Abstract
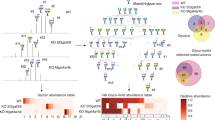
The marine bacterium Alcanivorax borkumensis degrades alkanes derived from phytoplankton, natural hydrocarbon seeps and oil spills. We study the biosynthesis and function of a glycine-glucolipid biosurfactant from A. borkumensis for alkane degradation and identify a gene cluster encoding a nonribosomal peptide synthetase, glycosyltransferase and phosphopantetheinyl transferase. Analyses of A. borkumensis mutants and expression studies reveal that the nonribosomal peptide synthetase catalyzes the synthesis of the aglycone (tetra-d-3-hydroxydecanoyl-glycine) from glycine and d-3-hydroxydecanoyl-CoA, to which a glucose moiety is added by the glycosyltransferase. Deficiency in glycine-glucolipid impairs the ability of mutant cells to attach to the oil–water interface, compromises growth on hexadecane and affects carbon storage. The glycine-glucolipid is essential for biofilm formation on oil droplets and uptake of alkanes. The high incidence of Alcanivorax at oil-polluted sites can in part be explained by the accumulation of the glycine-glucolipid on the cell surface, effectively making the cells themselves act as biosurfactants.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data generated in this study are provided within the article and in Supplementary Information. The complete genome sequence of A. borkumensis SK2, including the gene cluster ABO_1784, ABO_1783 and ABO_1782, can be retrieved at www.ncbi.nlm.nih.gov/nuccore/AM286690.1. The following databases were used in this study: Carbohydrate Active enZYmes (www.cazy.org) and PKS/NRPS Analysis website (https://nrps.igs.umaryland.edu/). Data are also available from the corresponding author upon request. Source data are provided with this paper.
References
Yakimov, M. M., Bargiela, R. & Golyshin, P. N. Calm and frenzy: marine obligate hydrocarbonoclastic bacteria sustain ocean wellness. Curr. Opin. Biotechnol. 73, 337–345 (2022).
Yakimov, M. M. et al. Alcanivorax borkumensis gen. nov., sp. nov., a new, hydrocarbon-degrading and surfactant-producing marine bacterium. Int. J. Syst. Bacteriol. 48, 339–348 (1998).
Kasai, Y. et al. Predominant growth of Alcanivorax strains in oil-contaminated and nutrient-supplemented sea water. Environm. Microbiol. 4, 141–147 (2002).
Rezaei Somee, M. et al. Distinct microbial community along the chronic oil pollution continuum of the Persian Gulf converge with oil spill accidents. Sci. Rep. 11, 11316 (2021).
Lea-Smith, D. J. et al. Contribution of cyanobacterial alkane production to the ocean hydrocarbon cycle. Proc. Natl Acad. Sci. USA 112, 13591–13596 (2015).
Love, C. R. et al. Microbial production and consumption of hydrocarbons in the global ocean. Nat. Microbiol. 6, 489–498 (2021).
Prasad, M. et al. Alcanivorax borkumensis biofilms enhance oil degradation by interfacial tubulation. Science 381, 748–753 (2023).
Cui, J. et al. The glycine-glucolipid of Alcanivorax borkumensis is resident to the bacterial cell wall. Appl. Environm. Microbiol. 88, e0112622 (2022).
Zenati, B. et al. A non-toxic microbial surfactant from Marinobacter hydrocarbonoclasticus SdK644 for crude oil solubilization enhancement. Ecotoxicol. Environ. Saf. 154, 100–107 (2018).
Lan, L.-H., Zhao, H., Chen, J.-C. & Chen, G.-Q. Engineering Halomonas spp. as a low-cost production host for production of bio-surfactant protein PhaP. Biotechnol. J. 11, 1595–1604 (2016).
Karmainski, T. et al. High-quality physiology of Alcanivorax borkumensis SK2 producing glycolipids enables efficient stirred-tank bioreactor cultivation. Front. Bioeng Biotechnol. 11, 1325019 (2023).
Burger, M. M., Glaser, L. & Burton, R. M. The enzymatic synthesis of a rhamnose-containing glycolipid by extracts of Pseudomonas aeruginosa. J. Biol. Chem. 238, 2595–2602 (1963).
Ochsner, U. A., Fiechter, A. & Reiser, J. Isolation, characterization, and expression in Escherichia coli of the Pseudomonas aeruginosa rhlAB genes encoding a rhamnosyltransferase involved in rhamnolipid biosurfactant synthesis. J. Biol. Chem. 269, 19787–19795 (1994).
