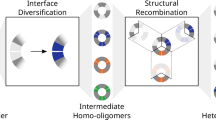
Abstract
Peroxiredoxins are thiol peroxidases, which detoxify peroxides, relay redox signals and act as chaperones. In eukaryotes, multiple peroxiredoxin-1 (Prx1)/AhpC-type isoforms frequently co-exist in the same subcellular compartment, yet have been assumed to assemble only as homo-oligomeric complexes. Here we show that hetero-oligomerization is a conserved and functionally relevant property of Prx1/AhpC-type peroxiredoxins. Using biochemical reconstitution, native mass photometry, electron microscopy and live-cell assays, we demonstrate formation of heterodimers and heterodecamers, with diverse subunit stoichiometries, in peroxiredoxin pairs from different eukaryotic kingdoms. In Saccharomyces cerevisiae, oxidative challenge induces Tsa1–Tsa2 heterodecamerization with substoichiometric Tsa2 incorporation sufficing to stabilize the decameric state. Functional hetero-oligomers are also observed forming among human, plant and Leishmania peroxiredoxins. Our findings provide new insights into peroxiredoxin structural plasticity with broad implications for redox biology, stress responses and cellular adaptation, and also challenge the long-held paradigm of peroxiredoxin homo-oligomerization.

Similar content being viewed by others
Main
Peroxiredoxins are highly efficient enzymes found in nearly all living organisms. As some of the most abundant cellular proteins, they have key roles in peroxide scavenging, redox signaling and as molecular chaperones1,2,3,4,5 (Extended Data Fig. 1). These versatile enzymes are classified into various groups based on their enzymatic mechanisms or structural features.
Peroxiredoxin-1 (Prx1)/AhpC-type peroxiredoxins are typically found in a dynamic equilibrium between homodimers and homodecamers3,6,7. Homodimers interact across the B-interface, whereas homodecamers are of the (α2)5 type, in which five dimers associate through their A-type dimer interfaces8(Extended Data Fig. 1). The dimer–decamer equilibrium is influenced by factors including protein concentration, the redox state of the catalytic cysteine residues, pH and various post-translational modifications6,9,10,11,12,13,14,15,16. All characterized Prx1/AhpC-type peroxiredoxins are mechanistically typical 2-Cys peroxiredoxins, containing a peroxidatic cysteine (CP) and a resolving cysteine (CR) on each subunit, both of which are essential for catalysis8,17. Dimers arise from a ‘head-to-tail’ arrangement of subunits, which enables intermolecular disulfide bond formation between the CP and CR of opposing monomers at both ends of the dimer (Extended Data Fig. 1).
Many organisms harbor two or more Prx1/AhpC-type peroxiredoxins, often with two highly homologous isoforms located in the same subcellular compartment. In the budding yeast Saccharomyces cerevisiae, Tsa1 and Tsa2 share 86% sequence identity and are both cytosolic18,19,20. Likewise, human PRDX1 and PRDX2 and Leishmania infantum LiPRX1 and LiPRX2 co-exist in the cytosol and share 78% and 87% sequence identity, respectively21,22,23. Finally, in Arabidopsis thaliana, AtBAS1A and AtBAS1B, sharing 96% sequence identity (excluding the chloroplast-targeting transit peptides), both reside in the chloroplast stroma. Nonetheless, these isoforms frequently exhibit marked differences in biophysical and biochemical properties, including their dimer–decamer equilibria, isoelectric points, enzyme kinetics and susceptibility to various post-translational modifications10,24,25,26,27,28.
Despite their high sequence identity and very similar quaternary structures, Prx1/AhpC-type peroxiredoxins have traditionally been considered to assemble exclusively into homo-oligomeric complexes. The first challenge to this view emerged from the characterization of peroxiredoxin-based H2O2 sensors29,30. We showed that the genetically encoded roGFP2-Tsa2ΔCR probe forms enzymatically active hetero-oligomeric complexes with endogenous Tsa1 in yeast29. More recently, human PRDX1 and PRDX2 were also shown to form hetero-oligomers in vitro, although functionality, activity and in vivo relevance were not assessed31. These observations prompted us to investigate whether peroxiredoxin hetero-oligomerization is limited to specialized experimental contexts or represents a broader, biologically relevant feature of eukaryotic Prx1/AhpC-type peroxiredoxins.
In this study, we demonstrate that eukaryotic peroxiredoxins can assemble into hetero-oligomers with a broad range of subunit stoichiometries, with hetero-oligomer formation typically modulating the dimer–decamer equilibrium. In yeast, hetero-oligomerization between Tsa1 and Tsa2 is inducible upon oxidative challenge, coincident with the upregulation of TSA2 expression. Even substoichiometric incorporation of Tsa2 strongly stabilizes the decameric state. We detect human PRDX1–PRDX2 hetero-oligomers in HEK293T cells and show that Arabidopsis and Leishmania peroxiredoxins likewise assemble into functional hetero-oligomers. Given the link between peroxiredoxin oligomeric state and function, hetero-oligomerization likely represents a widespread and conserved regulatory mechanism that fine-tunes peroxiredoxin activity across different compartments, cell types and species throughout the domain Eukaryota.
Results
Tsa1 and Tsa2 form hetero-oligomers in Escherichia coli
We first asked whether Tsa1 and Tsa2 can form hetero-oligomers when purified from E. coli (Fig. 1 and Supplementary Fig. 1). Recombinant N-terminally His6-tagged or Strep-tagged Tsa1 and His6-tagged Tsa2 were produced and purified individually (Supplementary Fig. 1a) and then mixed at a 1:1 molar ratio. After a tandem-affinity purification using Ni-NTA agarose followed by StrepTactin agarose beads, no interaction between Strep-Tsa1 and His6-Tsa2 was detected. This suggests that individually purified recombinant Tsa1 and Tsa2 form stable homo-oligomeric complexes and do not exchange subunits under these conditions.
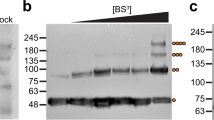
a, Purification scheme and WB analysis of the eluates from tandem-affinity copurifications of Strep-Tsa1 and His6-Tsa2 with Ni-NTA agarose, followed by StrepTactin agarose or vice versa. Eluate samples were separated by reducing SDS–PAGE. The calculated molecular masses of Strep-Tsa1 and His6-Tsa2 are 22.9 kDa and 23.8 kDa, respectively. Uncropped blots are shown in Supplementary Fig. 1. b, Tandem-affinity copurification of Strep-Tsa1ΔCR and His6-Tsa2. Protein samples were treated with NEM to block free thiols and separated by nonreducing SDS–PAGE to preserve intersubunit disulfide bonds, as shown in b. In a and b, gels are representative of those obtained during three experimental repeats, each yielding similar results. M, marker; FT, flow through; W, wash; E, eluate.
