Abstract
The organization of the genome in three-dimensional space is highly dynamic, yet how these dynamics are regulated and the role they play in genome function is poorly understood. Here we utilized acute depletion of NIPBL to characterize cohesin-mediated loop extrusion in vivo. We find that many chromatin loops are rapidly diminished upon loss of NIBPL, but some cohesin-dependent chromatin loops persist for multiple hours. These persistent loops required NIPBL for their establishment during mitotic exit, were associated with distinct chromatin states and were preferentially dependent on STAG1 for their persistence. Furthermore, by depleting NIPBL from multiple cell types, we find that NIPBL specifically regulates cell identity genes by supporting a unique local genome conformation defined by greater spatial proximity to nearby super-enhancers and weaker transcription start site insulation of genomic contacts. Overall, we show that NIPBL-mediated loop extrusion is critical to genome organization and transcription regulation in vivo.
Similar content being viewed by others
Main
In mammals, the genome is organized into hierarchical structures across three-dimensional (3D) space. This organization has been reported to play important roles in transcription regulation and DNA repair, with its dysfunction particularly associated with developmental disorders and cancer1,2. At the sub-chromosomal level, 3D genome organization is composed of chromatin loops and topologically associating domains, which form through the process of loop extrusion by cohesin3,4,5,6. Current models for cohesin-mediated loop extrusion suggest that this process can be highly dynamic, yet we understand relatively little about cohesin dynamics in vivo and how they influence transcription7.
Recent live-cell imaging approaches at single loci have suggested that chromatin loops persist for only 10–20 min (refs. 8,9), consistent with the ~20-min residence time of cohesin on chromatin10. In vitro, cohesin-mediated loop extrusion requires the accessory protein NIPBL, which was originally described as a cohesin loading factor11,12. NIPBL likely facilitates loop extrusion through its ability to activate the ATPase activity of cohesin13. However, the contribution of NIPBL to 3D genome organization is under-explored in mammalian cells14,15,16, particularly at the short time-scales necessary for understanding chromatin looping dynamics. Despite the association of 3D genome organization with transcriptional regulation, disruption to this organization has surprisingly minimal effects on gene expression17,18. Proposed reasons for this disconnect include a preferential role of cohesin in gene activation over maintenance19,20 and the presence of residual enhancer–promoter connectivity with cohesin depletion21. Regardless, it is not clear why or how only a subset of genes are regulated by cohesin. Given that enhancer–promoter contacts can be highly dynamic22, understanding how these dynamics contribute to transcriptional regulation is critical.
To address these questions, we established a system for the acute depletion of NIPBL to disrupt cohesin dynamics. While many loops were rapidly lost upon NIPBL depletion, a subset of cohesin-dependent loops persist in the absence of NIPBL. Notably, these loops require NIPBL for their establishment during mitotic exit, suggesting distinct roles of NIPBL in the formation versus maintenance of chromatin loops in vivo and raising the possibility that chromatin loop longevity may vary genome-wide. Furthermore, given the essentiality of cohesin in sister chromatid cohesion23 and a dispensability for NIPBL for cohesion during mitotic exit, we used NIPBL depletion to test the role of post-mitotic genome reorganization in transcriptional reactivation24,25,26,27. We found that NIPBL facilitates expression of lineage-defining genes by promoting strong contacts with nearby super-enhancer elements and by enabling extensive connectivity between regions up- and downstream of the transcription start site (TSS). Surprisingly, we found that this role of cohesin is not unique to mitotic exit and that cohesin regulates distinct sets of lineage-specific genes across human induced pluripotent stem cell (hiPSC)-derived lineages through similar patterns of spatial organization. Taken together, our data illuminate the contribution of NIPBL to cohesin-mediated loop extrusion dynamics in vivo and reveal how disruption to these dynamics affects the regulation of transcription.
Results
Acute depletion of NIPBL from hTERT RPE-1 cells
To understand cohesin-mediated loop extrusion in vivo, we employed the dTAG system28,29 for acute depletion of NIPBL in hTERT RPE-1 cells (Fig. 1a). We knocked-in an FKBP12F36V tag to the N terminus of NIPBL, generating two independent clonal cell lines (NIPBL-A2 and NIPBL-D7; Extended Data Fig. 1a–c). Degradation of HA-NIPBL began within 1 h of treatment with dTAGV-1 and had limited effects on other cohesin subunits, but was accompanied by the loss of MAU2, an interaction partner of NIPBL that is used as an independent measure of NIPBL loss30,31 (Fig. 1b and Extended Data Fig. 1d). We similarly engineered cells for depletion of the core cohesin subunit RAD21 (Extended Data Fig. 1e–g) to compare the consequences of perturbing all cohesin-dependent genome organization with disrupting only their formation.
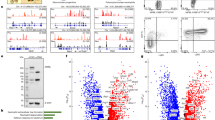
a, Schematic of the cohesin loading cycle, with depletion of NIPBL aiming to impair the ability of new chromatin loops to be established. b, NIPBL-D7 cells were treated with DMSO for 24 h or dTAGV-1 for varying lengths of time, with the degree of HA-NIPBL depletion determined by western blotting. Also shown are the effects of NIPBL depletion on MAU2. α-Tubulin is shown as a loading control. Two biological replicates were performed with similar outcomes. c, Heatmap showing max-normalized balanced counts of chromatin loops that we classified as RAD21-independent or RAD21-dependent based on a twofold decrease in chromatin loop strength (balanced counts) in response to RAD21 depletion (top). Also shown are the consequences of NIPBL depletion on the strength of RAD21-dependent and RAD21-independent chromatin loops (bottom). d, Heatmap showing max-normalized balanced counts of k-means clustering of RAD21-dependent chromatin loops from asynchronous cells depleted of NIPBL. Cluster names refer to NIPBL-dependency: minimum-dependency (‘min.-dep.’), low-dependency (‘low-dep.’), medium-dependency (‘med.-dep.’), mixed-dependency (‘mixed-dep.’), high-dependency (‘high-dep.’) and maximum-dependency (‘max.-dep.’). e, RAD21 differential binding following NIPBL depletion at loop anchors in the different clusters. For each chromatin loop anchor, we identified the RAD21 peak that had the smallest −log2(FC) and found the mean of the two anchors for a given loop. Chromatin loops with no intersecting RAD21 peaks were excluded, but we kept chromatin loops with only one anchor intersecting an RAD21 peak. P value shown is from a Kruskal–Wallis test (min.-dep. n = 1,018, low-dep. n = 2,715, med.-dep. n = 3,476, mixed-dep. n = 1,670, high-dep. n = 3,746, max.-dep. n = 2,487). The center line marks the median, the box shows the IQR and the whiskers extend to 1.5 × IQR. f, Observed/expected pile-up analysis at a 10-kb resolution of the mixed-dep. cluster in cells depleted of NIPBL (top), RAD21 (middle) or CTCF (bottom). g, As in f, but with NIPBL depletion in the max.-dep. cluster. h, Sizes of chromatin loops from each of the clusters. P value shown is from a Kruskal–Wallis test (min.-dep. n = 1,171, low-dep. n = 3,164, med.-dep. n = 3,931, mixed-dep. n = 1,757, high-dep. n = 4,124, max.-dep. n = 2,662). The center line marks the median, the box shows the IQR and the whiskers extend to 1.5 × IQR. i, As in f, but with acute depletion of NIPBL from synchronized cells during mitotic exit at 2 h following release from nocodazole. j, Example chromatin loop that falls into the mixed-dep. cluster, with Hi-C heatmaps from NIPBL, RAD21 and CTCF depletion from asynchronous cells and NIPBL depletion during mitotic exit. Below the heatmaps are a selection of genes that fall into the region and scale factor-normalized BigWig tracks from RAD21 ChIP–seq in asynchronous NIPBL-D7 cells treated with DMSO or dTAGV-1 for 4 or 24 h. The chromatin loop of interest is marked by an arrow, while its anchor regions are marked by gray bars in the tracks below. IB, immunoblot; FC, fold-change; obs/exp, observed/expected.
We next characterized the consequences of short- and long-term depletion of NIPBL on chromatin association of cohesin using chromatin fractionation and chromatin immunoprecipitation followed by sequencing (ChIP–seq). By chromatin fractionation, we observed only a modest reduction in chromatin-associated RAD21 (Extended Data Fig. 2a). In comparison, RAD21 was progressively lost from its binding sites, with almost all ChIP–seq peaks showing a degree of reduction at both 4 and 24 h of dTAGV-1 (Extended Data Fig. 2b). Retained and diminished peaks had similar enrichment of HA-NIPBL, but diminished peaks had weaker RAD21 binding (Extended Data Fig. 2c). Together, these findings suggest that cohesin may retain nonspecific association with chromatin in the absence of NIPBL. We observed a G1 cell cycle arrest with NIBPL depletion, a mix of G1 and 4N cells with RAD21 depletion, but no cell cycle changes with dTAGV-1 treatment of parental hTERT RPE-1 cells (Extended Data Fig. 2d,e). Thus, residual cohesive cohesin from cells in S/G2/M, when cohesin is more stable on chromatin10, may account for the fewer differential ChIP–seq peaks at 4 h of NIPBL depletion compared with 24 h.
NIPBL-dependence defines distinct classes of chromatin loops
To understand the impact of NIPBL-mediated loop extrusion, we compared the consequences of NIPBL and RAD21 depletion, sorting 2N cells from RAD21 control and dTAGV-1-treated cells to account for cell cycle perturbations (Extended Data Fig. 3a). Overall, the effect of NIPBL depletion on genome organization was less severe compared with depletion of RAD21, but chromatin loops, stripes and compartments were all affected by both NIPBL and RAD21 depletion (Extended Data Fig. 3b–e). However, we found that genomic regions differed in their sensitivity to NIPBL depletion (Extended Data Fig. 3f). To understand this, we first classified 16,860 chromatin loops as cohesin-dependent based on RAD21 depletion (Fig. 1c and Extended Data Fig. 4a–c). K-means clustering of these cohesin-dependent loops identified six clusters with variable loss of strength following NIPBL depletion (from minimum to maximum dependency, Fig. 1d and Extended Data Fig. 4d). Chromatin loops that were less dependent on NIPBL retained more RAD21 by ChIP–seq with NIPBL depletion than more NIPBL-dependent loops (Fig. 1e and Extended Data Fig. 4e). Most clusters showed a similar loss of loop strength at both 4 and 24 h (Fig. 1d and Extended Data Fig. 4d), with the exception of a ‘mixed-dependency’ cluster that had stronger reduction in loop strength at 24 h than at 4 h (Fig. 1f,g, compare top row of Fig. 1f with Fig. 1g, and Extended Data Fig. 4f). We also engineered a CTCF degron in hTERT RPE-1 cells and found that all clusters were perturbed by CTCF depletion (Fig. 1f and Extended Data Fig. 4g–j). Together, these findings indicate that these clusters represent bona fide cohesin/CTCF-dependent chromatin loops that have variable persistence in the absence of ongoing loop establishment by NIPBL.
We next examined features of each cluster to understand their differential dependence on NIPBL. We found that more NIPBL-dependent loops were longer and less NIPBL-dependent loops were shorter, although the mixed-dependency cluster had the largest median loop size of 450 kilobases (kb) (Fig. 1h). Structural loops (cohesin/CTCF at both anchors)24 were common in each cluster (ranging from 38.9% for low-dependency to 53.6% for mixed-dependency), but less NIPBL-dependent loops were more likely to be classified as cis regulatory element loops (enhancer/promoter at both anchors, ranging from 9.7% for maximum-dependency to 25.2% for minimum-dependency; Extended Data Fig. 4k). Overall, our findings suggest genomic context, such as association with enhancers or cis regulatory elements, is modestly associated with differential dependence of chromatin loops on NIPBL for their maintenance.
