Abstract
Targeted protein degradation (TPD) is an emerging therapeutic strategy that would benefit from new chemical entities with which to recruit a wider variety of ubiquitin E3 ligases to target proteins for proteasomal degradation. Here we describe a TPD strategy involving the recruitment of FBXO22 to induce degradation of the histone methyltransferase and oncogene NSD2. UNC8732 facilitates FBXO22-mediated degradation of NSD2 in acute lymphoblastic leukemia cells harboring the NSD2 gain-of-function mutation p.E1099K, resulting in growth suppression, apoptosis and reversal of drug resistance. The primary amine of UNC8732 is metabolized to an aldehyde species, which engages C326 of FBXO22 to recruit the SCFFBXO22 Cullin complex. We further demonstrate that a previously reported alkyl amine-containing degrader targeting XIAP is similarly dependent on SCFFBXO22. Overall, we present a potent NSD2 degrader for the exploration of NSD2 disease phenotypes and a new FBXO22-recruitment strategy for TPD.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All supporting data for this study can be found within the article, its extended data figures and Supplementary Information documents. All mass spectrometry data are available at massive.ucsd.edu—accession MSV000093206. The sequence of the pCDF-BirA vector is available at GenBank (accession JF914075.1). Source data are provided with this paper.
Code availability
Analysis code for further BioID data processing and visualization is available via Zenodo at https://doi.org/10.5281/zenodo.10930672 (ref. 52).
References
Békés, M., Langley, D. R. & Crews, C. M. PROTAC targeted protein degraders: the past is prologue. Nat. Rev. Drug Discov. 21, 181–200 (2022).
Humphreys, L. M., Smith, P., Chen, Z., Fouad, S. & D’Angiolella, V. The role of E3 ubiquitin ligases in the development and progression of glioblastoma. Cell Death Differ. 28, 522–537 (2021).
Kramer, L. T. & Zhang, X. Expanding the landscape of E3 ligases for targeted protein degradation. Curr. Res. Chem. Biol. 2, 100020 (2022).
Schapira, M., Calabrese, M. F., Bullock, A. N. & Crews, C. M. Targeted protein degradation: expanding the toolbox. Nat. Rev. Drug Discov. 18, 949–963 (2019).
Sun, J. et al. Downregulation of miR-21 inhibits the malignant phenotype of pancreatic cancer cells by targeting VHL. Onco Targets Ther. 12, 7215–7226 (2019).
Hu, J. et al. Tumor heterogeneity in VHL drives metastasis in clear cell renal cell carcinoma. Signal Transduct. Target Ther. 8, 155 (2023).
Gopalsamy, A. Selectivity through targeted protein degradation (TPD). J. Med. Chem. 65, 8113–8126 (2022).
Hanley, R. P. et al. Discovery of a potent and selective targeted NSD2 degrader for the reduction of H3K36me2. J. Am. Chem. Soc. 145, 8188 (2023).
Meng, F. et al. Discovery of a first-in-class degrader for nuclear receptor binding SET domain protein 2 (NSD2) and Ikaros/Aiolos. J. Med. Chem. 65, 10611–10625 (2022).
LegaardAndersson, J. et al. Discovery of NSD2‐degraders from novel and selective DEL hits. ChemBioChem 24, e202300515 (2023).
Kuo, A. J. et al. NSD2 links dimethylation of Histone H3 at Lysine 36 to oncogenic programming. Mol. Cell 44, 609–620 (2011).
Jaffe, J. D. et al. Global chromatin profiling reveals NSD2 mutations in pediatric acute lymphoblastic leukemia. Nat. Genet. 45, 1386–1391 (2013).
Sengupta, D. et al. NSD2 dimethylation at H3K36 promotes lung adenocarcinoma pathogenesis. Mol. Cell 81, 4481–4492.e9 (2021).
Yuan, S. et al. Global regulation of the histone mark H3K36me2 underlies epithelial plasticity and metastatic progression. Cancer Discov. 10, 854–871 (2020).
den Besten, W. et al. Primary amine tethered small molecules promote the degradation of X-linked inhibitor of apoptosis protein. J. Am. Chem. Soc. 143, 10571–10575 (2021).
Dilworth, D. et al. A chemical probe targeting the PWWP domain alters NSD2 nucleolar localization. Nat. Chem. Biol. 18, 56–63 (2022).
Swaroop, A. et al. An activating mutation of the NSD2 histone methyltransferase drives oncogenic reprogramming in acute lymphocytic leukemia. Oncogene 38, 671–686 (2019).