We then coexpressed genes encoding Strep-Tsa1 and His6-Tsa2 from a single plasmid in E. coli and performed tandem-affinity purifications with Ni-NTA agarose followed by StrepTactin agarose, or vice versa. Under these conditions, Strep-Tsa1 and His6-Tsa2 were successfully copurified irrespective of the order of the tandem-affinity purification, supporting an interaction between recombinant Tsa1 and Tsa2 in E. coli (Fig. 1a and Supplementary Fig. 1b,c). Semiquantitative western blotting calibrated with individually purified proteins revealed similar levels of copurified Tsa1 and Tsa2, suggesting a ~1:1 ratio of Strep-Tsa1 and His6-Tsa2 (Supplementary Fig. 1d).
To rule out other possibilities for copurification, for example, the formation of heterogenous stacks of homodecameric complexes, we performed tandem-affinity purification with the resolving cysteinyl mutant Strep-Tsa1ΔCR, which cannot form disulfide-linked Tsa1ΔCR–Tsa1ΔCR homodimers, and His6-Tsa2, which can form either Tsa2–Tsa2 homodimers or disulfide-linked Tsa1ΔCR–Tsa2 heterodimers (Fig. 1b). Nonreducing SDS–PAGE and western blotting detected two disulfide-linked dimer species for His6-Tsa2, whereas a single disulfide-linked dimer species was detected for Strep-Tsa1ΔCR, indicating that the proteins form heterodimers through the B-type interface (Fig. 1b). The formation of Tsa1–Tsa2 heterodimers was also supported by in silico analysis using HADDOCK-based molecular docking (Supplementary Fig. 2). In conclusion, recombinant yeast Tsa1 and Tsa2 readily interact through their B-type interface, and probably also through their A-type interface, forming hetero-oligomeric complexes.
Tsa1 and Tsa2 form decamers with varied stoichiometries
We then sought to visualize Tsa1–Tsa2 hetero-oligomers by negative-stain electron microscopy (EM). To this end, we coexpressed Strep-Tsa1 and His6-Tsa2-EPEA, in which Tsa2 additionally contains a C-terminal EPEA epitope that binds with high affinity to the nanobody Nbsyn2.20 (refs. 32,33). Recombinant Strep-Tsa1–His6-Tsa2-EPEA hetero-oligomers were produced and purified using tandem Ni-NTA and StrepTactin affinity chromatography, and three different fractions were collected for analysis. Strep-Tsa1 homo-oligomers and His6-Tsa2-EPEA homo-oligomers were purified separately as controls (Supplementary Fig. 3a–d). Native PAGE analysis showed that the Tsa1–Tsa2 hetero-oligomers run at a mass consistent with a decamer, between the bands observed for the Tsa1 and Tsa2 homo-oligomers (Supplementary Fig. 3e). Matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry (MS) analysis revealed that all Tsa1–Tsa2 hetero-oligomers contain disulfide-linked dimers, consistent with the lack of a reducing agent in the purification protocol (Fig. 2a). Notably, all three hetero-oligomer populations contained disulfide-linked Tsa1–Tsa2 heterodimers as well as Tsa2–Tsa2 homodimers. One of the hetero-oligomer fractions also contained disulfide-linked Tsa1–Tsa1 homodimers (Fig. 2a). All dimers were reducible with dithiothreitol (DTT), and no contamination with E. coli AhpC was detected (Supplementary Fig. 4 and Supplementary Table 1). Liquid chromatography (LC)–MS/MS analysis of the three hetero-oligomer populations revealed distinct Tsa1:Tsa2 molar ratios; 0.25 mol mol−1 (most Tsa2), 0.42 mol mol−1 and 0.88 mol mol−1 (least Tsa2; PRIDE accession PXD060819), indicating hetero-oligomer populations with different subunit stoichiometries. Finally, negative-stain EM imaging of Tsa1 and Tsa2 homo-oligomers and the 0.88 mol mol−1 Tsa1–Tsa2 hetero-oligomer fraction revealed donut-shaped particles in all samples, consistent with decamers of similar diameter (Supplementary Fig. 5).
a, MALDI-TOF MS analysis of Strep-Tsa1 and His6-Tsa2-EPEA homo-oligomers and Strep-Tsa1–His6-Tsa2-EPEA hetero-oligomers. The Tsa1:Tsa2 molar ratio of each hetero-oligomer sample is displayed above each panel. The signal intensity (a.u.) is plotted against different m/z ratios (n = 2 independent experimental repeats). Expected masses are listed in Supplementary Table 1. b, Negative staining EM analysis of Nbsyn2.20 binding to the 0.88 mol mol−1 Strep-Tsa1–His6-Tsa2-EPEA heterodecamer at multiple positions. Protein particles appear bright against the 2% uranyl-acetate stain, revealing a characteristic decameric ‘donut-like’ structure. Nbsyn2.20 positions are marked with white arrows. Scale bars, 200 Å. Images are representative of those obtained during three experimental repeats, each yielding similar results. c, NanoDSF analysis of Strep-Tsa1 and His6-Tsa2-EPEA homo-oligomers and Strep-Tsa1–His6-Tsa2-EPEA hetero-oligomers. Temperature was increased to 100 °C (2 °C min−1; n = 3 technical replicates). Tm, melting temperature. d, CD analysis of Strep-Tsa1 and His6-Tsa2-EPEA homo-oligomers, and Strep-Tsa1–His6-Tsa2-EPEA hetero-oligomer samples (n = 5 technical replicates). e, Mass photometry analysis of Strep-Tsa1, His6-Tsa2-EPEA and the 0.88 mol mol−1 Strep-Tsa1–His6-Tsa2-EPEA sample at 20 °C, 30 °C, 40 °C and 45 °C, respectively. Data were acquired for 60 s, and counts of individual molecules were plotted against their molecular weight (n = 3 technical replicates with separate aliquots of proteins obtained from 1 purification). Data are presented as mean ± s.d.
We then used the Nbsyn2.20 nanobody to visualize the position of Tsa2 within the hetero-oligomers. We first assessed the specificity of Nbsyn2.20 binding to the EPEA tag. Biolayer interferometry (BLI) and mass photometry confirmed specific interaction with EPEA-tagged proteins, with no binding observed to Tsa1 homo-oligomers (Supplementary Fig. 6a). Mass photometry further revealed that at least two Nbsyn2.20 molecules bind to a Tsa1–Tsa2 heterodecamer. Multiple populations of Tsa1–Tsa2 heterodecamers were detected, as reflected by an asymmetric molecular-weight peak with a tail. The Tsa2 homo-oligomer control showed various populations with different numbers of Nbsyn2.20 molecules bound, likely due to the dynamic nature of the interaction (Supplementary Fig. 6b). Tsa1 homodecamers did not bind Nbsyn2.20, confirming the specificity of the nanobody–EPEA interaction.
