Abstract
Transcription factors (TFs) are instrumental in kickstarting embryonic genome activation (EGA) in many species, yet their regulatory roles in human embryos remain poorly understood. Here, we show that OTX2, a maternally provided PRD-like homeobox TF, is required for proper human EGA and early development. At the four-cell stage, OTX2 promotes activation of key EGA genes, including TPRX1 and TPRX2, and the EGA-associated repeat HERVL-int and MLT2A1. At EGA targets, OTX2 directly binds promoters and putative enhancers, many of which overlap with Alu and MaLR repetitive elements containing the OTX2 motif, and promotes chromatin accessibility. The transcriptome and developmental defects upon OTX2 knockdown are partially rescued by overexpression of TPRX1 and TPRX2. Finally, joint knockdown of OTX2 and TPRXL, encoding another maternal PRD-like homeobox TF, exacerbates chromatin opening and EGA defects at the 8C stage. These findings establish OTX2 as a crucial maternal TF that awakes the genome at the beginning of human life.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive at the National Genomics Data Center103,104, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (https://ngdc.cncb.ac.cn/gsa-human, accession number HRA006621). The processed data reported in this paper have been deposited in the OMIX103,104, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (https://ngdc.cncb.ac.cn/omix, accession number OMIX006476).
Code availability
All data were analyzed using published pipelines with parameters described in the Methods. All custom codes are available at Zenodo (https://doi.org/10.5281/zenodo.15761671)105.
References
Jukam, D., Shariati, S. A. M. & Skotheim, J. M. Zygotic genome activation in vertebrates. Dev. Cell 42, 316–332 (2017).
Lee, M. T., Bonneau, A. R. & Giraldez, A. J. Zygotic genome activation during the maternal-to-zygotic transition. Annu. Rev. Cell Dev. Biol. 30, 581–613 (2014).
Schulz, K. N. & Harrison, M. M. Mechanisms regulating zygotic genome activation. Nat. Rev. Genet. 20, 221–234 (2019).
Tadros, W. & Lipshitz, H. D. The maternal-to-zygotic transition: a play in two acts. Development 136, 3033–3042 (2009).
Liu, B. et al. The landscape of RNA Pol II binding reveals a stepwise transition during ZGA. Nature 587, 139–144 (2020).
Abe, K. I. et al. Minor zygotic gene activation is essential for mouse preimplantation development. Proc. Natl Acad. Sci. USA 115, E6780–E6788 (2018).
Schultz, R. M., Stein, P. & Svoboda, P. The oocyte-to-embryo transition in mouse: past, present, and future. Biol. Reprod. 99, 160–174 (2018).
Mendez, R. & Richter, J. D. Translational control by CPEB: a means to the end. Nat. Rev. Mol. Cell Biol. 2, 521–529 (2001).
Liang, H. L. et al. The zinc-finger protein Zelda is a key activator of the early zygotic genome in Drosophila. Nature 456, 400–403 (2008).
Gaskill, M. M., Gibson, T. J., Larson, E. D. & Harrison, M. M. GAF is essential for zygotic genome activation and chromatin accessibility in the early Drosophila embryo. eLife 10, e66668 (2021).
Duan, J. et al. CLAMP and Zelda function together to promote Drosophila zygotic genome activation. eLife 10, e69937 (2021).
Lee, M. T. et al. Nanog, Pou5f1 and SoxB1 activate zygotic gene expression during the maternal-to-zygotic transition. Nature 503, 360–364 (2013).
Leichsenring, M., Maes, J., Mössner, R., Driever, W. & Onichtchouk, D. Pou5f1 transcription factor controls zygotic gene activation in vertebrates. Science 341, 1005–1009 (2013).
Miao, L. et al. The landscape of pioneer factor activity reveals the mechanisms of chromatin reprogramming and genome activation. Mol. Cell 82, 986–1002 (2022).
Whiddon, J. L., Langford, A. T., Wong, C. J., Zhong, J. W. & Tapscott, S. J. Conservation and innovation in the DUX4-family gene network. Nat. Genet. 49, 935–940 (2017).
Hendrickson, P. G. et al. Conserved roles of mouse DUX and human DUX4 in activating cleavage-stage genes and MERVL/HERVL retrotransposons. Nat. Genet. 49, 925–934 (2017).
De Iaco, A. et al. DUX-family transcription factors regulate zygotic genome activation in placental mammals. Nat. Genet. 49, 941–945 (2017).
