Abstract
CRISPR–Cas9 has yielded a plethora of effectors, including targeted transcriptional activators, base editors and prime editors. Current approaches for inducibly modulating Cas9 activity lack temporal precision and require extensive screening and optimization. We describe a versatile, chemically controlled and rapidly activated single-component DNA-binding Cas9 switch, ciCas9, which we use to confer temporal control over seven Cas9 effectors, including two cytidine base editors, two adenine base editors, a dual base editor, a prime editor and a transcriptional activator. Using these temporally controlled effectors, we analyze base editing kinetics, showing that editing occurs within hours and that rapid early editing of nucleotides predicts eventual editing magnitude. We also reveal that editing at preferred nucleotides within target sites increases the frequency of bystander edits. Thus, the ciCas9 switch offers a simple, versatile approach to generating chemically controlled Cas9 effectors, informing future effector engineering and enabling precise temporal effector control for kinetic studies.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequencing data are available in the Sequence Read Archive under accession number PRJNA879077. Source data are provided with this paper. Data for supplementary figures are available as supplementary datasets.
Code availability
Scripts written for parsing data and plotting figures are available on Github (https://github.com/cindytxwei/ciCas9effectors). Scripts written for the permutation analysis based on the chi-squared test statistic are also available on Github (https://github.com/omripel/BEAnalysis).
References
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR–Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 337, 816–821 (2020).
Nakamura, M., Gao, Y., Dominguez, A. A. & Qi, L. S. CRISPR technologies for precise epigenome editing. Nat. Cell Biol. 23, 11–22 (2021).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Gaudelli, N. M. et al. Programmable base editing of A·T to G·C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Newald, J. G. X. et al. A dual-deaminase CRISPR base editor enables concurrent adenine and cytosine editing. Nat. Biotechnol. 38, 861–864 (2020).
Sakata, R. C. et al. Base editors for simultaneous introduction of C-to-T and A-to-G mutations. Nat. Biotechnol. 38, 865–869 (2020).
Zhang, X. et al. Dual base editor catalyzes both cytosine and adenine base conversions in human cells. Nat. Biotechnol. 38, 856–860 (2020).
Chen, L. et al. Programmable C:G to G:C genome editing with CRISPR–Cas9-directed base excision repair proteins. Nat. Commun. 12, 1384 (2021).
Zhao, D. et al. New base editors change C to A in bacteria and C to G in mammalian cells. Nat. Biotechnol. 39, 35–40 (2021).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Gangopadhyay, S. A. et al. Precision control of CRISPR–Cas9 using small molecules and light. Biochemistry 58, 234–244 (2019).
Dow, L. E. et al. Inducible in vivo genome editing with CRISPR–Cas9. Nat. Biotechnol. 33, 390–394 (2015).
González, F. et al. An iCRISPR platform for rapid, multiplexable, and inducible genome editing in human pluripotent stem cells. Cell Stem Cell 15, 215–226 (2014).
Kleinjan, D. A., Wardrope, C., Nga Sou, S. & Rosser, S. J. Drug-tunable multidimensional synthetic gene control using inducible degron-tagged dCas9 effectors. Nat. Commun. 8, 1191 (2017).
Maji, B. et al. Multidimensional chemical control of CRISPR–Cas9. Nat. Chem. Biol. 13, 9–11 (2017).
Senturk, S. et al. Rapid and tunable method to temporally control gene editing based on conditional Cas9 stabilization. Nat. Commun. 8, 14370 (2017).
Liu, K. I. et al. A chemical-inducible CRISPR–Cas9 system for rapid control of genome editing. Nat. Chem. Biol. 12, 980–987 (2016).
Nguyen, D. P. et al. Ligand-binding domains of nuclear receptors facilitate tight control of split CRISPR activity. Nat. Commun. 7, 12009 (2016).
Oakes, B. L. et al. Profiling of engineering hotspots identifies an allosteric CRISPR–Cas9 switch. Nat. Biotechnol. 34, 646–651 (2016).