Persistent loops require NIPBL for establishment post mitosis
We next investigated how NIPBL contributes to the establishment of different classes of chromatin loops using the intrinsic reorganization of the 3D genome that occurs during mitotic exit24,27,32. Cells were synchronized to mitosis using double thymidine block and nocodazole, depleted of NIPBL and released into G1. Both DMSO- and dTAGV-1-treated cells exited mitosis with similar kinetics and were in G1 at 8 h post release, with only a subset of DMSO-treated cells progressing out of G1 after 16 h (Extended Data Fig. 5a–c). Critically, HA-NIPBL was successfully depleted with dTAGV-1 treatment of mitotic cells (Extended Data Fig. 5d). We then performed Hi-C at 30-min intervals following mitotic release (Extended Data Fig. 5e). Substantial structural reorganization occurred under both conditions (Extended Data Fig. 5f), but the consequences of NIPBL depletion were stronger during mitotic exit than in asynchronous cells (Extended Data Fig. 5g). NIPBL was also essential for the establishment of the mixed-dependency chromatin loops during mitotic exit, contrasting strongly with the effect of NIPBL depletion on mixed-dependency loops from asynchronous cells (Fig. 1i versus Fig. 1f). Indeed, the consequences of NIPBL depletion on the mixed- and maximum-dependency loops were more similar during mitotic exit than in asynchronous cells (Fig. 1j and Extended Data Fig. 5h). Overall, all classes of loops were perturbed by NIPBL depletion during mitotic exit, but only the less-dependent clusters regained a degree of their strength at later timepoints (Extended Data Fig. 5i), possibly through NIPBL-independent mechanisms. Our finding that the mixed-dependency cluster depends on NIPBL for their establishment, but not maintenance, suggests that these represent a unique class of chromatin loop.
Loop persistence is altered by cohesin subunit composition
STAG1 and STAG2 are mutually exclusive cohesin subunits, with STAG1 thought to support longer and STAG2 shorter chromatin loops33. Consistent with this, STAG1 was enriched at the more NIPBL-dependent loops, while STAG2 was higher at the less-dependent loops (Fig. 2a). SMC3 acetylation can enhance the residence time of cohesin on chromatin33, and we find that the less NIPBL-dependent loops have higher levels of SMC3ac (Extended Data Fig. 6a). Additionally, the levels of HA-NIPBL were lowest and PDS5A highest at the mixed-dependency cluster (Extended Data Fig. 6a), consistent with evidence that NIPBL and PDS5A compete for binding to cohesin13,34. The cohesin unloading factor WAPL showed only a mild relationship with the different clusters (Extended Data Fig. 6a), suggesting that the differential stability of chromatin contacts was not driven by altered WAPL distribution.
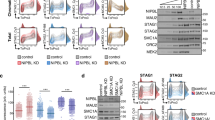
a, Boxplot showing the RAD21-normalized RPKM for STAG1 and STAG2 ChIP–seq at the different clusters from Fig. 1d. For each chromatin loop, the mean RPKM was taken from all RAD21 ChIP–seq peaks across both anchors and normalized to the mean RPKM of RAD21 at the same anchors to account for differences in RAD21 ChIP–seq signal. Differences in chromatin binding at the clusters were tested using a Kruskal–Wallis test (min.-dep. n = 1,037, low-dep. n = 2,780, med.-dep. n = 3,533, mixed-dep. n = 1,678, high-dep. n = 3,820, max.-dep. n = 2,523). The center line marks the median, the box shows the IQR and the whiskers extend to 1.5 × IQR. b, Observed/expected pile-up analysis at a 10-kb resolution of the mixed-dep. cluster in cells depleted of NIPBL, with either wild-type STAG1/2, STAG1 knockout or STAG2 knockout. c, Quantification of loop strength across the different clusters and how this is affected by NIPBL depletion, STAG1 knockout and STAG2 knockout. P values were determined using two-sided Wilcoxon rank-sum tests (see Supplementary Table 6 for exact values; min.-dep. n = 1,171, low-dep. n = 3,164, med.-dep. n = 3,931, mixed-dep. n = 1,757, high-dep. n = 4,124, max.-dep. n = 2,662). **P ≤ 0.01, ****P ≤ 0.0001. The center line marks the median, the box shows the IQR and the whiskers extend to 1.5 × IQR. d, Example mixed-dep. chromatin loop, showing effects of STAG1 and STAG2 knockout. Below the heatmaps are Virtual 4C tracks using the left loop anchor as ‘bait’. e, Mean FPKM of nascent transcripts whose promoter-proximal regions (TSS −1 kb/+0.5 kb) intersect chromatin loop anchors from the different clusters. Nascent transcript levels are from SLAM-seq performed on DMSO-treated NIPBL-D7 cells. P value shown is from a Kruskal–Wallis test (min.-dep. n = 284, low-dep. n = 932, med.-dep. n = 1,264, mixed-dep. n = 519, high-dep. n = 1,350, max.-dep. n = 737). The center line marks the median, the box shows the IQR and the whiskers extend to 1.5 × IQR. f, Enrichment of ChromHMM chromatin states in cluster loop anchors and loop spans. For loop anchors, both anchors were included in analysis. Fold-enrichment is calculated by ChromHMM based on the proportions of bases in the chromatin loop anchors/spans that fall into each state compared with the proportion of bases in the whole genome that fall into that state. g, RPKM-normalized ChIP signal for example histone marks used in identifying ChromHMM chromatin states centered at RAD21 ChIP–seq peaks that overlap anchors of chromatin loops in each of the clusters. E., enhancer; KO, knockout; NS, nonsignificant; P., promoter; RPKM, reads per kilobase million; txn, transcriptional; WT, wild type.
To explore whether the STAG subunits influence loop persistence, we knocked out STAG1 or STAG2 in NIPBL-D7 cells (Extended Data Fig. 6b). Upon STAG1 knockout, the strength of the mixed-dependency cluster in the DMSO-treated condition is not significantly changed (Fig. 2b–d), indicating that cohesinSTAG2 is similarly capable to cohesinSTAG1 at establishing chromatin loops within this cluster. The shorter minimum- and low-dependency clusters were also unperturbed by STAG1 knockout (Fig. 2c). Upon depletion of NIPBL, however, we found that the mixed-dependency cluster was comparatively weaker in the absence of STAG1, but stronger in the absence of STAG2 (Fig. 2b). This outcome suggests that cohesinSTAG1 is responsible for the persistence of the mixed-dependency cluster in the absence of NIPBL. This contrasts with the maximum-dependency cluster, which is weakened by STAG1 knockout in both control and dTAG-treated conditions (Fig. 2c and Extended Data Fig. 6c,d). Our findings are consistent with cohesinSTAG1 having a longer residence time on chromatin and enhancing the stability of chromatin loops33.
Chromatin state predicts loop persistence
Genes associated with the loop anchors of each cluster showed distinct levels of nascent transcription (Fig. 2e). Based on this, we used ChromHMM to classify chromatin states (Extended Data Fig. 6e) to determine whether this is associated with loop stability. All clusters were strongly enriched for insulator state (Fig. 2f), which is driven by CTCF (Extended Data Fig. 6e). Enhancer chromatin states were strongly associated with the anchors and span of less-dependent loops (Fig. 2f). More-dependent clusters tended to have promoter chromatin states at their anchors but enhancer states within their loops (Fig. 2f). In contrast, the anchors of the mixed-dependency cluster showed low enrichment of promoter- or enhancer-associated chromatin states (Fig. 2f). Instead, the anchors and span of the mixed-dependency cluster showed slight enrichment in repressive chromatin (1.7-fold-enrichment versus 0.6-fold for the maximum-dependency cluster at loop anchors; Fig. 2f), associated with the presence of the H3K27me3 mark (Extended Data Fig. 6e). Indeed, repressive chromatin is the majority state for ~65% of loop anchors within the mixed-dependency cluster compared with ~20% of the maximum-dependency cluster (Extended Data Fig. 6f). While RAD21 peaks themselves are devoid of H3K27me3, the mixed-dependency cluster shows particularly strong signal flanking the peak (Fig. 2g and Extended Data Fig. 6g). In summary, acute depletion of NIPBL has locus-dependent consequences on genome organization related to chromatin loop size, cohesin subunit composition and chromatin state.
NIPBL is necessary for activation of lineage-defining genes
Concurrent with the genome reorganization that occurs during mitotic exit, there is widespread transcriptional activation25,26,27. Whether 3D genome organization is required for post-mitotic gene activation is unknown. We tested the consequences of NIPBL depletion on transcription during mitotic exit using thiol(SH)-linked alkylation for the metabolic sequencing of RNA (SLAM-seq)35, with both longer 4-thiouridine (4sU) labeling for differential analysis (mitotic exit approach B) and shorter labeling for the kinetics of nascent transcription (mitotic exit approach A) (Extended Data Fig. 7a).
Depletion of NIPBL during mitotic exit resulted in the differential expression of 549 genes, including 457 decreased genes (Fig. 3a,b). Genes that failed to activate with NIBPL depletion were enriched in processes related to cell behavior, including cell migration and shape (Fig. 3c). Similarly, gene set enrichment analysis showed a negative correlation with genes involved in epithelial to mesenchymal transition and KRAS signaling (Fig. 3d), highlighting a possible role of NIPBL in the regulation of lineage-defining genes. hTERT RPE-1 cells are dedifferentiated and fibroblast-like compared with primary epithelial pigment cells36,37. Indeed, with prolonged depletion of NIPBL, cells shift from a fibroblastic to a more epithelial-like morphology (NIPBL-D7 (Fig. 3e); NIPBL-A2 (Extended Data Fig. 7b)). Taken together, these results suggest that NIPBL may play a role in controlling gene regulatory programs that are critical for maintaining cell identity.
a, Volcano plot from differential expression analysis of nascent transcripts between DMSO- and dTAGV-1-treated NIPBL-D7 cells based on a 120-min 4sU labeling period during mitotic exit. Significantly changed (|log2FC| > 0.585 and Padj < 0.1) nascent transcripts are highlighted. P values were calculated from two-sided Wald tests using 3–4 independent replicates and adjusted (Padj) for multiple testing using the Benjamini–Hochberg method. b, Scatter plot showing the log10-transformed FPKM values for nascent transcripts from a. Significantly changed transcripts (|log2FC| > 0.585 and Padj < 0.1) are highlighted. A kernel density estimate plot is shown on each axis to represent the distribution of the datapoints on that axis. To each FPKM value, 0.001 was added, before log-transformation to allow for plotting of zero FPKM values. c, Biological process gene ontology analysis was performed on the significantly decreased (top) and increased (bottom) nascent transcripts between DMSO and dTAGV-1 following a 120-min 4sU labeling period during mitotic exit. The top five categories are shown for each analysis, with the P values derived using the ‘elim’ algorithm and a Fisher’s exact test using TopGO. d, Gene set enrichment analysis of genes ranked by log2FC from differential gene expression analysis of nascent transcripts with 120-min 4sU labeling period. Two ‘Hallmark’ gene sets are shown. e, Morphology of NIPBL-D7 cells treated with DMSO or dTAGV-1 for 72 h. Cells were stained with phalloidin (actin; green) and DAPI (DNA; blue). Two biological replicates were performed with similar outcomes. f, Line plots showing the mean and standard error of FC in FPKM over t = 0 min for each timepoint for NIPBL-D7 cells treated with DMSO or dTAGV-1 during mitotic exit (approach A, Extended Data Fig. 7a). Significantly decreased (top) and increased (bottom) genes identified from differential analysis of nascent transcripts with 120-min 4sU labeling period (approach B, Extended Data Fig. 7a) are shown. Some significantly changed transcripts were not detected with the shorter labeling period and are excluded. g, Box plots showing the FPKM values for nascent transcripts from 15-min 4sU labeling period at late timepoints following treatment of NIPBL-D7 cells with DMSO or dTAGV-1 during mitotic exit (approach A, Extended Data Fig. 7a). Significantly decreased (top) and increased (bottom) genes identified from differential analysis of nascent transcripts with 120-min 4sU labeling period (approach B, Extended Data Fig. 7a) are shown. Kruskal–Wallis tests were performed first, before pairwise comparisons were made using two-sided Wilcoxon rank-sum tests. The center line marks the median, the box shows the IQR and the whiskers extend to 1.5 × IQR. h, Comparison of differential analysis of nascent transcripts with a 120-min 4sU labeling period performed with DMSO or dTAGV-1 treatment administered during mitotic exit (‘mitosis’; mitotic exit approach A, Extended Data Fig. 7a) or 4 h post release from nocodazole (‘mid-G1’; mid-G1 approach B, Extended Data Fig. 7c). Shown are all significantly changed genes during mitotic exit, ranked by mitosis log2FC. Nascent transcripts that also met the criteria for significance (|log2FC| > 0.585 and Padj < 0.1) with the mid-G1 approach are denoted with a black line. i, Example line plots showing the mean FPKM and standard error for nascent transcripts from 15-min 4sU labeling period for NIPBL-D7 cells treated with DMSO or dTAGV-1 during mitotic exit (‘mitosis’; mitotic exit approach B, Extended Data Fig. 7a) or 4 h post release from nocodazole (‘mid-G1’; mid-G1 approach A, Extended Data Fig. 7c). SOX9 is an example of a gene differentially expressed only during mitotic exit, SNAI2 is differentially expressed during both mitotic exit and mid-G1, and ASNS is nonsignificant during both mitotic exit and mid-G1. For ‘mitosis’, the x axis is time post release from nocodazole, while for ‘mid-G1’, the x axis is time post addition of DMSO/dTAGV-1. Additional examples can be found in Extended Data Fig. 7g. FDR, false discovery rate; NES, normalized enrichment score.