Finally, negative-stain EM imaging of Tsa1–Tsa2 heterodecamers in the presence of Nbsyn2.20 showed a diverse population with different numbers of nanobodies binding to different particles. This is likely explained by the existence of different stoichiometries of Tsa1 and Tsa2 in solution (Fig. 2b). In agreement with the BLI and mass photometry analyses, no interaction was observed between Tsa1 homo-oligomers and Nbsyn2.20, whereas Tsa2 homo-oligomers displayed up to ten Nbsyn2.20 molecules bound at distinct positions (Fig. 2b). In summary, our structural analysis of recombinant Tsa1–Tsa2 hetero-oligomers reveals a donut-shaped organization in which Tsa1 and Tsa2 likely assemble into multiple, distinct, heterodecameric species.
Tsa2 stabilizes the decameric state of the hetero-oligomers
We then evaluated the decamer stability and dimer–decamer equilibrium of Tsa1–Tsa2 hetero-oligomers in comparison to their homo-oligomeric counterparts, focusing here on the disulfide-linked oxidation form. NanoDSF analysis of the Tsa1 homo-oligomer revealed two inflection points at 39.5 °C and 71.2 °C (Fig. 2c). In contrast, Tsa2 homo-oligomers displayed only one inflection point at 78.5 °C. The three purified Tsa1–Tsa2 hetero-oligomer populations also presented only one inflection point at 76.4 °C (0.25 mol mol−1 population), 74.6 °C (0.42 mol mol−1 population) and 70.7 °C (0.88 mol mol−1 population; Fig. 2c). These data indicate that increasing Tsa2 content enhances oligomer stability, as reflected by progressively higher inflection temperatures. We then performed circular dichroism (CD) measurements for the Tsa1 homo-oligomer, Tsa2 homo-oligomer and the 0.88 mol mol−1 Tsa1–Tsa2 hetero-oligomer sample at temperatures of 25 °C, 39.5 °C, the respective inflection point temperature for each sample and 95 °C (Fig. 2d). The CD data showed no change in secondary structure at 39.5 °C in any sample. For all three proteins, the second inflection point near 75 °C in Fig. 2c corresponds to a loss of some secondary structural features; however, none of the protein samples fully denatured at any temperature tested, suggesting that the Tm values lie above 95 °C (Fig. 2d).
To gain further insight into the molecular basis of the inflection points observed by nanoDSF, we again used mass photometry (Fig. 2e and Supplementary Table 2). Measurements were performed at 20 °C, 30 °C, 40 °C and 45 °C for the Tsa1 homo-oligomer, Tsa2 homo-oligomer and the 0.88 mol mol−1 Tsa1–Tsa2 hetero-oligomer samples, respectively. The relative abundance (%) of low-molecular-weight (LMW) species and decamers was quantified under each condition. Tsa1 showed a clear loss of the decameric state and dissociation to dimers with increasing temperature, with a pronounced shift at 40 °C. This supports the interpretation that the 39.5 °C inflection point observed by nanoDSF reflects dissociation of Tsa1 homodecamers into homodimers. In contrast, the Tsa2 homo-oligomer and 0.88 mol mol−1 Tsa1–Tsa2 hetero-oligomer samples displayed far less dissociation into LMW oligomers (Fig. 2e).
To examine potential effects of peptide tags on oligomer stability, we used mass photometry to monitor the temperature-dependent dimer–decamer ratios of Tsa1, Tsa2 and Tsa1–Tsa2 hetero-oligomers with N-terminal and C-terminal green fluorescent protein (GFP) tags (Supplementary Fig. 7 and Supplementary Table 2). An N-terminal GFP tag destabilized the decameric state, particularly for Tsa1. Nonetheless, the increased decamer stability of the hetero-oligomeric complexes, and especially for Tsa2, was consistently observed regardless of tag identity or tag position. Notably, in the His6-GFP-TEV-Tsa2 homo-oligomer and in the Tsa1–Tsa2-TEV-GFP-His6 hetero-oligomer samples, we detected some AhpC contamination (Supplementary Figs. 8 and 9; LC–MS data—PRIDE accession PXD060819).
In summary, our data support that the Tsa1 homodecamer is less stable than the Tsa2 homodecamer, consistent with previous reports27. Intriguingly, the incorporation of substoichiometric amounts of Tsa2 strongly stabilizes the decameric state of the resultant Tsa1–Tsa2 heterodecamers, suggesting that hetero-oligomerization may serve as a mechanism for regulating peroxiredoxin oligomeric state dynamics.
H2O2-induced Tsa1–Tsa2 assembly stabilizes decamers in yeast
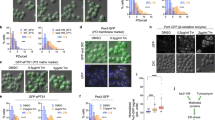
Under standard growth conditions, Tsa1 is highly abundant in yeast cells, whereas Tsa2 levels are much lower34. However, TSA2 expression is known to be inducible under oxidative challenge35,36. To investigate whether Tsa1 and Tsa2 hetero-oligomers form when expressed from their native promoters, we replaced either TSA1 or TSA2 at their endogenous genomic loci with roGFP2-tagged versions, that is, RoGFP2-TSA1 and RoGFP2-TSA2, while maintaining the native promoter and 3′ untranslated regions. Fluorescence measurements confirmed that the roGFP2-Tsa1 level was approximately tenfold higher than that of roGFP2-Tsa2 (Fig. 3a). Upon exposure to 1 mM H2O2, the roGFP2-Tsa1 fluorescence remained unchanged, whereas the roGFP2-Tsa2 fluorescence increased approximately fivefold within 90 min, confirming induction of TSA2 expression but not TSA1 upon oxidative challenge (Fig. 3a).
a, Graph showing the GFP fluorescence intensity of roGFP2-Tsa1 and roGFP2-Tsa2 constructs in yeast cells at the indicated time points after the treatment with 1 mM exogenous H2O2. The constructs were expressed from genes integrated into the TSA1 and TSA2 genomic loci, respectively, remaining under the control of the native promoters and 3′ untranslated regions. Experiments were repeated thrice with independent yeast cultures. Data represent mean ± s.d. b, Clear native PAGE gels, monitored for GFP fluorescence, of lysates taken from the cultures of the indicated yeast strains at the indicated time points following the addition of 1 mM exogenous H2O2. Wedges indicate hetero-oligomers containing an increasing number of Tsa2 subunits (first panel from the left) and an increasing number of roGFP2-Tsa2 subunits (third panel from the left). Two further experimental repeats are shown in Supplementary Fig. 10.
To further explore whether TSA2 induction promotes hetero-oligomer formation, we performed clear native gel electrophoresis. Under standard conditions, roGFP2-Tsa1 migrated predominantly as a dimer. Within 30 min after H2O2 addition to Δtsa1::RoGFP2-TSA1 cells, we observed additional bands at a molecular mass consistent with roGFP2-Tsa1–Tsa2 heterodecamers, and these bands intensified during the time-course of the experiment (Fig. 3b and Supplementary Fig. 10). We observed a ladder pattern of decamer bands, consistent with the formation of roGFP2-Tsa1–Tsa2 heterodecamers containing different subunit stoichiometries (Fig. 3b). No decameric bands were observed in Δtsa1::RoGFP2-TSA1 Δtsa2 cells in which TSA2 is deleted (Fig. 3b). As an additional control, we monitored the hyperoxidation status of the peroxidatic cysteines, which could potentially stabilize the decameric state (Supplementary Fig. 11a). No hyperoxidation was detected under our experimental conditions.
