Abstract
Anaphase chromosome segregation depends on forces exerted by spindle microtubules. Current models propose two force-generating mechanisms: kinetochore–microtubule (kMT) depolymerization pulls chromosomes toward spindle poles (anaphase A), while antiparallel microtubule sliding in the central spindle further separates sister chromosomes by elongating the spindle (anaphase B). Experimental evidence in cells supports the sliding mechanism but contributions of the depolymerization mechanism remain unclear. We show that kMT depolymerization limits spindle elongation rather than moving chromosomes apart. We developed a chemical optogenetic approach to recruit microtubule depolymerases to kinetochores at anaphase onset, thereby increasing kMT depolymerization rates without perturbing earlier stages of mitosis. We find that increased depolymerization slows the velocity at which spindle poles move apart without changing kinetochore separation velocities. Our findings support a model in which kinetochores selectively couple to central spindle microtubules parallel to their kMTs, such that antiparallel sliding drives chromosome segregation while kMT depolymerization pulls poles inward.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Plasmids generated during the current study are available through Addgene. Source data are provided with this paper.
References
Maiato, H. & Lince-Faria, M. The perpetual movements of anaphase. Cell. Mol. Life Sci. 67, 2251–2269 (2010).
Alberts, B. et al. Cells in their social context. In Molecular Biology of the Cell. 7th edn (W. W. Norton & Company, 2022).
Asbury, C. L. Anaphase A: disassembling microtubules move chromosomes toward spindle poles. Biology 6, 15 (2017).
McIntosh, J. R. Anaphase A. Semin. Cell Dev. Biol. 117, 118–126 (2021).
Scholey, J. M. Civelekoglu-Scholey, G. & Brust-Mascher, I. Anaphase B. Biology 5, 51 (2016).
Vukusic, K. & Tolic, I. M. Anaphase B: long-standing models meet new concepts. Semin. Cell Dev. Biol. 117, 127–139 (2021).
Wadsworth, P. The multifunctional spindle midzone in vertebrate cells at a glance. J. Cell Sci. 134, jcs250001 (2021).
Kapitein, L. C. et al. The bipolar mitotic kinesin Eg5 moves on both microtubules that it crosslinks. Nature 435, 114–118 (2005).
Wijeratne, S. & Subramanian, R. Geometry of antiparallel microtubule bundles regulates relative sliding and stalling by PRC1 and Kif4A. eLife 7, e32595 (2018).
Lim, W. M. et al. Regulation of minimal spindle midzone organization by mitotic kinases. Nat. Commun. 15, 9213 (2024).
Yu, C. H. et al. Central-spindle microtubules are strongly coupled to chromosomes during both anaphase A and anaphase B. Mol. Biol. Cell 30, 2503–2514 (2019).
Vukusic, K. et al. Microtubule sliding within the bridging fiber pushes kinetochore fibers apart to segregate chromosomes. Dev. Cell 43, 11–23 (2017).
Ris, H. The anaphase movement of chromosomes in the spermatocytes of the grasshopper. Biol. Bull. 96, 90–106 (1949).
Nicklas, R. B. Chromosome movement and spindle birefringence in locally heated cells: interaction versus local control. Chromosoma 74, 1–37 (1979).
Pamula, M. C. et al. High-resolution imaging reveals how the spindle midzone impacts chromosome movement. J. Cell Biol. 218, 2529–2544 (2019).
Vukusic, K., Ponjavic, I., Buda, R., Risteski, P. & Tolic, I. M. Microtubule-sliding modules based on kinesins EG5 and PRC1-dependent KIF4A drive human spindle elongation. Dev. Cell 56, 1253–1267 (2021).
Orr, B. et al. An anaphase surveillance mechanism prevents micronuclei formation from frequent chromosome segregation errors. Cell Rep. 37, 109783 (2021).
Brust-Mascher, I., Sommi, P., Cheerambathur, D. K. & Scholey, J. M. Kinesin-5-dependent poleward flux and spindle length control in Drosophila embryo mitosis. Mol. Biol. Cell 20, 1749–1762 (2009).