Based on the reported ‘waves’ of transcriptional activation that occur during the M–G1 transition25,26,27, we used the mitotic exit approach A to understand the behavior of differentially expressed genes during this period. In control conditions, genes decreased by NIPBL depletion progressively increased expression beginning at 30–60 min post release from nocodazole (Fig. 3f, top), at which point cells are transitioning from mitosis to G1 (Extended Data Fig. 5a). NIPBL depletion did not blunt this initial activation, but activation after 60 min was weakened (Fig. 3f, top). Genes increased with NIPBL depletion had a stronger spike with dTAGV-1 at 60–90 min compared with the DMSO condition (Fig. 3f, bottom). When looking at longer timepoints following release, decreased genes did not recover their expression, whereas increased genes returned to DMSO-like levels (Fig. 3g).
Lastly, we asked whether these alterations in gene expression were unique to mitotic exit. In asynchronous cells, fewer genes were dysregulated with NIPBL depletion compared with mitotic exit (Fig. 3h and Extended Data Fig. 7c–e). This may be due to the distinct severity of the consequences of NIPBL depletion on the establishment versus maintenance of genome organization. Many genes were disrupted in both mitotic exit and G1 (Fig. 3h), with similar gene ontology categories observed (Extended Data Fig. 7f). However, some genes show distinct responses to NIPBL depletion during mitotic exit and G1 (Fig. 3i and Extended Data Fig. 7g). For example, SOX9 and SNAI2 were significantly affected by NIPBL depletion during mitotic exit, but only SNAI2 was also significantly perturbed in G1 (Fig. 3i). Overall, these findings suggest that NIPBL is essential for lineage-defining genes to reach their full transcriptional potential during mitotic exit.
A highly connected enhancer neighborhood at dependent genes
Despite NIPBL depletion causing substantial disruption to genome organization during mitotic exit, only a fraction of the transcriptome was affected (Fig. 3a), leading us to ask what makes these genes sensitive. We compared the nonsignificant and significant genes using both all genes and by sub-sampling nonsignificant genes using nearest neighbor analysis based on gene and enhancer density, and fragments per kilobase million (FPKM) (Extended Data Fig. 8a,b). Both decreased and increased genes were more likely to loop to typical- and super-enhancers (Extended Data Fig. 8c) and exhibited stronger contacts with super- and typical-enhancers within 1 megabase (Mb) compared with nonsignificant genes (Fig. 4a,b and Extended Data Fig. 8d). For decreased genes, the heightened enhancer–promoter connectivity and its dependence on NIPBL was restricted to within 500 kb (Fig. 4b and Extended Data Fig. 8e). The strength and NIPBL-dependence of contacts between genes and their enhancers was evident at individual genes that were sensitive to NIPBL depletion (Fig. 4c and Extended Data Fig. 9a,b). These findings indicate that significantly decreased genes show a higher degree of contact with nearby enhancers and their ability to contact these enhancers is more dependent on NIPBL.
a, Off-diagonal pile-up analysis of Hi-C data at a 10-kb resolution from DMSO- and dTAG-treated cells collected 90 min post release from nocodazole. Shown is the log-transformed mean observed/expected contact frequency between promoter-proximal regions (TSS −1 kb/+500 bp) and all neighborhood super-enhancers called from H3K27ac ChIP–seq with a TSS exclusion value of 2,500 bp and a stitch value of 12,500 bp. Enhancers within 10 kb of promoter-proximal regions were excluded. b, Loess-smoothened relationship with 95% confidence interval at a 5-kb resolution between distance and contact frequency between all enhancers (typical and super) and promoter-proximal regions at 90 min post release from nocodazole. Only nearest neighbors are shown for the nonsignificant genes. c, MYC is an example of a gene that is significantly decreased in response to NIPBL depletion during mitotic exit. Shown are Hi-C heatmaps (top row: DMSO; bottom row: dTAG), sequencing-depth normalized BigWig tracks of read pairs whereby at least one read contains a T→C conversion (that is, nascent reads) and Virtual 4C anchored at MYC promoter-proximal regions (TSS −1 kb/+500 bp) for both DMSO- and dTAG-treated samples at t = 30 min, 60 min, 90 min and 120 min. The marked TE and SE sites are the same across all plots. The arrow on the Hi-C heatmaps marks the MYC TSS, and the gray bar on the tracks below marks the gene location. d, On-diagonal pile-up analysis of Hi-C data at a 10-kb resolution from DMSO- and dTAG-treated cells collected 90 min post release from nocodazole. Pile-up was centered on the TSS, with all genes oriented in the same direction (right-facing). NN, nearest neighbors; SE, super-enhancers; TE, typical-enhancers.
Local genome conformation at dependent genes
Genes can function as barriers to loop extrusion38,39, leading to insulation separating regions upstream and downstream of a TSS. Curiously, we noticed that NIPBL-dependent genes had a degree of permissiveness to this insulation (Extended Data Fig. 10a), with stronger chromatin contacts between regions up- and downstream of TSSs compared with nonsignificantly changed genes (Fig. 4d). Furthermore, decreased genes had NIPBL-dependent stripe-like features emanating outwards from the TSS (Fig. 4d). The contact domains on either side of the TSS were perturbed by NIPBL depletion for all genes, but the cross-TSS contacts between the domains were lost only at decreased genes (Extended Data Fig. 10b–d). A similar cross-TSS contact pattern has previously been recapitulated genome-wide by WAPL depletion39, suggesting that cohesin removal by WAPL may normally limit the processivity of cohesin across the TSS. Indeed, we found that the ratio of NIPBL to WAPL, but not NIPBL to PDS5, was higher at decreased genes (Extended Data Fig. 10e). The imbalance of NIPBL and WAPL may reflect distinct cohesin dynamics at these loci.
Enhanced cohesin dynamics at a given locus may promote the formation of multi-way contacts between a gene and numerous enhancers40,41,42. Indeed, we found that both super- and typical-enhancers near genes sensitive to NIPBL depletion showed strong, NIPBL-dependent engagement with enhancers on the opposite side of the TSS (Extended Data Fig. 10f,g). Thus, we conclude that NIPBL-mediated loop extrusion regulates gene activity by facilitating a combination of enhancer–promoter and enhancer–enhancer contacts across 3D space, possibly by enabling the formation of higher-order structures.
NIPBL regulates neuronal and cardiomyocyte lineage genes
A surprising aspect of recent studies on cohesin’s role in gene regulation is that relatively few genes are affected by acute depletion of cohesin17,21. Despite this, several studies have linked cohesin to regulation of cell-type-specific genes, including in neurons20, macrophages19 and mouse embryonic stem cells43,44,45,46. Given these observations and our data in hTERT RPE-1 cells showing that NIPBL activates lineage-defining genes, we wanted to test whether cohesin may have had a more widespread impact on gene regulation if we considered its cell-type-specific contributions, and whether NIPBL-dependent genes showed a similar unique promoter conformation across lineages. To test this, we engineered WTC11 hiPSCs with the same NIPBL degron system we used in hTERT RPE-1 cells (Extended Data Fig. 1a, mGreenLantern). HA-NIPBL and MAU2 were both rapidly depleted upon dTAGV-1 treatment of hiPSCs and hiPSC-derived cardiomyocytes or excitatory neurons (Fig. 5a,b and Extended Data Fig. 10h). NIPBL depletion resulted in differential expression of 602 genes in cardiomyocytes and 580 genes in neurons, with the vast majority of these downregulated (Fig. 5c). By comparing the differentially expressed genes between hTERT RPE-1, cardiomyocytes and neurons, we found that dysregulated genes were largely cell-type-specific (Fig. 5d). Collectively, NIPBL depletion led to altered expression of 1,392 genes, with 78.4% (1,091 of 1,392) of these being significant in only one of the three lineages. Gene ontology analysis of the genes unique to either cardiomyocytes or neurons showed enrichment of categories related to the function of these cell types (Fig. 5e). Based on this, we conclude that NIPBL is essential for the accurate expression of lineage-defining genes across different cell types.
a, hiPSCs engineered for acute depletion of NIPBL were treated with DMSO or dTAGV-1 for 4 h, either as undifferentiated cells or following differentiation into cardiomyocytes or excitatory neurons. Two biological replicates were performed for undifferentiated cells and neurons with similar outcomes, and one biological replicate for cardiomyocytes. b, Immunofluorescence of hiPSC-derived neurons following treatment with DMSO or dTAGV-1 for 4 h. NeuN and TUBB3 are commonly used markers for neuronal differentiation. Two biological replicates were performed with similar outcomes. c, Volcano plot from differential expression analysis of nascent transcripts between DMSO- and dTAGV-1-treated hiPSC-derived cardiomyocytes and neurons based on a 120-min 4sU labeling period. Significantly changed (|log2FC| > 0.585 and Padj < 0.1) nascent transcripts are highlighted. P values were calculated from two-sided Wald tests using two independent replicates and adjusted (Padj) for multiple testing using the Benjamini–Hochberg method. d, Heatmap showing FC (log2FC) of differentially expressed nascent genes (2-h 4sU SLAM-seq) with NIPBL depletion from RPE-1 (R), hiPSC-derived cardiomyocytes (C) and hiPSC-derived neurons (N). Genes are grouped by the cell types in which changes are significant (|log2FC| > 0.585 and Padj < 0.1). e, Biological process gene ontology analysis of genes that are significantly changed only in hiPSC-derived cardiomyocytes or only in hiPSC-derived neurons. The top five categories are shown for each analysis, with the P values derived using the ‘elim’ algorithm and a Fisher’s exact test using TopGO. f, Off-diagonal pile-up (left) and on-diagonal pile-up (TSS-centered; right) for nonsignificant and decreased genes with NIPBL depletion from iPSC-derived neurons (top) and cardiomyocytes (bottom). Off-diagonal pile-up is of contacts between super-enhancers within 1 Mb of the TSS and promoter-proximal regions. Shown are pile-ups for all nonsignificantly changed and attribute-matched nearest neighbors. g, Hi-C heatmap of the CLSTN2 locus in DMSO- and dTAG-treated hiPSC-derived neurons (top) and the CASZ1 locus in DMSO- and dTAG-treated hiPSC-derived cardiomyocytes (bottom). Below each heatmap are a selection of genes from the region, super-enhancers, typical-enhancers and Virtual 4C anchored at CLSTN2 or CASZ1 promoter-proximal regions (TSS −1 kb/+500 bp) for both DMSO- and dTAG-treated samples. The arrow on the Hi-C heatmaps marks the CLSTN2 or CASZ1 TSS. CLSTN2 expression is decreased with NIPBL depletion from neurons, while CASZ1 expression is decreased with NIPBL depletion from cardiomyocytes.
We next explored whether NIPBL-sensitive genes in hiPSC-derived cells also had the genomic organization we observed in hTERT RPE-1. Based on enhancer data from published hiPSCs-derived cells47,48, we found that decreased genes in neurons and cardiomyocytes showed greater contact with neighborhood super- and typical-enhancers than nonsignificant genes, with these contacts dependent on NIPBL (Fig. 5f, left, and Extended Data Fig. 10i–k). Additionally, we observed reduced TSS insulation at sensitive genes, but not insensitive genes (Fig. 5f, right). Compared with hTERT RPE-1 cells, NIPBL-sensitive genes in hiPSC-derived cells had strong directionality (towards gene) of stripes from the TSS, but these stripes were still unique to the significant genes (Fig. 5f, right). Surprisingly, in both the neurons and cardiomyocytes the TSS insulation remained somewhat stable in the absence of NIPBL, but the stripes were more strongly perturbed (Fig. 5f, right). This may be a consequence of more stable and NIPBL-independent genome organization in post-mitotic cells. Regardless, genes that were perturbed had strong spatial organization that is sensitive to NIPBL depletion (Fig. 5g). In summary, we have demonstrated that NIPBL promotes the formation of unique 3D genomic structures at certain cell identity genes that is necessary for their expression.
