Abstract
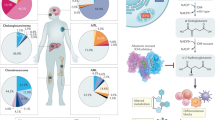
Gain-of-function mutations of isocitrate dehydrogenase 1 (IDH1) lead to oncometabolite (R)-2-hydroxyglutarate production, contributing to the tumorigenesis of multiple human cancers. While fatty acid biosynthesis is critical for IDH1-mutant tumor growth, the underlying mechanisms remain unclear. Here, leveraging chemical probes and chemoproteomic profiling, we identified that oncogenic IDH1-R132H is uniquely autopalmitoylated at C269, which is not observed in wild-type IDH1. This modification responds to fatty acids and regulates R132H enzymatic activity by enhancing substrate and cofactor binding, as well as dimerization. Loss of C269 palmitoylation reverses IDH1-R132H-induced metabolic reprogramming and hypermethylation phenotypes and impairs cell transformation. Interestingly, C269 autopalmitoylation occurs within a hydrophobic pocket, targeted by a clinical IDH1-mutant inhibitor (LY3410738). Our study reveals that autopalmitoylation, conferred by the IDH1R132H mutation, links fatty acid metabolism to the regulation of IDH1 mutant activity and represents a druggable vulnerability in IDH1-mutant cancers.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw RNA-seq data were deposited to the National Center Biotechnology Information Gene Expression Omnibus under accession number GSE260937. The raw proteomics data were deposited to the ProteomeXchange Consortium through the PRIDE partner repository with the dataset identifier PXD070770. The atomic models and cryo-EM density maps were deposited to the Protein Data Bank and EM Data Bank under the following accession numbers: IDH1-R132H, PDB 9YHA and EMD-72964; mtC269S, PDB 9YHB and EMD-72965. Statistical source data for the supplementary figures are available in Supplementary Data 3. Initial and final configurations of all the structures generated in the MD simulations are available in Supplementary Data 4 and 5, respectively. All other data are available in the main text or Supplementary Information. Source data are provided with this paper.
Change history
16 January 2026
In the version of the article initially published, the author of ref. 30 was given as “Dixon, S. J.” but should have been “Kahlson, M. A. et al”. This correction has been made to the HTML and PDF versions of the article.
References
Waitkus, M. S., Diplas, B. H. & Yan, H. Biological role and therapeutic potential of IDH mutations in cancer. Cancer Cell 34, 186–195 (2018).
Pirozzi, C. J. & Yan, H. The implications of IDH mutations for cancer development and therapy. Nat. Rev. Clin. Oncol. 18, 645–661 (2021).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009).
Ward, P. S. et al. The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting α-ketoglutarate to 2-hydroxyglutarate. Cancer Cell 17, 225–234 (2010).
Raineri, S. & Mellor, J. IDH1: linking metabolism and epigenetics. Front. Genet. 9, 1–8 (2018).
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010).
Lu, C. et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483, 474–478 (2012).
Turcan, S. et al. Mutant-IDH1-dependent chromatin state reprogramming, reversibility, and persistence. Nat. Genet. 50, 62–72 (2018).
Badur, M. G. et al. Oncogenic R132 IDH1 mutations limit NADPH for de novo lipogenesis through (D)2-hydroxyglutarate production in fibrosarcoma sells. Cell Rep. 25, 1018–1026 (2018).
Thomas, D. et al. Dysregulated lipid synthesis by oncogenic IDH1 mutation is a targetable synthetic lethal vulnerability. Cancer Discov. 13, 496–515 (2023).
Thomas, D. et al. IDH1 mutant AML is susceptible to targeting de novo lipid synthesis independent of 2-hydroxyglutarate and has a distinct metabolic profile from IDH2 mutant AML. Blood 132, 440–440 (2018).
Sinha, S. et al. Systematic discovery of mutation-specific synthetic lethals by mining pan-cancer human primary tumor data. Nat. Commun. 8, 1–13 (2017).
Fan, J. et al. Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510, 298–302 (2014).
Chen, B., Sun, Y., Niu, J., Jarugumilli, G. K. & Wu, X. Protein lipidation in cell signaling and diseases: function, regulation, and therapeutic opportunities. Cell Chem. Biol. 25, 817–831 (2018).
Jiang, H. et al. Protein lipidation: occurrence, mechanisms, biological functions, and enabling technologies. Chem. Rev. 118, 919–988 (2018).
Rana, M. S. et al. Fatty acyl recognition and transfer by an integral membrane S-acyltransferase. Science 359, eaao6326 (2018).