Vuoristo, S. et al. DUX4 is a multifunctional factor priming human embryonic genome activation. iScience 25, 104137 (2022).
Liu, Y. et al. Efficient silencing of the multicopy DUX4 gene by ABE-mediated start codon mutation in human embryos. J. Genet. Genomics 49, 982–985 (2022).
Chen, Z. & Zhang, Y. Loss of DUX causes minor defects in zygotic genome activation and is compatible with mouse development. Nat. Genet. 51, 947–951 (2019).
Guo, M. et al. Precise temporal regulation of Dux is important for embryo development. Cell Res. 29, 956–959 (2019).
Zou, Z. et al. Translatome and transcriptome co-profiling reveals a role of TPRXs in human zygotic genome activation. Science 378, eabo7923 (2022).
Ji, S. et al. OBOX regulates mouse zygotic genome activation and early development. Nature 620, 1047–1053 (2023).
Lewin, T. D., Royall, A. H. & Holland, P. W. H. Dynamic molecular evolution of mammalian homeobox genes: duplication, loss, divergence and gene conversion sculpt PRD class repertoires. J. Mol. Evol. 89, 396–414 (2021).
Madissoon, E. et al. Characterization and target genes of nine human PRD-like homeobox domain genes expressed exclusively in early embryos. Sci. Rep. 6, 28995 (2016).
Tohonen, V. et al. Novel PRD-like homeodomain transcription factors and retrotransposon elements in early human development. Nat. Commun. 6, 8207 (2015).
Maeso, I. et al. Evolutionary origin and functional divergence of totipotent cell homeobox genes in eutherian mammals. BMC Biol. 14, 45 (2016).
Maclean, J. A. et al. Rhox: a new homeobox gene cluster. Cell 120, 369–382 (2005).
Galliot, B., de Vargas, C. & Miller, D. Evolution of homeobox genes: Q50 Paired-like genes founded the Paired class. Dev. Genes Evol. 209, 186–197 (1999).
Simeone, A., Acampora, D., Gulisano, M., Stornaiuolo, A. & Boncinelli, E. Nested expression domains of four homeobox genes in developing rostral brain. Nature 358, 687–690 (1992).
Plouhinec, J. L. et al. The mammalian Crx genes are highly divergent representatives of the Otx5 gene family, a gnathostome orthology class of orthodenticle-related homeogenes involved in the differentiation of retinal photoreceptors and circadian entrainment. Mol. Biol. Evol. 20, 513–521 (2003).
Puelles, E. et al. Otx2 regulates the extent, identity and fate of neuronal progenitor domains in the ventral midbrain. Development 131, 2037–2048 (2004).
Beurdeley, M. et al. Otx2 binding to perineuronal nets persistently regulates plasticity in the mature visual cortex. J. Neurosci. 32, 9429–9437 (2012).
Beby, F. & Lamonerie, T. The homeobox gene Otx2 in development and disease. Exp. Eye Res. 111, 9–16 (2013).
Zhang, J. et al. OTX2 restricts entry to the mouse germline. Nature 562, 595–599 (2018).
Yang, S. H. et al. Otx2 and Oct4 drive early enhancer activation during embryonic stem cell transition from naive pluripotency. Cell Rep. 7, 1968–1981 (2014).
Buecker, C. et al. Reorganization of enhancer patterns in transition from naive to primed pluripotency. Cell Stem Cell 14, 838–853 (2014).
Acampora, D., Di Giovannantonio, L. G. & Simeone, A. Otx2 is an intrinsic determinant of the embryonic stem cell state and is required for transition to a stable epiblast stem cell condition. Development 140, 43–55 (2013).
Martinez-Morales, J. R., Signore, M., Acampora, D., Simeone, A. & Bovolenta, P. Otx genes are required for tissue specification in the developing eye. Development 128, 2019–2030 (2001).
Acampora, D. et al. Forebrain and midbrain regions are deleted in Otx2−/− mutants due to a defective anterior neuroectoderm specification during gastrulation. Development 121, 3279–3290 (1995).
Ang, S.-L. et al. A targeted mouse Otx2 mutation leads to severe defects in gastrulation and formation of axial mesoderm and to deletion of rostral brain. Development 122, 243–252 (1996).
Gregory, L. C. et al. The phenotypic spectrum associated with OTX2 mutations in humans. Eur. J. Endocrinol. 185, 121–135 (2021).
Larsen, K. B., Lutterodt, M. C., Møllgård, K. & Møller, M. Expression of the homeobox genes OTX2 and OTX1 in the early developing human brain. J. Histochem. Cytochem. 58, 669–678 (2010).