Discussion
Here we characterized the role of cohesin dynamics in regulating 3D genome organization and transcription by targeting the cohesin accessory protein NIPBL. We found that loop extrusion facilitated by NIPBL is more critical to the establishment of chromatin loops upon mitotic exit than their maintenance in asynchronous cells. We also identified a subset of chromatin loops that were persistent when NIPBL was depleted from asynchronous cells but failed to form when NIPBL was depleted during mitotic exit. Additionally, NIPBL depletion enabled us to characterize the role of 3D genome organization in transcription activation during mitotic exit. Consistent with a recent preprint49, we find that the majority of cohesin-anchored loops are perturbed with NIPBL depletion during mitotic exit. Further, NIPBL preferentially activated lineage-defining genes during mitotic exit, with sensitive genes having a unique spatial organization. We extended our observations to hiPSC-derived cardiomyocytes and neurons, finding that NIPBL depletion results in perturbed expression of distinct, cell-type-specific sets of genes. Overall, our findings provide comprehensive insight into the role of NIPBL in transcription regulation and genome organization across multiple contexts.
Consistent with our study, earlier work employing conditional knockout of NIPBL from mouse hepatocytes found that the loss of NIPBL resulted in reduced association of cohesin with chromatin, extensive and widespread disruption to topologically associating domains and chromatin loops, and modest changes in gene expression16. However, the slow time course (10 d) of conditional knockouts is limiting for studying the more acute role of NIPBL in regulating cohesin dynamics. Indeed, we found that depletion of NIPBL from cells during both steady-state and mitotic exit revealed a differential role of NIPBL in genome organization ‘maintenance’ versus ‘formation’. NIPBL depletion during mitotic exit was more severe than when NIPBL was depleted from asynchronous cells. We also identified chromatin loops that are dependent on RAD21 and CTCF but differed in their persistence when NIPBL was depleted. Of particular interest are the mixed-dependency loops, which are anchored in repressive chromatin and were highly dependent on NIPBL for formation during mitotic exit. In contrast to the findings that chromatin loops can persist for only 10–20 min (refs. 8,9), this set of chromatin loops may represent a distinct type of loop whose longevity is greater than other loops of a similar size. Previous research has shown that cohesinSTAG1 is more stable on chromatin than cohesinSTAG2 (ref. 33). In our study, cohesinSTAG1 or cohesinSTAG2 were equally capable of forming loops in the mixed-dependency cluster, but cohesinSTAG1 was critical for the persistence of these loops upon NIPBL depletion. This difference in persistence and STAG1-dependence may be due to SMC3 acetylation, which is substantially higher at the anchors of the mixed-dependency cluster than the maximum-dependency cluster, with this modification suggested to stabilize cohesinSTAG1 during the G1 phase of the cell cycle33. Given the association of the mixed-dependency cluster with repressive chromatin, it is worth considering that SMC3 acetylation may occur preferentially here. Lastly, it is also possible that an unidentified factor may also promote cohesin processivity during G1, with this factor potentially compensating to maintain certain chromatin loops when NIPBL is depleted.
Three-dimensional genome organization was first proposed as a mechanism to enable transcriptional regulation, specifically by facilitating distal regulatory elements coming into close spatial proximity with gene promoters7. However, the role of cohesin-dependent chromatin looping in transcriptional regulation has come into doubt over recent years, with multiple studies demonstrating that depletion of the 3D genome machinery has only minor effects on gene expression17,18,19,20,21. Despite this, studies employing cohesin disruption in distinct systems have also found that the limited number of genes affected by cohesin perturbation have lineage-specific functions19,20, including in polymorphonuclear neutrophils15 and mouse embryonic stems cells43,44. Collectively these works suggest that cohesin may play particularly important roles in regulating cell-type-specific gene expression. Indeed, we depleted NIPBL across different cell types, leading to the identification of >1,000 cell-type-specific genes that are regulated by NIPBL. This suggests that cohesin-mediated genome organization may modulate thousands of genes genome-wide, but is responsible for regulating only a few hundred of these in any given cell type. These findings are particularly interesting when considering that Cornelia de Lange syndrome, which is largely driven by NIPBL mutations, is a multi-system disorder that manifests as distinct, tissue-specific phenotypes50.
Enhancer–promoter connectivity is increasingly viewed as a more complex, higher-order structure than a single enhancer proximal to a single promoter. This complexity may include condensates or hubs involving multiple distal elements51. While Hi-C is mostly ineffective for detecting these complex structures, we did observe a distinct local 3D genome architecture at NIPBL-dependent genes. Aspects of this unique organization resemble ‘promoter–enhancer stripes’ that were recently observed in single-cell micro-C data and proposed to facilitate transcriptional bursting52. We posit that cohesin dynamics at sensitive genes are enhanced by higher levels of NIPBL, which may facilitate ongoing loop extrusion at certain genes despite collisions with RNA polymerase, which are thought to be responsible for gene-centric insulation39. Furthermore, this unique organization may also reflect the formation of higher-order 3D genome structures, including simultaneous association of enhancers both up- and downstream of sensitive genes with their promoters. Indeed, an imaging-based approach, ORCA, has recently been used to demonstrate multi-way contacts by SOX9 (ref. 41), which we found to be sensitive to NIPBL depletion during mitotic exit. Similarly, a three-way ‘kissing’ model for cohesin-dependent transcriptional bursting has been found to regulate Sox2 expression22. Further, recent evidence has found CTCF-dependent multi-way hubs form at tissue-specific genes to help promote their regulation42. While this still begs the question as to why the whole genome is organized by cohesin, with only lineage-defining genes carrying these unique structural features, our findings demonstrate a definitive role of NIPBL and cohesin in regulating cell-type-specific gene expression programs through this organization.
Methods
Cell culture
hTERT RPE-1 cells were obtained from ATCC (CRL-4000) and maintained in DMEM/F12 (Thermo Fisher cat. no. 11330-057) supplemented with 10% FBS (Gibco cat. no. A5256701) and 1% penicillin-streptomycin (Gibco cat. no. 15140122). NIH/3T3 cells were obtained from ATCC (CRL-1658) and maintained in DMEM with 4.5 g l−1 glucose, L-glutamine and sodium pyruvate (Corning cat. no. 10-013-CV) supplemented with 10% FBS (Gibco cat. no. A5256701) and 1% penicillin-streptomycin (Gibco cat. no. 15140122). WTC11 hiPSCs containing inducible neurogenin-2 (iNgn2)53 were a gift from Y. Shen47. Before differentiation, these cells were maintained on Cultrex-coated plates in either Essential 8 Flex Medium (Gibco cat. no. A2858501) or mTeSR Plus (StemCell Technologies cat. no. 100-0276).
DMSO (Sigma-Aldrich cat. no. D2650) or 500 nM dTAGV-1 (Tocris cat. no. 6914) was added to cells for the indicated periods of time.
Engineering of dTAG and STAG1/2 knockout cell lines was performed using CRISPR–Cas9. See Supplementary Information for detailed methods. Single guide RNA and genotyping primers are shown in Supplementary Table 1.
Differentiation of human induced pluripotent stem cells into neurons or cardiomyocytes
hiPSC)s engineered for acute depletion of NIPBL (iPSC-NIPBL) were differentiated into ventricular cardiomyocytes using the STEMdiff Ventricular Cardiomyocyte Differentiation Kit (StemCell Technologies cat. no. 05010) and according to the manufacturer’s instructions. Beating cardiomyocytes were treated on Day 15 for 4 h with DMSO or dTAGV-1 and collected for immunofluorescence, western blot, SLAM-seq or Hi-C.
Differentiation of iPSC-NIPBL cells into glutamatergic neurons was performed based on SMAD inhibition54. iPSC-NIPBL cells were seeded as single cells at ~18,000 cells per cm2 in medium supplemented with ROCKi (Y-27632; Tocris cat. no. 1254), with medium replaced the following day. At 3 d following initial plating (Day 0), medium was replaced with Neural Induction Medium (DMEM/F12 + GlutaMax (Gibco cat. no. 10565-018), Neurobasal-A (Gibco cat. no. 10888-022), 1 × NEAA (Gibco cat. no. 11140-050), 1 × B-27 (Gibco cat. no. 17504-044), 1 × N-2 (Gibco cat. no. 17502-048), 3 μM CHIR99021, 10 μM SB431542, 5 μM DMH-1, 100 μg ml−1 dibutyryl cAMP and 2 μg ml−1 laminin). Neural Induction Medium was completely replaced every 2 d. On Day 10, cells were trypsinized and replated separately for eventual DMSO/dTAG treatment. Cells were replated in Neural Induction Medium supplemented with ROCKi, with medium replaced the following day. Neural Induction Medium was ~50% replaced every 2 d. On Day 15, medium was replaced with Neural Maturation Medium (BrainPhys Neuronal Medium (StemCell Technologies cat. no. 05790), 1 × B-27 (Gibco cat. no. 17504-044), 1 × N-2 (Gibco cat. no. 17502-048), 20 ng ml−1 BDNF, 20 ng ml−1 GDNF, 100 μg ml−1 dibutyryl cAMP and 2 μg ml−1 laminin). Neural Maturation Medium was completely replaced every 2 d. Differentiated neurons were treated on Day 21 for 4 h with DMSO or dTAGV-1 and collected for immunofluorescence, western blot, SLAM-seq or Hi-C.
Whole-cell extracts for western blot
Cells were washed in ice-cold Dulbecco’s phosphate-buffered saline (DPBS; Gibco cat. no. 14190144) and collected, and cell pellets resuspended in Kischkel buffer (50 mM Tris pH 8.0, 150 mM NaCl, 5 mM EDTA, 1% Triton X-100) supplemented with 1 × cOmplete, EDTA-free Protease Inhibitor Cocktail (PIC; Roche cat. no. COEDTAF-RO) and 1 mM PMSF. Cells were sonicated at 25% for 15 s using an Active Motif Epishear Probe Sonicator. Debris was removed by centrifugation at 20,000g for 10 min at 4 °C. Total protein was quantified using 1 × Protein Assay Reagent (BioRad cat. no. 5000006) against a BSA standard curve. Samples were typically balanced in a new tube to a final concentration of 1 mg ml−1 using Kischkel buffer and Laemmli buffer (300 mM Tris pH 6.8, 32% glycerol, 8% SDS, bromophenol blue) supplemented with BME, before incubation at 95 °C for 5 min.
Chromatin fractionation
Cells were subjected to chromatin fractionation as described in ref. 55. Briefly, cells were washed with DPBS, collected and resuspended in ice-cold Buffer A (10 mM HEPES pH 7.9, 10 mM KCl, 1.5 mM MgCl2, 0.34 M sucrose, 10% glycerol) supplemented with 1 mM DTT, 1 × PIC and 1 mM PMSF. Triton X-100 was added to a final concentration of 0.1%, and cells were incubated on ice for 8 min. The reaction was centrifuged at 1,700g for 5 min at 4 °C. The supernatant (fraction S1) was transferred to a new tube and clarified by centrifugation (20,000g for 10 min at 4 °C), resulting in the cytosolic fraction (fraction S2). The pelleted nuclei (fraction P1) were gently washed with Buffer A, resuspended in Buffer B (3 mM EDTA, 0.2 mM EGTA) supplemented with 1 mM DTT, 1 × PIC and 1 mM PMSF, and incubated on ice for 30 min. The reaction was centrifuged at 1,700g for 5 min at 4 °C, with the resulting supernatant (fraction S3; nucleoplasm) transferred to a new tube. The insoluble pellet (fraction P3; chromatin) was gently washed with Buffer B, resuspended in 1 × Laemmli Buffer supplemented with BME and sonicated at 25% for 15 s using an Active Motif Epishear Probe Sonicator. Laemmli buffer supplemented with BME was added to 1× to fractions S2 and S3. All samples were incubated at 95 °C for 5 min.
Western blot
Details of western blotting can be found in Supplementary Information. For western blotting of NIPBL, rabbit polyclonal antibodies from Bethyl (cat. no. A301-779A) or Proteintech (cat. no. 18792-1-AP) were used independently at 1:1,000 or combined at 1:1,000 each. Other primary antibodies used for western blotting are anti-RAD21 C terminus (Bethyl cat. no. A300-080A; 1:1,000), anti-RAD21 N terminus (Abcam cat. no. ab154769; 1:1,000), anti-SMC3 (Bethyl cat. no. A300-060A; 1:25,000), anti-α-Tubulin (Millipore cat. no. CP06100UG; 1:5,000), anti-Histone H3 (Cell Signaling cat. no. 4499; 1:2,000), anti-HA (Cell Signaling cat. no. 3724; 1:1,000), anti-MAU2/SCC4 (Abcam cat. no. ab183033; 1:1,000), anti-STAG1 N terminus (Invitrogen cat. no. PA5-57115; 1:1,000), anti-STAG1 C terminus (Invitrogen cat. no. MA5-27807; 1:1,000), anti-STAG2 N terminus (Cell Signaling cat. no. 5882; 1:1,000), anti-STAG2 C terminus (Santa Cruz cat. no. sc-81852; 1:500) and anti-CTCF (Active Motif cat. no. 61932; 1:1,000). HRP-conjugated primary antibody used was anti-GAPDH (Cell Signaling cat. no. 8884; 1:5,000). HRP-conjugated secondary antibodies used were goat anti-mouse IgG (Jackson cat. no. 115-035-071; 1:5,000) and goat anti-rabbit IgG (Invitrogen cat. no. 31463; 1:5,000).