Chan, P. et al. Autopalmitoylation of TEAD proteins regulates transcriptional output of the Hippo pathway. Nat. Chem. Biol. 12, 282–289 (2016).
Pobbati, A. V. et al. Targeting the central pocket in human transcription factor TEAD as a potential cancer therapeutic strategy. Structure 23, 2076–2086 (2015).
Turnbull, A. P. et al. Structure of palmitoylated Bet3: Insights into TRAPP complex assembly and membrane localization. EMBO J. 24, 875–884 (2005).
Cleary, J. M. et al. Secondary IDH1 resistance mutations and oncogenic IDH2 mutations cause acquired resistance to ivosidenib in cholangiocarcinoma. NPJ Precis. Oncol. 6, 1–8 (2022).
Salama, V. Preclinical Studies of a Novel IDH1 Inhibitor in Acute Myeloid Leukemia (AML). MSc thesis, The University of Texas MD Anderson Cancer (2020).
Salama, V. et al. Abstract 6417: LY3410738, a novel inhibitor of mutant IDH1 is more effective than Ivosidenib and potentiates antileukemic activity of standard chemotherapy in preclinical models of acute myeloid leukemia (AML). Cancer Res. 80, 6417 (2020).
Zheng, B. et al. 2-Bromopalmitate analogues as activity-based probes to explore palmitoyl acyltransferases. J. Am. Chem. Soc. 135, 7082–7085 (2013).
Rana, M. S., Lee, C. J. & Banerjee, A. The molecular mechanism of DHHC protein acyltransferases. Biochem. Soc. Trans. 47, 157–167 (2018).
Rivas, G. & Minton, A. P. Macromolecular crowding in vitro, in vivo, and in between. Trends Biochem. Sci. 41, 970–981 (2016).
Cisar, J. S. & Cravatt, B. F. Fully functionalized small-molecule probes for integrated phenotypic screening and target identification. J. Am. Chem. Soc. 134, 10385–10388 (2012).
Abo, M., Li, C. & Weerapana, E. Isotopically-labeled iodoacetamide-alkyne probes for quantitative cysteine-reactivity profiling. Mol. Pharm. 15, 743–749 (2018).
Blanc, M. et al. SwissPalm: protein palmitoylation database. F1000Res. 4, 261 (2015).
Zhang, S. et al. Long-chain fatty acyl coenzyme A inhibits NME1/2 and regulates cancer metastasis. Proc. Natl Acad. Sci. USA 119, 1–8 (2022).
Kahlson, M. A. et al. Functional dissection of the zDHHC palmitoyltransferase 5–golgin A7 palmitoylation complex. J. Biol. Chem. 301, 110694 (2025).
Jennings, B. C. & Linder, M. E. DHHC protein S-acyltransferases use similar ping-pong kinetic mechanisms but display different Acyl-CoA specificities. J. Biol. Chem. 287, 7236–7245 (2012).
Brooks, E. et al. Identification and characterization of small-molecule inhibitors of the R132H/R132H mutant isocitrate dehydrogenase 1 homodimer and R132H/wild-type heterodimer. J. Biomol. Screen. 19, 1193–1200 (2014).
Noland, C. L. et al. Palmitoylation of TEAD transcription factors is required for their stability and function in Hippo pathway signaling. Structure 24, 179–186 (2016).
Kim, Y. G. et al. Crystal structure of Bet3 reveals a novel mechanism for Golgi localization of tethering factor TRAPP. Nat. Struct. Mol. Biol. 12, 38–45 (2005).
Xie, X. et al. Allosteric mutant IDH1 inhibitors reveal mechanisms for IDH1 mutant and isoform selectivity. Structure 25, 506–513 (2017).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Chen, D. et al. Mutant and wild-type isocitrate dehydrogenase 1 share enhancing mechanisms involving distinct tyrosine kinase cascades in cancer. Cancer Discov. 9, 756–777 (2019).
Intlekofer, A. M. et al. Acquired resistance to IDH inhibition through trans or cis dimer-interface mutations. Nature 559, 125–129 (2018).
Reinbold, R. et al. Resistance to the isocitrate dehydrogenase 1 mutant inhibitor ivosidenib can be overcome by alternative dimer-interface binding inhibitors. Nat. Commun. 13, 1–12 (2022).
Lyu, J. et al. Disabling uncompetitive inhibition of oncogenic IDH mutations drives acquired resistance. Cancer Discov. 13, 170–193 (2023).
Hay, N. Reprogramming glucose metabolism in cancer: can it be exploited for cancer therapy? Nat. Rev. Cancer 16, 635–649 (2016).
