Abstract
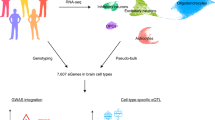
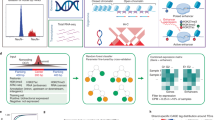
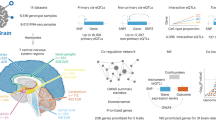
Most genetic risk variants for neurological diseases are located in noncoding regulatory regions, where they often act as expression quantitative trait loci (eQTLs), modulating gene expression and influencing disease susceptibility. However, eQTL studies in bulk brain tissue or cell lines fail to capture the brain’s cellular diversity. Single-nucleus RNA sequencing (snRNA-seq) allows high-resolution mapping of eQTLs across diverse brain cell types. Here we performed a meta-analysis by integrating snRNA-seq and genotype data from four cohorts, totaling 5.8 million nuclei from 983 individuals of European ancestry. We mapped cis-eQTLs and trans-eQTLs across major brain cell types and subtypes, including disease-specific and sex-specific eQTLs, and applied colocalization and Mendelian randomization to identify genes that mediate neurological disease risk. We observed up to tenfold more cis-eQTLs and uncovered cell-type-specific genes linked to neurological disease. SingleBrain is a comprehensive single-cell eQTL resource that provides insights into the genetic mechanism of brain disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Genotypes from refs. 11,14 are available at https://doi.org/10.7303/syn10901595. snRNA-seq data from ref. 11 are available at https://www.synapse.org/Synapse:syn31512863. snRNA-seq data from ref. 14 are available at https://www.synapse.org/Synapse:syn52293417. snRNA-seq and genotype data from ref. 15 are available at https://www.synapse.org/Synapse:syn26223298. snRNA-seq data and genotype data for the Roche cohort, which is part of ref. 12, have been deposited in the European Genome-Phenome Archive, which is hosted by the European Bioinformatics Institute and the Center for Genomic Regulation, under accession EGAS00001006345. GTEx Consortium. The GTEx Project (v10) is available at https://gtexportal.org. The massive parallel reporter assay-SCZ dataset was downloaded from https://github.com/thewonlab/schizophrenia-MPRA. Epigenomic data from purified human microglia, neurons, astrocytes and oligodendrocytes19 were downloaded from https://github.com/nottalexi/brain-cell-type-peak-files. SingleBrain eQTL browser is accessible at https://singlebrain.nygenome.org. All cis-eQTL summary statistics are accessible through Zenodo at https://doi.org/10.5281/zenodo.14908182 (ref. 63). Bonferroni-corrected significant trans-eQTL summary statistics are accessible through Zenodo at https://doi.org/10.5281/zenodo.15860673 (ref. 64). All SAIGE-QTL summary statistics are accessible through Zenodo at https://doi.org/10.5281/zenodo.15860973 (ref. 65). All disease-specific and sex-specific eQTL summary statistics are accessible through Zenodo at https://doi.org/10.5281/zenodo.16051904 (ref. 66).
Code availability
QTL preparation and meta-analysis pipeline can be found on Zenodo at https://doi.org/10.5281/zenodo.18332620 (ref. 67). SAIGE-QTL pipeline is available through Zenodo at https://doi.org/10.5281/zenodo.18239115 (ref. 68). Downstream-QTL analysis pipeline is available through Zenodo at https://doi.org/10.5281/zenodo.18320789 (ref. 69). All codes used to produce analysis for figures are available through Zenodo at https://doi.org/10.5281/zenodo.18239090 (ref. 70).
References
Ng, B. et al. An xQTL map integrates the genetic architecture of the human brain’s transcriptome and epigenome. Nat. Neurosci. 20, 1418–1426 (2017).
De Klein, N. et al. Brain expression quantitative trait locus and network analyses reveal downstream effects and putative drivers for brain-related diseases. Nat. Genet. 55, 377–388 (2023).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Lopes, K. P. et al. Genetic analysis of the human microglial transcriptome across brain regions, aging and disease pathologies. Nat. Genet. 54, 4–17 (2022).