In Δtsa2::RoGFP2-TSA2 cells, the predominant species were decamers and a band that we interpret as unassembled monomers. Upon H2O2 treatment and the resulting induction of roGFP2-Tsa2 expression, a series of higher-molecular-mass bands appeared above the original decamer band. This suggests that as roGFP2-Tsa2 levels increase, hetero-oligomers containing increasing proportions of roGFP2-Tsa2 subunits are formed (Fig. 3b). Consistent with our previous results, we found no evidence of hyperoxidation in these cells during the time-course of this experiment (Supplementary Fig. 11b).
In conclusion, our findings demonstrate that Tsa1–Tsa2 hetero-oligomerization is inducible upon oxidative challenge and that incorporation of only one or two Tsa2 subunits is sufficient to stabilize a heterodecameric complex.
Hetero-oligomerization allows high structural plasticity
Our data support the existence of Tsa1–Tsa2 interactions across both the A-type and B-type interfaces, prompting us to calculate the theoretical maximum number of possible different hetero-oligomeric configurations. This analysis revealed that two distinct monomer types can assemble into 120 unique decamers. These are defined as structures that are not rotations of the same decamer around the fivefold symmetry axis perpendicular to the plane of the decamer or across the five twofold symmetry axes parallel to this plane (Extended Data Fig. 2 and Supplementary Note). These 120 distinct heterodecamers include 2 distinct homodecameric structures, 2 decamers with a 1:9 or 9:1 stoichiometric ratio, 14 decamers with 2:8 or 8:2 ratios, 24 decamers with 3:7 or 7:3 ratios, 52 decamers with 4:6 or 6:4 ratios and 26 decamers with 5:5 ratio.
The potential number of distinct hetero-oligomers increases dramatically when the multiple structural states of each monomer are considered. Each monomer switches among at least three distinct structural states during the catalytic cycle, and these states multiply several-fold when known post-translational modifications are also considered37. Consequently, the number of possible unique heterodecamer configurations scales with the 10th power of the number of monomer states (Extended Data Fig. 2b and Supplementary Note). This number would far exceed the number of peroxiredoxin decamers present in a cell, even if each monomer type had only four states. The potential configurations increase substantially when more realistic state counts are considered. Therefore, only a very small fraction of the possible peroxiredoxin hetero-oligomer configurations can exist in a cell at any given moment.
Tsa1–Tsa2 hetero-oligomers show homo-oligomer-like kinetics
We recently analyzed the catalytic cycle of recombinant His6-Tsa1 using stopped-flow kinetic measurements, revealing three distinct reaction phases for the H2O2-dependent oxidation of reduced Tsa1, that is, Tsa1(SH)2, and three phases for the yeast Trx1-dependent reduction of Tsa1 disulfide, that is, Tsa1(S2)38. To compare the catalytic activity of Tsa1–Tsa2 hetero-oligomers to that of their homo-oligomeric counterparts, we analyzed individually purified His6-Tsa1 and His6-Tsa2, and Tsa1–Tsa2 hetero-oligomers (Fig. 4, Supplementary Fig. 1 and Supplementary Table 3).
a, Reaction scheme for the oxidation and reduction of recombinant Tsa1 and/or Tsa2. Only the dimeric enzyme species are shown for simplicity. b, Representative changes of tryptophan fluorescence during the H2O2-dependent oxidation of individually purified His6-Tsa1(SH)2 (left) or His6-Tsa2(SH)2 (right). c, Secondary plots of the observed rate constants (kobs) from exponential fits of the three reaction phases at variable H2O2 concentrations from b. d, Representative changes of tryptophan fluorescence during the yeast Trx1-dependent reduction of individually purified His6-Tsa1(S2) (left) or His6-Tsa2(S2) (right). e, Secondary plots of the kobs values from exponential fits of the two or three reaction phases at variable Trx1 concentrations from d. f, Representative changes of tryptophan fluorescence during the H2O2-dependent oxidation of copurified His6-Tsa1(SH)2 and His6-Tsa2(SH)2 (left) and secondary plot of the kobs values from exponential fits of the first reaction phase (right). g, Representative changes of tryptophan fluorescence during the yeast Trx1-dependent reduction of copurified His6-Tsa1(S2) and His6-Tsa2(S2) (Supplementary Fig. 1). Quadruple exponential fits (black lines) were calculated using the kobs values of the first phase of individually purified His6-Tsa1 and His6-Tsa2 at the corresponding Trx1 concentration as input. All stopped-flow measurements were performed at pH 7.4 and 25 °C. Ten times higher enzyme concentrations were used to highlight the second and third oxidation phases (dashed lines). Data from c and e were from technical triplicates of independent biological triplicate protein purifications and measurements. Data from f and fits in g were from technical triplicates of a single measurement. Error bars in all panels represent the s.d. Rate constants are summarized in Supplementary Table 3.
As previously reported for Tsa1 (refs. 27,38,39), three distinct phases were observed during the oxidation of individually purified His6-Tsa1(SH)2 and His6-Tsa2(SH)2. First, a decrease in tryptophan fluorescence reflects the rapid formation of the sulfenic acid species at higher H2O2 concentrations. Second, an increase in fluorescence indicates a conformational change from the fully folded to the locally unfolded state. Third, fluorescence decreases again due to the formation of an intermolecular disulfide bond at the B-type dimer interface (Fig. 4a,b). The second-order rate constant k1ox of 1.3 × 107 M−1 s−1 for the sulfenic acid formation in His6-Tsa2 was tenfold lower than that for His6-Tsa1, whereas the first-order rate constants (k2ox and k3ox) for the H2O2-independent phases were similar for both enzymes (Fig. 4c and Supplementary Table 3). Control experiments with Strep-Tsa1 revealed that the Strep-tag had no major effect on the oxidation kinetics compared to His6-Tsa1 (Supplementary Table 3).
For the yeast Trx1-dependent reduction, three phases were observed for His6-Tsa1(S2) and two phases for His6-Tsa2(S2) (Fig. 4a,d). The second-order rate constant k1red of 3.9 × 106 M−1 s−1 for the formation of the mixed disulfide between His6-Tsa2 and yeast Trx1 was slightly higher than that for the mixed disulfide of His6-Tsa1 (Fig. 4e and Supplementary Table 3). Similar y-axis intercepts, 8.7 s−1 for His6-Tsa1 and 10.8 s−1 for His6-Tsa2, suggest either a reverse reaction or a conformational change. At higher Trx1 concentrations, the second and third phases exhibited pseudo-first-order reaction kinetics, with k2red values around 10 s−1 for both enzymes (Fig. 4e and Supplementary Table 3). The second phase likely corresponds to the formation of reduced peroxiredoxins and Trx1(S2). The third phase with k3red (0.6 s−1) for His6-Tsa1 may indicate a Trx1-induced decamer dissociation, a process not observed for His6-Tsa2.