Li, J. et al. PRC2 inhibitors overcome glucocorticoid resistance driven by NSD2 mutation in pediatric acute lymphoblastic leukemia. Cancer Discov. 12, 186–203 (2022).
Sato, K. et al. Structural basis of the regulation of the normal and oncogenic methylation of nucleosomal histone H3 Lys36 by NSD2. Nat. Commun. 12, 6605 (2021).
Li, W. et al. Molecular basis of nucleosomal H3K36 methylation by NSD methyltransferases. Nature 590, 498–503 (2021).
Mandadapu, S. R. et al. Inhibition of norovirus 3CL protease by bisulfite adducts of transition state inhibitors. Bioorg. Med. Chem. Lett. 23, 62–65 (2013).
Gingras, A.-C., Abe, K. T. & Raught, B. Getting to know the neighborhood: using proximity-dependent biotinylation to characterize protein complexes and map organelles. Curr. Opin. Chem. Biol. 48, 44–54 (2019).
Baek, K. et al. Systemwide disassembly and assembly of SCF ubiquitin ligase complexes. Cell 186, 1895–1911.e21 (2023).
Horn-Ghetko, D. et al. Ubiquitin ligation to F-box protein targets by SCF–RBR E3–E3 super-assembly. Nature 590, 671–676 (2021).
Machleidt, T. et al. NanoBRET—a novel BRET platform for the analysis of protein–protein interactions. ACS Chem. Biol. 10, 1797–1804 (2015).
Vu, V., Szewczyk, M. M., Nie, D. Y., Arrowsmith, C. H. & Barsyte-Lovejoy, D. Validating small molecule chemical probes for biological discovery. Annu. Rev. Biochem. 91, 61–87 (2022).
Masson, G. R. et al. Recommendations for performing, interpreting and reporting hydrogen deuterium exchange mass spectrometry (HDX-MS) experiments. Nat. Methods 16, 595–602 (2019).
James, E. I., Murphree, T. A., Vorauer, C., Engen, J. R. & Guttman, M. Advances in hydrogen/deuterium exchange mass spectrometry and the pursuit of challenging biological systems. Chem. Rev. 122, 7562–7623 (2022).
Bateman, A. et al. UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res. 51, D523–D531 (2023).
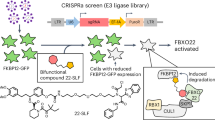
Basu, A. A. et al. A CRISPR activation screen identifies FBXO22 as an E3 ligase supporting targeted protein degradation. Preprint at bioRxiv https://doi.org/10.1101/2023.09.15.557708 (2023).
Lignitto, L. et al. Nrf2 activation promotes lung cancer metastasis by inhibiting the degradation of Bach1. Cell 178, 316–329.e18 (2019).
Lin, M. et al. Fbxo22 promotes cervical cancer progression via targeting p57Kip2 for ubiquitination and degradation. Cell Death Discov. 13, 805 (2022).
Ge, M.-K. et al. FBXO22 degrades nuclear PTEN to promote tumorigenesis. Nat. Commun. 11, 1720 (2020).
Cheng, J. et al. Emerging role of FBXO22 in carcinogenesis. Cell Death Discov. 6, 66 (2020).
Chen, S. et al. Pan-cancer analyses reveal oncogenic role and prognostic value of F-Box only protein 22. Front. Oncol. 11, 790912 (2022).
Kim, D. I. et al. Probing nuclear pore complex architecture with proximity-dependent biotinylation. Proc. Natl Acad. Sci. USA 111, E2453–E2461 (2014).
Soucy, T. A. et al. An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature 458, 732–736 (2009).
Meier, F. et al. Parallel accumulation–serial fragmentation (PASEF): multiplying sequencing speed and sensitivity by synchronized scans in a trapped ion mobility device. J. Proteome Res. 14, 5378–5387 (2015).
Zhang, X. et al. Proteome-wide identification of ubiquitin interactions using UbIA-MS. Nat. Protoc. 13, 530–550 (2018).
Roux, K. J., Kim, D. I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Coyaud, E. et al. BioID-based identification of Skp Cullin F-box (SCF)β-TrCP1/2 E3 ligase substrates*. Mol. Cell. Proteomics 14, 1781–1795 (2015).
Kessner, D., Chambers, M., Burke, R., Agus, D. & Mallick, P. ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24, 2534–2536 (2008).