Young, A. M. H. et al. A map of transcriptional heterogeneity and regulatory variation in human microglia. Nat. Genet. 53, 861–868 (2021).
Kosoy, R. et al. Genetics of the human microglia regulome refines Alzheimer’s disease risk loci. Nat. Genet. 54, 1145–1154 (2022).
Humphrey, J. et al. Long-read RNA sequencing atlas of human microglia isoforms elucidates disease-associated genetic regulation of splicing. Nat. Genet. 57, 604–615 (2025).
Jerber, J. et al. Population-scale single-cell RNA-seq profiling across dopaminergic neuron differentiation. Nat. Genet. 53, 304–312 (2021).
McAfee, J. C. et al. Systematic investigation of allelic regulatory activity of schizophrenia-associated common variants. Cell Genom. 3, 100404 (2023).
Fujita, M. et al. Cell subtype-specific effects of genetic variation in the Alzheimer’s disease brain. Nat. Genet. 56, 605–614 (2024).
Bryois, J. et al. Cell-type-specific cis-eQTLs in eight human brain cell types identify novel risk genes for psychiatric and neurological disorders. Nat. Neurosci. 25, 1104–1112 (2022).
Emani, P. S. et al. Single-cell genomics and regulatory networks for 388 human brains. Science 384, eadi5199 (2024).
Mathys, H. et al. Single-cell atlas reveals correlates of high cognitive function, dementia, and resilience to Alzheimer’s disease pathology. Cell 186, 4365–4385 (2023).
Gabitto, M. I. et al. Integrated multimodal cell atlas of Alzheimer’s disease. Nat. Neurosci. 27, 2366–2383 (2024).
Zeng, B. et al. Multi-ancestry eQTL meta-analysis of human brain identifies candidate causal variants for brain-related traits. Nat. Genet. 54, 161–169 (2022).
Dimas, A. S. et al. Common regulatory variation impacts gene expression in a cell type-dependent manner. Science 325, 1246–1250 (2009).
Gamazon, E. R. et al. Using an atlas of gene regulation across 44 human tissues to inform complex disease- and trait-associated variation. Nat. Genet. 50, 956–967 (2018).
Nott, A. et al. Brain cell type-specific enhancer–promoter interactome maps and disease-risk association. Science 366, 1134–1139 (2019).
Qi, T. et al. Identifying gene targets for brain-related traits using transcriptomic and methylomic data from blood. Nat. Commun. 9, 2282 (2018).
Green, G. S. et al. Cellular communities reveal trajectories of brain ageing and Alzheimer’s disease. Nature 633, 634–645 (2024).
Zhou, W. et al. Efficient and accurate mixed model association tool for single-cell eQTL analysis. Preprint at medRxiv https://doi.org/10.1101/2024.05.15.24307317 (2024).
Bellenguez, C. et al. New insights into the genetic etiology of Alzheimer’s disease and related dementias. Nat. Genet. 54, 412–436 (2022).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Marioni, R. E. et al. GWAS on family history of Alzheimer’s disease. Transl. Psychiatry 8, 99 (2018).
Nalls, M. A. et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 18, 1091–1102 (2019).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
Leonard, H. L. & Global Parkinson’s Genetics Program (GP2). Novel Parkinson’s disease genetic risk factors within and across European populations. Preprint at medRxiv https://doi.org/10.1101/2025.03.14.24319455 (2025).
International Multiple Sclerosis Genetics Consortium Multiple sclerosis genomic map implicates peripheral immune cells and microglia in susceptibility. Science 365, eaav7188 (2019).
Van Rheenen, W. et al. Common and rare variant association analyses in amyotrophic lateral sclerosis identify 15 risk loci with distinct genetic architectures and neuron-specific biology. Nat. Genet. 53, 1636–1648 (2021).