Zhu, K. & Rock, C. O. RhlA converts β-hydroxyacyl-acyl carrier protein intermediates in fatty acid synthesis to the β-hydroxydecanoyl-β-hydroxydecanoate component of rhamnolipids in Pseudomonas aeruginosa. J. Bacteriol. 190, 3147–3154 (2008).
Rahim, R. et al. Cloning and functional characterization of the Pseudomonas aeruginosa rhlC gene that encodes rhamnosyltransferase 2, an enzyme responsible for di-rhamnolipid biosynthesis. Mol. Microbiol. 40, 708–718 (2001).
Abbasi, A., Bothun, G. D. & Bose, A. Attachment of Alcanivorax borkumensis to hexadecane-in-artificial sea water emulsion droplets. Langmuir 34, 5352–5357 (2018).
Godfrin, M. P., Sihlabela, M., Bose, A. & Tripathi, A. Behavior of marine bacteria in clean environment and oil spill conditions. Langmuir 34, 9047–9053 (2018).
Katsuyama, Y. & Miyanaga, A. Recent advances in the structural biology of modular polyketide synthases and nonribosomal peptide synthetases. Curr. Opin. Chem. Biol. 71, 102223 (2022).
Denaro, R. et al. Alcanivorax borkumensis produces an extracellular siderophore in iron-limitation condition maintaining the hydrocarbon-degradation efficiency. Mar. Genomics 17, 43–52 (2014).
Schneiker, S. et al. Genome sequence of the ubiquitous hydrocarbon-degrading marine bacterium Alcanivorax borkumensis. Nat. Biotechnol. 24, 997–1004 (2006).
Stachelhaus, T., Mootz, H. D. & Marahiel, M. A. The specificity-conferring code of adenylation domains in nonribosomal peptide synthetases. Chem. Biol. 6, 493–505 (1999).
Challis, G. L., Ravel, J. & Townsend, C. A. Predictive, structure-based model of amino acid recognition by nonribosomal peptide synthetase adenylation domains. Chem. Biol. 7, 211–224 (2000).
Hoffmann, D., Hevel, J. M., Moore, R. E. & Moore, B. S. Sequence analysis and biochemical characterization of the nostopeptolide A biosynthetic gene cluster from Nostoc sp. GSV224. Gene 311, 171–180 (2003).
May, J. J., Wendrich, T. M. & Marahiel, M. A. The dhb operon of Bacillus subtilis encodes the biosynthetic template for the catecholic siderophore 2,3-dihydroxybenzoate-glycine-threonine trimeric ester bacillibactin. J. Biol. Chem. 276, 7209–7217 (2001).
Hojati, Z. et al. Structure, biosynthetic origin, and engineered biosynthesis of calcium-dependent antibiotics from Streptomyces coelicolor. Chem. Biol. 9, 1175–1187 (2002).
Pospiech, A., Bietenhader, J. & Schupp, T. Two multifunctional peptide synthetases and an O-methyltransferase are involved in the biosynthesis of the DNA-binding antibiotic and antitumour agent saframycin Mx1 from Myxococcus xanthus. Microbiology 142, 741–746 (1996).
Frueh, D. P. et al. Dynamic thiolation-thioesterase structure of a non-ribosomal peptide synthetase. Nature 454, 903–906 (2008).
Etchegaray, A., Silva-Stenico, M. E., Moon, D. H. & Tsai, S. M. In silico analysis of nonribosomal peptide synthetases of Xanthomonas axonopodis pv. citri: identification of putative siderophore and lipopeptide biosynthetic genes. Microbiol. Res. 159, 425–437 (2004).
Rausch, C., Hoof, I., Weber, T., Wohlleben, W. & Huson, D. H. Phylogenetic analysis of condensation domains in NRPS sheds light on their functional evolution. BMC Evol. Biol. 7, 78 (2007).
Imker, H. J., Krahn, D., Clerc, J., Kaiser, M. & Walsh, C. T. N-Acylation during glidobactin biosynthesis by the tridomain nonribosomal peptide synthetase module GlbF. Chem. Biol. 17, 1077–1083 (2010).
Kraas, F. I., Helmetag, V., Wittmann, M., Strieker, M. & Marahiel, M. A. Functional dissection of surfactin synthetase initiation module reveals insights into the mechanism of lipoinitiation. Chem. Biol. 17, 872–880 (2010).
Gehring, A. M., Mori, I. & Walsh, C. T. Reconstitution and characterization of the Escherichia coli enterobactin synthetase from EntB, EntE, and EntF. Biochemistry 37, 2648–2659 (1998).