The oxidation kinetics of copurified recombinant Strep-Tsa1–His6-Tsa2 hetero-oligomers were broadly similar to those of the individual enzymes (Fig. 4f) but characterization of the first oxidation phase required a quadruple exponential fit. The rate constants, k1oxA and k1oxB, had intermediate values of 6.1 × 107 M−1 s−1 and 2.8 × 107 M−1 s−1 compared to the k1ox values of the individual enzymes (Fig. 4f and Supplementary Table 3). The second and third phases displayed similar kinetics to those observed for the homo-oligomeric enzymes, yielding k2ox and k3ox values of 45 and 3.6 s−1, respectively. Premixed 1:1 mixtures of recombinant His6-Tsa1 and His6-Tsa2, which do not exchange subunits and thus do not assemble into hetero-oligomers, showed similar oxidation kinetics as the individual enzymes, with k1oxA and k1oxB values from a double exponential fit of 9.8 × 107 M−1 s−1 and 1.2 × 107 M−1 s−1 (Supplementary Table 3). Thus, the oxidation kinetics of Strep-Tsa1 and His6-Tsa2 mixtures reflect the superposition of the individual enzyme activities, whereas Strep-Tsa1–His6-Tsa2 hetero-oligomers exhibit intermediate macroscopic rate constants (k1ox) for the reaction with H2O2.
The reduction kinetics of Strep-Tsa1–His6-Tsa2 hetero-oligomers resembled those of His6-Tsa1 but required a quadruple exponential fit with two preset rate constants (Fig. 4g). The best fit was obtained using kobs values for the first reduction phase from the individually purified enzymes at the corresponding Trx1 concentration. The remaining kobs values were consistent with those observed for His6-Tsa1. This suggests that the reduction kinetics of Strep-Tsa1–His6-Tsa2 hetero-oligomers can be approximated by superimposing the reduction profiles of the individually purified enzymes.
To examine potential concentration-dependent effects on the rate constants, we also established a coupled steady-state peroxidase assay containing NADPH, TrxR, Trx1, H2O2 and nanomolar peroxidase concentrations (Extended Data Fig. 3a,b). The assays displayed a high degree of variability and required exact mixing routines as well as fixed volumes and dilutions of stock solutions, consistent with sensitive temperature-dependent and concentration-dependent effects on the oligomeric state. Variability was highest for His6-Tsa1, consistent with the results shown in Fig. 2. However, in contrast to previous reports that suggested rate constants around 104 M−1s−1 based on apparent kcat and Km values for recombinant Tsa1 and Tsa2 (ref. 27), our steady-state assays gave approximate rate oxidation constants in the range of 107 – 108 M−1 s−1 (Extended Data Fig. 3c–e and Supplementary Table 4), differing from the stopped-flow values by less than one order of magnitude (Supplementary Table 3). Approximate rate constants for the Trx1-dependent reduction were around 106‒107 M−1s−1 and also agreed within one order of magnitude (Extended Data Fig. 3f and Supplementary Tables 3 and 5).
One notable difference between the second-order rate constants from the stopped-flow and steady-state measurements was observed for His6-Tsa1, which showed a tenfold lower kcatapp/Kmapp value in the steady-state assays (Supplementary Tables 3 and 4). A likely explanation is the higher sensitivity of the threonine-containing active site of Tsa1 to hyperoxidation compared to the serine-containing active site of Tsa2 (ref. 40). The inactivation became relevant and detectable at micromolar-to-nanomolar peroxide-to-Tsa1 ratios but was reduced or even absent for Strep-Tsa1–His6-Tsa2 hetero-oligomers (Extended Data Fig. 3c,e).
Another notable difference between the stopped-flow and steady-state measurements was that the k3ox or k2red in Supplementary Table 3 were smaller than the v/(E) values shown in Extended Data Fig. 3b–f. One explanation is that nanomolar enzyme concentrations in the steady-state assay favor substantial populations of homodimeric or heterodimeric species with distinct kinetic properties compared with the predominantly decameric species present at micromolar concentrations in the stopped-flow measurements. Different dimer concentrations also explain the v/(E) values for 10 nM Strep-Tsa1–His6-Tsa2 hetero-oligomers, which were about four times higher than for 50 nM His6-Tsa1 and also 1.5 times higher than for 10 nM His6-Tsa2 (Extended Data Fig. 3c–e). Overall, His6-Tsa1, His6-Tsa2 and Strep-Tsa1–His6-Tsa2 hetero-oligomers exhibit similar second-order rate constants for H2O2 and Trx1 at both micromolar and nanomolar peroxiredoxin concentrations. However, nanomolar enzyme concentrations resulted in higher v/(E) values, likely reflecting an increased dimer abundance. In the steady-state assays, very high peroxide-to-enzyme ratios caused hyperoxidation of His6-Tsa1 but not of His6-Tsa2 or Strep-Tsa1–His6-Tsa2 hetero-oligomers.
In summary, analyses of oxidation and reduction kinetics of homo-oligomers, mixtures of Tsa1 and Tsa2 homo-oligomers, and Tsa1–Tsa2 hetero-oligomers revealed that the hetero-oligomers exhibit enzyme kinetics largely comparable to those of their homo-oligomeric counterparts. Although the formation of the sulfenic acid species in His6-Tsa1 was approximately tenfold faster than in His6-Tsa2, copurified Tsa1–Tsa2 hetero-oligomers displayed intermediate macroscopic rate constants for the reaction with H2O2 in the stopped-flow experiments. Our findings suggest that hetero-oligomerization and/or heterodimerization might protect Tsa1 from hyperoxidation at low enzyme concentrations at the cost of a slightly reduced reactivity with H2O2 at high enzyme concentrations.
Tsa1–Tsa2 hetero-oligomers are enzymatically active in yeast
We previously demonstrated that a roGFP2-Tsa2ΔCR probe can form enzymatically active hetero-oligomers with endogenous Tsa1 in the yeast cytosol29. To further investigate the assembly and enzymatic activity of Tsa1–Tsa2 hetero-oligomers in yeast, we expressed roGFP2-Tsa1 variants in Δtsa1Δtsa2 yeast cells together with either wild-type (WT) or catalytically inactive Tsa2. Specifically, cells were transformed with plasmids encoding roGFP2-Tsa1ΔCPΔCR, in which both catalytic cysteine residues were mutated to serine, and either Tsa2wt or Tsa2ΔCPΔCR. Cells were then exposed to increasing concentrations of H2O2 (0–500 µM), and roGFP2 oxidation was quantified using a fluorescence plate-reader assay (Supplementary Fig. 12). In this assay, roGFP2 oxidation in response to H2O2 is efficiently catalyzed in trans, that is, by a partner peroxiredoxin in the dimer or decamer. Notably, in trans oxidation is only possible if the two peroxiredoxins are physically associated in a complex.