Kinoshita, M. et al. Capture of mouse and human stem cells with features of formative pluripotency. Cell Stem Cell 28, 453–471 (2021).
Tsankov, A. M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015).
Wu, J. et al. Chromatin analysis in human early development reveals epigenetic transition during ZGA. Nature 557, 256–260 (2018).
Zhu, H.-Y. et al. Histone demethylase KDM4A overexpression improved the efficiency of corrected human tripronuclear zygote development. Mol. Hum. Reprod. 27, gaab012 (2021).
Xia, W. et al. Resetting histone modifications during human parental-to-zygotic transition. Science 365, 353–360 (2019).
Clift, D., So, C., McEwan, W. A., James, L. C. & Schuh, M. Acute and rapid degradation of endogenous proteins by Trim-Away. Nat. Protoc. 13, 2149–2175 (2018).
Xue, Z. et al. Genetic programs in human and mouse early embryos revealed by single-cell RNA sequencing. Nature 500, 593–597 (2013).
Leng, L. et al. Single-cell transcriptome analysis of uniparental embryos reveals parent-of-origin effects on human preimplantation development. Cell Stem Cell 25, 697–712 (2019).
Asami, M. et al. Human embryonic genome activation initiates at the one-cell stage. Cell Stem Cell 29, 209–216 (2022).
Liu, Y., Nie, H., Liu, H. & Lu, F. Poly(A) inclusive RNA isoform sequencing (PAIso−seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails. Nat. Commun. 10, 5292 (2019).
Liu, Y. et al. Remodeling of maternal mRNA through poly(A) tail orchestrates human oocyte-to-embryo transition. Nat. Struct. Mol. Biol. 30, 200–215 (2023).
Franke, V. et al. Long terminal repeats power evolution of genes and gene expression programs in mammalian oocytes and zygotes. Genome Res. 27, 1384–1394 (2017).
Gifford, W. D., Pfaff, S. L. & Macfarlan, T. S. Transposable elements as genetic regulatory substrates in early development. Trends Cell Biol. 23, 218–226 (2013).
Garcia-Perez, J. L., Widmann, T. J. & Adams, I. R. The impact of transposable elements on mammalian development. Development 143, 4101–4114 (2016).
Göke, J. et al. Dynamic transcription of distinct classes of endogenous retroviral elements marks specific populations of early human embryonic cells. Cell Stem Cell 16, 135–141 (2015).
Pontis, J. et al. Hominoid-specific transposable elements and KZFPs facilitate human embryonic genome activation and control transcription in naive human ESCs. Cell Stem Cell 24, 724–735 (2019).
Hashimoto, K. et al. Embryonic LTR retrotransposons supply promoter modules to somatic tissues. Genome Res. 31, 1983–1993 (2021).
Skene Peter, J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Gifford, C. A. et al. Transcriptional and epigenetic dynamics during specification of human embryonic stem cells. Cell 153, 1149–1163 (2013).
Bentsen, M. et al. ATAC-seq footprinting unravels kinetics of transcription factor binding during zygotic genome activation. Nat. Commun. 11, 4267 (2020).
Boulay, G. et al. OTX2 activity at distal regulatory elements shapes the chromatin landscape of group 3 medulloblastoma. Cancer Discov. 7, 288–301 (2017).
Zhang, X. O., Pratt, H. & Weng, Z. Investigating the potential roles of SINEs in the human genome. Annu. Rev. Genomics Hum. Genet. 22, 199–218 (2021).
Polak, P. & Domany, E. Alu elements contain many binding sites for transcription factors and may play a role in regulation of developmental processes. BMC Genomics 7, 133 (2006).
Müller, P., Yanze, N., Schmid, V. & Spring, J. The homeobox gene Otx of the jellyfish Podocoryne carnea: role of a head gene in striated muscle and evolution. Dev. Biol. 216, 582–594 (1999).
Royall, A. H., Frankenberg, S., Pask, A. J. & Holland, P. W. H. Of eyes and embryos: subfunctionalization of the CRX homeobox gene in mammalian evolution. Proc. Biol. Sci. 286, 20190830 (2019).
Vignali, R. et al. Xotx5b, a new member of the Otx gene family, may be involved in anterior and eye development in Xenopus laevis. Mech. Dev. 96, 3–13 (2000).
Li, L. et al. Lineage regulators TFAP2C and NR5A2 function as bipotency activators in totipotent embryos. Nat. Struct. Mol. Biol. 31, 950–963 (2024).