Lin, T. P., Chen, C. L., Chang, L. K., Tschen, J. S. & Liu, S. T. Functional and transcriptional analyses of a fengycin synthetase gene, fenC, from Bacillus subtilis. J. Bacteriol. 181, 5060–5067 (1999).
Konz, D., Doekel, S. & Marahiel, M. A. Molecular and biochemical characterization of the protein template controlling biosynthesis of the lipopeptide lichenysin. J. Bacteriol. 181, 133–140 (1999).
Belecki, K. & Townsend, C. A. Biochemical determination of enzyme-bound metabolites: preferential accumulation of a programmed octaketide on the enediyne polyketide synthase CalE8. J. Am. Chem. Soc. 135, 14339–14348 (2013).
Abraham, W.-R., Meyer, H. & Yakimov, M. Novel glycine containing glucolipids from the alkane using bacterium Alcanivorax borkumensis. Biochim. Biophys. Acta 1393, 57–62 (1998).
Passeri, A. et al. Marine biosurfactants. IV. Production, characterization and biosynthesis of an anionic glucose lipid from the marine bacterial strain MM1. Appl. Microbiol. Biotechnol. 37, 281–286 (1992).
Kem, M. P., Zane, H. K., Springer, S. D., Gauglitz, J. M. & Butler, A. Amphiphilic siderophore production by oil-associating microbes. Metallomics 6, 1150–1155 (2014).
Jung, J., Bashiri, G., Johnston, J. M. & Baker, E. N. Mass spectral determination of phosphopantetheinylation specificity for carrier proteins in Mycobacterium tuberculosis. FEBS Open Bio 6, 1220–1226 (2016).
Mootz, H. D., Finking, R. & Marahiel, M. A. 4′-Phosphopantetheine transfer in primary and secondary metabolism of Bacillus subtilis. J. Biol. Chem. 276, 37289–37298 (2001).
Jacketti, M., Beegle-Krause, C. J. & Englehardt, J. D. A review on the sinking mechanisms for oil and successful response technologies. Mar. Pollut. Bull. 160, 111626 (2020).
Boufadel, M. C., Sharifi, Y., van Aken, B., Wrenn, B. A. & Lee, K. Nutrient and oxygen concentrations within the sediments of an Alaskan beach polluted with the Exxon Valdez oil spill. Environ. Sci. Technol. 44, 7418–7424 (2010).
Anderlei, T. & Büchs, J. Device for sterile online measurement of the oxygen transfer rate in shaking flasks. Biochem. Eng J. 7, 157–162 (2001).
Rosenberg, M., Gutnick, D. & Rosenberg, E. Adherence of bacteria to hydrocarbons: a simple method for measuring cell-surface hydrophobicity. FEMS Microbiol. Lett. 9, 29–33 (1980).
Déziel, E., Lépine, F., Milot, S. & Villemur, R. rhlA is required for the production of a novel biosurfactant promoting swarming motility in Pseudomonas aeruginosa: 3-(3-hydroxyalkanoyloxy)alkanoic acids (HAAs), the precursors of rhamnolipids. Microbiology 149, 2005–2013 (2003).
Tiso, T. et al. Designer rhamnolipids by reduction of congener diversity: production and characterization. Microb. Cell Fact. 16, 225 (2017).
Kalscheuer, R. et al. Analysis of storage lipid accumulation in Alcanivorax borkumensis: evidence for alternative triacylglycerol biosynthesis routes in bacteria. J. Bacteriol. 189, 918–928 (2007).
Manilla-Pérez, E., Lange, A. B., Luftmann, H., Robenek, H. & Steinbüchel, A. Neutral lipid production in Alcanivorax borkumensis SK2 and other marine hydrocarbonoclastic bacteria. Eur. J. Lipid Sci. Technol. 113, 8–17 (2011).
Omarova, M. et al. Biofilm formation by hydrocarbon-degrading marine bacteria and its effects on oil dispersion. ACS Sustain. Chem. Eng 7, 14490–14499 (2019).
Sabirova, J. S. et al. Mutation in a ‘tesB-like’ hydroxyacyl-coenzyme A-specific thioesterase gene causes hyperproduction of extracellular polyhydroxyalkanoates by Alcanivorax borkumensis SK2. J. Bacteriol. 188, 8452–8459 (2006).
Matsuyama, T., Tanikawa, T. & Nakagawa, Y. in Biosurfactants (ed Soberón-Chávez, G.) 93–120 (Springer, 2011).