Cell cycle synchronization
hTERT RPE-1 cells were synchronized to prometaphase using a double thymidine block and nocodazole as previously described27. Briefly, cells were treated with 2 mM thymidine (Sigma-Aldrich cat. no. T1895) for 14 h, released for 10 h, then treated again with 2 mM thymidine for 14 h. Cells were then released for 8 h before treatment with 20 ng ml−1 nocodazole (Sigma-Aldrich cat. no. M1404) for 2 h. For experiments aimed at studying post-mitotic transcriptional reactivation, treatments (DMSO or dTAGV-1) were initiated simultaneously with nocodazole. Mitotic cells were then collected by shake-off and washed once, before being released into fresh media with or without treatment. For experiments aimed at determining gene expression changes within the G1 phase, cells were released from nocodazole for 4 h before addition of DMSO or dTAGV-1. For all experiments, cells were collected at the stated timepoints post nocodazole release for propidium iodide staining, western blotting, SLAM-seq or Hi-C.
Cell cycle analysis by propidium iodide staining
Cells treated with DMSO or dTAGV-1 were collected at the stated times, immediately fixed in ice-cold 90% ethanol and incubated at −20 °C for at least 2 h. Cells were brought to room temperature and washed once with DPBS. Fixed cells were then resuspended in propidium iodide staining buffer (10 μg ml−1 propidium iodide (Abcam cat. no. ab14083), 100 μg ml−1 RNase (Thermo Scientific cat. no. EN0531), 2 mM MgCl2, DPBS) and incubated overnight at 4 °C. Flow cytometry analysis was performed using either a BD LSR II or BD FACSymphony A3 in the Salk Flow Cytometry Core. An example of the gating strategy used for these experiments is shown in Extended Data Fig. 2e.
Cell imaging
hTERT RPE-1 cells were grown on glass coverslips for 24 h before being treated with DMSO or dTAGV-1 for 72 h. Coverslips were transferred to a 24-well plate, washed twice with DPBS and fixed in 4% methanol-free formaldehyde for 15 min at room temperature. Following two washes with DPBS, cells were permeabilized with 0.1% Triton X-100 for 15 min at room temperature and then washed once in DPBS. Cells were then stained using 1 × Alexa Fluor 488 Phalloidin (Invitrogen cat. no. A12379) in DMSO for 45 min at room temperature. DAPI (Thermo Scientific cat. no. D1306) was then added to a final concentration of 0.35 ng μl−1, incubating for a further 15 min at room temperature. Coverslips were washed twice with DPBS and then mounted to slides using Prolong Diamond Antifade Mountant (Invitrogen cat. no. P36965). Slides were then imaged using the Olympus VS-120 Virtual Slide Scanning Microscope.
For cardiomyocytes, iPSC-NIPBL cells were plated directly on a Cultrex-coated Ibidi Chamber Slide before differentiation. For neurons, cells were plated on a Cultrex-coated Ibidi Chamber Slide on Day 10 of differentiation. For both cardiomyocytes (Day 15) and neurons (Day 21), cells were washed twice with DPBS and fixed in 4% methanol-free formaldehyde for 20 min at room temperature. Cells were then permeabilized with 0.2% Triton X-100 for 15 min at room temperature, washed twice with DPBS and blocked in 1% BSA in DPBS for 60 min at room temperature. Cardiomyocytes were incubated in Cardiac Troponin T (cTnT) antibody (Proteintech cat. no. 68300-1-Ig; 1:400) in 1% BSA in DPBS overnight at 4 °C, washed three times with DPBS, then incubated with goat anti-mouse conjugated to Alexa Fluor 488 (Invitrogen cat. no. A-11001; 1:1,000). Neurons were incubated in anti-FOX3 (NeuN) antibody (BioLegend cat. no. 834502; 1:1,000) in 1% BSA in DPBS overnight at 4 °C, washed three times with DPBS, then incubated with goat anti-mouse conjugated to Alexa Fluor 488 (Invitrogen cat. no. A-11001; 1:1,000). Neurons were subsequently washed three times with DPBS and incubated with anti-Tubulin β3 (TUBB3) antibody conjugated to Alexa Fluor 594 (BioLegend cat. no. 801207; 1:1,000). Both cardiomyocytes and neurons were then washed three times with DPBS, incubated in 500 ng ml−1 DAPI in DPBS at room temperature for 15 min and washed three times with DPBS, and coverslips were mounted onto the slides using Prolong Diamond Antifade Mountant (Invitrogen cat. no. P36965). Slides were then imaged using the Zeiss Axioscan 7 Slide Scanner.
ChIP–seq
For RAD21 ChIP–seq with NIPBL depletion, ChIP–seq in NIPBL-A2 and ChIP–seq in NIPBL-D7 were treated as two biological replicates. Chromatin preparation for ChIP–seq using antibodies against CTCF, RAD21, PDS5A, WAPL, STAG1 and STAG2 was performed by fixing ~10 million cells with 1% methanol-free formaldehyde (Thermo Scientific cat. no. 28908) for 10 min, then quenching with 125 mM glycine for an additional 10 min. For HA-NIPBL ChIP–seq, ~10 million cells were fixed with 2 mM DSG (Santa Cruz cat. no. sc-285455) for 45 min, washed three times with DPBS, fixed further with 1% methanol-free formaldehyde for 10 min and quenched with 125 mM glycine for an additional 10 min. Cells were washed twice in DPBS, collected, flash-frozen and stored at −80 °C until proceeding. Cells were then lysed for 15 min on ice using formaldehyde lysis buffer (FALB; 50 mM HEPES pH 7.5, 140 mM NaCl, 1 mM EDTA, 1% Triton X-100) supplemented with 1% SDS and 1 × PIC, and sonicated for 9 min (10 s on/10 s off) at 25% power using an Active Motif Epishear Probe Sonicator. Debris was removed by centrifugation at 21,000g for 10 min at 4 °C. For immunoprecipitation, 30 μl of Protein A Dynabeads (Invitrogen cat. no. 10002D) were washed three times with FALB (no SDS) and blocked using 10 μg of BSA. Antibody and beads were incubated together for 6–8 h, rotating at 4 °C, after which the beads were washed three times in FALB (no SDS). An ‘input’ sample of the prepared chromatin was taken out before the remaining chromatin was diluted out tenfold with FALB to lower the SDS concentration, and antibody-bound beads were added, incubating overnight, rotating at 4 °C. Following incubation, beads were washed once for 5 min, rotating at 4 °C, in each of the following ice-cold buffers: high-salt wash buffer (20 mM Tris pH 8.0, 500 mM NaCl, 2 mM EDTA, 0.1% SDS, 1% Triton X-100), lithium chloride wash buffer (10 mM Tris pH 8.0, 250 mM LiCl, 1 mM EDTA, 1% IGEPAL) and TE wash buffer (10 mM Tris pH 8.0, 1 mM EDTA, 50 mM NaCl). Beads were incubated in elution buffer (50 mM Tris pH 8, 1 mM EDTA, 1% SDS) for 1 h at 65 °C, with the supernatant transferred to a new tube. ‘Input’ and chromatin immunoprecipitation (ChIP) samples were supplemented with NaCl to 200 mM and 40 μg of proteinase K (NEB cat. no. P8107S), and incubated for 30 min at 55 °C and then overnight at 65 °C. The following day, samples were treated with RNAse A (Thermo Scientific cat. no. EN0531) for 2 h at 37 °C, before purification using the Zymo ChIP DNA Clean & Concentrator kit (cat. no. D5205).
For experiments that included a 10% spike-in of NIH/3T3 cells (RAD21 ChIP), the NIH/3T3 cells were fixed, then lysed and sonicated using the required buffer and conditions. Chromatin prepared from the NIH/3T3 cells was then combined with the chromatin prepared from the engineered hTERT RPE-1 cells (~10% NIH/3T3 + ~90% hTERT RPE-1 by cell number) before dilution and immunoprecipitation.
For CTCF and HA-NIPBL ChIP–seq, ‘input’ and ChIP samples were subjected to a dual-sided size selection using AMPure XP beads (Beckman Coulter cat. no. A63880) before sequencing libraries were prepared using 2S Plus DNA Library Kit (IDT cat. no. 10009878). For RAD21, WAPL, PDS5A, STAG1 and STAG2 ChIP–seq, sequencing libraries of input and ChIP samples were prepared using Diagenode Microplex v3 (cat. no. C05010001) and unique dual indexes (Set I: cat. no. C05010008; Set 2: cat. no. C05010009). Final libraries were quantified by quantitative PCR (qPCR) using KAPA Illumina Library Quantification Kit (Roche cat. no. KK4854), with Agilent TapeStation used to estimate individual library sizes. Paired-end 50-base pair (bp) or 100-bp sequencing was carried out by the Salk Next-Generation Sequencing Core on an Illumina NextSeq 2000 or NovaSeqX. The number of total and deduplicated sequencing reads is shown in Supplementary Table 2. For ChIP–seq analysis, see Supplementary Information.
Antibodies
Abflex anti-CTCF (Active Motif cat. no. 91285; 4 μl per ChIP), anti-RAD21 (Abcam cat. no. ab992; 5 μl per ChIP), anti-STAG1 (Invitrogen cat. no. PA5-57115; 5 μl per ChIP), anti-STAG2 (Cell Signaling cat. no. 5882; 5 μl per ChIP), anti-SCC-112/PDS5A (Bethyl cat. no. A300-089A; 10 μl per ChIP), anti-HA (Cell Signaling cat. no. 3724; 6 μl per ChIP) and anti-WAPL (Proteintech cat. no. 16370-1-AP; 5 μl per ChIP).
In situ Hi-C
All Hi-C experiments were performed using the Arima Genomics Hi-C+ Kit, according to the manufacturer’s instructions. Hi-C experiments for hTERT RPE-1 cells were performed in independent clones presented, which were used as two biological replicates (that is, NIPBL-A2 and NIPBL-D7; RAD21-B1 and RAD21-B3; STAG1-A7 and STAG1-C5; STAG2-A2 and STAG2-B8), with the exception of the mitotic synchronization Hi-C experiments which were done as two biological replicates both using the NIPBL-D7 cell line. Briefly, cells were resuspended in DPBS and crosslinked with 2% methanol-stabilized formaldehyde (Fisher Scientific cat. no. F79500) for 10 min at room temperature. Stop Solution 1 was added to quench the formaldehyde, incubating first for 5 min at room temperature and then for 15 min on ice. Fixed cells were centrifuged, washed once with DPBS and then flash-frozen for storage at −80 °C. Cells were lysed and nuclei permeabilized to allow for restriction digest of DNA before biotin fill-in ligation of proximal DNA ends. Samples were decrosslinked, digested with proteinase K and DNA purified using Solid Phase Reverse Immobilization (SPRI) beads (Sera-Mag SpeedBead Carboxylate-Modified (E7) Magnetic Particles, Cytiva cat. no. 45152105050250). Purified DNA was sonicated to ~400 bp using a Covaris M220 with 50-W power, 10% duty factor and 200 cycles per burst for 78 s, and then size-selected using SPRI beads. Based on quantification of DNA by Qubit, library preparation was either performed by KAPA HyperPrep (Roche cat. no. KK8502; for high input) based on Arima-provided instructions, or using the 2S Plus DNA Library Kit (IDT cat. no. 10009878; for low input).
For KAPA HyperPrep, biotin-labeled DNA was bound to Arima Enrichment Beads and used as input for end repair, dA-tailing and ligation of TruSeq DNA Single Indexes (Illumina cat. no. 20015960). Libraries were first quantified by qPCR using the KAPA Illumina Library Quantification Kit (Roche cat. no. KK4854) to estimate the required number of PCR cycles, before performing PCR amplification and SPRI bead clean-up. The final libraries were again quantified by qPCR using KAPA Illumina Library Quantification Kit (Roche cat. no. KK4854), and Agilent TapeStation was used to estimate individual library sizes.