Li, L. et al. Multifaceted SOX2–chromatin interaction underpins pluripotency progression in early embryos. Science 382, eadi5516 (2023).
Lai, F. et al. NR5A2 connects zygotic genome activation to the first lineage segregation in totipotent embryos. Cell Res. 33, 952–966 (2023).
Guo, Y. et al. Obox4 promotes zygotic genome activation upon loss of Dux. eLife 13, e95856 (2024).
Wilming, L. G., Boychenko, V. & Harrow, J. L. Comprehensive comparative homeobox gene annotation in human and mouse. Database 2015, bav091 (2015).
Royall, A. H., Maeso, I., Dunwell, T. L. & Holland, P. W. H. Mouse Obox and Crxos modulate preimplantation transcriptional profiles revealing similarity between paralogous mouse and human homeobox genes. EvoDevo 9, 2 (2018).
Yao, T. & Ndoja, A. Regulation of gene expression by the ubiquitin–proteasome system. Semin. Cell Dev. Biol. 23, 523–529 (2012).
Mark, K. G. & Rape, M. Ubiquitin-dependent regulation of transcription in development and disease. EMBO Rep. 22, e51078 (2021).
Frietze, S. & Farnham, P. J. Transcription factor effector domains. Subcell. Biochem. 52, 261–277 (2011).
Soto, L. F. et al. Compendium of human transcription factor effector domains. Mol. Cell 82, 514–526 (2022).
Liang, D. et al. Distinct dynamics of parental 5-hydroxymethylcytosine during human preimplantation development regulate early lineage gene expression. Nat. Cell Biol. 26, 1458–1469 (2024).
Zhang, T. et al. Maternal transcription factor OTX2 directly induces SETD1A and promotes embryonic genome activation in human pre-implantation embryos. Sci. China Life Sci. 68, 2058–2072 (2025).
Wu, K., et al. Dynamics of histone acetylation during human early embryogenesis. Cell Discov. 9, 29 (2023).
Li, S. et al. Cistrome-GO: a web server for functional enrichment analysis of transcription factor ChIP–seq peaks. Nucleic Acids Res. 47, W206–W211 (2019).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Roy, T. K., Bradley, C. K., Bowman, M. C. & McArthur, S. J. Single-embryo transfer of vitrified-warmed blastocysts yields equivalent live-birth rates and improved neonatal outcomes compared with fresh transfers. Fertil. Steril. 101, 1294–1301 (2014).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Theunissen, T. W. et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 15, 471–487 (2014).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
Huang, H. et al. CTCF mediates dosage- and sequence-context-dependent transcriptional insulation by forming local chromatin domains. Nat. Genet. 53, 1064–1074 (2021).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Boratyn, G. M., Thierry-Mieg, J., Thierry-Mieg, D., Busby, B. & Madden, T. L. Magic-BLAST, an accurate RNA-seq aligner for long and short reads. BMC Bioinformatics 20, 405 (2019).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Chen, T. et al. The Genome Sequence Archive family: toward explosive data growth and diverse data types. Genomics Proteomics Bioinformatics 19, 578–583 (2021).
CNCB-NGDC Members and Partners. Database resources of the National Genomics Data Center, China National Center for Bioinformation in 2025. Nucleic Acids Res. 53, D30–D44 (2025).
Wang, Q. Scripts used in ‘Maternal factor OTX2 regulates human embryonic genome activation and early development’. Zenodo https://doi.org/10.5281/zenodo.15761672 (2025).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Acknowledgements
We are grateful to members of the Xie laboratory and the Chen laboratory for discussions and comments during the OTX2 study and the preparation of the manuscript. We thank the Animal Center and Biocomputing Facility at Tsinghua University for support. We thank J. Na for providing the primed H9 hESCs, L. Wang and Y. Ye for advice and help, and B. Liu and L. Li for help with data analysis. This work was funded by the National Key R&D Program of China (2021YFA1100102 to W.X.), the National Natural Science Foundation of China (31988101 and 31830047 to W.X., 82192874 to H.Z. and 323B2027 to C.Z.), the Excellence Research Group Program of the NSFC (32588201 to Z.-J.C.), the Tsinghua–Peking Center for Life Sciences (to W.X.) and the Fundamental Research Funds of Shandong University (2023QNTD004 to K.W.). W.X. is an HHMI International Research Scholar and a New Cornerstone Investigator.