Sieber, S. A. & Marahiel, M. A. Molecular mechanisms underlying nonribosomal peptide synthesis: approaches to new antibiotics. Chem. Rev. 105, 715–738 (2005).
Jenner, M. et al. An unusual Burkholderia gladioli double chain-initiating nonribosomal peptide synthetase assembles ‘fungal’ icosalide antibiotics. Chem. Sci. 10, 5489–5494 (2019).
Hai, Y., Jenner, M. & Tang, Y. Fungal siderophore biosynthesis catalysed by an iterative nonribosomal peptide synthetase. Chem. Sci. 11, 11525–11530 (2020).
Gaitatzis, N., Kunze, B. & Müller, R. In vitro reconstitution of the myxochelin biosynthetic machinery of Stigmatella aurantiaca Sg a15: biochemical characterization of a reductive release mechanism from nonribosomal peptide synthetases. Proc. Natl Acad. Sci. USA 98, 11136–11141 (2001).
Trauger, J. W., Kohli, R. M., Mootz, H. D., Marahiel, M. A. & Walsh, C. T. Peptide cyclization catalysed by the thioesterase domain of tyrocidine synthetase. Nature 407, 215–218 (2000).
Chen, J. et al. Biosynthesis and gene regulation of rhamnolipid congeners. Curr. Microbiol. 80, 302 (2023).
Germer, A. et al. Exploiting the natural diversity of RhlA acyltransferases for the synthesis of the rhamnolipid precursor 3-(3-hydroxyalkanoyloxy)alkanoic acid. Appl. Environm. Microbiol. 86, e02317-19 (2020).
Blecher, M. Synthesis of long-chain fatty acyl-coA thioesters using N-hydroxysuccinimide esters. Meth. Enzymol. 72, 404–408 (1981).
Rehm, B. H., Krüger, N. & Steinbüchel, A. A new metabolic link between fatty acid de novo synthesis and polyhydroxyalkanoic acid synthesis. The PHAG gene from Pseudomonas putida KT2440 encodes a 3-hydroxyacyl-acyl carrier protein-coenzyme a transferase. J. Biol. Chem. 273, 24044–24051 (1998).
Blin, K. et al. antiSMASH 7.0: new and improved predictions for detection, regulation, chemical structures and visualisation. Nucl. Acids Res. 51, W46–W50 (2023).
Bachmann, B. O. & Ravel, J. in Complex Enzymes in Microbial Natural Product Biosynthesis, Part A: Overview Articles and Peptides, Vol. 458 (ed Hopwood, D. A.) 181–217 (Elsevier, 2009).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Drula, E. et al. The carbohydrate-active enzyme database: functions and literature. Nucl. Acids Res. 50, D571–D577 (2022).
Semeniuk, A., Sohlenkamp, C., Duda, K. & Hölzl, G. A bifunctional glycosyltransferase from Agrobacterium tumefaciens synthesizes monoglucosyl and glucuronosyl diacylglycerol under phosphate deprivation. J. Biol. Chem. 289, 10104–10114 (2014).
Hobson, C. et al. Diene incorporation by a dehydratase domain variant in modular polyketide synthases. Nat. Chem. Biol. 18, 1410–1416 (2022).
Lipphardt, A., Karmainski, T., Blank, L. M., Hayen, H. & Tiso, T. Identification and quantification of biosurfactants produced by the marine bacterium Alcanivorax borkumensis by hyphenated techniques. Anal. Bioanal. Chem. 415, 7067–7084 (2023).
Acknowledgements
We thank the Microscopy Core Facility of the Medical Faculty at the University of Bonn for providing support and instrumentation funded by the German Research Foundation (grant 388171357). We thank L. Sundermeyer for support with the initial isolation of the gglsA–gglsB–gglsC gene cluster and F. Diaz and L.-M. Kirschen for their experimental support. This work was supported by the German Ministry of Education and Research (BioProMare, GlycoX; grant 161B0866A to K.-E.J., S.T. and S.K.; 161B0866B to L.M.B. and T.K.; 161B0866C to P.D.) and by the German Research Foundation (grant EXC-2070–390732324, PhenoRob to P.D.; EXC-2186, Fuel Science Center FSC to L.M.B.).
Author information
Authors and Affiliations
Contributions
K.-E.J., S.T., L.M.B. and P.D. conceived the study. J.C., M.F., V.V., M.M.D., G.H., T.K., T.T., S.K. and S.T. designed and performed most experiments. S.T., K.-E.J., L.M.B. and P.D. acquired funding and supervised the project. K.-E.J., S.T., S.K., L.M.B., T.T., T.K. and P.D. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Noah Youssef and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The protein sequence of the nonribosomal peptide synthetase (NRPS) GglsA (ABO_1784).
