected 459 rearrangements, which enhanced phylogenetic resolution and resolved the third pandemic’s polytomy. Rearrangements are primarily mediated by four common insertion sequences, with IS1661 and IS100 showing the highest activity. These rearrangements are under strong positive selection, evidenced by 43 hotspots and convergent evolution in the rpsO-pnp operon, whose disruptions and reconnections altered gene expressions and temperature stress responses. We also identified unique structural alterations in human avirulent phylogroups, inactivating three genes and reordering 17 intergenic regions, some affecting virulence-related genes. This study provides a fresh perspective on Y. pestis evolution, revealing experimental targets and establishing a methodology for microbes with frequent rearrangements.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The complete genome assemblies and long-read sequencing data generated in this study have been deposited in GenBank and the Sequence Read Archive under BioProject no. PRJNA1190511 and in the NMDC under BioProject no. NMDC10018925. Accession numbers for each genome are listed in Supplementary Table 1. Publicly available complete genomes and the RNA-seq data used in this study are provided in Supplementary Tables 2 and 10, respectively. Insertion sequence types were identified using sequences from the ISfinder database. Functional annotations were assigned using the online eggNOG database. Source data are provided with this paper.
Code availability
The code relevant to this manuscript is publicly available via GitHub at github.com/WuYarong/YP_Rearrangement_Analysis and via Zenodo at https://doi.org/10.5281/zenodo.15152926 (ref. 54). It was implemented in Python v.3.7.10, using the packages SciPy (v.1.5.2), NumPy (v.1.18.1) and statsmodels (v.0.11.1). The insertion sequence copy number was calculated using a custom script based on the BLASTn v.2.13.0+ output format 6.
References
Sun, S., Ke, R., Hughes, D., Nilsson, M. & Andersson, D. I. Genome-wide detection of spontaneous chromosomal rearrangements in bacteria. PLoS ONE 7, e42639 (2012).
Periwal, V. & Scaria, V. Insights into structural variations and genome rearrangements in prokaryotic genomes. Bioinformatics 31, 1–9 (2015).
Ho, S. S., Urban, A. E. & Mills, R. E. Structural variation in the sequencing era. Nat. Rev. Genet. 21, 171–189 (2020).
Raeside, C. et al. Large chromosomal rearrangements during a long-term evolution experiment with Escherichia coli. mBio 5, e01377-14 (2014).
Yebra, G. et al. Radical genome remodelling accompanied the emergence of a novel host-restricted bacterial pathogen. PLoS Pathog. 17, e1009606 (2021).
Irvine, S. et al. Genomic and transcriptomic characterization of Pseudomonas aeruginosa small colony variants derived from a chronic infection model. Microb. Genom. 5, e000262 (2019).
Le, V. V. H., León-Quezada, R. I., Biggs, P. J. & Rakonjac, J. A large chromosomal inversion affects antimicrobial sensitivity of Escherichia coli to sodium deoxycholate. Microbiology 168, 001232 (2022).
Achtman, M. et al. Yersinia pestis, the cause of plague, is a recently emerged clone of Yersinia pseudotuberculosis. Proc. Natl Acad. Sci. USA 96, 14043–14048 (1999).
Yang, R. et al. Yersinia pestis and plague: some knowns and unknowns. Zoonoses 3, 5 (2023).
Morelli, G. et al. Yersinia pestis genome sequencing identifies patterns of global phylogenetic diversity. Nat. Genet. 42, 1140–1143 (2010).
Cui, Y. et al. Historical variations in mutation rate in an epidemic pathogen, Yersinia pestis. Proc. Natl Acad. Sci. USA 110, 577–582 (2013).
Wu, Y. et al. Hotspots of genetic change in Yersinia pestis. Nat. Commun. 16, 388 (2025).
Wu, Y. et al. Small insertions and deletions drive genomic plasticity during adaptive evolution of Yersinia pestis. Microbiol. Spectr. 10, e0224221 (2022).
Parkhill, J. et al. Genome sequence of Yersinia pestis, the causative agent of plague. Nature 413, 523–527 (2001).