Rogers, G. C. et al. Two mitotic kinesins cooperate to drive sister chromatid separation during anaphase. Nature 427, 364–370 (2004).
Ganem, N. J., Upton, K. & Compton, D. A. Efficient mitosis in human cells lacking poleward microtubule flux. Curr. Biol. 15, 1827–1832 (2005).
Zhang, D., Rogers, G. C., Buster, D. W. & Sharp, D. J. Three microtubule severing enzymes contribute to the ‘Pacman-flux’ machinery that moves chromosomes. J. Cell Biol. 177, 231–242 (2007).
Rath, U. et al. The Drosophila kinesin-13, KLP59D, impacts Pacman- and flux-based chromosome movement. Mol. Biol. Cell 20, 4696–4705 (2009).
Mukherjee, S. et al. Human Fidgetin is a microtubule severing the enzyme and minus-end depolymerase that regulates mitosis. Cell Cycle 11, 2359–2366 (2012).
Guerreiro, A. et al. WDR62 localizes katanin at spindle poles to ensure synchronous chromosome segregation. J. Cell Biol. 220, e202007171 (2021).
Moore, A. T. et al. MCAK associates with the tips of polymerizing microtubules. J. Cell Biol. 169, 391–397 (2005).
Wordeman, L., Wagenbach, M. & von Dassow, G. MCAK facilitates chromosome movement by promoting kinetochore microtubule turnover. J. Cell Biol. 179, 869–879 (2007).
Domnitz, S. B., Wagenbach, M., Decarreau, J. & Wordeman, L. MCAK activity at microtubule tips regulates spindle microtubule length to promote robust kinetochore attachment. J. Cell Biol. 197, 231–237 (2012).
Anjur-Dietrich, M. I., Kelleher, C. P. & Needleman, D. J. Mechanical mechanisms of chromosome segregation. Cells 10, 465 (2021).
Zimyanin, V. & Redemann, S. Microtubule length correlates with spindle length in C. elegans meiosis. Cytoskeleton 81, 356–368 (2024).
Okumura, M., Natsume, T., Kanemaki, M. T. & Kiyomitsu, T. Dynein–dynactin–NuMA clusters generate cortical spindle-pulling forces as a multi-arm ensemble. eLife 7, e36559 (2018).
Jagrić, M., Risteski, P., Martinčić, J., Milas, A. & Tolić, I. M. Optogenetic control of PRC1 reveals its role in chromosome alignment on the spindle by overlap length-dependent forces. eLife 10, e61170 (2021).
Canman, J. C. et al. Determining the position of the cell division plane. Nature 424, 1074–1078 (2003).
Su, K. C. et al. A regulatory switch alters chromosome motions at the metaphase-to-anaphase transition. Cell Rep. 17, 1728–1738 (2016).
Fuller, B. G. et al. Midzone activation of aurora B in anaphase produces an intracellular phosphorylation gradient. Nature 453, 1132–1136 (2008).
Kapoor, T. M., Mayer, T. U., Coughlin, M. L. & Mitchison, T. J. Probing spindle assembly mechanisms with monastrol, a small molecule inhibitor of the mitotic kinesin, Eg5. J. Cell Biol. 150, 975–988 (2000).
Khodjakov, A., Copenagle, L., Gordon, M. B., Compton, D. A. & Kapoor, T. M. Minus-end capture of preformed kinetochore fibers contributes to spindle morphogenesis. J. Cell Biol. 160, 671–683 (2003).
Ballister, E. R., Aonbangkhen, C., Mayo, A. M., Lampson, M. A. & Chenoweth, D. M. Localized light-induced protein dimerization in living cells using a photocaged dimerizer. Nat. Commun. 5, 5475 (2014).
Zhang, H. et al. Optogenetic control of kinetochore function. Nat. Chem. Biol. 13, 1096–1101 (2017).
Chen, G. Y. et al. Tension promotes kinetochore–microtubule release by Aurora B kinase. J. Cell Biol. 220, e202007030 (2021).
Chen, G. Y. & Lampson, M. A. Chemical tools for dissecting cell division. Nat. Chem. Biol. 17, 632–640 (2021).