For 2S Plus DNA Library preparation, end repair and adapter ligation were performed according to manufacturer instructions. For biotin pull-down, 50 μl of Dynabeads MyOne Streptavidin T1 beads (Invitrogen cat. no. 65601) were washed once with Tween Washing Buffer (5 mM Tris pH 8, 0.5 mM EDTA, 1 M NaCl, 0.05% Tween-20) and resuspended in 300 μl of 2 × binding buffer (10 mM Tris pH 8, 1 mM EDTA, 2 M NaCl) for addition to the end-prepped/adapter-ligated DNA, with incubation for 15 min at room temperature. Beads were then washed twice at 55 °C for 2 min using Tween Washing Buffer, washed once briefly using Low EDTA TE Buffer (from 2S Plus DNA Library Kit) and resuspended in Low EDTA TE Buffer. Unique combinatorial dual indexes (IDT cat. no. 10009909) were added to each reaction before PCR amplification and SPRI bead clean-up. As above, final libraries were quantified by qPCR using KAPA Illumina Library Quantification Kit (Roche cat. no. KK4854), with Agilent TapeStation used to estimate individual library sizes.
To account for the accumulation of 4N cells with RAD21 depletion, we performed Hi-C mostly as described above. However, before decrosslinking, samples were incubated in propidium iodide staining buffer (10 μg ml−1 propidium iodide (Abcam cat. no. ab14083), 100 μg ml−1 RNase (Thermo Scientific cat. no. EN0531), 2 mM MgCl2, DPBS), and the G1/2N populations were sorted in bulk using BD Influx Cell Sorters in the Salk Flow Cytometry Core (see Extended Data Fig. 3a for gating strategy used). The sorted samples then reentered the Hi-C protocol starting with decrosslinking and proteinase K digestion.
Sequencing libraries first underwent a quality-control paired-end 37-bp run on Illumina MiniSeq, before paired-end 42 bp, 50 bp or 100 bp on Illumina NovaSeq6000 or NovaSeqX. Sequencing was carried out by the Salk Next-Generation Sequencing Core. The numbers of total and deduplicated sequencing reads, and cis/trans ratios, are shown in Supplementary Table 3. For Hi-C analysis, see Supplementary Information.
SLAM-seq
Cells were treated with 100 μM 4sU for the amount of time mentioned (15 or 120 min), before being collected in TRIzol (Invitrogen cat. no. 15596018). RNA was prepared under red light using the Direct-zol RNA Miniprep Kit (Zymo cat. no. R2052) and then treated with iodoacetamide using the SLAM-seq Kinetics Kit – Anabolic Kinetics Module (Lexogen cat. no. 061), according to the manufacturer’s instructions. For the mitosis synchronization experiments, the RNA was then purified by ethanol precipitation, while for asynchronous depletion experiments, we used the RNA Clean & Concentrator-5 kit (Zymo cat. no. R1013). RNA integrity was determined using Agilent TapeStation. To maximize the recovery of nascent RNA, library preparation was performed using Zymo-Seq RiboFree Total RNA Library Kit (Zymo cat. no. R3003), according to the manufacturer’s instructions. Library quantification was performed by qPCR using KAPA Illumina Library Quantification Kit (Roche cat. no. KK4854), with library sizes estimated using Agilent TapeStation.
Next-generation sequencing was performed by the Salk Next-Generation Sequencing Core on an Illumina NovaSeq6000 using paired-end 50 bp or 100 bp or on Element AVITI paired-end 75 bp. The number of total and deduplicated sequencing reads is shown in Supplementary Table 4. For the experiments utilizing 15-min 4sU labeling periods during mitotic exit, most samples had four biological replicates. Samples for DMSO t = 0 min R1, DMSO t = 30 min R1, dTAG t = 30 min R4 and dTAG t = 120 min R3 were lost during processing, leaving these treatments/timepoints with only three biological replicates. For the experiments utilizing 120-min 4sU labeling periods during mitotic exit and 15- or 120-min labeling periods in mid-G1, we had three biological replicates. For the experiments looking at longer-term consequences of NIPBL depletion during mitotic exit with 15-min labeling periods, two biological replicates were performed. For SLAM-seq analysis, see Supplementary Information.
Statistics and reproducibility
Two-sided t-tests, two-sided Wilcoxon rank-sum, Kruskal–Wallis and chi-squared statistical tests were carried out using SciPy in Python56. See figure legends for the specific tests used. Data met the assumptions of the statistical tests used. Differential expression and binding analyses were performed using DESEQ2, with Wald tests and multiple testing correction of P values using the Benjamini–Hochberg method. The center line in box plots marks the median, the box shows the interquartile range (IQR) and the whiskers extend to 1.5 × IQR. Replicate measurements were always taken from distinct samples (‘biological replicates’). No statistical method was used to predetermine sample size. No data were excluded from the analyses. Data collection and analysis were not performed blind to the conditions of the experiments. Relevant information for all statistical tests conducted is shown in Supplementary Table 6.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Hi-C, SLAM-seq and ChIP–seq data collected for the purposes of this study are available at the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) with the accession number GSE277721. All datasets generated in this study are presented in Supplementary Tables 2–4. Datasets accessed from the Sequence Read Archive (SRA) are listed in Supplementary Table 5. Source data are provided with this paper.
Code availability
Code used for this paper is available at the Dixon Lab GitHub page, and archived versions of the alignment of processing of Hi-C and SLAM-seq sequencing data are available via Zenodo at https://doi.org/10.5281/zenodo.13835562 (ref. 57) and https://doi.org/10.5281/zenodo.13835596 (ref. 58).
References
Waldman, T. Emerging themes in cohesin cancer biology. Nat. Rev. Cancer 20, 504–515 (2020).
Liu, J. & Krantz, I. D. Cohesin and human disease. Annu. Rev. Genomics Hum. Genet. 9, 303–320 (2008).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Popay, T. M. & Dixon, J. R. Coming full circle: on the origin and evolution of the looping model for enhancer–promoter communication. J. Biol. Chem. 298, 102117 (2022).
Gabriele, M. et al. Dynamics of CTCF- and cohesin-mediated chromatin looping revealed by live-cell imaging. Science 376, 496–501 (2022).
Mach, P. et al. Cohesin and CTCF control the dynamics of chromosome folding. Nat. Genet. 54, 1907–1918 (2022).
Hansen, A. S., Pustova, I., Cattoglio, C., Tjian, R. & Darzacq, X. CTCF and cohesin regulate chromatin loop stability with distinct dynamics. eLife 6, e25776 (2017).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
Petela, N. J. et al. Scc2 is a potent activator of cohesin’s ATPase that promotes loading by binding Scc1 without Pds5. Mol. Cell 70, 1134–1148 (2018).
Alonso-Gil, D., Cuadrado, A., Giménez-Llorente, D., Rodríguez-Corsino, M. & Losada, A. Different NIPBL requirements of cohesin-STAG1 and cohesin-STAG2. Nat. Commun. 14, 1326 (2023).
Patta, I. et al. Nuclear morphology is shaped by loop-extrusion programs. Nature 627, 196–203 (2024).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Cuartero, S. et al. Control of inducible gene expression links cohesin to hematopoietic progenitor self-renewal and differentiation. Nat. Immunol. 19, 932–941 (2018).
Calderon, L. et al. Cohesin-dependence of neuronal gene expression relates to chromatin loop length. eLife 11, e76539 (2022).
Hsieh, T.-H. S. et al. Enhancer-promoter interactions and transcription are largely maintained upon acute loss of CTCF, cohesin, WAPL or YY1. Nat. Genet. 54, 1919–1932 (2022).
Du, M. et al. Direct observation of a condensate effect on super-enhancer controlled gene bursting. Cell 187, 331–344 (2024).
Peters, J.-M., Tedeschi, A. & Schmitz, J. The cohesin complex and its roles in chromosome biology. Genes Dev. 22, 3089–3114 (2008).
Zhang, H. et al. Chromatin structure dynamics during the mitosis-to-G1 phase transition. Nature 576, 158–162 (2019).
Palozola, K. C. et al. Mitotic transcription and waves of gene reactivation during mitotic exit. Science 358, 119–122 (2017).
Hsiung, C. C.-S. et al. A hyperactive transcriptional state marks genome reactivation at the mitosis-G1 transition. Genes Dev. 30, 1423–1439 (2016).
Kang, H. et al. Dynamic regulation of histone modifications and long-range chromosomal interactions during postmitotic transcriptional reactivation. Genes Dev. 34, 1–18 (2020).
Nabet, B. et al. The dTAG system for immediate and target-specific protein degradation. Nat. Chem. Biol. 14, 431–441 (2018).
Nabet, B. et al. Rapid and direct control of target protein levels with VHL-recruiting dTAG molecules. Nat. Commun. 11, 4687 (2020).
Zuin, J. et al. A cohesin-independent role for NIPBL at promoters provides insights in CdLS. PLoS Genet. 10, e1004153 (2014).
Watrin, E. et al. Human Scc4 is required for cohesin binding to chromatin, sister-chromatid cohesion, and mitotic progression. Curr. Biol. 16, 863–874 (2006).
Brunner, A. et al. Quantitative imaging of loop extruders rebuilding interphase genome architecture after mitosis. J. Cell Biol. 224, e202405169 (2025).
Wutz, G. et al. ESCO1 and CTCF enable formation of long chromatin loops by protecting cohesinSTAG1 from WAPL. eLife 9, e52091 (2020).
Kikuchi, S., Borek, D. M., Otwinowski, Z., Tomchick, D. R. & Yu, H. Crystal structure of the cohesin loader Scc2 and insight into cohesinopathy. Proc. Natl Acad. Sci. USA 113, 12444–12449 (2016).
Herzog, V. A. et al. Thiol-linked alkylation of RNA to assess expression dynamics. Nat. Methods 14, 1198–1204 (2017).
Alge, C. S., Hauck, S. M., Priglinger, S. G., Kampik, A. & Ueffing, M. Differential protein profiling of primary versus immortalized human RPE cells identifies expression patterns associated with cytoskeletal remodeling and cell survival. J. Proteome Res. 5, 862–878 (2006).
Rambhatla, L., Chiu, C.-P., Glickman, R. D. & Rowe-Rendleman, C. In vitro differentiation capacity of telomerase immortalized human RPE cells. Invest. Ophthalmol. Vis. Sci. 43, 1622–1630 (2002).
Hsieh, T.-H. S. et al. Resolving the 3D landscape of transcription-linked mammalian chromatin folding. Mol. Cell 78, 539–553 (2020).
Banigan, E. J. et al. Transcription shapes 3D chromatin organization by interacting with loop extrusion. Proc. Natl Acad. Sci. USA 120, e2210480120 (2023).
Hafner, A. et al. Loop stacking organizes genome folding from TADs to chromosomes. Mol. Cell 83, 1377–1392 (2023).
Chen, L.-F. et al. Structural elements promote architectural stripe formation and facilitate ultra-long-range gene regulation at a human disease locus. Mol. Cell 83, 1446–1461 (2023).
Karpinska, M. A. et al. CTCF depletion decouples enhancer-mediated gene activation from chromatin hub formation. Nat. Struct. Mol. Biol. 32, 1268–1281 (2025).
Liu, N. Q. et al. WAPL maintains a cohesin loading cycle to preserve cell-type-specific distal gene regulation. Nat. Genet. 53, 100–109 (2021).
Hansen, K. L. et al. Synergy between regulatory elements can render cohesin dispensable for distal enhancer function. Science https://doi.org/10.1126/science.adt4221 (2025).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Dowen, J. M. et al. Control of cell identity genes occurs in insulated neighborhoods in mammalian chromosomes. Cell 159, 374–387 (2014).
Song, M. et al. Mapping cis-regulatory chromatin contacts in neural cells links neuropsychiatric disorder risk variants to target genes. Nat. Genet. 51, 1252–1262 (2019).
Sakamoto, T. et al. The nuclear receptor ERR cooperates with the cardiogenic factor GATA4 to orchestrate cardiomyocyte maturation. Nat. Commun. 13, 1991 (2022).
Aboreden, N. G. et al. Cis-regulatory chromatin contacts form de novo in the absence of loop extrusion. Preprint at bioRxiv https://doi.org/10.1101/2025.01.12.632634 (2025).
Kline, A. D. et al. Diagnosis and management of Cornelia de Lange syndrome: first international consensus statement. Nat. Rev. Genet. 19, 649–666 (2018).
Uyehara, C. M. & Apostolou, E. 3D enhancer-promoter interactions and multi-connected hubs: organizational principles and functional roles. Cell Rep. 42, 112068 (2023).
Wu, H., Zhang, J., Tan, L. & Xie, X. S. Single-cell Micro-C profiles 3D genome structures at high resolution and characterizes multi-enhancer hubs. Nat. Genet. 1786, 1777 (2025).