Liang, Y. et al. Genome rearrangements of completely sequenced strains of Yersinia pestis. J. Clin. Microbiol. 48, 1619–1623 (2010).
Darling, A. E., Miklós, I. & Ragan, M. A. Dynamics of genome rearrangement in bacterial populations. PLoS Genet. 4, e1000128 (2008).
Li, Y. et al. Different region analysis for genotyping Yersinia pestis isolates from China. PLoS ONE 3, e2166 (2008).
Anisimov, A. P., Lindler, L. E. & Pier, G. B. Intraspecific diversity of Yersinia pestis. Clin. Microbiol. Rev. 17, 434–464 (2004).
Li, Y. et al. The GntR-like transcriptional regulator HutC involved in motility, biofilm-forming ability, and virulence in Vibrio parahaemolyticus. Micro. Pathog. 167, 105546 (2022).
Koebnik, R., Hantke, K. & Braun, V. The TonB-dependent ferrichrome receptor FcuA of Yersinia enterocolitica: evidence against a strict co-evolution of receptor structure and substrate specificity. Mol. Microbiol. 7, 383–393 (1993).
Islam, M. M., Kim, K., Lee, J. C. & Shin, M. LeuO, a LysR-type transcriptional regulator, is involved in biofilm formation and virulence of Acinetobacter baumannii. Front. Cell. Infect. Microbiol. 11, 738706 (2021).
Jiao, W. B. & Schneeberger, K. Chromosome-level assemblies of multiple Arabidopsis genomes reveal hotspots of rearrangements with altered evolutionary dynamics. Nat. Commun. 11, 989 (2020).
Crow, K. D., Wagner, G. P. & SMBE Tri-National Young Investigators Proceedings of the SMBE Tri-National Young Investigators’ Workshop 2005. What is the role of genome duplication in the evolution of complexity and diversity? Mol. Biol. Evol. 23, 887–892 (2006).
Hawkey, J., Monk, J. M., Billman-Jacobe, H., Palsson, B. & Holt, K. E. Impact of insertion sequences on convergent evolution of Shigella species. PLoS Genet. 16, e1008931 (2020).
Tenaillon, O. et al. The molecular diversity of adaptive convergence. Science 335, 457–461 (2012).
Seferbekova, Z. et al. High rates of genome rearrangements and pathogenicity of Shigella spp. Front. Microbiol. 12, 628622 (2021).
Zabelkin, A., Yakovleva, Y., Bochkareva, O. & Alexeev, N. PaReBrick: PArallel REarrangements and BReaks identification toolkit. Bioinformatics 38, 357–363 (2022).
Régnier, P. & Portier, C. Initiation, attenuation and RNase III processing of transcripts from the Escherichia coli operon encoding ribosomal protein S15 and polynucleotide phosphorylase. J. Mol. Biol. 187, 23–32 (1986).
Clarke, D. J. & Dowds, B. C. The gene coding for polynucleotide phosphorylase in Photorhabdus sp. strain K122 is induced at low temperatures. J. Bacteriol. 176, 3775–3784 (1994).
Goverde, R. L., Huis in’t Veld, J. H., Kusters, J. G. & Mooi, F. R. The psychrotrophic bacterium Yersinia enterocolitica requires expression of pnp, the gene for polynucleotide phosphorylase, for growth at low temperature (5 °C). Mol. Microbiol. 28, 555–569 (1998).
Bralley, P., Gatewood, M. L. & Jones, G. H. Transcription of the rpsO-pnp operon of Streptomyces coelicolor involves four temporally regulated, stress responsive promoters. Gene 536, 177–185 (2014).
Zhang, Q. et al. Yersinia pestis biovar Microtus strain 201, an avirulent strain to humans, provides protection against bubonic plague in rhesus macaques. Hum. Vaccin. Immunother. 10, 368–377 (2014).
Song, Y. et al. Complete genome sequence of Yersinia pestis strain 91001, an isolate avirulent to humans. DNA Res. 11, 179–197 (2004).