Craig, R. & Beavis, R. C. TANDEM: matching proteins with tandem mass spectra. Bioinformatics 20, 1466–1467 (2004).
Eng, J. K., Jahan, T. A. & Hoopmann, M. R. Comet: an open-source MS/MS sequence database search tool. Proteomics 13, 22–24 (2013).
Deutsch, E. W. et al. A guided tour of the Trans‐Proteomic Pipeline. Proteomics 10, 1150–1159 (2010).
Liu, G. et al. ProHits: integrated software for mass spectrometry–based interaction proteomics. Nat. Biotechnol. 28, 1015–1017 (2010).
Teo, G. et al. SAINTexpress: improvements and additional features in Significance Analysis of INTeractome software. J. Proteomics 100, 37–43 (2014).
Hutchinson, A. & Seitova, A. Production of recombinant PRMT proteins using the Baculovirus expression vector system. J. Vis. Exp. 2021, e62510 (2021).
Wu, T. et al. Three essential resources to improve differential scanning fluorimetry (DSF) experiments. Preprint at bioRxiv https://doi.org/10.1101/2020.03.22.002543 (2020).
Scott, D. C. et al. Two distinct types of E3 ligases work in unison to regulate substrate ubiquitylation. Cell 166, 1198–1214.e24 (2016).
Duda, D. M. et al. Structural insights into NEDD8 activation of Cullin-RING ligases: conformational control of conjugation. Cell 134, 995–1006 (2008).
d0minicO/NSD2_BioID: NSD2 BioID. Zenodo https://doi.org/10.5281/zenodo.10930672 (2024).
Acknowledgements
The authors thank the members of the James laboratory and Stephen Frye for helpful discussions and input throughout the project. The authors thank P. H. Buttery and J. L. R. Sanchez for the review of experimental data. This work is supported by grants from the Canadian Institutes of Health Research (CIHR) (FDN154328, OGB190363) and the Princess Margaret Cancer Foundation to C.H.A., from the National Institutes of Health (NCI) (R01CA242305) to L.I.J., a Leukemia and Lymphoma Society Specialized Center for Research and Florida Department of Health Grant 22L03 to J.D.L., and grants from the CIHR (PJT156093) and Princess Margaret Cancer Foundation to B.R. D.Y.N. is supported by a Canada Graduate Scholarship – Doctoral Research Award from CIHR (494204) and a Doctoral Training Scholarship from Fonds de recherche du Québec – Santé (320128). J.R.T. is supported by the UNC Lineberger Comprehensive Cancer Center Cancer Epigenetics Training Program (5T32CA217824-05). D.B.-L. is supported by CRS grant 25418. D.W. is supported by the NSERC Discovery (RGPIN-480432) and NSERC Collaborative Research and Development (CRDPJ-504037) grants. Portions of this work have been supported by certain funds managed by Deerfield Management Company, L.P. Deerfield Management Company is a healthcare-focused investment management firm. The Structural Genomics Consortium is a registered charity (no. 1097737) that receives funds from Bayer AG, Boehringer Ingelheim, BristolMyersSquibb, Genentech, Genome Canada through Ontario Genomics Institute (OGI-196), EU/EFPIA/OICR/McGill/KTH/Diamond Innovative Medicines Initiative 2 Joint Undertaking (EUbOPENGrant875510), Janssen, Merck KGaA (also known as EMD in Canada and the United States), Pfizer and Takeda. This material is based in part upon work supported by the National Science Foundation under grant no. CHE-1726291. The authors thank the University of North Carolina’s Department of Chemistry Mass Spectrometry Core Laboratory for their assistance with mass spectrometry analysis. The authors thank Syngene International for support of portions of this work.
Author information
Authors and Affiliations
Contributions
D.Y.N., J.L., A.P., M.S., A.J.L. and M.M. designed, performed and analyzed cellular experiments. J.R.T., R.P.H. and L.I.J. designed and synthesized the compounds. S.S., M.K., T.M.G.K., F.L. and D.Y.N. designed, performed and analyzed biophysical experiments. S.D. and M.K. performed cloning and protein purification. J.S.-G., M.E.R.M., D.D.G.O. and D.Y.N. designed, performed and analyzed BioID and global proteomics experiments. E.W. and M.K. designed, performed and analyzed HDX-MS experiments. E.P. performed in vitro ubiquitination experiments. L.Z.P., D.B.-L., N.G.B., A.M.B., A.W.S., J.L.C., D.J.W., B.R., J.D.L., L.I.J. and C.H.A. provided supervision and/or funding. D.Y.N., J.R.T., L.I.J. and C.H.A. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.D.G.O. is an employee of Amphista Therapeutics, a company that is developing TPD therapeutic platforms. A.M.B. and A.W.S. are employees of Deerfield Management Company, a healthcare-focused investment management firm. The remaining authors declare no competing interests.