Gross, S. et al. Cancer-associated metabolite 2-hydroxyglutarate accumulates in acute myelogenous leukemia with isocitrate dehydrogenase 1 and 2 mutations. J. Exp. Med. 207, 339–344 (2010).
Zhao, H. et al. Myeloid-derived itaconate suppresses cytotoxic CD8+ T cells and promotes tumour growth. Nat. Metab. 4, 1660–1673 (2022).
Zhao, S. et al. Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1α. Science 324, 261–265 (2009).
Hao, Z. et al. Idh1 mutations contribute to the development of T-cell malignancies in genetically engineered mice. Proc. Natl Acad. Sci. USA 113, 1387–1392 (2016).
Gunn, K. et al. (R)-2-Hydroxyglutarate inhibits KDM5 histone lysine demethylases to drive transformation in IDH-mutant cancers. Cancer Discov. 13, 1478–1497 (2023).
Losman, J. A. et al. (R)-2-Hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science 340, 1621–1625 (2013).
DiNardo, C. D. et al. Abstract CT026: a first-in-human phase 1 study of LY3410738, a covalent inhibitor of mutant IDH, in advanced myeloid malignancies. Cancer Res. 83, CT026 (2023).
Rodon, J. et al. Abstract CT098: a first-in-human phase 1 study of LY3410738, a covalent inhibitor of mutant IDH, in advanced IDH-mutant cholangiocarcinoma and other solid tumors. Cancer Res. 83, CT098 (2023).
Bauer, R. A. et al. 7-Phenylethylamino-4h-pyrimido[4,5-d][1,3]oxazin-2-one compounds as mutant IDH1 and IDH2 inhibitors. WIPO patent WO2018111707A1 (2018).
Wakimoto, H. et al. Targetable signaling pathway mutations are associated with malignant phenotype in IDH-mutant gliomas. Clin. Cancer Res. 20, 2898–2909 (2014).
Vinograd, J. & Lebowitz, J. Estimation of globular protein secondary structure from circular dichroism. Biochemistry 20, 33–37 (1981).
Lobley, A., Whitmore, L. & Wallace, B. A. DICHROWEB: an interactive website for the analysis of protein secondary structure from circular dichroism spectra. Bioinformatics 18, 211–212 (2002).
Sreerama, N. & Woody, R. W. Estimation of protein secondary structure from circular dichroism spectra: comparison of CONTIN, SELCON, and CDSSTR methods with an expanded reference set. Anal. Biochem. 287, 252–260 (2000).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Kagami, L., Wilter, A., Diaz, A. & Vranken, W. The ACPYPE web server for small-molecule MD topology generation. Bioinformatics 39, 4–6 (2023).
Hoover, W. Canonical dynamics: equilibrium phase-space distributions. Phys. Rev. A 31, 1695–1697 (1985).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: A LINEAR CONSTRAINT SOLVER FOR MOLECULAR SIMULATIONS. J. Comput. Chem. 18, 1463–1472 (1997).
Mitternacht, S. FreeSASA: an open source C library for solvent accessible surface area calculations. F1000Res. 5, 1–11 (2016).
Yuan, S. et al. Conformational dynamics of the activated GLP-1 receptor–Gs complex revealed by cross-linking mass spectrometry and integrative structure modeling. ACS Cent. Sci. 9, 992–1007 (2023).
Mark, P. & Nilsson, L. Structure and dynamics of the TIP3P, SPC, and SPC/E water models at 298 K. J. Phys. Chem. A 105, 9954–9960 (2001).
Curry, H. B. The method of steepest descent for non-linear minimization problems. Q. Appl. Math. 2, 258–261 (1944).
Rendina, A. R. et al. Mutant IDH1 enhances the production of 2-hydroxyglutarate due to its kinetic mechanism. Biochemistry 52, 4563–4577 (2013).
Balss, J. et al. Enzymatic assay for quantitative analysis of (d)-2-hydroxyglutarate. Acta Neuropathol. 124, 883–891 (2012).
Acknowledgements
We thank MGH Executive Committee on Research (ECOR) Interim Support Funding to X.W. for supporting the work. L.H. is partly supported by a postdoctoral fellowship from the Antidote Health Foundation for Cure of Cancer and the MGH ECOR Fund for Medical Discovery Fundamental Research Fellowship Award. Y.S. is partly supported by Natural Science Foundation of China grant 8251101517. The work in the S.S. lab was supported by an award from the Canada Excellence Research Chair program. We thank Q. Xu, C. Braithwaite and V. Alvarado for technical assistance, the MGH confocal image core for fluorescence imaging and the Taplin mass spec core at Harvard Medical School for chemoproteomic profiling.