Zhao, C. et al. HIT-Cas9: a CRISPR–Cas9 genome-editing device under tight and effective drug control. Mol. Ther. Nucleic Acids 13, 208–219 (2018).
Zetsche, B., Volz, S. E. & Zhang, F. A split-Cas9 architecture for inducible genome editing and transcription modulation. Nat. Biotechnol. 33, 139–142 (2015).
Davis, K. M., Pattanayak, V., Thompson, D. B., Zuris, J. A. & Liu, D. R. Small molecule-triggered Cas9 protein with improved genome-editing specificity. Nat. Chem. Biol. 11, 316–318 (2015).
Rose, J. C. et al. Rapidly inducible Cas9 and DSB-ddPCR to probe editing kinetics. Nat. Methods 14, 891–896 (2017).
Liu, Y. et al. Very fast CRISPR on demand. Science 368, 1265–1269 (2020).
Park, J. et al. Recording of elapsed time and temporal information about biological events using Cas9. Cell 184, 1047–1063.e23 (2021).
Brinkman, E. K. et al. Kinetics and fidelity of the repair of Cas9-induced double-strand DNA breaks. Mol. Cell 70, 801–813.e6 (2018).
Kundert, K. et al. Controlling CRISPR–Cas9 with ligand-activated and ligand-deactivated sgRNAs. Nat. Commun. 10, 2127 (2019).
Gao, Y. et al. Complex transcriptional modulation with orthogonal and inducible dCas9 regulators. Nat. Methods 13, 1043–1049 (2016).
Foight, G. W. et al. Multi-input chemical control of protein dimerization for programming graded cellular responses. Nat. Biotechnol. 37, 1209–1216 (2019).
Polstein, L. R. & Gersbach, C. A. A light-inducible CRISPR–Cas9 system for control of endogenous gene activation. Nat. Chem. Biol. 11, 198–200 (2015).
Nihongaki, Y., Yamamoto, S., Kawano, F., Suzuki, H. & Sato, M. CRISPR–Cas9-based photoactivatable transcription system. Chem. Biol. 22, 169–174 (2015).
Berríos, K. N. et al. Controllable genome editing with split-engineered base editors. Nat. Chem. Biol. 17, 1262–1270 (2021).
Rose, J. C., Stephany, J. J., Wei, C. T., Fowler, D. M. & Maly, D. J. Rheostatic control of Cas9-mediated DNA double strand break (DSB) generation and genome editing. ACS Chem. Biol. 13, 438–442 (2018).
Zalatan, J. G. et al. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds. Cell 160, 339–350 (2015).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Lapinaite, A. et al. DNA capture by a CRISPR–Cas9-guided adenine base editor. Science 369, 566–571 (2020).
Jang, H.-K. et al. High-purity production and precise editing of DNA base editing ribonucleoproteins. Sci. Adv. 7, eabg2661 (2021).
Tao, Z.-F. et al. Discovery of a potent and selective BCL-XL inhibitor with in vivo activity. ACS Med. Chem. Lett. 5, 1088–1093 (2014).
Wilson, W. H. et al. Navitoclax, a targeted high-affinity inhibitor of BCL-2, in lymphoid malignancies: a phase 1 dose-escalation study of safety, pharmacokinetics, pharmacodynamics, and antitumour activity. Lancet Oncol. 11, 1149–1159 (2010).
Newby, G. A. et al. Base editing of haematopoietic stem cells rescues sickle cell disease in mice. Nature 595, 295–302 (2021).
Hanna, R. E. et al. Massively parallel assessment of human variants with base editor screens. Cell 184, 1064–1080.e20 (2021).
Boersma, M. D., Sadowsky, J. D., Tomita, Y. A. & Gellman, S. H. Hydrophile scanning as a complement to alanine scanning for exploring and manipulating protein–protein recognition: application to the Bim BH3 domain. Protein Sci. 17, 1232–1240 (2008).
Arbeithuber, B., Makova, K. D. & Tiemann-Boege, I. Artifactual mutations resulting from DNA lesions limit detection levels in ultrasensitive sequencing applications. DNA Res. 23, 547–559 (2016).