Ellis, M. J., Trussler, R. S., Charles, O. & Haniford, D. B. A transposon-derived small RNA regulates gene expression in Salmonella typhimurium. Nucleic Acids Res. 45, 5470–5486 (2017).
Ellis, M. J., Trussler, R. S. & Haniford, D. B. A cis-encoded sRNA, Hfq and mRNA secondary structure act independently to suppress IS200 transposition. Nucleic Acids Res. 43, 6511–6527 (2015).
Hu, F., Lin, Y. & Tang, J. MLGO: phylogeny reconstruction and ancestral inference from gene-order data. BMC Bioinformatics 15, 354 (2014).
Drillon, G., Champeimont, R., Oteri, F., Fischer, G. & Carbone, A. Phylogenetic reconstruction based on synteny block and gene adjacencies. Mol. Biol. Evol. 37, 2747–2762 (2020).
Bohnenkämper, L., Braga, M. D. V., Doerr, D. & Stoye, J. Computing the rearrangement distance of natural genomes. J. Comput. Biol. 28, 410–431 (2021).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, e1005595 (2017).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Hunt, M. et al. Circlator: automated circularization of genome assemblies using long sequencing reads. Genome Biol. 16, 294 (2015).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–522 (2018).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Argimón, S. et al. Microreact: visualizing and sharing data for genomic epidemiology and phylogeography. Microb. Genom. 2, e000093 (2016).
Popescu, A.-A., Huber, K. T. & Paradis, E. ape 3.0: new tools for distance-based phylogenetics and evolutionary analysis in R. Bioinformatics 28, 1536–1537 (2012).
Wu, Y. WuYarong/YP_Rearrangement_Analysis: v1.0.0. Zenodo https://doi.org/10.5281/zenodo.15152926 (2025).
Siguier, P., Perochon, J., Lestrade, L., Mahillon, J. & Chandler, M. ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res. 34, D32–D36 (2006).
Minkin, I. & Medvedev, P. Scalable multiple whole-genome alignment and locally collinear block construction with SibeliaZ. Nat. Commun. 11, 6327 (2020).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Tonkin-Hill, G. et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21, 180 (2020).
Goel, M., Sun, H., Jiao, W.-B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277 (2019).
Goel, M. & Schneeberger, K. plotsr: visualizing structural similarities and rearrangements between multiple genomes. Bioinformatics 38, 2922–2926 (2022).
Sedlazeck, F. J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat. Methods 15, 461–468 (2018).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (no. 2022YFC2305304 to Y.C.), the National Natural Science Foundation of China (no. 31970002 to Y.C.) and the State Key Laboratory of Pathogen and Biosecurity (no. SKLPBS2215 to Y.W.).
Author information
Authors and Affiliations
Contributions
Y.C. designed the study. Y.G. conducted the DNA library construction and long-read sequencing. K.M. performed the complete genome assembly. Y.W. and C.Y. analyzed the genomic rearrangements. Y.W. and Y.C. drafted the manuscript. Y.S. provided valuable advice and insights. Y.C. and R.Y. supervised the project. All authors read, revised and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Hendrik Poinar and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Pipeline for identifying rearrangement events in Y. pestis phylogroups.
a, Synteny block construction without a reference genome using SibellaZ and maf2synteny tools. The adjacency marks synteny block neighbors, with ‘h’ for the head and ‘t’ for the tail of blocks. For instance, ‘2h-3t’ illustrates that the head of block no. 2 is adjacent to the tail of block no. 3. b, Pairwise rearrangement detection based on reference genomes with SyRI software. SYN denotes syntenic regions, INV for inversions, TRANS for translocations, INVTR for inverted translocations, and DUP for duplications. Our focus of genomic rearrangements was on INV, TRANS, and INVTR. c, Synteny block analysis combined with pairwise rearrangement identification. d, Determining rearrangement events in the last common ancestor (LCA) of phylogroups. e, Detecting rearrangement events within specific phylogroups.