In Δtsa1Δtsa2 cells coexpressing roGFP2-TSA1ΔCPΔCR and TSA2wt, roGFP2 oxidation was detected at exogenous H2O2 concentrations as low as 10 µM, indicating a highly sensitive response. In contrast, no roGFP2 oxidation was observed in cells coexpressing roGFP2-TSA1ΔCPΔCR and TSA2ΔCPΔCR, even at the highest H2O2 concentration. Because roGFP2 is only very poorly oxidized directly by H2O2, these results confirm that Tsa1 and Tsa2 assembles into functional hetero-oligomers in the yeast cytosol29,30,41,42. In a complementary experiment, roGFP2-TSA2ΔCPΔCR was coexpressed with either TSA1wt or TSA1ΔCPΔCR, resulting in robust roGFP2 oxidation only in the presence of Tsa1wt, with almost no oxidation with Tsa1ΔCPΔCR (Supplementary Fig. 12).
In summary, our findings confirm that Tsa1 and Tsa2 can form catalytically active hetero-oligomers in the yeast cytosol and function cooperatively to mediate roGFP2 oxidation.
Native PRDX1 and PRDX2 form hetero-oligomers in HEK293 cells
To determine whether peroxiredoxin hetero-oligomerization extends beyond yeast, we investigated human PRDX1 and PRDX2, which were recently shown to form hetero-oligomers in vitro31. Despite sharing 78% sequence identity, PRDX1 and PRDX2 have distinct isoelectric points (7.80 and 5.71, respectively), allowing to use ion-exchange chromatography to separate them in Flp-In T-Rex HEK293 cell lysates (Fig. 5 and Supplementary Fig. 13). Western blot (WB) analysis of collected fractions revealed that, although PRDX1 and PRDX2 primarily eluted at different positions along the NaCl gradient, they also eluted across a broad range of partially overlapping fractions (Fig. 5a,b).
a,b, WB analysis (a) and quantification of AEC fractions (b) from HEK293 WT and PRDX1-KO and PRDX2-KO cell lysates (n = 3 experimental repeats, the 2 further experimental repeats are shown in Supplementary Fig. 13). Proteins from the three cell lines were separated by AEC and fractions subsequently analyzed by WB for the presence of PRDX1 and PRDX2. Lysates from the three cell lines were prepared and fractionated independently, with no mixing of lysates at any stage. WBs (a) and quantification (b) show that in the PRDX-KO cell lines, the PRDX1 and PRDX2 signals show no overlap in the middle fractions, whereas in the WT lysate, there is a clear overlap in the middle fractions, indicating a shift in the isoelectric point of the protein complex species, suggesting the presence of hetero-oligomers. c, Graphs showing change in OxD of the indicated roGFP2-PRDX fusion constructs in Δtsa1Δtsa2 yeast cells treated with the indicated concentrations of H2O2. Experiments were repeated thrice with independent yeast cultures. Data are presented as mean ± s.d. AEC, anion exchange chromatography; pl, isoelectric point; OxD, degree of oxidation.
To further explore this interaction, we repeated the experiment using lysates from CRISPR–Cas9 knockout (KO) cells lacking either PRDX1 or PRDX2 (ref. 43). In PRDX1 KO and PRDX2 KO cells, the remaining PRDX isoform eluted in a narrower range of fractions, with no overlap observed (Fig. 5a,b). This shift in elution pattern suggests that PRDX1 and PRDX2 influence each other’s chromatographic behavior and provides strong evidence for their interaction as native hetero-oligomers.
To assess whether PRDX1–PRDX2 hetero-oligomers are enzymatically active, we used a yeast-based roGFP2 assay. Heterologous roGFP2-PRDX1ΔCPΔCR was expressed in Δtsa1Δtsa2 yeast cells together with either PRDX2 or PRDX2ΔCPΔCR and roGFP2 oxidation was measured upon H2O2 exposure (Fig. 5c). Consistent with our observation with yeast Tsa1 and Tsa2, roGFP2 oxidation was observed only when PRDX2 was present, indicating that PRDX1 requires an active PRDX2 partner to mediate roGFP oxidation. Likewise, roGFP2-PRDX2ΔCPΔCR only supported roGFP2 oxidation in the presence of PRDX1, but not PRDX1ΔCPΔCR (Fig. 5c).
In summary, these findings demonstrate that PRDX1 and PRDX2 form enzymatically active hetero-oligomers in both HEK293T cells and yeast and support the presence of peroxiredoxin hetero-oligomerization across species.
Diverse eukaryotic peroxiredoxins form hetero-oligomers
To further assess whether hetero-oligomerization is a conserved feature of peroxiredoxins across eukaryotes, we extended our analysis to peroxiredoxins from non-opisthokont species using the yeast-based roGFP2 system. We generated roGFP2 fusion constructs with LiPRX1 and LiPRX2 from the cytosol of the kinetoplastid parasite L. infantum23, as well as AtBAS1A and AtBAS1B from the chloroplast stroma of the plant A. thaliana44,45. Each construct was expressed in Δtsa1Δtsa2 yeast cells together with either the WT or double cysteine mutant of the corresponding partner peroxiredoxin isoform (Fig. 6a). In all cases, roGFP2 oxidation was observed exclusively in the presence of the WT partner and not with the cysteine mutant, consistent with our observations for yeast and human peroxiredoxins.
a, Graphs showing the change in the OxD in response to 1 mM H2O2 of roGFP2-AtBAS1BΔCPΔCR and roGFP2-AtBAS1AΔCPΔCR constructs (left) or roGFP2-LiPrx1ΔCPΔCR and roGFP2-LiPrx2ΔCPΔCR constructs (right) expressed in Δtsa1Δtsa2 yeast together with either a WT or cysteine-less (ΔCPΔCR) variant of the corresponding partner peroxiredoxin. Experiments were repeated thrice with independent yeast cultures. Data are presented as mean ± s.d. b, Clear native PAGE analysis of lysates from Δtsa1Δtsa2 yeast strains expressing the indicated combination of constructs. Gels were imaged for GFP fluorescence. Two further experimental repeats are shown in Supplementary Fig. 14. c, Graph showing the percentage of sequenced eukaryotic genomes in which there are predicted to be two or more Prx1/AhpC-type peroxiredoxins present in the indicated subcellular compartments. d, Model illustrating the impact of peroxiredoxin hetero-oligomerization on oligomeric state. Substoichiometric subunit incorporation is capable of strongly shifting the dimer–decamer equilibrium. HD, homodecamer and heterodecamer.