Hunter, A. W. et al. The kinesin-related protein MCAK is a microtubule depolymerase that forms an ATP-hydrolyzing complex at microtubule ends. Mol. Cell 11, 445–457 (2003).
Lan, W. et al. Aurora B phosphorylates centromeric MCAK and regulates its localization and microtubule depolymerization activity. Curr. Biol. 14, 273–286 (2004).
Mollinari, C. et al. PRC1 is a microtubule binding and bundling protein essential to maintain the mitotic spindle midzone. J. Cell Biol. 157, 1175–1186 (2002).
Lansky, Z. et al. Diffusible crosslinkers generate directed forces in microtubule networks. Cell 160, 1159–1168 (2015).
Gaska, I., Armstrong, M. E., Alfieri, A. & Forth, S. The mitotic crosslinking protein PRC1 acts like a mechanical dashpot to resist microtubule sliding. Dev. Cell 54, 367–378 (2020).
Braun, M. et al. Adaptive braking by Ase1 prevents overlapping microtubules from sliding completely apart. Nat. Cell Biol. 13, 1259–1264 (2011).
Valdez, V. A., Neahring, L., Petry, S. & Dumont, S. Mechanisms underlying spindle assembly and robustness. Nat. Rev. Mol. Cell Biol. 24, 523–542 (2023).
Oriola, D., Needleman, D. J. & Brugues, J. The physics of the metaphase spindle. Annu. Rev. Biophys. 47, 655–673 (2018).
Redemann, S. et al. C. elegans chromosomes connect to centrosomes by anchoring into the spindle network. Nat. Commun. 8, 15288 (2017).
Carlini, L., Renda, F., Pamula, M. C., Khodjakov, A. & Kapoor, T. M. Coupling of microtubule bundles isolates them from local disruptions to set the structural stability of the anaphase spindle. Proc. Natl Acad. Sci. USA 119, e2204068119 (2022).
Deng, C. et al. Conditional localization pharmacology manipulates the cell cycle with spatiotemporal precision. Preprint at bioRxiv https://doi.org/10.1101/2024.09.12.612697 (2024).
Janson, M. E. et al. Crosslinkers and motors organize dynamic microtubules to form stable bipolar arrays in fission yeast. Cell 128, 357–368 (2007).
Mastronarde, D. N., McDonald, K. L., Ding, R. & McIntosh, J. R. Interpolar spindle microtubules in PTK cells. J. Cell Biol. 123, 1475–1489 (1993).
Brust-Mascher, I., Civelekoglu-Scholey, G., Kwon, M., Mogilner, A. & Scholey, J. M. Model for anaphase B: role of three mitotic motors in a switch from poleward flux to spindle elongation. Proc. Natl Acad. Sci. USA 101, 15938–15943 (2004).
Brust-Mascher, I. & Scholey, J. M. Microtubule flux and sliding in mitotic spindles of Drosophila embryos. Mol Biol Cell 13, 3967–3975 (2002).
Steblyanko, Y. et al. Microtubule poleward flux in human cells is driven by the coordinated action of four kinesins. EMBO J. 39, e105432 (2020).
Desai, A., Maddox, P. S., Mitchison, T. J. & Salmon, E. D. Anaphase A chromosome movement and poleward spindle microtubule flux occur At similar rates in Xenopus extract spindles. J. Cell Biol. 141, 703–713 (1998).
Murray, A. W., Desai, A. B. & Salmon, E. D. Real time observation of anaphase in vitro. Proc. Natl Acad. Sci. USA 93, 12327–12332 (1996).
Laband, K. et al. Chromosome segregation occurs by microtubule pushing in oocytes. Nat. Commun. 8, 1499 (2017).
Dumont, J., Oegema, K. & Desai, A. A kinetochore-independent mechanism drives anaphase chromosome separation during acentrosomal meiosis. Nat. Cell Biol. 12, 894–901 (2010).
Nahaboo, W., Zouak, M., Askjaer, P. & Delattre, M. Chromatids segregate without centrosomes during Caenorhabditis elegans mitosis in a Ran- and CLASP-dependent manner. Mol. Biol. Cell 26, 2020–2029 (2015).