Extended Data Fig. 2 Synteny blocks dynamics in Y. pestis.
a, Clustered heatmap of 113 accessory blocks in Y. pestis strains, based on their copy number variations. Color intensity reflects copy number, with dendrogram showing clustering. Names of 242 Y. pestis strains and Y. pseudotuberculosis outgroup are listed on the right. b, Close-up on the phylogroup-specific distribution of 43 synteny blocks, a subset derived from panel a, using the same heatmap color legend. Blocks are arranged by variations across phylogroups in the heatmap, highlighted with purple outlines. Parallelism scores from PareBrick analysis are shown above, while blue lines below mark the merging of adjacent blocks into larger segments, with prophage and ribosomal RNA (rRNA) related blocks labeled.
Extended Data Fig. 3 Principal component analysis (PCA) of phylogroup clustering in Y. pestis and comparison of synteny diversity with Y. pseudotuberculosis.
a, PCA cumulative variance plot of 178 synteny block adjacencies associated with rearrangements. The first six principal components (PCs) accounted for over 60.2% of the variance. b, PCA cumulative variance plot of 3,185 single nucleotide polymorphisms (SNPs) and clustering of Y. pestis phylogroups on PC1 and PC2. The first six PCs accounted for 69.8% of the variance. The shaded area represents the cluster of early-diverging 0.PE phylogroups, including 0.PE7, 0.PE2, 0.PE4A, 0.PE4B, and 0.PE4C. c, PCA cumulative variance plot of 2,023 indels and clustering of Y. pestis phylogroups on PC1 and PC2. The first six PCs accounted for 58.6% of the variance. The shaded area represents the cluster of early-diverging 0.PE phylogroups, including 0.PE7, 0.PE2, 0.PE4A, 0.PE4B, and 0.PE4C. d, Synteny diversity comparison between 242 Y. pestis and 25 Y. pseudotuberculosis strains. A sliding window analysis was performed using the IP32953 chromosome as the reference. Synteny diversity in 50-kb windows (10-kb steps) is visualized in purple for Y. pestis and orange for Y. pseudotuberculosis, with 5-kb windows (1-kb steps) in grey for both.
Extended Data Fig. 4 Phylogroup LCA rearrangement events with varied reference genomes.
a, Rearrangements in the LCAs of branch 0 phylogroups, using the Y. pseudotuberculosis IP32953 chromosome as a reference. Synteny diversity (πsyn) for each phylogroup’s strains (5 kb sliding windows with a 1 kb step-size) is aligned with this reference genome. b, Rearrangements in the LCAs of phylogroups within branches 2-4, along with 1.ANT1 and 1.IN1 from branch 1, using the chromosome of strain 43005 (from 0.ANT3) as a reference. c, Rearrangements in 1.IN1-1.IN5 and 1.ORI1 LCAs, referenced against the chromosome of strain 15002 (from 1.IN2). d, Rearrangements in 1.ORI2 and 1.ORI3 LCAs, with the chromosome of strain El-Dorado (from 1.ORI1) as the reference. To be noted, 0.PE2, 0.PE4B, 0.PE4C, and 0.ANT3 (represented as 0.PE4-0.ANT1) have been presented in Fig. 3. In order to comprehensively depict the various populations of Y. pestis, they are listed here again.
Extended Data Fig. 5 Comparative synteny diversity in Y. pestis phylogroups.
a, Synteny diversity (πsyn) in IP32953 reference alignments. This panel illustrates a prominent inversion in the Y. pestis phylogenetic stem between 0.PE4 and 0.ANT1, along with a smaller-scale translocation, demarcated by shaded areas. b, Synteny diversity (πsyn) in 43005 reference alignments. This section highlights stepwise genomic rearrangements within branch 1, as indicated in color-coded shaded regions.
Extended Data Fig. 6 Heatmap of average genomic synteny proportion between pairwise strains across Y. pestis phylogroups.
Color gradient from blue to red represents increasing genomic collinearity from low to high.