The domains of the NRPS GglsA are depicted in different colors (C domain, yellow; A domain, bright blue; T domain, golden; TE domain, pink). The two serines, Ser1039 (attachment site of phosphopantetheine group) and Ser1177 (active site of TE domain) are shown in bold red.
Extended Data Fig. 2 GglsA containing the A domain of E. coli EntF catalyzes the synthesis of the serine-glucolipid.
a, The GglsA protein carrying the A domain of the E. coli EntF protein (GglsA-EntFA) was expressed in E. coli, and lipids measured by LC-MS/MS. The MS/MS spectrum displays the parental mass at 803.5788 m/z indicative for the aglycone H(O-10:0)4Ser which differs from the aglycone of the glycine-glucolipid H(O-10:0)4Gly (773.5905 m/z) (Fig. 1c) by 29.9883 m/z equivalent to one HC-OH unit (difference between Ser/Gly). The same mass difference was observed between the fragment peak of H(O-10:0)4Ser (106.0436 m/z) and H(O-10:0)4Gly (76.0386 m/z). The structure of the serine-containing aglycone on the right shows the calculated masses. b, The chimeric GglsA-EntFA protein containing the A domain of E. coli EntF catalyzes the synthesis of the aglycone H(O-10:0)4Ser carrying a serine moiety instead of glycine.
Extended Data Fig. 3 Reaction mechanism of GglsA.
a, The GglsA protein carrying the Ser1177Ala mutation to disable thioesterase activity was expressed in E. coli. Glycine-glucolipid intermediates bound to the phosphopantetheine group were released with cysteamine and analyzed by LC-MS. The MS/MS spectrum of the peak eluting at 48.3 min (inset) shows the fragmentation of the proton adduct of the di-cysteamine derivative of the aglycone H(O-10:0)4Gly carrying four acyl groups. b, The GglsA construct containing the C-A-T domains but lacking the TE domain was expressed in E. coli. The aglycones produced were released by cysteamine cleavage. LC-MS analysis revealed the synthesis of four different aglycones with four, three, two or one 3-hydroxydecanoyl groups with different retention times and MS/MS spectra. The insets show the LC-MS chromatograms for the parental ions (x axis, time in min).
Extended Data Fig. 4 Nonpolar lipid accumulation in A. borkumensis ΔgglsA and ΔgglsB mutant cells.
Cells were grown in pyruvate or hexadecane medium. Lipid extracts equivalent to the same amount of total lipids (measured by gas chromatography of fatty acid methyl esters) were separated by thin-layer chromatography (TLC) and stained with iodine vapor. Storage lipids were identified by co-migration with standards or by isolation and mass spectrometry. TAG, triacylglycerol; WE, wax ester.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–4 and Tables 1–4.
Supplementary Data 1 (download JPG )
Source photo of SDS gel for Supplementary Fig. 1.
Supplementary Data 2 (download XLSX )
Source gel photos for Supplementary Fig. 2.
Supplementary Data 3 (download XLSX )
Source glycine-glucolipid contents for Supplementary Fig. 3.
Supplementary Data 4 (download XLSX )
Source glycine-glucolipid contents for Supplementary Fig. 4.
Source data
Source Data Fig. 1 (download XLSX )
Source data for MS spectra and enzyme assay.
Source Data Fig. 3 (download XLSX )
Source data for MS spectra and enzyme assay.
Source Data Fig. 4 (download XLSX )
Source data for growth curves and OTR curves.
Source Data Fig. 5 (download XLSX )
Source data for BATH assay, emulsification and oil binding.
Source Data Fig. 6 (download XLSX )
Statistical source data and raw images for Fig. 6.
Source Data Extended Data Fig. 2 (download XLSX )
Data for MS spectra.
Source Data Extended Data Fig. 3 (download XLSX )
Data for MS spectra.
Source Data Extended Data Fig. 4 (download JPG )
Photo of thin-layer chromatography (TLC) plate.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cui, J., Fassl, M., Vasanthakumaran, V. et al. Biosurfactant biosynthesis by Alcanivorax borkumensis and its role in oil biodegradation. Nat Chem Biol 21, 1631–1641 (2025). https://doi.org/10.1038/s41589-025-01908-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-025-01908-1
This article is cited by
-
Bioremediation of oil contaminants from the marine ecosystem by Alcanivorax borkumensis: an overview
Archives of Microbiology (2026)