Mullins, N. et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat. Genet. 53, 817–829 (2021).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Yao, D. W., O’Connor, L. J., Price, A. L. & Gusev, A. Quantifying genetic effects on disease mediated by assayed gene expression levels. Nat. Genet. 52, 626–633 (2020).
Paolicelli, R. C. et al. Microglia states and nomenclature: a field at its crossroads. Neuron 110, 3458–3483 (2022).
Sun, N. et al. Human microglial state dynamics in Alzheimer’s disease progression. Cell 186, 4386–4403 (2023).
Tuddenham, J. F. et al. A cross-disease resource of living human microglia identifies disease-enriched subsets and tool compounds recapitulating microglial states. Nat. Neurosci. 27, 2521–2537 (2024).
Thrupp, N. et al. Single-nucleus RNA-seq is not suitable for detection of microglial activation genes in humans. Cell Rep. 32, 108189 (2020).
Zhou, H. J., Li, L., Li, Y., Li, W. & Li, J. J. PCA outperforms popular hidden variable inference methods for molecular QTL mapping. Genome Biol. 23, 210 (2022).
Álvarez Jerez, P. et al. African ancestry neurodegeneration risk variant disrupts an intronic branchpoint in GBA1. Nat. Struct. Mol. Biol. 31, 1955–1963 (2024).
The Synapse Engineering Team. Synapse Python Client. https://pypi.org/project/synapseclient/ (2026).
Pedersen, B. S. et al. Somalier: rapid relatedness estimation for cancer and germline studies using efficient genome sketches. Genome Med. 12, 62 (2020).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Stegle, O., Parts, L., Durbin, R. & Winn, J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLoS Comput. Biol. 6, e1000770 (2010).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Horton, R. et al. Gene map of the extended human MHC. Nat. Rev. Genet. 5, 889–899 (2004).
Saha, A. & Battle, A. False positives in trans-eQTL and co-expression analyses arising from RNA-sequencing alignment errors. F1000Res. 7, 1860 (2018).
Tang, Y., Ghosal, S. & Roy, A. Nonparametric Bayesian estimation of positive false discovery rates. Biometrics 63, 1126–1134 (2007).
Liu, Y. et al. ACAT: a fast and powerful p value combination method for rare-variant analysis in sequencing studies. Am. J. Hum. Genet. 104, 410–421 (2019).
Lee, C.-Y. et al. Regulatory landscape of Alzheimer’s disease variants in human microglia. Preprint at medRxiv https://doi.org/10.1101/2025.10.13.25337902 (2025).
Collins, R. L. et al. A structural variation reference for medical and population genetics. Nature 581, 444–451 (2020).
Myers, T. A., Chanock, S. J. & Machiela, M. J. LDlinkR: an R package for rapidly calculating linkage disequilibrium statistics in diverse populations. Front. Genet. 11, 157 (2020).
Hemani, G. et al. The MR-base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Hartwig, F. P., Davey Smith, G. & Bowden, J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int. J. Epidemiol. 46, 1985–1998 (2017).
Bowden, J., Davey Smith, G., Haycock, P. C. & Burgess, S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40, 304–314 (2016).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Cooler, S. & Schwartz, G. W. An offset ON-OFF receptive field is created by gap junctions between distinct types of retinal ganglion cells. Nat. Neurosci. 24, 105–115 (2021).
Fang, R. et al. Mapping of long-range chromatin interactions by proximity ligation-assisted ChIP–seq. Cell Res. 26, 1345–1348 (2016).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Jang, B., Won, H.-H. & Raj, T. SingleBrain: single-nucleus expression QTL meta-analysis summary statistics across multiple human brain cohorts. Zenodo https://doi.org/10.5281/zenodo.14908182 (2025).
Jang, B., Hong, W. H. & Raj, T. SingleBrain: single-nucleus trans expression QTL meta-analysis summary statistics across multiple human brain cohorts. Zenodo https://doi.org/10.5281/zenodo.15860673 (2025).