Author information
Authors and Affiliations
Contributions
L.H. contributed to project conceptualization, synthesis, biochemical and cellular data curation, analysis and manuscript preparation. A.M.B, K.S.T., X.Z. and S.S. contributed to cryo-EM sample preparation, data processing and analysis. H.-S.S and S.D.-P. contributed to ITC data acquisition and analysis. P.L. and Y.S. contributed to RNA-seq data analysis. L.N., X.B. and X.L. contributed to CD spectrum acquisition and analysis. L.S., J.L., D.T. and Y.S. contributed to structural analysis, MD simulations and manuscript preparation. J.Z. tested the expression levels of ZDHHC14/23 in knockdown experiments. H.W. and D.P.C. contributed to experiments using MGG152 cells and revised the manuscript. J.M.A contributed to metabolomic profiling and analysis. X.W. contributed to project conceptualization, design, analysis and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
L.H. and X.W. are inventors of a patent application covering chemical probes for chemoproteomic profiling. D.P.C. has received financial compensation from Lilly (2020), Boston Pharmaceuticals (2020), GlaxoSmithKline (2021), Pyramid Biosciences (equity interest 2021), the Massachusetts Institute of Technology (2022), Incephalo (2023), Servier (2023) and Boston Scientific (2023) for advisory input. D.P.C. has also received financial compensation and travel reimbursement from Merck for invited lectures (2016) and from the US National Institutes of Health and Department of Defense for clinical trial and grant review (2023). S.S. is the founder and chief executive officer of Gandeeva Therapeutics. X.W. has financial interest in Tasca Therapeutics, which is developing small-molecule modulators of TEAD palmitoylation and transcription factors. X.W.’s interests were reviewed and are managed by MGH and Mass General Brigham in accordance with their conflict of interest policies. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Xin Jin, Christal Sohl, Hui Ye and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 B4 is an activity-based probe for the autopalmitoylated proteome.
a-b, Comparison of labeling efficiency of different probes in HEK293A lysate (a) or live cells (b). A21, A63, B1, B4: 10 µM; alkyne 2-BP: 50 µM. c, Competition of B4 (0.1 µM) labeling of His-TEAD2 by palmitoyl-CoA. d, Inhibition of His-TEAD2 autopalmitoylation by non-tag-B4. Alkyne palmitoyl-CoA: 1 µM. e, Competition of B4 (0.1 µM) labeling of His-Bet3 by palmitoyl-CoA. f, Inhibition of His-Bet3 autopalmitoylation by non-tag-B4. Alkyne palmitoyl-CoA: 1 µM. g, Mutation of C359 blocked the labeling of Myc-TEAD1 by B4 (10 µM) in HEK293A cells. h, Labeling of endogenous ZDHHC7 by B4 (10 µM) in HEK293A cells. i, Mutation of C160 inhibited the labeling of HA-ZDHHC7 by B4 (10 µM). j-k, Gel-based profiling of autopalmitoylated proteome with B4 (10 µM) in HEK293A cells (j) or lysate (k) at different time points. The experiments in a-k were repeated independently three times with similar results and representative results are presented.
Extended Data Fig. 2 Both IDH1 WT and R132H are S-fatty acylated.
a, Streptavidin pulldown and western blot analysis showing that exogenous IDH1wt and IDH1mt were palmitoylated in HEK293A cells. Alk-16C: 50 µM. Relative palmitoylation levels of IDH1 were normalized to Input and indicated. b, Streptavidin pulldown and western blot analysis showing that endogenous IDH1wt and IDH1mt were palmitoylated in HEK293A and MGG152 cells, respectively. c, Fatty acid preference of IDH1wt/mt lipidation using chemical probes (50 µM) with different carbon-chain lengths. Relative lipidation levels of IDH1 were normalized to Input and indicated. d, Treatment of hydroxylamine (1 M) abolished palmitoylation signals of both IDH1wt and IDH1mt. e, ABE assay showing that endogenous IDH1wt and IDH1mt were palmitoylated at cysteine residues in HEK293A and MGG152 cells, respectively. f, ABE assay showing that exogenous IDH1wt and IDH1mt were palmitoylated at cysteine residues. g, Analysis of palmitoylation levels of IDH1wt and IDH1mt using various site-directed mutants. Relative palmitoylation levels of IDH1 were normalized to Input and indicated. h, Measurement of Km value of His-IDH1wt autopalmitoylation in vitro (n = 3 independent experiments). Top, representative western blot images. Bottom, the plot for the Km value calculation. Data were shown as mean ± SEM. i-j, Biotin switch to assess disulfide bond (i) and S-sulfenylation (j) of Flag-IDH1mt and Flag-IDH1mt C269S in HEK293A cells. TCEP, 10 mM; m-Arsenite, 200 mM. The experiments in a-g and i-j were repeated independently three times with similar results, and representative results are presented.