Alseth, I., Dalhus, B. & Bjørås, M. Inosine in DNA and RNA. Curr. Opin. Genet. Dev. 26, 116–123 (2014).
Wang, Q. et al. A general theoretical framework to design base editors with reduced bystander effects. Nat. Commun. 12, 6529 (2021).
Ede, C., Chen, X., Lin, M.-Y. & Chen, Y. Y. Quantitative analyses of core promoters enable precise engineering of regulated gene expression in mammalian cells. ACS Synth. Biol. 5, 395–404 (2016).
Matreyek, K. A., Stephany, J. J., Chiasson, M. A., Hasle, N. & Fowler, D. M. An improved platform for functional assessment of large protein libraries in mammalian cells. Nucleic Acids Res. 11, 1782–1712 (2019).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Acknowledgements
This work was supported by the NIH, grant nos. RM1HG010461 (NHGRI) and R01GM109110 (NIGMS) to D.M.F., and grant no. R01GM145011 (NIGMS) to D.J.M. O.P. was supported in part by a fellowship from the Edmond J. Safra Center for Bioinformatics at Tel Aviv University. E.B. is a Faculty Fellow of the Edmond J. Safra Center for Bioinformatics at Tel Aviv University.
Author information
Authors and Affiliations
Contributions
C.T.W., D.J.M. and D.M.F. conceived the work and wrote the paper. C.T.W. performed all base and prime editing experiments and data analysis. C.T.W. performed the transcriptional activation experiments with dciCas9-VPR in Fig. 1 and Supplementary Fig. 3. N.A.P. and R.L.P. performed the transcriptional activation experiments with dciCas9-VPR in Extended Data Fig. 1. O.P. and E.B. designed and performed the chi-squared test for the base editing dependency analysis.
Corresponding authors
Ethics declarations
Competing interests
C.T.W. is a current employee at the Novartis Institutes for BioMedical Research Inc.
Peer review
Peer review information
Nature Chemical Biology thanks Chase Beisel and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
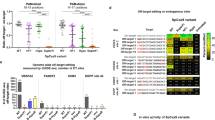
Extended Data Fig. 1 Transcriptional activation of a diverse range of target sequences using dciCas9-VPR.
Activation of an EGFP reporter locus downstream of the indicated target sequence using dCas9-VPR or dciCas9-VPR targeted to this synthetic locus (see Methods) in the presence or absence of 1 μM A115. Cells were treated with A115 for 48 hr prior to flow cytometry analysis. Bars represent the geometric mean EGFP fluorescence ± SEM of three cell culture replicates. Three cell culture replicates are shown in the blue circles overlapping each bar.
Extended Data Fig. 2 Chemically-controlled cytidine base editors without codon optimization.
a) Schematic of the domain arrangements in the unmodified BE4max and AncBE4max base editors and the chemically-controlled BE4max and AncBE4max base editors without codon optimization and using the ciCas9(L22) variant. 3 different versions of ciCas9 were used, ciCas9(L22), ciCas9(L22) without a Flag-tag (ΔFlag), and ciCas9(L22) without a Flag-tag and additional SV40-NLS (ΔFlag/ΔNLS). b-c) C-to-T editing frequency with BE4max and BE4max-ciCas9 at the EMX1 (b) and HEK3 (c) target sites. d-e) C-to-T editing frequency with AncBE4max and AncBE4max-ciCas9 at the EMX1 (d) and HEK3 (e) target sites. BE4max and AncBE4max editing were measured at 48 and 72 hr after co-transfection of BE4max and sgRNA. BE4max-ciCas9 and AncBE4max-ciCas9 editing were measured at 24 and 72 hr after 1 μM A115 addition. C-to-T editing is shown at the 2 nucleotides in each target site with highest editing frequency with the Cas9 version of base editors (BE4max or AncBE4max). The 2 different nucleotides are indicated by color in the target sequence. Bars show mean editing frequency ± SEM of 3 cell culture replicates with white circles showing individual replicates.