Jang, B., Hong, W. H. & Raj, T. SAIGE-QTL: single-nucleus expression QTL summary statistics using human brain cohort. Zenodo https://doi.org/10.5281/zenodo.15860973 (2025).
Jang, B., Hong, W. H. & Raj, T. SingleBrain: single-nucleus disease-/sex-specific expression QTL meta-analysis summary statistics across multiple human brain cohorts. Zenodo https://doi.org/10.5281/zenodo.16051904 (2025).
Kailash, B. P., Humphrey, J., Jang, B., Brophy, E. & Naito, T. RajLabMSSM/mmQTL-Pipeline: publication_v2.0. Zenodo https://doi.org/10.5281/zenodo.18332620 (2026).
Jang, B. RajLabMSSM/SAIGEQTL-Pipeline: RajLabMSSM/SAIGEQTL-Pipeline: publication version. Zenodo https://doi.org/10.5281/zenodo.18239115 (2026).
Humphrey, J., Jang, B. & Kailash, B. P. RajLabMSSM/downstream-QTL: publication_v2.1. Zenodo https://doi.org/10.5281/zenodo.18320789 (2026).
Jang, B. RajLabMSSM/SingleBrain: RajLabMSSM/SingleBrain: publication version. Zenodo https://doi.org/10.5281/zenodo.18239090 (2026).
Acknowledgements
The authors thank the patients and their families who donated the materials for these studies. T.R. and J.B. received funding from Calico Life Sciences. T.R., B.J., W.H.D., T.N. and J.H. were supported by National Institutes of Health (NIH; grants NIA U01-AG068880, NIA R21-AG063130, NIA R01-AG054005, NIA RF1-AG065926, NIA R01-AG065926, NIA R56-AG088669, NIA R21-AG091272, NIA P30-AG066514, NINDS U54-NS123743 and NINDS R01-NS116006). H.-H.W. was supported by the National Research Foundation of Korea grant supported by the Korea government (MSIT; RS-2023-00223277 and RS-2023-00262527) and the Future Medicine 2030 Project of the Samsung Medical Center (SMX1250081). This work was supported in part by the computational resources and staff expertise provided by Scientific Computing at the Icahn School of Medicine at Mount Sinai and by the Clinical and Translational Science Awards grant (UL1TR004419) from the National Center for Advancing Translational Sciences. This research was supported by the Office of Research Infrastructure of the NIH (awards S10OD026880 and S10OD030463). The funders had no role in the design and conduct of the study; the collection, management, analysis and interpretation of the data; the preparation, review or approval of the manuscript; or the decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
T.R. conceived and designed the study. B.J. analyzed the data and performed statistical analyses with assistance from K.B.P., W.H.D., A.R., S.-H.J., T.N., B.K., M.S.K., M.C., M.-S.P., M.R. and J.H., supervised by H.-H.W. and T.R. A.T. constructed its website. B.J., J.B., H.-H.W. and T.R. interpreted the results. B.J. and T.R. wrote the manuscript. J.H., D.A.K. and H.-H.W. critically reviewed this study. All the authors have read and approved the final manuscript. T.R. and H.-H.W. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
T.R. served as a scientific advisor for Merck and a consultant for Curie.Bio. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Julien Bryois, Sarah Gagliano Taliun, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SAIGE-QTL and colocalization compared to SingleBrain.