Author information
Authors and Affiliations
Contributions
Q.W., Z.Z., Z.-J.C., H.Z. and W.X. conceived and designed the project. C.Z., C.L. and S.M. collected human embryos. C.Z. and H.L. collected human oocytes. K.W. and C.Z. performed human 3PN zygote injection and embryo IF with the help of X.M. and Z.Y. Q.W. performed RNA-seq. Q.W. performed CUT&RUN on hESCs. Q.W. performed ChIP–seq on hESCs with help from J.S. Q.W. and Y.D. performed CUT&RUN on human embryos. Q.W. and C.Z. performed ATAC-seq on human embryos. Q.W., Z.Z. and Y.D. conducted pilot experiments with the help of L.W. Z.L. and J.S. helped with various test experiments including cell line construction and RNA-seq. Q.W. analyzed data with help from Z.Z. Q.W. and W.X. prepared figures and wrote the manuscript with help from all authors. C.Z. and H.Z. also participated in the preparation of methods. Z.-J.C., H.Z. and W.X. supervised related experiments and the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Todd Macfarlan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 OTX2 is required for proper human early development.
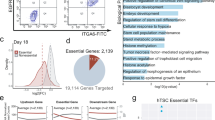
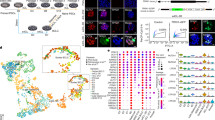
(a) Heatmaps exhibiting OTX2 translation and RNA levels in human and mouse oocytes, early embryos, and ESCs. RPF, ribosome-protected fragment. (b) TF motifs identified from distal ATAC-seq peaks at each stage in human embryos (2-cell, 4-cell, 8-cell, and ICM). Sizes of circles indicate –Log P-values. RPF levels of TFs are color-coded. (c) Bar plot showing OTX2 mRNA levels in control and α-amanitin treated embryos at the 2C and 8C stage46. Mean and SEM are shown. (d) 2PN (N = 19) and 3PN (N = 14) embryo developmental rates on day 3 after IVF. P-value, one-sided chi-square test. (e) Heatmaps showing the mRNA levels of 4C to ICM stage-specifically activated genes in 2PN and 3PN embryos across stages22. (f) Immunostaining of OTX2 in control and OTX2 KD (without Trim-Away) embryos (maximum projections of Z-stacks) and quantification of OTX2 protein levels on day 2. Each dot in the jitter plot represents a single blastomere. Mean and SD are shown. P-value, one-sided t-test. Scale bar, 20 μm. (g) Control embryo developmental rates in a previous study22 (injected with control siRNA and morpholino) (N = 34) and this study (injected with control siRNA and morpholino as well as the Trim21 mRNA and IgG) (N = 35). MO, morula. P-value, one-sided chi-square test. (h) Scatter plots comparing gene expression between the control embryos of OTX2 KD (injected with control siRNA and morpholino as well as the Trim21 mRNA and IgG) and normal embryos on day 2 and day 322. Pearson correlation coefficients are also shown. (i) Immunostaining results of OTX2 in control and OTX2 KD (with Trim-Away) embryos on day 3 (maximum projections of Z-stacks). Red and white arrows, nuclei showing presence and absense of OTX2 proteins, respectively. Scale bar, 20 μm.
Extended Data Fig. 2 OTX2 KD in human embryos on day 2.
(a) Embryo developmental rates of control and OTX2 KD embryos from day 2 to day 4 from two independent batches. N, embryo numbers; MO, morula. (b) Volcano plot showing the differentially expressed genes upon OTX2 KD on D2. The x-axis and y-axis refer to the gene expression fold changes and the p-values computed by DESeq2, respectively. The up-regulated and down-regulated genes are marked in red and blue, respectively. (c) Heatmaps showing the single-embryo mRNA levels (FPKM followed by row z-score normalization) of DEGs upon OTX2 KD on day 2. (d) Violin plots showing the numbers of the OTX2 motif in nearby proximal or distal open chromatin regions for 4C-activated genes down-regulated and unaffected in OTX2 KD embryos on day 2. White center dot, median; black lines, 25th and 75th percentiles; whiskers, 1.5 × interquartile range; P-value, two-sided t-test, same for (i). (e) Immunostaining of TPRX1 in control and OTX2 KD embryos on day 2. Scale bar, 20 μm. (f) PCA of transcriptomes of FGO/MII/1C/2C/4C/8C/ICM (reference data)22, D2 control (black dots), and D2 OTX2 KD (red dots) embryos. (g) Box plots showing mRNA levels of maternal genes defined by Zou et al. that undergo decay at the 4C stage (4C FPKM/ 1C FPKM < 0.5) in normal development22, as well as in OTX2 KD and control embryos on day 2. Center line, median; box, 25th and 75th percentiles; whiskers, 1.5 × interquartile range. P-value, two-sided t-test. (h) Box plots showing the average mRNA levels of 4C-activated (top) and all expressed (bottom) genes in each D2 control and OTX2 KD embryos. The selected control and KD embryos of similar developmental stages are marked in blue and red, respectively. Center line, median; box, 25th and 75th percentiles; whiskers, 1.5 × interquartile range. (i) Violin plots showing the numbers of the OTX2 motif in nearby proximal or distal open chromatin regions for 4C-activated genes down-regulated and unaffected in OTX2 KD embryos selected from (h) on day 2. (j) Volcano plot showing the differentially expressed genes upon OTX2 KD in embryos selected from (h).