Extended Data Fig. 7 Rearrangement patterns in branch 2 for the 2.ANT and 2.MED phylogroups.
Numerical labels represent rearrangement counts, with orange points highlighting events that delineate phylogenetic splits. Given the complexity of determining synteny block order for the common ancestor of the 2.ANT and 2.MED phylogroups, our analysis was limited to their rearrangements compared with strain 43005 from the 0.ANT3 phylogroup, as marked by asterisks. The same rearrangement variations occurring across different phylogroups were independently counted within each respective population.
Extended Data Fig. 8 Distribution of four common insertion sequences in various states across Y. pestis chromosomes.
Both single elements (for example, IS100 or IS1541) and combined elements (for example, ‘IS100-IS1541’, representing adjacent insertion sequences treated as a single unit) are shown (n = 242 chromosomes). Combined elements are counted separately and not included in single-element counts. Counts reflect the number of genomic fragments mapped to each insertion sequence type, irrespective of fragment length, which may result in slight differences from the copy number accumulation analyses described in the Supplementary Notes. In boxplots, the top, middle, and bottom lines indicate the 75th percentile, median, and 25th percentile, respectively. Whiskers extend to 1.5 times the interquartile range (IQR), and points beyond this range are shown as outliers.
Extended Data Fig. 9 Comparison of two methods for identifying rearrangement hotspots and reassessment using an alternative reference.
a, Comparison of the methods using strain 43005 as reference. Linear regression was performed between the -log10-transformed corrected p-values from the Poisson distribution method (y-axis) and the parallelism break scores from PareBrick (x-axis) in R, with hotspot 391t-42h (unfilled circle) excluded as an outlier. The solid line represents the fitted regression line (predicted mean response), with the shaded area indicating the 95% confidence interval (CI) of the fitted values. Adjusted r² and P value for the slope coefficient are shown. Horizontal and vertical dashed lines mark the detection thresholds of the Poisson method (corrected P = 0.05, -log10 scale) and PareBrick (score=1), respectively. Red diamonds indicate hotspots identified by the Poisson method only, while green highlights those identified by PareBrick but lacking statistical significance in the Poisson method. b, Profiling rearrangement hotspots using IP32953 chromosome as reference. See Fig. 5 for legend details. c, Comparison of the methods using strain IP32953 as reference. The figure legend is the same as in panel a.
Extended Data Fig. 10 Evolutionary changes of the rpsO-pnp operon within Y. pestis.
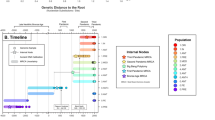
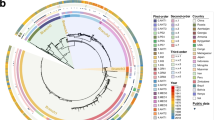
The left panel displays the phylogenetic tree of 242 Y. pestis strains, with Y. pseudotuberculosis IP32953 as the outgroup. The tree highlights five main branches of Y. pestis in distinct colors, with strain names positioned near the tree. The phylogenetic positions of the three global pandemics are annotated. Y. pestis strains from the first two, as ancient DNA without complete genomes, are not included in this study and marked by dashed arrows. The colored bar to the right of the tree represents designated phylogroups. The second bar denotes the rpsO-pnp operon’s state as disrupted or connected. The third bar, normalized to an average sequencing depth of 100, indicates the number of long reads supporting the operon’s connected state. The final bar shows whether there is an IS1661 insertion between the rpsO and pnp genes, with ‘IS1661-free’ marking the absence of such insertion.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–4 and Figs. 1–4.
Supplementary Tables 1–17 (download XLSX )
Supplementary Tables 1–17.
Supplementary Data (download XLSX )
Statistical source data for Supplementary Figs. 1–4.
Source data
Source data for Figs. 1–6 and Extended Data Figs 2–10 (download XLSX )
Statistical source data for Figs. 1–6 and Extended Data Figs. 2–10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, Y., Yang, C., Mu, K. et al. Insights into Yersinia pestis evolution through rearrangement analysis of 242 complete genomes. Nat Genet 57, 1994–2003 (2025). https://doi.org/10.1038/s41588-025-02264-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02264-5