a, Comparison of the number of eGenes (q-value < 0.05) identified in this study (SingleBrain, n = 983), SAIGE-QTL of ROS/MAP-Columbia (n = 397), and eQTL summary statistics11. b,c, Pairwise sharing of genes with significant cis-eQTLs across major cell types. The numbers shown are Storey’s π1 for each cell type pair. π1 is an estimate of the proportion of true alternative hypotheses in the replication cell type, derived from the distribution of P-values. The columns are used for brain major eQTL lead single-nucleotide polymorphisms (SNPs) identified in SAIGE-QTL, and the replication rates were obtained from (b) ROS/MAP-Columbia11 and (c) this study. d, Scatterplots of effect size estimates from SAIGE-QTL (x-axis) and this study (y-axis) are shown for each of the seven cell types. Top variants in eGenes identified in SAIGE-QTL and eQTL variants of eGenes in SingleBrain were included. The Rb statistic20 indicates the correlation of the effect size estimates between matched SNP–gene pairs. e, Number of AD and PD GWAS loci with a PP4 > 0.8. The dark bars represent SingleBrain, and the light bars represent SAIGE-QTL. f,g, The SCAF11 PD GWAS locus and PTK2B AD GWAS locus at astrocytes ANO6 eQTL and oligodendrocytes PTK2B eQTL in SAIGE-QTL and this study. Reported P values are two-sided and not corrected for multiple testing. The x-axis and the y-axis represent the −log10(P value) for the association of variants with AD and PD, and gene expression, respectively. Single-nucleotide polymorphisms (SNPs) are colored by linkage disequilibrium (LD) with the lead GWAS SNP. MG, microglia; Ext, excitatory neuron; IN, inhibitory neuron; OD, oligodendrocyte; Ast, astrocyte; OPC, oligodendrocyte progenitor cell; End, endothelial cell.
Extended Data Fig. 2 SCZ GWAS27 loci with a PP4 > 0.8 and significant MR association in SingleBrain and MiGA.
The circle represents expression quantitative trait loci (eQTLs) with a PP4 > 0.8. Triangles denote significant genes in the Mendelian randomization (MR) results, with the orientation of the triangle corresponding to the direction of the effect. COLOC, colocalization; MR, Mendelian randomization; MiGA, Microglia Genomic Atlas; MG, microglia; Ext, excitatory neuron; IN, inhibitory neuron; Ast, astrocyte; OD, oligodendrocyte; OPC, oligodendrocyte progenitor cell; End, endothelial cell.
Extended Data Fig. 3 ALS, BDP, and MS GWAS loci with a PP4 > 0.8 and significant MR association in SingleBrain and MiGA.
a–c, The circle represents expression quantitative trait loci (eQTLs) with a PP4 > 0.8. Triangles denote significant genes in the MR results, with the orientation of the triangle corresponding to the direction of the effect—(a) amyotrophic lateral sclerosis (ALS)30, (b) bipolar disorder (BPD)31 and (c) multiple sclerosis (MS)29. ALS, amyotrophic lateral sclerosis; BPD, bipolar disorder; MS, multiple sclerosis; COLOC, colocalization; MR, Mendelian randomization; MiGA, Microglia Genomic Atlas; MG, microglia; Ext, excitatory neuron; IN, inhibitory neuron; Ast, astrocyte; OD, oligodendrocyte; OPC, oligodendrocyte progenitor cell; End, endothelial cell.
Extended Data Fig. 4 AD loci from three separate AD GWAS with a PP4 > 0.8 and significant MR association in SingleBrain and MiGA.
a–c, The circle represents expression quantitative trait loci (eQTLs) with a PP4 > 0.8. Triangles denote significant genes in the MR results, with the orientation of the triangle corresponding to the direction of the effect24,25,32. COLOC, colocalization; MR, Mendelian randomization; MiGA, Microglia Genomic Atlas; MG, microglia; Ext, excitatory neuron; IN, inhibitory neuron; Ast, astrocyte; OD, oligodendrocyte; OPC, oligodendrocyte progenitor cell; End, endothelial cell.
Extended Data Fig. 5 PD (proxy and clinical cases) and PD (clinical only) GWAS28 loci with a PP4 > 0.8 and significant MR association in SingleBrain and MiGA.
a,b, The circle represents expression quantitative trait loci (eQTLs) with a PP4 > 0.8. Triangles denote significant genes in the MR results, with the orientation of the triangle corresponding to the direction of the effect. PD GWAS including proxy cases (a). PD GWAS with clinically defined cases (b). PD, Parkinson’s disease; COLOC, colocalization; MR, Mendelian randomization; MiGA, Microglia Genomic Atlas; MG, microglia; Ext, excitatory neuron; IN, inhibitory neuron; Ast, astrocyte; OD, oligodendrocyte; OPC, oligodendrocyte progenitor cell; End, endothelial cell.