Extended Data Fig. 3 OTX2 CUT&RUN in 4C human embryos.
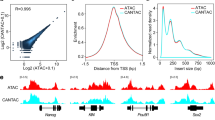
(a) UCSC browser snapshots showing OTX2 CUT&RUN signals in OTX2-overexpressed hESCs (as OTX2 endogenous expression level is low) (100 or 5000 cells) and OTX2 ChIP-seq signals (21 million cells) in OTX2-overexpressed hESCs. The p-values and ranks of the OTX2 motif enriched in the corresponding peaks are also shown on the right. (b) Scatter plot comparing the signals of OTX2 ChIP-seq and CUT&RUN using various numbers of cells. Pearson correlation coefficients are also shown. (c) Scatter plots comparing the signals of 4C OTX2 CUT&RUN between the two replicates. Pearson correlation coefficients are also shown. (d) Venn diagram showing the comparison of OTX2 binding peaks between two replicates. (e) The genomic distribution of OTX2 binding in 4C embryos and hESCs. (f) Bar chart showing percentages of genes with OTX2 binding at promoters (TSS ± 2.5 kb) for 4C-activated genes and all non-maternally expressed genes. (g) The cumulative curve showing distributions of 4C-activated and non-maternally expressed genes with defined distances (x-axis) between their TSSs and nearest 4C OTX2 binding peaks. (h) Line plots showing average enrichment of OTX2 binding signals at the up-regulated genes, down-regulated genes upon OTX2 KD, and non-maternally expressed genes in day 2 embryos. (i) Heatmap showing enrichment of promoter OTX2 binding signals in 4C embryos, primed hESCs (ESC), derived ectoderm-like cells (dEC), and OTX2 motif densities in different clusters. GO analysis results for nearby genes by Cistrome and example genes are shown on the right. (j) TF motifs enriched in 4C-specific, ESC-specific, and dEC-specific promoter OTX2 binding peaks are shown. Sizes of circles indicate –Log P-values. Expression levels of TFs are color-coded.
Extended Data Fig. 4 ATAC-seq analyses in day 2 OTX2 KD embryos.
(a) Heat maps showing OTX2 binding signals and ATAC-seq signals in human 4C embryos, OTX2 motif densities, and ATAC-seq signals in human 1C/2C/8C/ICM embryos[30] in promoter and distal open regions with/without OTX2 binding. (b) UCSC browser snapshots showing ATAC-seq signals of control and OTX2 KD embryos on day 2 (rep 1 and rep 2), ATAC-seq signals in untreated 4C and 8C46 embryos, and OTX2 binding on day 2. (c) Scatter plots comparing the rep 1 and rep 2 ATAC-seq signals in D2 control and OTX2 KD embryos. Pearson correlation coefficients are also shown. (d) Volcano plot showing the TF footprints in differentially accessible regions in ATAC-seq upon OTX2 KD on day 2. Footprints in regions with increasing and decreasing accessibilities are colored in red and blue, respectively. P-values for their enrichment are also shown on the y-axis. (e) Left, heatmap showing enrichment of ATAC-seq signals in promoter regions in 2C46 and 4C embryos, OTX2 binding signals in 4C embryos, OTX2 motif densities, ATAC-seq signals in control and OTX2 KD embryos at 4C. ATAC-seq signals are Z-scored. Right, line plots showing average enrichment of ATAC-seq signals in control and OTX2 KD 4C embryos at the 2C-4C-shared and 4C-specific distal ATAC-seq peaks with or without OTX2 binding. N, peak numbers. (f) Box plots showing RNA changes of genes surrounding the 4C-specific promoter ATAC-seq peaks with/without OTX2 binding in (e) upon OTX2 KD. Center line, median; box, 25th and 75th percentiles; whiskers, 1.5 × interquartile range. P-values, two-sided t-test.