Extended Data Fig. 3 C269 palmitoylation was observed in other IDH1 mutants.
a, Mutation of C269 substantially decreased palmitoylation signals of IDH1 R132G and R132S. b, Mutation of C269 blocked the labeling of IDH1 R132G and R132S by B4 (10 µM). c, Mutation of C269 did not decrease palmitoylation signals of IDH1 R132C. d, Mutation of C269 decreased the labeling of IDH1 R132C by B4 (10 µM). The experiments in a-d were repeated independently three times with similar results, and representative results are presented.
Extended Data Fig. 4 C269 palmitoylation of mutant IDH1 was not mediated by ZDHHC family enzymes.
a-c, Screening of mouse Zdhhc-PATs that are responsible for IDH1mt palmitoylation. HEK293A cells were co-transfected with Flag-IDH1mt and HA-Zdhhcs. Palmitoylation signals of IDH1mt were determined by streptavidin pulldown and western blot analysis. d, Analysis of palmitoylation levels of IDH1mt 3CS (C73S/C114S/C379S) after knocking down endogenous ZDHHC14 or 23 using shRNA. Bar graph showing knockdown efficiency of ZDHHC14 or ZDHHC23 at the mRNA level (n = 3 independent experiments). Data were shown as mean ± SEM. Statistical significance was assessed using an unpaired, two-sided Student’s t-test. e-f, Analysis of palmitoylation levels of IDH1mt 3CS after knocking down endogenous ZDHHC7 (e) or 9 (f) using siRNAs. g, Analysis of palmitoylation levels of IDH1mt 3CS after individually overexpressing Zdhhc7/9/14/23. h, Co-IP assays showing there were no interactions between IDH1mt 3CS and Zdhhc7, 9, 14 or 23. The experiments in a-c and e-h were repeated independently three times with similar results, and representative results are presented.
Extended Data Fig. 5 IDH1mt autopalmitoylation induced transcriptional reprogramming in IDH-mut cells.
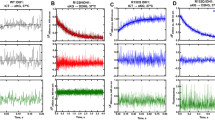
a, 3D PCA plot with genes plotted in three dimensions using their projections onto the first three principal components, and nine samples plotted using their weights for the components (n = 3 independent experiments). Green, control group. Red, mtC269S group. Blue, IDH1mt group. b, Heatmap analysis of global gene expression in control, IDH1mt and mtC269S U87 cells (n = 3 independent experiments). c, GO enrichment analysis with differential genes in IDH1mt and mtC269S cells showing the most significant 10 upregulated biological processes. d, DO enrichment analysis with differential genes in IDH1mt and mtC269S cells showing the most significant 20 DO Terms. Statistical significance in c-d was assessed using a hypergeometric test and adjusted for multiple comparisons using the Benjamini-Hochberg false discovery rate (FDR) method.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–4, Figs. 1–16, Note and Source data for blots in supplementary figures.
Supplementary Data 1 (download XLSX )
List of proteins identified in the chemoproteomic profiling with B4.
Supplementary Data 2 (download XLSX )
List of metabolites identified in the metabolic profiling of U87 cells.
Supplementary Data 3 (download XLSX )
Statistical source data for supplementary figures.
Supplementary Data 4 (download TXT )
Initial configurations for all structures generated in MD simulations.
Supplementary Data 5 (download TXT )
Final configurations for all structures generated in MD simulations.
Supplementary Table 5 (download XLSX )
Sequences of primers.
Supplementary Video 1 (download MP4 )
Supplementary videos for MD simulations.
Source data
Source Data Fig. 1 (download PDF )
Unprocessed western blots and/or gels.
Source Data Fig. 2 (download PDF )
Unprocessed western blots and/or gels.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed western blots and/or gels.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download PDF )
Unprocessed western blots and/or gels.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed western blots and/or gels.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download PDF )
Unprocessed western blots and/or gels.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, L., Lin, J., Sun, L. et al. Autopalmitoylation of IDH1-R132H regulates its neomorphic activity in cancer cells. Nat Chem Biol (2026). https://doi.org/10.1038/s41589-025-02131-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41589-025-02131-8