Wang, C. et al. Scalable production of iPSC-derived human neurons to identify tau-lowering compounds by high-content screening. Stem Cell Rep. 9, 1221–1233 (2017).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Méndez, J. & Stillman, B. Chromatin association of human origin recognition complex, cdc6, and minichromosome maintenance proteins during the cell cycle: assembly of prereplication complexes in late mitosis. Mol. Cell. Biol. 20, 8602–8612 (2000).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Dixon, J. R. Basic Hi-C alignment tools. Zenodo https://doi.org/10.5281/zenodo.13835562 (2024).
Popay, T. M. SLAM-seq analysis tools. Zenodo https://doi.org/10.5281/zenodo.13835596 (2024).
Acknowledgements
We thank C.-Z. Zhang for providing a comprehensive list of SNPs in hTERT RPE-1 cells, Y. Shen for the WTC11 hiPSCs, B. Ren for assisting with hiPSC engineering, and J. Jones and F. Gage for advice and protocols for the differentiation of hiPSCs into neurons. This work was supported by the Salk Excellerators Fellowship to T.M.P. and the 4D Nucleome grant to J.R.D. (grant no. U01-CA260700). J.R.D. is also supported as a Rita Allen Foundation Scholar and as a Pew Biomedical Scholar. The Flow Cytometry Core Facility of the Salk Institute (RRID:SCR_014839) is supported by funding from NIH-NCI CCSG grant no. P30 CA01495, and Shared Instrumentation Grants no. S10-OD023689 (Aria Fusion cell sorter) and no. S10-OD034268 (Thermo Fisher Bigfoot). This work was supported by the Razavi Newman Integrative Genomics and Bioinformatics Core Facility of the Salk Institute (RRID:SCR_014842 and SCR_014846) with funding from NIH-NCI CCSG grant no. P30 CA014195, NIH-NIA San Diego Nathan Shock Center grant no. P30 AG068635, the NIH-NIA Liver Cancer grant no. P01 AG073084-04, the Howard and Maryam Newman Family Foundation and the Helmsley Trust. This work was supported by the Single Cell & Spatial Omics Core of the Salk Institute (RRID:SCR_026396) with funding from NIH-NCI CCSG grant no. P30 CA014195, NIH-NIA San Diego Nathan Shock Center grant no. P30 AG068635, the Chapman Foundation and the Helmsley Charitable Trust. The Waitt Advanced Biophotonics Core Facility of the Salk Institute is supported by funding from NIH-NCI CCSG grant no. P30 CA01495, NIH-NlA San Diego Nathan Shock Center grant no. P30 AG068635 and the Waitt Foundation. This work was supported by the Stem Cell Core Facility of the Salk Institute (RRID:SCR_014850) with funding from the NIH-NIA San Diego Nathan Shock Center grant no. P30 AG068635, the NIH-NIA Alzheimer’s Disease Research Center grant no. P30 AG062429, the AHA Allen Initiative, the California Institute for Regenerative Medicine and the Helmsley Charitable Trust.
Author information
Authors and Affiliations
Contributions
Conceptualization: T.M.P. and J.R.D. Formal analysis: T.M.P. Funding acquisition: J.R.D. Investigation: T.M.P., A.P., F.M., M.T., M.E.B. and N.H. Methodology: T.M.P. and J.R.D. Supervision: J.R.D. Visualization: T.M.P. Writing—original draft: T.M.P. and J.R.D. Writing—review and editing: T.M.P., A.P., F.M., M.T., M.E.B., N.H. and J.R.D.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Anton Goloborodko and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Acute depletion of NIPBL and RAD21 from hTERT RPE-1 cells.
a, Schematics of N-terminal FKBP12F36V-NIPBL fusions engineered into hTERT RPE-1 cells using CRISPR/Cas9-directed homologous recombination. N-terminal tagging was employed to maximize degradation of NIPBL isoforms (A and B). The NIPBL-A2 and NIPBL-D7 clones are homozygous for a P2A-linked mCherry and P2A-linked mGreenLantern, respectively. b, Example gating strategy used for fluorescence activated cell sorting of hTERT RPE-1 cells transfected with Cas9/NIPBL sgRNA, with or without a targeting vector containing mGreenLantern-P2A-FKBP12F36V flanked by NIPBL homology arms. mGreenLantern (mGL)-positive cells were sorted based on absence in sgRNA-only and presence in sgRNA + targeting vector. c, Whole cell extracts were harvested from parental hTERT RPE-1 and the engineered NIPBL-A2 and NIPBL-D7 cell lines treated with DMSO or dTAGV-1 for 24 hours. The shift in molecular weight of NIPBL in the engineered cell lines is demonstrated using two commercially available polyclonal antibodies against NIPBL (PTG: 18792-1-AP; Bethyl: A301-779A). Two biological replicates were performed with similar outcomes. d, Western blot of whole-cell lysates from NIPBL-A2 cells treated with DMSO for 24 hours or dTAGV-1 for varying lengths of time to monitor HA-NIPBL depletion. α-Tubulin is shown as a loading control. Two biological replicates were performed with similar outcomes. e, Schematic of C-terminal RAD21-FKBP12F36V fusion engineered in hTERT RPE-1 cells using CRISPR/Cas9-directed homologous recombination. The RAD21-B1 and RAD21-B3 clonal cell lines are homozygous for T2A-linked mGreenLantern. f, Western blot of parental hTERT RPE-1, RAD21-B1, and RAD21-B3 cells treated with DMSO or dTAGV-1 for 24 hours. Two biological replicates were performed with similar outcomes. g, Lysates for western blot were harvested from RAD21-B1 (top) and RAD21-B3 (bottom) cells treated with DMSO for 24 hours or dTAGV-1 for increasing lengths of time. Four biological replicates were performed with similar outcomes.
Extended Data Fig. 2 Chromatin binding of cohesin and cell cycle defects with NIPBL depletion.
a, Chromatin fractionation was used to separate out cytoplasmic (S2), nucleoplasmic (S3), and chromatin (P3) fractions from DMSO- or dTAGV-1-treated NIPBL-A2 (left) and NIPBL-D7 (right) cells. Two different commercially available polyclonal antibodies raised against NIPBL (PTG: 18792-1-AP; Bethyl: A301-779A) were combined to improve detection of NIPBL in western blotting. Fractionation success was determined by the absence of α-Tubulin and presence of H3 in the P3 fraction, and vice versa in the S2 and S3 fractions. Two biological replicates were performed with similar outcomes. b, Volcano plots from differential analysis of high-confidence RAD21 ChIP-seq peaks with NIPBL depletion for 4 hours (left) and 24 hours (right). RAD21 ChIP-seq from NIPBL-A2 and NIPBL-D7 were treated as biological replicates. No significantly increased peaks were observed, but significantly decreased peaks (FDR < 0.05) are highlighted in blue. P-values were calculated from two-sided Wald tests and adjusted (padj) for multiple testing using the Benjamini-Hochberg method. c, Scale factor-normalized RAD21 ChIP signal at significantly decreased and non-significantly changed peaks from differential analysis of RAD21 ChIP-seq. Additionally shown is RPKM-normalized ChIP signal for HA-NIPBL and CTCF. d, Quantification (top) and examples (bottom) of cell cycle distributions by propidium iodide staining of cells treated with DMSO or dTAGV-1, as measured by flow cytometry. n = 2. e, Example flow cytometry gating strategy used for determining cell cycle distribution by propidium iodide staining. Cells were separated from debris by gating in SSC-A vs. FSC-A, then single cells were selected based PE-Texas Red-A vs. PE-Texas Red-H. Depending on the flow cytometer used, propidium iodide was detected using either the PE-Texas Red or PE-CF594 channel.
Extended Data Fig. 3 Consequences of NIPBL and RAD21 depletion on genome organization.
a, Gating strategy used for fluorescence activated cell sorting of 2 N RAD21-B1/B3 cells for Hi-C. Cells first underwent initial steps for Hi-C, including crosslinking, restriction digestion, and proximity ligation, before being stained with propidium iodide for sorting. Post-sorting, cells were decrosslinked and sequencing libraries prepared. b, Relationship between distance and contact frequency at a 10 kb resolution based on Hi-C performed in cells depleted of NIPBL (top) or RAD21 (bottom; G1 sorted) for 4 or 24 hours. For each timepoint, Hi-C experiments performed on independent clonal cell lines for NIPBL (NIPBL-A2 and NIPBL-D7) and RAD21 (RAD21-B1 and RAD21-B3) were treated as replicates, with the resulting datasets merged for plotting. c, Pile-up analysis showing log-transformed observed/expected contact frequency from Hi-C at chromatin loops (left) and stripes (right) called from DMSO-treated FKBP12F36V-NIPBL cells. d, Saddleplots showing genome-wide changes in compartments with depletion of NIPBL (left) or RAD21 (right) for 4 or 24 hours. e, Quantification of saddle strength based on average signal strength in the saddleplot corners (top left: BB; bottom right: AA; top right: BA; bottom left: AB). f, Example heatmaps showing the consequences of NIPBL (top) and RAD21 (bottom) depletion for 4 and 24 hours. Beneath the NIPBL heatmaps are scale factor-normalized RAD21 ChIP-seq tracks for the corresponding treatment in NIPBL-D7 cells. A chromatin loop of interest is highlighted with an arrow in the heatmaps, while the loop anchors in the ChIP-seq tracks are marked a grey box.
Extended Data Fig. 4 Chromatin loops show differential dependence on NIPBL.
a, Scatter plot showing the log-transformed balanced counts between DMSO and dTAGV-1-treated conditions with RAD21 depletion. Chromatin loops that showed a fold-change of at least two are highlighted. A kernel density estimate (KDE) plot is shown on each axis to represent the distribution of the datapoints on that axis. To each balanced count value, 0.0001 was added prior to log-transformation to allow for plotting of zero values. b, Observed/expected pile-up analysis at a 10 kb resolution at RAD21-dependent chromatin loops (RAD21 FC > 2) from Hi-C performed on DMSO- and dTAGV-1-treated FKBP12F36V-NIPBL and RAD21-FKBP12F36V cells. c, As in a, but showing consequences of NIPBL depletion on RAD21-dependent chromatin loops (RAD21 FC > 2). d, As in Fig. 1f, but for all of the clusters with NIPBL depletion at 4 and 24 hours and RAD21 depletion at 4 hours. e, Relationship between clusters and differential RAD21 ChIP-seq peaks with depletion of NIPBL for 4 hours. f, Scatter plot comparing how NIPBL depletion differentially affects chromatin loops from the mixed- and max.-dependency clusters from Fig. 1d. g, Schematic of C-terminal CTCF-FKBP12F36V fusion engineered in hTERT RPE-1 cells using CRISPR/Cas9-directed homologous recombination. h, Western blot of parental hTERT RPE-1, CTCF-C1, and CTCF-C8 cells treated with DMSO or dTAGV-1 for 24 hours. Two biological replicates were performed with similar outcomes. i, Western blot of lysates harvested from CTCF-C1 (top) and CTCF-C8 (bottom) cells treated with DMSO or dTAGV-1 for varying lengths of time. Four biological replicates were performed with similar outcomes. j, As in Fig. 1f, but for all of the clusters with CTCF depletion. k, Classification of chromatin loops based on their association with RAD21/CTCF ChIP-seq peaks (structural), cis regulatory element (CRE), or a mix of both. P-value is from a chi-squared test.
Extended Data Fig. 5 Depletion of NIPBL during mitotic exit.
a, Quantification (left) and examples (right) of cell cycle distributions by propidium iodide (PI) staining of NIPBL-D7 cells treated with DMSO or dTAGV-1 during mitotic exit, as measured by flow cytometry. Cells were collected at t = 0 min following a 2-hour nocodazole synchronization with DMSO/dTAGV-1 treatment, and at 30-minute intervals following release. n = 2. b, c, Quantification (left) and examples (right) of PI staining of NIPBL-D7 cells treated with DMSO or dTAGV-1 during mitotic exit. Cells were harvested 8 hours (b) and 16 hours (c) post-release from nocodazole. n = 2. d, Western blot of cell lysates harvested at t = 0 min and t = 120 min following nocodazole synchronization and release with DMSO/dTAGV-1 treatment of the NIPBL-D7 cell line. Two biological replicates were performed with similar outcomes. e, Schematic outlining the approach to perform Hi-C with depletion of NIPBL during mitotic exit. Cells were synchronized to prometaphase using a double thymidine block and nocodazole treatment. DMSO/dTAGV-1 was added simultaneous with nocodazole. f, Genome-wide contact frequency over genomic distance at a 10 kb resolution from Hi-C for DMSO- and dTAGV-1 NIPBL-D7 cells during mitotic exit. Cells were collected at 30-minute intervals starting with nocodazole-arrested cells (t = 0 min). Data from replicate experiments were combined for this analysis. The arrows highlight the region of these plots that reflect chromatin loops. g, Observed/expected pile-up analysis of Hi-C data at a 10 kb resolution of cohesin-dependent chromatin loops identified in Extended Data Fig. 4a. Asynchronous pile-up is from Extended Data Fig. 4b. h, As in Fig. 1j, but for an example max.-dependency chromatin loop. i, Observed/expected pile-up analysis at a 10 kb resolution in cells depleted of NIPBL during mitotic exit, both at short (t = 120 minutes) and long (t = 8 hours) timepoints following release from nocodazole.