Extended Data Fig. 5 TPRX1/2 over-expression in OTX2 KD embryos.
(a) Heatmaps showing average mRNA levels of TPRX1 and TPRX2 in control, OTX2 KD, and TPRX1/2 rescued OTX2 KD D2 embryos. (b) Violin plots showing the numbers of the ‘TAATCC’ motif in nearby proximal or distal open chromatin regions for TPRX1/2 rescued or not-rescued genes among those were down-regulated upon OTX2 KD embryos on day 2. White center dot, median; black lines, 25th and 75th percentiles; whiskers, 1.5 × interquartile range). P-value, two-sided t-test. (c) Top, bar charts showing mRNA levels of HESX1, MED4, and RPF2 in control, OTX2 KD, and TPRX1/2 rescued OTX2 KD embryos on day 2. Mean and SEM are shown. P-value, two-sided Wald test calculated by DESeq2. Bottom, UCSC browser snapshots showing TPRX1/2 binding in hESCs and OTX2 binding in 4C embryos at example rescued genes. (d) Developmental rates of control, OTX2 KD, and TPRX1/2 rescued OTX2 KD embryos on day 4 (two batches). MO, morula. N, embryo number.
Extended Data Fig. 6 OTX2 over-expression in hESCs.
(a) Volcano plot showing DEGs (computed by DESeq2) upon OTX2 OE in primed hESCs. (b) The cumulative curve showing distributions of down-regulated, up-regulated, and all expressed genes upon OTX2 OE with defined distances (x-axis) between their TSSs and nearest OTX2 binding peaks in the OTX2 OE hESCs. P-value, two-sided Kolmogorov–Smirnov test. (c) Left, heatmap showing enrichment of ATAC-seq signals in hESCs before and after OTX2 OE, OTX2 motif densities, OTX2 binding signals in OTX2 OE hESCs and 4C embryos. Right, cumulative curves showing distributions of down-regulated, up-regulated, and all expressed genes upon OTX2 OE in hESCs with defined distances (x-axis) between their TSSs and nearest ATAC-seq peaks. ATAC-seq signals are Z-scored. N, peak numbers. P-values (Kolmogorov–Smirnov test) are shown. (d) Box plots showing RNA changes of genes surrounding the promoter ATAC-seq peaks in (c) upon OTX2 OE. Center line, median; box, 25th and 75th percentiles; whiskers, 1.5 × interquartile range. P-values, two-sided t-test. (e) UCSC browser snapshots showing ATAC-seq signals before and after OE OTX2 in hESCs, OTX2 binding signals in OTX2 OE hESCs, and 4C embryos at targeted genes. mRNA levels are shown in the heatmaps. (f) Venn diagram comparing the OTX2 and TPRX1/2 OE up-regulated genes in primed hESCs and 4C/8C-activated EGA genes. P-values (Fisher’s test) are shown. (g) GO terms for the up-regulated/down-regulated EGA genes upon OTX2 OE in primed hESCs (left) and naïve hESCs (right). (h) Venn diagram showing the OTX2 OE up-regulated genes in primed /naïve hESCs and 4C/8C-activated genes. P-values (Fisher’s test) are shown.
Extended Data Fig. 7 OTX2 occupies and opens repetitive elements in 4C embryos.
(a) Scatter plot comparing the relative enrichment (observed number/expected number from randomized peaks, Obs./Exp.) of each repeat family (>5000 copies) in 4C ATAC peaks and 4C OTX2 binding peaks near minor EGA genes (TSS ± 50 kb). (b) Schematic of ‘TAATCC’ motif positions in the consensus sequence83 of repeat elements. (c) TF motifs identified from 4C-open Alu elements with or without OTX2 binding. Sizes of circles indicate –Log P-values. Expression levels of TFs are color-coded. (d) TF motifs identified from 4C-open MaLR elements with or without OTX2 binding. Sizes of circles indicate –Log P-values. Expression levels of TFs are color-coded. (e) UCSC browser snapshots showing ATAC-seq signals of control and OTX2 KD and OTX2 binding on day 2 at MaLR elements near DUXB genes. Heatmaps showing the average mRNA levels of DUXB in control and OTX2 KD embryos on day 2.
Extended Data Fig. 8 OTX2 KD in day 3 embryos.