Extended Data Fig. 3 Heatmaps of base editing by chemically-controlled base editors compared to unmodified base editors.
a–b) Heatmaps of BE4max, ciBE4max (a) and AncBE4max, ciAncBE4max (b) C-to-T base editing as a percentage of the highest edited nucleotide for each editor throughout the entire indicated target sites. c-d) Heatmaps of ABEmax, ciABEmax (c) and ABE8e, ciABE8e (d) A-to-G base editing as a percentage of the highest edited nucleotide for each editor throughout the entire indicated target sites. Editing is represented as a percentage of the highest edited nucleotide to allow comparison of the spatial distribution of edits between base editors. Each row shows an individual cell culture replicate. Editing frequencies of the unmodified base editors were quantified at 72 hr after transfection for the HEK3 target site and 48 hr after transfection for the ABE9 and HEK2 target sites. Chemically-controlled base editing frequencies were quantified at 72 hr after 1 μM A115 addition to HEK293T cells for the HEK3 target site and 24 hr after 1 μM A115 addition to HEK293T cells for the ABE9 and HEK2 target sites. The control shows untransfected cells harvested at the same time as the chemically-controlled base editors. The numbers below the heatmaps show the position of the nucleotide from the most PAM-distal nucleotide.
Extended Data Fig. 4 Early time points in time courses of base editing with the chemically-controlled base editors.
Early time courses of chemically-controlled base editing using ciBE4max (a), ciABEmax (b), and ciABE8e (c) activated using 1 μM A115 at the indicated target sites. Time courses shown for the nucleotide colored in the target sequences shown. Numbers underneath the target sequence show the position of the nucleotide from the most PAM-distal nucleotide. Bars show mean editing ± SEM of 3 cell culture replicates with white circles showing individual replicates. Significance of editing at different time points were compared to editing frequency at 0 hr using a One-way ANOVA, statistical values shown in Supplementary Table 2. In (a), **P = 0.0033, ***P = 0.0005, ****P = < 0.0001. In (b), ****P = < 0.0001, ABE9 ***P = 0.0003, ABE16 ***P = 0.0002. In (c), ****P = < 0.0001, ***P = 0.0007, ABE16 *P = 0.0315, HEK3 *P = 0.0228.
Extended Data Fig. 5 Time courses of base editing with the chemically-controlled base editors.
a) Time course of chemically-controlled cytidine base editing by ciBE4max at the ABE9, EMX1, HEK2, and HEK3 target sites. ciBE4max was activated with 1 μM A115. Cells were harvested and editing was quantified at specified time points after activation. Colors of lines represent the corresponding nucleotide within the target site. Numbers underneath the target sequence show the position of the nucleotide from the most PAM-distal nucleotide. b,c) Time course of chemically-controlled adenine base editing by ciABEmax (b) and ciABE8e (c) at the ABE9, ABE16, HEK2, and HEK3 target sites. ciABEmax and ciABE8e were activated with 1 μM A115. Cells were harvested and editing was quantified at specified time points after activation. Colors of lines represent the corresponding nucleotide within the target site. Numbers underneath the target sequence show the position of the nucleotide from the most PAM-distal nucleotide. Data represented as mean editing ± SEM of 3 cell culture replicates. Time courses shown for all nucleotides where base editing frequency was greater than 0.5% at 24 hr after A115 addition.
Extended Data Fig. 6 Time courses of ciBE4max base editing allele outcomes.
Time course of allele formation by ciBE4max after activation with 1 μM A115 or DMSO. Black lines and circles show editing with 1 μM A115, gray lines and circles show editing with DMSO. Data represented as mean allele frequency ± SEM of 3 cell culture replicates.
Extended Data Fig. 7 Time courses of ciABE8e base editing allele outcomes.
Time course of allele formation by ciABE8e after activation with 1 μM A115 or DMSO. Black lines and circles show editing with 1 μM A115, gray lines and circles show editing with DMSO. Data represented as mean allele frequency ± SEM of 3 cell culture replicates.
Extended Data Fig. 8 Time course of measured and expected allele frequencies by ciBE4max.