Spurck, T. P. et al. UV microbeam irradiations of the mitotic spindle. II. Spindle fiber dynamics and force production. J. Cell Biol. 111, 1505–1518 (1990).
Spurck, T., Forer, A. & Pickett-Heaps, J. Ultraviolet microbeam irradiations of epithelial and spermatocyte spindles suggest that forces act on the kinetochore fibre and are not generated by its disassembly. Cell Motil. Cytoskeleton 36, 136–148 (1997).
Nicklas, R. B. The motor for poleward chromosome movement in anaphase is in or near the kinetochore. J. Cell Biol. 109, 2245–2255 (1989).
Suresh, P., Long, A. F. & Dumont, S. Microneedle manipulation of the mammalian spindle reveals specialized, short-lived reinforcement near chromosomes. eLife 9, e53807 (2020).
Elting, M. W., Prakash, M., Udy, D. B. & Dumont, S. Mapping load-bearing in the mammalian spindle reveals local kinetochore fiber anchorage that provides mechanical isolation and redundancy. Curr Biol 27, 2112–2122 (2017).
Kajtez, J. et al. Overlap microtubules link sister k-fibres and balance the forces on bi-oriented kinetochores. Nat. Commun. 7, 10298 (2016).
Sikirzhytski, V. et al. Microtubules assemble near most kinetochores during early prometaphase in human cells. J. Cell Biol. 217, 2647–2659 (2018).
Renda, F. et al. Non-centrosomal microtubules at kinetochores promote rapid chromosome biorientation during mitosis in human cells. Curr. Biol. 32, 1049–1063 (2022).
Lecland, N. & Luders, J. The dynamics of microtubule minus ends in the human mitotic spindle. Nat. Cell Biol. 16, 770–778 (2014).
Petry, S., Groen, A. C., Ishihara, K., Mitchison, T. J. & Vale, R. D. Branching microtubule nucleation in Xenopus egg extracts mediated by augmin and TPX2. Cell 152, 768–777 (2013).
Almeida, A. C. et al. Augmin-dependent microtubule self-organization drives kinetochore fiber maturation in mammals. Cell Rep. 39, 110610 (2022).
Stimac, V., Koprivec, I., Manenica, M., Simunic, J. & Tolic, I. M. Augmin prevents merotelic attachments by promoting proper arrangement of bridging and kinetochore fibers. eLife 11, e83287 (2022).
Uehara, R. et al. Augmin shapes the anaphase spindle for efficient cytokinetic furrow ingression and abscission. Mol. Biol. Cell 27, 812–827 (2016).
Haren, L., Gnadt, N., Wright, M. & Merdes, A. NuMA is required for proper spindle assembly and chromosome alignment in prometaphase. BMC Res. Notes 2, 64 (2009).
Valdez, V. A., Ma, M., Gouveia, B., Zhang, R. & Petry, S. HURP facilitates spindle assembly by stabilizing microtubules and working synergistically with TPX2. Nat. Commun. 15, 9689 (2024).
Yang, Z., Tulu, U. S., Wadsworth, P. & Rieder, C. L. Kinetochore dynein is required for chromosome motion and congression independent of the spindle checkpoint. Curr. Biol. 17, 973–980 (2007).
Risteski, P. et al. Length-dependent poleward flux of sister kinetochore fibers promotes chromosome alignment. Cell Rep. 40, 111169 (2022).
Sturgill, E. G. et al. Kinesin-12 Kif15 targets kinetochore fibers through an intrinsic two-step mechanism. Curr. Biol. 24, 2307–2313 (2014).
Yoshida, M. W., Yamada, M. & Goshima, G. Moss kinesin-14 KCBP accelerates chromatid motility in anaphase. Cell Struct. Funct. 44, 95–104 (2019).
Almeida, A. C. & Maiato, H. Chromokinesins. Curr. Biol. 28, R1131–R1135 (2018).
Schmidt, J. C. et al. The kinetochore-bound Ska1 complex tracks depolymerizing microtubules and binds to curved protofilaments. Dev. Cell 23, 968–980 (2012).