Extended Data Fig. 6 AD GWAS23 loci colocalized with disease- and sex-specific eQTLs in SingleBrain (PP4 > 0.8).
The shape represents each type of expression quantitative trait loci (eQTLs) with PP4 greater than 0.8. Disease-specific eQTL (left) and sex-specific eQTL (right). MG, microglia; Ext, excitatory neuron; IN, inhibitory neuron; Ast, astrocyte; OD, oligodendrocyte; OPC, oligodendrocyte progenitor cell; End, endothelial cell.
Extended Data Fig. 7 Analysis of the PBX1 excitatory neuron eQTL with BPD GWAS NUF2 locus.
a, PBX1 expression is associated with the rs4442348 genotype, specifically in excitatory neurons. Residual expression was PEER-adjusted. The nominal P-value and β from the linear regression model in the brain cell type expression quantitative trait loci (eQTLs) analysis are indicated above the box plots (n = 983). The box plots show the median, the box spans from the first to the third quartiles, and the whiskers extend 1.5× interquartile range (IQR) from the box. b, Locus zoom and fine-mapping of the NUF2 BPD genome-wide association study (GWAS) and PBX1 excitatory neurons eQTL. Labels refer to lead single-nucleotide polymorphisms (SNPs) and fine-mapping SNPs with P < 1 × 10−4. Reported P values are two-sided and not corrected for multiple testing. The x-axis represents chromosomal position, and the y-axis represents the −log10(P value) for the association of variants with BPD and gene expression. SNPs are colored by the linkage disequilibrium (LD) with the lead GWAS SNP. Fine-mapped SNPs with PIP > 0.95 from GWAS and eQTL are presented under the plot. The red vertical line indicates the position of GWAS, eQTL lead SNPs, and fine-mapped SNPs with PIP > 0.95 and P < 1 × 10−4. c, Fine-mapping of the NUF2 locus and combination with the GWAS lead SNP, eQTL lead SNP, fine-mapping SNPs, credible SNPs (from colocalization analaysis), and consensus SNPs (defined as SNPs in LD [r2 = 1] with GWAS lead SNP). SNPs are colored by the LD with the lead GWAS SNP, overlapped with cell-type-specific enhancers and promoters were defined in ref. 19. Genomic plots (hg19) of lead SNPs and fine-mapping SNPs with P < 1 × 10−4 and epigenomic data from the microglia chromatin immunoprecipitation (ChIP)–seq and proximity ligation-assisted ChIP (PLAC)–seq junctions. BPD, bipolar disorder; Ext, excitatory neuron; IN, inhibitory neuron; OD, oligodendrocyte; Ast, astrocyte; OPC, oligodendrocyte progenitor cell; MG, microglia; End, endothelial cell.
Extended Data Fig. 8 Colocalization between SingleBrain eQTLs and GTEx brain cortex sQTLs.
a,b, All (a) AD and (b) PD GWAS loci with a colocalization PP4 > 0.8 at GTEx brain cortex sQTLs3 with SingleBrain eQTLs. The shape indicates eQTL and sQTL; opacity and size of the shape were scaled to PP4. Numbers next to the circles represent PP4 values.
Supplementary information
Supplementary Information (download PDF )
Supplementary Note and Supplementary Figs. 1–22.
Supplementary Tables (download XLSX )
Supplementary Tables 1–21.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jang, B., BP, K., Tokolyi, A. et al. A meta-analysis of single-nucleus expression quantitative trait loci linking genetic risk to brain disorders. Nat Genet (2026). https://doi.org/10.1038/s41588-026-02541-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41588-026-02541-x