Peer review
Reviewer Recognition
Nature Chemical Biology thanks Milka Kostic and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
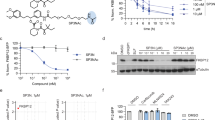
Extended Data Fig. 1 UNC8732 inhibits cell growth and restores glucocorticoid sensitivity of NSD2 p.E1099K mutant ALL cells.
(a): Total proteome analysis (label-free quantification) of U2OS cells treated with 2 µM UNC8732 or an equivalent volume of DMSO for 3 hours. Volcano plot of all quantified proteins (8240) with log2 fold change shown on the x-axis (dashed line equivalent to a 2-fold change) and -log10 adjusted p values shown on the y-axis (dashed line equivalent to p.adjusted = 0.05). Adjusted p-values were derived from Limma and DEP packages (see methods) and were adjusted using the Benjamini-Hochberg procedure. N=5 independent experiments. (b-d): Viability of isogenic RCH-ACV determined by CellTiter-Glo assay after treatment with varying concentrations of UNC8732 and UNC8884 for (b) 12 days, (c) 15 days, or (d) 21 days. (e-f): Apoptosis of isogenic RCH-ACV detected using annexin V/PI staining by flow cytometry after treatment with varying concentrations of UNC8732 and UNC8884 for (e) 15 days or (f) 21 days. (g-h): Viability of NSD2 mutant RCH-ACV cells determined by CellTiter-Glo after the pretreatment of varying concentrations of UNC8732 and UNC8884 for (g) 12 days or (h) 15 days followed by dexamethasone (1 µM) for 72 hours. (i-j): Apoptosis of NSD2 mutant RCH-ACV cell line detected using annexin V/PI staining by flow cytometry after the pretreatment of varying concentrations of UNC8732 and UNC8884 for (i) 12 days or (j) 15 days followed by dexamethasone (1 µM) for 72 hours. Data represents the mean ± SEM from three biological replicates. The statistical significance was evaluated using the Two-way ANOVA test and the p-values are displayed. WT, NSD2 WT; Mut, NSD2 p.E1099K; Dex, Dexamethasone. See Supplementary Figs. 3, 4 for representative flow cytometric analysis plots.
Extended Data Fig. 2 UNC8732 metabolism to the corresponding aldehyde drives NSD2 degradation.
(a): Structure of UNC8153 prodrug and its corresponding aldehyde metabolite. (b): MS peak area ratio representing the relative levels of UNC8153 and its associated aldehyde species in cell-free DMEM + 10% FBS at indicated time points. * For Extended Data Fig. 2b–d, the experimental treatment was designed for 0h. However, due to the sample preparation process for MS, there’s a brief period of simultaneous presence of the compounds in their respective conditions. (c): MS peak area ratio representing the relative levels of UNC8153 and its associated aldehyde species in cell-free DMEM with no FBS at indicated time points. (d): MS peak area ratio representing the relative levels of UNC8732 and its associated aldehyde species in cell-free DMEM + 10% FBS + 10 mM aminoguanidine (AG) at indicated time points. (e): Structure of UNC8153 prodrug and its corresponding aldehyde metabolite. (f): MS peak area ratio representing the relative level of the aldehyde adduct of UNC9801 upon addition of UNC9801 to DMEM + 10% FBS or DMEM only at indicated time points in hours.
Extended Data Fig. 3 FBXO22 is responsible for compound-mediated degradation of NSD2.