Sivakumar, S. et al. The human SKA complex drives the metaphase-anaphase cell cycle transition by recruiting protein phosphatase 1 to kinetochores. eLife 5, e12902 (2016).
Yoshida, S. et al. PRC1-rich kinetochores are required for error-free acentrosomal spindle bipolarization during meiosis I in mouse oocytes. Nat. Commun. 11, 2652 (2020).
Liu, B. & Lee, Y. J. Spindle assembly and mitosis in plants. Annu. Rev. Plant Biol. 73, 227–254 (2022).
Hu, C. K., Coughlin, M., Field, C. M. & Mitchison, T. J. KIF4 regulates midzone length during cytokinesis. Curr. Biol. 21, 815–824 (2011).
Kunchala, P. et al. Plasticity of the mitotic spindle in response to karyotype variation. Curr. Biol. 34, 3416–3428 (2024).
Uehara, R. et al. Aurora B and Kif2A control microtubule length for assembly of a functional central spindle during anaphase. J. Cell Biol. 202, 623–636 (2013).
Rizk, R. S., Discipio, K. A., Proudfoot, K. G. & Gupta, M. L. Jr. The kinesin-8 Kip3 scales anaphase spindle length by suppression of midzone microtubule polymerization. J. Cell Biol. 204, 965–975 (2014).
Straight, A. F., Sedat, J. W. & Murray, A. W. Time-lapse microscopy reveals unique roles for kinesins during anaphase in budding yeast. J. Cell Biol. 143, 687–694 (1998).
Nam, S. & Chaudhuri, O. Mitotic cells generate protrusive extracellular forces to divide in three-dimensional microenvironments. Nat. Phys. 14, 621–628 (2018).
Arquint, C., Gabryjonczyk, A. M. & Nigg, E. A. Centrosomes as signalling centres. Philos. Trans. R. Soc. Lond. B 369, 20130464 (2014).
Pollard, T. D. Nine unanswered questions about cytokinesis. J. Cell Biol. 216, 3007–3016 (2017).
Dumont, S. & Mitchison, T. J. Force and length in the mitotic spindle. Curr. Biol. 19, R749–R761 (2009).
Khandelia, P., Yap, K. & Makeyev, E. V. Streamlined platform for short hairpin RNA interference and transgenesis in cultured mammalian cells. Proc. Natl Acad. Sci. USA 108, 12799–12804 (2011).
Andrews, P. D. et al. Aurora B regulates MCAK at the mitotic centromere. Dev. Cell 6, 253–268 (2004).
Subramanian, R., Ti, S. C., Tan, L., Darst, S. A. & Kapoor, T. M. Marking and measuring single microtubules by PRC1 and kinesin-4. Cell 154, 377–390 (2013).
Subramanian, R. et al. Insights into antiparallel microtubule crosslinking by PRC1, a conserved nonmotor microtubule binding protein. Cell 142, 433–443 (2010).
Gillingham, A. K. & Munro, S. The PACT domain, a conserved centrosomal targeting motif in the coiled-coil proteins AKAP450 and pericentrin. EMBO Rep. 1, 524–529 (2000).
Acknowledgements
We thank the Philly ChromoClub, the Penn Center for Genome Integrity and A. Khodjakov (Wadsworth Center) for insightful discussions. This work was supported by the National Institutes of Health (GM122475 and P01-CA265794) and the Basser Center for BRCA (Early Career Award).
Author information
Authors and Affiliations
Contributions
All authors contributed to development of chemical optogenetics probes and experimental design. C.D. and D.M.C. synthesized and characterized the chemical optogenetic probes. G.-Y.C. conducted the cell biology experiments and wrote the paper. M.A.L. edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Recruitment of an MCAK catalytic mutant to kinetochores modestly shortens the metaphase spindle.
MCAK depolymerase activity was suppressed by generating a mutant (MCAKMUT) with combined mutations at S192E and the hypir triple substitution (H536A/R540A/K543A). Pole-to-pole distances were measured for each cell and averaged at every time point (n = 29 MCAK cells pooled from four independent experiments; n = 8 MCAKMUT cells pooled from two independent experiments). For MCAK cells, data are replotted from Fig. 1f. Error bars: mean ± SEM.