Measured and expected allele frequencies over time created by ciBE4max that show a dependent model of base editing for multiply-edited alleles. Black lines and solid circles show measured allele frequencies, gray lines and open circles show expected allele frequencies. Measured data represented as mean editing frequency ± SEM of 3 cell culture replicates. Expected editing frequency represented as mean expected editing frequency ± relative error. Calculations for expected frequency and relative error described in the methods.
Extended Data Fig. 9 Time course of measured and expected allele frequencies by ciABE8e.
Measured and expected allele frequencies over time created by ciABE8e that show a dependent model of base editing for multiply-edited alleles. Black lines and solid circles show measured allele frequencies, gray lines and open circles show expected allele frequencies. Measured data represented as mean editing frequency ± SEM of 3 cell culture replicates. Expected editing frequency represented as mean expected editing frequency ± relative error. Calculations for expected frequency and relative error described in the Methods.
Extended Data Fig. 10 Heatmaps of SPACE and ciSPACE base editing.
Heatmaps of SPACE and ciSPACE editing through the entire HEK2 (top) and HEK3 (bottom) target sites. A-to-G base editing is shown in pink, C-to-T base editing is shown in blue. Editing is shown as a percentage of the highest edited nucleotide for each editor for that target site. Editing is represented as a percentage of the highest edited nucleotide to allow comparison of the spatial distribution of edits between base editors. Each row shows an individual cell culture replicate. SPACE editing frequencies were quantified at 72 hr after transfection and ciSPACE editing frequencies were quantified at 72 hr after 1 μM A115 addition to HEK293T cells. The control shows untransfected cells harvested at the same time as ciSPACE. The numbers below the heatmaps show the position of the nucleotide from the most PAM-distal nucleotide.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–22 and Tables 1–7.
Supplementary Data (download XLSX )
Supplementary Datasets 1–15 corresponding to the Supplementary Figs.
Source data
Source Data Fig. 1 (download XLSX )
Median fluorescence values from flow cytometry.
Source Data Fig. 2 (download XLSX )
Base editing frequencies for cytidine base editors.
Source Data Fig. 3 (download XLSX )
Base editing frequencies for adenine base editors.
Source Data Fig. 4 (download XLSX )
Base editing frequencies for time courses.
Source Data Fig. 5 (download XLSX )
Base editing allele frequencies for time courses.
Source Data Fig. 6 (download XLSX )
Base editing frequencies for SPACE/ciSPACE, prime editing frequencies for PE2/ciPE2.
Source Data Extended Data Fig. 1 (download XLSX )
Median fluorescence values from flow cytometry, additional target sites.
Source Data Extended Data Fig. 2 (download XLSX )
Base editing frequencies for non-codon-optimized cytidine base editors.
Source Data Extended Data Fig. 3 (download XLSX )
Heat maps of base editing frequencies by unmodified and chemically controlled base editors.
Source Data Extended Data Fig. 4 (download XLSX )
Base editing frequencies at early time points for chemically controlled base editors.
Source Data Extended Data Fig. 5 (download XLSX )
Base editing frequencies for full time courses.
Source Data Extended Data Fig. 6 (download XLSX )
Base editing allele frequencies for ciBE4max.
Source Data Extended Data Fig. 7 (download XLSX )
Base editing allele frequencies for ciABE8e.
Source Data Extended Data Fig. 8 (download XLSX )
Measured and expected allele frequency time courses for ciBE4max.
Source Data Extended Data Fig. 9 (download XLSX )
Measured and expected allele frequency time courses for ciABE8e.
Source Data Extended Data Fig. 10 (download XLSX )
Heat maps of base editing frequencies by unmodified and chemically controlled SPACE.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wei, C.T., Popp, N.A., Peleg, O. et al. A chemically controlled Cas9 switch enables temporal modulation of diverse effectors. Nat Chem Biol 19, 981–991 (2023). https://doi.org/10.1038/s41589-023-01278-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-023-01278-6
This article is cited by
-
Precision multiplexed base editing in human cells using Cas12a-derived base editors
Nature Communications (2025)
-
Synthetic macromolecular switches for precision control of therapeutic cell functions
Nature Reviews Bioengineering (2024)