To explore the oligomeric state of these hetero-oligomers, we coexpressed roGFP2-AtBAS1A and roGFP2-AtBAS1B with either an empty vector, AtBAS1A or AtBAS1B (Fig. 6b and Supplementary Fig. 14). While roGFP2-AtBAS1B predominantly formed a decamer, roGFP2-AtBAS1A was almost exclusively dimeric. Coexpression of AtBAS1A with roGFP2-AtBAS1B induced a shift from a decamer to a heterodimeric form, whereas coexpression of AtBAS1B with roGFP2-AtBAS1B maintained decamer stability, producing a series of heterodecamers with varying stoichiometries and a minor heterodimer band. Conversely, coexpression of AtBAS1B with roGFP2-AtBAS1A led to partial decamer formation, but only when AtBAS1A subunits were a minority, indicating that AtBAS1A destabilizes decamers. Coexpression of roGFP2-AtBAS1A and AtBAS1A resulted exclusively in heterodimers.
Similar results were obtained for L. infantum peroxiredoxins (Fig. 6b). In yeast lysates, roGFP2-LiPrx1 primarily assembled into decamers, whereas roGFP2-LiPrx2 showed little to no oligomerization. Coexpression of LiPRX2 with roGFP2-LiPRX1 disrupted decamer formation and promoted a shift toward putative heterodimeric species, whereas coexpression with LiPRX1 had no effect on decamer stability. Consistent with the apparent differences in decamer stability, coexpression of roGFP2-LiPRX2 with LiPRX1 induced a shift to decameric species, while coexpression with LiPRX2 did not.
In summary, our experiments suggest that strong differences in dimer–decamer equilibria are widespread among eukaryotic peroxiredoxins. Even limited incorporation of a second peroxiredoxin isoform can markedly shift the prevailing oligomeric state, highlighting the functional significance of hetero-oligomerization across diverse organisms.
Pairs of Prx1-type peroxiredoxins occur in most eukaryotes
Finally, we sought to determine the percentage of eukaryotes in which two or more Prx1/AhpC-type peroxiredoxin proteoforms are predicted to be present in the same subcellular compartment. An extensive sequence search identified 11,415 peroxiredoxin candidate sequences across 1,525 eukaryotic species representing the kingdoms Viridiplantae, Metazoa and Fungi. Of these species, 1,471 were predicted to contain at least one Prx1-type peroxiredoxin in the cytosol, and two or more Prx1-type cytosolic peroxiredoxins were predicted in 1,326 species (Fig. 6c and Supplementary Fig. 15). Mitochondria-targeted Prx1-type peroxiredoxins were predicted in 1,236 species, with 771 species predicted to contain two or more mitochondrial peroxiredoxins. Within the kingdom Viridiplantae, plastid-localized Prx1-type peroxiredoxins were predicted in 347 species, with two or more plastid-localized peroxiredoxins in 314 species.
Overall, our data indicate that peroxiredoxin hetero-oligomerization is likely to be relevant in more than 80% of sequenced eukaryotic species and probably occurs in several different subcellular compartments.
Discussion
Peroxiredoxins have been intensively studied since their discovery more than 30 years ago1,3,5,7,46,47. Nonetheless, Prx1/AhpC-type peroxiredoxins have until now been examined almost exclusively as homo-oligomeric complexes. Here we show that hetero-oligomerization is a common feature of Prx1/AhpC-type peroxiredoxins throughout the domain Eukaryota. Our findings therefore call for a careful re-evaluation of peroxiredoxin structural dynamics and functional diversity (Fig. 6d).
Regulation of the oligomeric state of peroxiredoxins appears to be closely linked to their functions. For example, the decameric state of Prx1/AhpC-type peroxiredoxins has been associated with protein chaperone activity48,49,50,51,52, whereas both dimers and decamers retain enzymatic activity6,11,53. In a possible example of functional specialization through structural adaptation, differences in dimer–decamer equilibria are common among Prx1/AhpC-type peroxiredoxins within an organism. For example, human PRDX1 is more stable as a decamer than PRDX2 (refs. 10,24). Likewise, yeast Tsa2 is more stable in the decameric form than Tsa1 (ref. 27). We observe similar differences in the dimer–decamer equilibria of A. thaliana chloroplast and L. infantum cytosolic peroxiredoxins. However, this simplistic binary view of peroxiredoxins as either ‘type-A’ or ‘type-B’ becomes difficult to sustain in light of hetero-oligomerization. In the absence of mechanisms to actively prevent hetero-oligomerization, of which we have found no evidence, cells are likely to contain a complex mixture of peroxiredoxin hetero-oligomers with diverse subunit stoichiometries, potentially each having its own dimer–decamer equilibrium (Fig. 6d). On the other hand, the observation that even substoichiometric incorporation of Tsa2 can impart Tsa2-like properties onto a Tsa1 decamer may reduce the complexity somewhat, at least in yeast. Finally, the possible role of hetero-oligomerization in the formation of higher-order assemblies, including stacked decamers and 12-decamer-containing dodecahedrons, remains largely unexplored, and the biological functions of these structures are still unclear13,54.
Hyperoxidation has been shown to be an important factor leading to decamer stabilization in several peroxiredoxins. However, the hyperoxidation sensitivity of Tsa1 is found to be approximately ~9× and ~100× lower than human PRDX1 and PRDX2, respectively39,40. Consistent with these findings and the lack of a strong signal for hyperoxidation in our WBs, we consider it unlikely that hyperoxidation is substantially affecting the stability of the hetero-oligomers formed in our yeast induction assays. However, given that changes in just one or two subunits can influence the properties of the entire decamer, hyperoxidation may still become relevant under other experimental conditions. For example, cell growth stage, carbon source and cell density are all important factors influencing hyperoxidation sensitivity in our hands.
Hetero-oligomerization not only influences oligomer stability but also brings together isoform-specific biochemical properties, including differences in isoelectric point, catalytic rates and susceptibility to hyperoxidation10,24,27, as well as modifications including limited proteolysis55, tyrosine nitration56, S-nitrosylation57, S-glutathionylation14 and phosphorylation58. By allowing for a graduated blending of properties, peroxiredoxin hetero-oligomerization may broaden functional outputs, for example, by modulating target protein specificity in redox signaling relays or altering the spectrum of potential chaperone clients. The structural plasticity of peroxiredoxins may therefore underpin functional plasticity, positioning them as potential hubs for integrating and transmitting signals within specific signaling pathways (Fig. 6d). This conceptual shift also raises new questions. If cells predominantly contain diverse hetero-oligomeric assemblies instead of pure homo-oligomeric complexes, how do individual isoforms maintain selective interactions with specific target proteins such as STAT3 (refs. 58,59,60) or ASK1 (ref. 61)? How signaling fidelity is preserved in the context of hetero-oligomeric mixtures remains unknown and should now be explored.