Extended Data Fig. 2 MCAK overexpression does not affect motions of chromosomes and poles.
Velocities of kMT depolymerization (a), kinetochore separation (b), and pole separation (c) were measured for cells expressing mScarlet-eDHFR (–MCAK, n = 40 cells) or MCAK-mScarlet-eDHFR (+MCAK, n = 58 cells) as in Fig. 2c, d. For +MCAK cells, data were replotted from –dimerizer cells in Fig. 2e–g. Each data point represents a single cell. Lines: mean ± SD. Statistics: two-tailed Student’s t-test.
Extended Data Fig. 3 Recruitment of an MCAK catalytic mutant to kinetochores modestly restricts spindle elongation.
Initial velocity analyses for kMT depolymerization (a), kinetochore separation (b), and pole separation (c). From left to right in each panel: n = 58 MCAK –dimerizer cells polled from six independent experiments, n = 30 MCAK +dimerizer cells polled from seven independent experiments, n = 18 MCAKMUT –dimerizer cells polled from two independent experiments, and n = 16 MCAKMUT +dimerizer cells polled from two independent experiments. For MCAK –dimerizer and MCAK +dimerizer cells, data are replotted from Fig. 2e–g. Each data point represents a single cell. Lines: mean ± SD.
Extended Data Fig. 4 Enhanced kMT depolymerization does not affect PRC1 bundle length reduction in anaphase.
(a) Schematic of PRC1 bundle length reduction during antiparallel sliding in anaphase. Sliding reduces the overlap length (blue, Vsliding), while polymerization of central spindle microtubules at plus-ends increases the overlap length (gray, Vpol), leading to the net rate of bundle length reduction (pink, VPRC1 = Vsliding - 2× Vpol). (b–d) Analysis of bundle reduction rate. Representative images (b) show 670nano3-PRC1 in cells also expressing Halo-GFP-SPC25, MCAK-mScarlet-eDHFR, and PACT-GFP. Time stamps (min:s) indicate time after first observing sister kinetochore separation at t = 0. Scale bars, 5 µm. Example snapshots of PRC1 intensity (c) were analyzed by line scans between the two spindle poles. Bundle lengths at t = 0 (black) and t = 2 min (red) are defined by their full widths at half-maximum. Example trace (d) shows changes in bundle length over time, with rate (dashed lines) measured in the time window defined as in Fig. 2d. (e–h) Initial velocities were calculated for kMT depolymerization (e), kinetochore separation (f), pole separation (g), and bundle length reduction (h) (n = 22 cells –dimerizer pooled from three independent experiments, 12 cells +dimerizer pooled from two independent experiments). Each data point represents a single cell. Lines: mean ± SD. Statistics: two-tailed Student’s t-test. We note that kinetochores separate faster than bundle length reduction (VKK > VPRC1) both with and without dimerizer, consistent with microtubule polymerization in the central spindle that increases bundle length during anaphase.
Extended Data Fig. 5 PRC1 morphology in monopolar spindles.
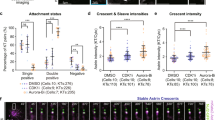
(a) Representative images of PRC1 bundles before (left) or after (right) anaphase onset. Time stamps indicate the time after uncaging. Scale bars, 5 µm. (b) An example of PRC1 intensity analyzed by line scan (box in a). Bundle length and intensity are defined by the full width at half-maximum and the peak height, respectively. (c, d) Analyses for bundle length (c) and intensity (d). Each dot represents a PRC1 bundle (n = 162 for pre-anaphase bundles and 148 for anaphase bundles from 23 cells). Lines: mean ± SD. Statistics: two-tailed Student’s t-test.
Extended Data Fig. 6 PRC1 overexpression and formation of brake complexes inhibit kMT depolymerization.