(a): U2OS cells were co-treated with DMSO control or 2 μM of UNC8732 and indicated concentrations of MLN4924 neddylation inhibitor for 24h. Representative immunoblot shown. Experiment repeated independently three times with consistent results. (b): U2OS cells were co-treated with DMSO control or 2 μM of UNC8732 and indicated concentrations of MG-132 proteosome inhibitor for 3h. Representative immunoblot shown. Experiment repeated independently three times with consistent results. (c): T-Rex 293 cells stable cell lines with tetracycline inducible NSD2-FLAG-miniTurbo was treated with 10 μM MG-132 and the indicated combination of DMSO, 5 μM UNC8732, tetracycline (1 μg/mL) and biotin (50 μM). (d): Principal Components analysis (PCA) of the BioID spectral counts data from DMSO (blue) or 5 μM UNC8732 (red) treated samples, each containing three independent experiments with 2 technical replicates for each independent experiment. (e): Uniform Manifold Approximation and Projection (UMAP) analysis of the BioID data of DMSO (blue) or 5 μM UNC8732 (red), each containing 3 independent experiments with 2 technical replicates for each independent experiment. (f): AlphaFold protein structure prediction of the SKP1-FBXO22 fusion protein. SKP1 is predicted to interact with the F-box domain of FBXO22 and the FIST_C domain remains some-what separate for putative substrate interactions. (g): SDS-PAGE analysis of the recombinant SKP1-FBXO22 fusion protein and the NSD2-PWWP1 post-purification. (h): Left: Recombinant proteins of SKP1-FBXO22 fusion apo or SKP1-FBXO22 fusion + NSD2-PWWP1 + UNC10088 analyzed using a Superdex 200 Increase 10/300 GL column in the indicated combinations. Right: SDS-PAGE gel showing the eluted proteins of the SKP1-FBXO22 + UNC10088 + NSD2-PWWP1 condition at the indicated peaks. (i): Left: Co-expressed SKP1/FBXO22 apo or co-expressed SKP1/FBXO22 + NSD2-PWWP1 + UNC10088 was analyzed using a Superdex 200 Increase 10/300 GL column in the indicated combinations. Right: SDS-PAGE gel showing the eluted proteins of the SKP1-FBXO22 + UNC10088 + NSD2-PWWP1 condition at the indicated peaks.
Extended Data Fig. 4 Aldehyde degraders bind FBXO22 in a cysteine 326-dependent manner.
(a): Representative BLI sensorgrams upon the addition of increasing concentrations of UNC10088 and a fixed concentration of NSD2-PWWP1 (2 µM). WT or C326A-mutant SKP1-FBXO22 was loaded on SA biosensors to an average response of 1 nm. Curves are shown as the average of three independent experiments. (b): In vitro NSD2-PWWP1 ubiquitination with the indicated sets of substrate priming machinery, UBCH5B or UBCH7 with HHARI. Each reaction also contained CUL1-RBX1 and CDC34B. The reaction mixture was analyzed by SDS-PAGE and fluorescence scanning. NSD2-PWWP1 was subject to a sortase reaction for fluorescent labeling. The gel presented is representative of three independent experiments. (c): Immunoblotting following transfection of empty vector pcDNA, WT FBXO22-HT, or C326A-mutant FBXO22-HT in U2OS cells for 24h. (d): Experimental flow chart for the designed ‘wash’ experiment to test for reversibility of the compound-FBXO22 interaction. *0 hour denotes the estimated concentration based on the series of dilutions performed. (e): Normalized fluorescence data from DSF for samples obtained from the experiment described in panel E. The data represents the mean ± SD of 3 technical replicates from one independent experiment. (f): Schematic of the reaction between the aldehyde-containing degrader compound and the C326 of FBXO22. (g): Percent Deuterium Exchange of C326-spanning peptides of co-purified SKP1/FBXO22 in the presence (orange) and absence (black) of UNC10088 over 12 hours. The data represents the mean ± 3σ (3 × SD) of a triplicate technical measurement (n=3).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–6 and note.
Supplementary Data 1 (download XLSX )
Global proteomics summary.
Supplementary Data 2 (download XLSX )
BioID SAINT PPI summary.
Source data
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nie, D.Y., Tabor, J.R., Li, J. et al. Recruitment of FBXO22 for targeted degradation of NSD2. Nat Chem Biol 20, 1597–1607 (2024). https://doi.org/10.1038/s41589-024-01660-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-024-01660-y
This article is cited by
-
High-throughput ligand diversification to discover chemical inducers of proximity
Nature Chemical Biology (2026)
-
NSD2 inhibitors rewire chromatin to treat lung and pancreatic cancers
Nature (2026)
-
F-box proteins in cancer: from cancer cells to the tumor microenvironment
Cell Communication and Signaling (2025)
-
Recent advances in targeting protein degradation for tumor immunotherapy
Journal of Hematology & Oncology (2025)
-
Implications of frequent hitter E3 ligases in targeted protein degradation screens
Nature Chemical Biology (2025)