Extended Data Fig. 6 Altering cohesin subunit composition by STAG1 or STAG2 knockout.
a, Boxplot showing the RPKM of cohesin accessory proteins at RAD21 peaks that intersect loop anchors. For each chromatin loop, the mean RPKM for each protein was taken from all RAD21 ChIP-seq peaks across both anchors and normalized to the mean RPKM of RAD21 at the same anchors to account for differences in RAD21 binding at different loops. Differences in chromatin binding at the clusters was tested using a Kruskal-Wallis test (min.-dep. n = 1037, low-dep. n = 2780, med.-dep. n = 3533, mixed-dep. n = 1678, high-dep. n = 3820, max.-dep. n = 2523). The center line marks the median, the box shows the IQR, and the whiskers extend to 1.5xIQR. b, Western blots showing STAG1 (left) and STAG2 (right) knockout engineered into the NIPBL-D7 degron background. Antibodies against the N- and C-termini are shown for both STAG1 and STAG2. Two biological replicates were performed with similar outcomes. c, As in Fig. 2d, but showing an example max.-dependency chromatin loop. d, Observed/expected pile-up analysis at a 10 kb resolution in STAG1/2 wild-type, STAG1 knockout, or STAG2 knockout cells depleted of NIPBL. e, Model of chromatin states in hTERT RPE-1 cells generated using ChromHMM. Mark probability is from the ChromHMM emission parameters and estimates the likelihood that a given mark would be found in that state. f, The chromatin state, excluding ‘Insulator’ and ‘Low signal’, that makes up the majority of the chromatin loop anchors for each of the clusters. g, Example mixed-dependency and max.-dependency chromatin loops showing distinct patterns of RPKM-normalized H3K27me3 ChIP-seq in hTERT RPE-1 cells. The chromatin loop of interest is marked by an arrow, while its anchor regions are marked by grey bars in the tracks below.
Extended Data Fig. 7 Approaches used to interrogate the contribution of NIPBL to gene regulation during mitotic exit and G1.
a, Schematic outlining the two approaches used to investigate NIPBL depletion during mitotic exit. For both approaches, cells first underwent a double thymidine block prior to nocodazole treatment. Approaches differ in the length of 4sU treatment for SLAM-seq, with approach A utilizing 15 min labeling periods and approach B 120 min labeling period. b, Morphology of NIPBL-A2 cells treated with DMSO or dTAGV-1 for 72 hours. Cells were stained with phalloidin (actin; green) and DAPI (DNA; blue). Two biological replicates were performed with similar outcomes. c, Schematic outlining the two approaches used to investigate NIPBL depletion during G1. Cells were synchronized with double thymidine block and 2 hour nocodazole, then released for 4 hours before addition of DMSO or dTAGV-1 for 2 hours. Cells were treated with 4sU for 15 min (approach A) or 2 hours (approach B). d, Volcano plot from differential expression analysis of nascent transcripts between DMSO- and dTAGV-1-treated NIPBL-D7 cells based on mid-G1 approach B. Significantly changed (|log2FC| > 0.585 and padj < 0.1) nascent transcripts are highlighted. P-values were calculated with two-sided Wald tests using three independent replicates and adjusted (padj) for multiple testing using the Benjamini-Hochberg method. e, Scatter plot showing the log-transformed FPKM values for nascent transcripts from (d). Significantly changed transcripts (|log2FC| > 0.585 and padj < 0.1) are highlighted. A kernel density estimate (KDE) plot is shown on each axis to represent the distribution of the datapoints on that axis. To each FPKM value, 0.001 was added prior to log-transformation to allow for plotting of zero FPKM values. f, Biological process gene ontology analysis was performed on decreased transcripts. The top 5 categories are shown, with the P-values derived using the ‘elim’ algorithm and a Fischer’s exact test in TopGO. g, Line plots for example differentially expressed genes, plotting the mean FPKM and standard error based on mitotic exit and mid-G1 approach A. All shown genes were significantly changed during mitotic exit, but only BDNF and VEGFA were also significantly changed in mid-G1. For ‘mitosis’, the x-axis is time post-release from nocodazole, while for ‘mid-G1’, the x-axis is time post-addition of DMSO/ dTAG V-1.
Extended Data Fig. 8 Connectivity with enhancer neighborhood is sensitive to NIPBL depletion.
a, For each of the significantly decreased (left) and increased (right) genes, nearest neighbor analysis was used to identify the closest non-significantly changed gene based on FPKM and the number of enhancers and TSSs within 1 Mb up- or down-stream. A grey line connects each significantly changed gene with its identified nearest neighbor. b, Comparison of FPKM, enhancer count, and gene count metrics between significantly changed genes and the non-significant nearest neighbors. P-values were determined using a two-sided Wilcoxon rank-sum test. The center line marks the median, the box shows the IQR, and the whiskers extend to 1.5xIQR. c, Stacked bar plot showing the percentage of non-significant, decreased, and significantly increased genes whose promoter-proximal regions (TSS −1 kb/+500 bp) intersect a chromatin loop anchor and whereby the distal loop anchor intersects with a super-enhancer (SE; purple), typical enhancer (TE; yellow), or no enhancer (loop:no E; magenta). Because promoter-proximal regions may intersect with multiple chromatin loop anchors, each gene is represented only once, with chromatin loops for a given gene preferentially selected based on intersection of the distal loop anchor with a super-enhancer, then typical enhancer, then no enhancer. P-values for chromatin loops called from the DMSO- and dTAGV-1-treated conditions were determined using a chi-squared test. d, As in Fig. 4a, but for the typical-enhancer neighborhood. e, Quantification of the difference in contact scores for enhancers within 500 kb of a TSS from Fig. 4b. P-values were determined using a two-sided Wilcoxon rank-sum test (non-sig. (decreased NN) n = 5276, decreased n = 5735, non-sig. (increased NN) n = 1037, increased n = 1012). NN = Nearest Neighbors. The center line marks the median, the box shows the IQR, and the whiskers extend to 1.5xIQR.
Extended Data Fig. 9 Examples of genes sensitive to NIPBL depletion during mitotic exit.
SOX9 (a) and JARID2 (b) are examples of genes that are significantly decreased with NIPBL depletion during mitotic exit. Shown are Hi-C heatmaps (top: DMSO; bottom: dTAG), sequencing-depth normalized BigWig tracks of read pairs whereby at least one read contains a T→C conversion (that is nascent reads), and Virtual 4 C anchored at promoter-proximal regions (TSS −1 kb/+500 bp) for both DMSO and dTAG-treated samples at t = 30 min, 60 min, 90 min, and 120 min. The marked TE and SE sites are the same across all plots. The arrow on the Hi-C heatmaps marks the transcription start site (TSS), and the grey bar on the tracks below marks the gene location.
Extended Data Fig. 10 NIPBL-dependence is associated with unique genomic structures across lineages.
a, Cross-TSS connectivity at example genes sensitive to NIPBL depletion during mitotic exit. The top-right corner of each box corresponds to the TSS of each gene. b, c, d, Quantification of Fig. 4d, specifically the mean observed/expected contact frequency for the flanking regions (b), central regions (c), and the ratio between flanking and central regions (d) for each TSS. P-values were determined by independent, two-sided t-tests for (b) and (c), and using two-sided Wilcoxon rank-sum tests for (d). (b) non-sig. DMSO n = 12964, non-sig. dTAG n = 12966, non-sig. (decreased NN) n = 450, decreased n = 450, non-sig. (increased NN) n = 87, and increased n = 91. (c & d) non-sig. DMSO n = 12889, non-sig. dTAG n = 12893, non-sig. (decreased NN) n = 450, decreased n = 449, non-sig. (increased NN) n = 87, and increased n = 91. The inset shows the regions that were quantified, with the mean observed/expected contact frequency for a given TSS taken from the shown regions for plotting. The center lines mark the median, the boxes show the IQR, and the whiskers extend to 1.5xIQR; all box plots in this figure use the same metrics. e, Ratio of NIPBL to PDS5 (left) and WAPL (right) at a 20 kb region centered on the TSSs of non-significant, decreased and increased genes. Two-sided Wilcoxon rank-sum tests were performed to give the shown p-values (non-sig. (decreased NN) n = 436, decreased n = 449, non-sig. (increased NN) n = 91, increased n = 92). Only non-significant nearest neighbors (NN) are shown. f, Balanced counts between super-enhancers (left) or typical-enhancers (right) and enhancers on the opposite side of a TSS. For each gene, the mean of different, non-zero enhancer-enhancer pairs was taken before plotting. P-values were determined using a two-sided Wilcoxon rank-sum test (SE-E: non-sig. n = 8204, non-sig. (decreased NN) n = 307, decreased n = 359, non-sig. (increased NN) n = 66, and increased n = 71; TE-E: non-sig. n = 11995, non-sig. (decreased NN) n = 431, decreased n = 440, non-sig. (increased NN) n = 87, and increased n = 91). All non-significantly changed genes and nearest neighbor (NN) non-significantly changed genes are shown. g, Examples of TSS insulation at a non-significantly changed gene (NECTIN3) and reduced TSS insulation at a significantly decreased gene (JARID2). For each gene we show Virtual 4 C anchored at two different super-enhancers (SE-A: blue, SE-B: pink) that are upstream of the TSS. The arrow marks the transcription start site (TSS). h, Immunofluorescence of hiPSC-derived cardiomyocytes following treatment with DMSO or dTAGV-1 for 4 hours. cTnT is a marker for cardiomyocyte differentiation. i, For the significantly decreased genes in hiPSC-derived neurons and cardiomyocytes, we performed nearest neighbor analysis to identify the closest non-significant genes based on FPKM and the number of enhancers and TSSs within 1 Mb up- or down-stream. A grey line connects each significantly changed gene with its nearest neighbor. j, Comparison of FPKM, enhancer count, and gene count metrics between significantly changed genes and the non-significant nearest neighbors. P-values were determined using a two-sided Wilcoxon rank-sum test. k, Observed/expected pile-up analysis at a 10 kb resolution between neighborhood (±1 Mb of TSS) typical-enhancers (TE) and non-significant genes, nearest neighbors, or decreased genes in hiPSC-derived neurons and cardiomyocytes.
Supplementary information
Supplementary Information (download PDF )
Supplementary methods.
Supplementary Tables (download XLSX )
Supplementary Table 1. sgRNA and genotyping primers for NIPBL, CTCF and RAD21 degrons, and genotyping primers for STAG1 and STAG2 knockout. Supplementary Table 2. ChIP–seq datasets generated in this study. Supplementary Table 3. Hi-C datasets generated in this study. Supplementary Table 4. SLAM-seq datasets generated in this study. Supplementary Table 5. GEO datasets utilized in this study. Supplementary Table 6. Statistical tests performed in this study.
Source data
Source Data Figs. 1 and 5 (download PDF )
Unprocessed western blots for Figs. 1b and 5a.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots for Extended Data Fig. 1c,d,f,g.
Source Data Figs. 1–5 (download XLSX )
Statistical source data for Figs. 1c–e,h, 2a,c,e,f, 3a–c,f–i, 4b and 5c–e.
Source Data Extended Data Figs. 2 and 4 (download PDF )
Unprocessed western blots for Extended Data Figs. 2a and 4h.
Source Data Extended Data Figs. 4–6 (download PDF )
Unprocessed western blots for Extended Data Figs. 4i, 5d and 6b.
Source Data Extended Data Figs. 2–8 and 10 (download XLSX )
Statistical source data for Extended Data Figs. 2b,d, 3b,e, 4a,c,e,f,k, 5a–c,f, 6a,e,f, 7d–g, 8a–c,e and 10b–f,i,j.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Popay, T.M., Pant, A., Munting, F. et al. Acute NIPBL depletion reveals in vivo dynamics of loop extrusion and its role in transcription activation. Nat Genet (2026). https://doi.org/10.1038/s41588-026-02516-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41588-026-02516-y