(a) Representative images of cells expressing CenpB-GFP and PACT-GFP, with or without expression of Halo-670nano3-PRC1. Time stamps (min:s) indicate time after first observing sister kinetochore separation at t = 0. Scale bars, 5 µm. (b–d) Dependence of pole separation (b), kinetochore separation (c), and kMT depolymerization (d) velocities on spindle-bound Halo-670nano3-PRC1. Colored data points show control cells without Halo-670nano3-PRC1 or mScarlet-eDHFR-PRC1M (gray), cells expressing Halo-670nano3-PRC1 (orange), cells expressing Halo-670nano3-PRC1 and mScarlet-eDHFR-PRC1M without dimerization (blue), and cells expressing Halo-670nano3-PRC1 and mScarlet-eDHFR-PRC1M with dimerization (pink). All cells express CenpB-GFP and PACT-GFP. Each data point represents a single cell.
Extended Data Fig. 7 Velocity relationships predicted by the sort-and-grip model and proposed mechanisms of parallel sorting.
(a) Schematic shows kinetochore velocity (VK), pole velocity (VP), and antiparallel sliding in the central spindle (Vsliding). Kinetochore and pole velocities are half of the velocities of kinetochore separation (VKK) and pole separation (VPP), respectively, as measured in Figs. 2 and 4. Kinetochores, poles, and fiducial marks that label antiparallel sliding are shown in green, black, and brown, respectively. In the sort-and-grip model, kinetochore velocity matches sliding velocity, and pole velocity is sliding velocity (moving poles outward) subtracting kMT depolymerization from either end (VPK, moving poles inward). Manipulations of kMT depolymerization affect pole velocity but not kinetochore velocity in this model. (b) Kinetochore-nucleated kMT fragments (red) have their plus-ends at kinetochores. Minus-end directed motors (gray) at the minus-ends of these short kMTs preferentially guide kMT growth toward the minus ends of parallel microtubules in the central spindle (blue) to biorient sister kinetochores. (c) Minus-end directed motors (gray) guide the minus ends of newly-grown microtubules in the central spindle toward spindle poles through kMTs. (d) A fraction of the central spindle is grown by microtubules branching from kMTs (gray). These branching microtubules form antiparallel bundles (bridging fibers) between the two sister kinetochores, bound by antiparallel couplers.
Supplementary information
Supplementary Information (download PDF )
Legends for Supplementary Videos 1–6.
Supplementary Video 1 (download AVI )
Max-intensity projection over 1 μm for the cell shown in Fig. 1e. Time stamps (min:s) indicate the time after uncaging at t = 0. White arrows represent the spindle pole. Scale bar, 10 μm.
Supplementary Video 2 (download AVI )
Max-intensity projection over 4 μm for the cells shown in Fig. 2b, with (right) or without (left) dimerization. Time stamps (min:s) indicate the time after uncaging at t = 0. White arrows represent the spindle pole. Scale bar, 5 μm.
Supplementary Video 3 (download AVI )
Max-intensity projection over 2 μm showing the off-bundle sister kinetochore pair (yellow arrow) indicated by the gray box in Fig. 3c. Time stamps (min:s) indicate the time after uncaging at t = 0. Scale bar, 5 μm.
Supplementary Video 4 (download AVI )
Max-intensity projection over 2 μm showing the on-bundle sister kinetochore pair (yellow arrow) indicated by the orange box in Fig. 3c. Time stamps (min:s) indicate the time after uncaging at t = 0. Scale bar, 5 μm.
Supplementary Video 5 (download AVI )
Max-intensity projection over the whole cell for the cells shown in Fig. 4c. Left, –dimerizer; right, +dimerizer. Time stamps (min:s) indicate the time after uncaging at t = 0. White arrows represent the spindle pole. Scale bar, 5 μm.
Supplementary Video 6 (download AVI )
Max-intensity projection over the whole cell for the cells shown in Extended Data Fig. 6a, with (right) or without (left) expression of Halo–670nano3–PRC1. Time stamps (min:s) indicate the time after first observing sister kinetochore separation at t = 0. White arrows represent the spindle pole. Scale bar, 5 μm.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, GY., Deng, C., Chenoweth, D.M. et al. Microtubule depolymerization at kinetochores restricts anaphase spindle elongation. Nat Chem Biol (2026). https://doi.org/10.1038/s41589-026-02143-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41589-026-02143-y