(a) GO terms analyzed by Metascape106 for genes down-regulated upon OTX2 KD on day 2 that still down-regulated (top) and recovered their expression (bottom) on day 3. (b) The cumulative curve showing distributions for ‘remain-down’ and ‘recovered’ genes, as well as for all expressed genes on day 3, with defined distances (x-axis) between their TSSs and nearest 4C OTX2 binding peaks. (c) Scatter plots comparing expression changes of repeat elements (log2 ratio computed by DESeq2) (y-axis) between OTX2 KD and control embryos on day 3, and repeat RNA changes for normal embryos between the 1C stage and the 8C stage (log2 ratio) (x-axis). Repeats up-regulated and down-regulated after KD are labeled in red and blue, respectively. (d) mRNA levels (single embryo) of OTX2 and TPRXL in control, OTX2 KD, and TPRXL KD embryos (left) as well as in control and OTX2/TPRXL dKD embryos (right) on day 3. Each dot in the jitter plot represents a single embryo. Mean and SD are shown. P-value, two-sided Wald test calculated by DESeq2. (e) Developmental rates of control and OTX2/TPRXL dKD embryos on day 3-4 (two batches). MO, morula. N, embryo number. (f) Scatter plots comparing mRNA changes (log2 ratio computed by DESeq2) between KD and control embryos (y-axis), and mRNA changes from 1C stage to 8C stage (normal embryos) (log2 ratio) (x-axis) are shown for OTX2 KD, TPRXL KD22, and OTX2/TPRXL dKD embryos. Maternal and 4C/8C-activated genes are labeled in yellow, green, and red, respectively. (g) Jitter plots showing day 2/3 mRNA levels (single embryo) of example genes upon OTX2 KD on day 2, and, OTX2 KD, TPRXL KD, and dKD on day 3. Each dot in the jitter plot represents a single embryo. Mean and SD are shown. P-value, two-sided Wald test calculated by DESeq2. (h) Venn diagram showing the overlap of down-regulated genes upon OTX2 KD, TPRXL KD, and dKD on day 3.
Extended Data Fig. 9 Joint OTX2 and TPRXL knockdown exacerbates defects of chromatin accessibility during major EGA.
(a) Hierarchical clustering of ATAC-seq peak signals from this study and the previous study46. Pearson correlation is used to measure distances. Blue, normal embryos; green, control embryos; red, KD embryos. (b) Volcano plot showing the TF footprints in differentially accessible regions in ATAC-seq upon OTX2 KD, TPRXL KD, and dKD on day 3. Footprints in regions with increasing and decreasing accessibilities are colored in red and blue, respectively. P values for their enrichment are also shown on the y-axis. (c) The cumulative curve showing distributions of down-regulated, up-regulated, and all expressed genes within defined distances (x-axis) from their TSSs to the nearest lost distal ATAC peaks upon OTX2 KD, TPRXL KD, and dKD. P-value, Kolmogorov–Smirnov test.
Extended Data Fig. 10 Impacts of OTX2/TPRXL dKD for gene expression on day 4.
(a) Heatmaps showing the average mRNA levels of 4C to ICM stage-specifically activated genes in normal embryos across stages22, as well as in the control and OTX2 single KD/dKD embryos on D4. Genes down-regulated (computed by DESeq2) and unaffected are labeled by the red and blue bars, respectively. (b) Venn diagram showing the overlapped genes that are down-regulated on day 3 and day 4 in dKD embryos. P-value (Fisher’s test) is shown. (c) Venn diagram showing the overlapped genes that are down-regulated in OTX2 single KD and dKD embryos on day 4. P-value (Fisher’s test) is shown.
Supplementary information
Supplementary Tables 1–7 (download XLSX )
Supplementary Table 1. siRNA and morpholino sequences used in this study. Supplementary Table 2. Human embryo numbers used in this study. Supplementary Table 3. Stage-activated gene list in this study. Supplementary Table 4. Human embryo OTX2-KD RNA counts. Supplementary Table 5. hESC OTX2-OE RNA counts. Supplementary Table 6. All differentially expressed gene lists in this study. Supplementary Table 7. OTX2 CUT&RUN peaks for the 4C stage.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Q., Zhang, C., Dang, Y. et al. Maternal factor OTX2 regulates human embryonic genome activation and early development. Nat Genet 57, 2772–2784 (2025). https://doi.org/10.1038/s41588-025-02350-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02350-8
This article is cited by
-
OTX2 awakens the genome of human embryos
Nature Genetics (2025)