The extent to which cells can regulate the formation and subunit composition of hetero-oligomers remains unclear. One obvious point of control is transcription. As shown here, this mechanism is prominent in yeast—under oxidative challenge, TSA2, normally expressed at much lower levels than TSA1, is strongly induced, leading to hetero-oligomers with a much broader range of subunit stoichiometries than those found in unstressed cells. Post-translational modifications may also modulate hetero-oligomer composition by altering interactions between different peroxiredoxin isoforms. Whether subunits can dynamically exchange between oligomers is unclear and may differ among isoforms depending on the stability of their dimeric and decameric states.
Are hetero-oligomers unique to Prx1-type enzymes? In principle, Prx1-type enzymes share a higher degree of structural similarity with Prx6-type enzymes in contrast to other peroxiredoxin subfamilies8. To explore this possibility, we recently analyzed the cytosolic Prx1-type and Prx6-type enzymes from the malaria parasite Plasmodium falciparum but found no evidence of interaction62. In summary, while further studies are needed to investigate potential interactions among other peroxiredoxin subfamilies, current evidence suggests that heterodimer and hetero-oligomer formation is likely restricted to members of the Prx1/AhpC-type subfamily.
Future work should clarify how hetero-oligomerization influences peroxiredoxin-mediated signaling and whether it contributes to diseases where redox homeostasis is disrupted. Given the increasing recognition of peroxiredoxins in aging, cancer and neurodegenerative diseases, understanding how hetero-oligomerization influences their function may reveal new principles of redox regulation and uncover therapeutic opportunities. Defining the structural basis of hetero-oligomerization will be essential: cryo-EM and crystallography can resolve the interaction interfaces governing stability, while targeted mutagenesis can identify critical residues that drive hetero-oligomer assembly and link these to functional consequences.
In summary, peroxiredoxin hetero-oligomerization remains a largely underexplored phenomenon that likely holds broad significance for cellular function and disease.
Methods
Materials
N,N,N′,N′-tetramethylazodicarboxamide (diamide), DTT, ethylenediaminetetraacetic acid (EDTA) and N-ethylmaleimide (NEM) were purchased from Sigma-Aldrich; diethylenetriaminepentaacetic acid (DTPA) was from Carl Roth; isopropyl-β-D-1-thiogalactopyranoside was from Serva; H2O2 was from VWR and desthiobiotin was from IBA Lifesciences. PCR primers were purchased from Metabion. All restriction enzymes, DNA polymerase and T4 DNA ligase were purchased from New England Biolabs.
Cloning and mutagenesis
The detailed information on the primers/constructs is presented in Supplementary Table 6.
TSA1 was PCR-amplified (Phusion HF) from pET15b/His-TSA1 and subcloned into pET45b/Strep (KpnI/AvrII), then into pCOLADuet MCS2 (NdeI/XhoI). TSA2 from pET15b/His6-TSA2 was inserted into pCOLADuet MCS1 (NcoI/BamHI) to yield pCOLADuet/His6-TSA2/Strep-TSA1. TSA1ΔCR (resolving Cys mutant) was generated by site-directed mutagenesis and cloned analogously. An EPEA tag was introduced into TSA2 to generate pCOLADuet/His6-TSA2-EPEA/Strep-TSA1 and pET15b/His6-TSA2-EPEA. All plasmids were verified by Sanger sequencing.
Protein expression and purification
Homo-oligomers Strep-TSA1 (pET45b plasmid), His6-TSA1 (pET15b plasmid), His6-TSA2 (pET15b plasmid) and His6-TSA2-EPEA (pET15b plasmid), as well as hetero-oligomers Strep-TSA1–His6-TSA2 (pCOLADuet-1 plasmid) and Strep-TSA1–His6-Tsa2-EPEA (pCOLADuet-1 plasmid), were expressed in E. coli strain SHuffle T7 Express in Luria Broth (LB) and induced with 0.5 mM IPTG. Eluted proteins were buffer-exchanged, assessed by SDS–PAGE, Clear/Blue Native gel and WB, and quantified at A280. Extinction coefficients: Strep-Tsa1 29,575 M−1 cm−1, His6-Tsa2-EPEA 24,075 M−1 cm−1, hetero-oligomer 26,825 M−1 cm−1, Nbsyn220 27,180 M−1 cm−1 and reduced Trx 9,970 M−1 cm−1—were calculated in the EXPASY webserver based on the sequence.
Tsa1 and Tsa2-EPEA homo-oligomers
pET45b plasmid, containing Strep-TSA1, and pET15b, containing His6-TSA2-EPEA, were independently expressed in E. coli strain SHuffle T7 Express in LB (100 μg ml−1 ampicillin) at 30 °C. Cells were induced with 0.5 mM IPTG, grown overnight at 30 °C and collected by centrifugation. Cells were lysed in 100 mM HEPES/NaOH pH 7.9, 300 mM NaCl, 20 mM MgCl2, 1 μg ml−1 DNaseI, 50 μg ml−1 leupeptin and 0.1 mg ml−1 AEBSF by sonication at 4 °C, and centrifuged at 39,846g for 25 min at 4 °C.
For His6-Tsa2-EPEA, supernatant was incubated with previously equilibrated Ni2+-Sepharose High Performance beads for 1 h at 4 °C in equilibration buffer (100 mM HEPES/NaOH pH 7.9, and 300 mM NaCl). Proteins were eluted using a stepwise gradient of 100 mM HEPES/NaOH pH 7.9, 300 mM NaCl and 1 M imidazole. For Strep-Tsa1, supernatant was loaded onto a StrepTactin column (IBA Lifesciences) equilibrated with 100 mM Tris–HCl, pH 8, 150 mM NaCl and 1 mM EDTA, and then eluted with the same buffer with 1 mM desthiobiotin in a one-step gradient. Fractions containing His-Tsa2-EPEA or Strep-Tsa1 were dialyzed against 100 mM Tris–HCl, pH 8, 150 mM NaCl and 1 mM EDTA, and stored at −80 °C.
Tsa1–Tsa2-EPEA hetero-oligomer
Strep-TSA1 and His6-TSA2-EPEA were expressed in pCOLADuet-1 plasmid and purified by tandem chromatography (Ni→Strep). Fractions from the Ni-NTA containing Strep-Tsa1 and His6-Tsa2-EPEA were dialyzed overnight at 4 °C against 100 mM Tris–HCl, pH 8, 150 mM NaCl and 1 mM EDTA with two buffer changes. The dialyzed sample was loaded onto a StrepTactin column. Fractions containing the hetero-oligomer were dialyzed against 100 mM Tris–HCl, pH 8, 150 mM NaCl and 1 mM EDTA, and stored at −80 °C.
Homo-oligomers and hetero-oligomers for kinetics
Cells were induced with 0.5 mM IPTG, grown for 4 h at 37 °C, and collected by centrifugation. Cell pellets were resuspended in 100 mM NaxHyPO4, pH 8.0, 20 mM imidazole, 300 mM NaCl at 4 °C and stirred on ice for 45 min with DNaseI and 10 mg l−1 lysozyme before sonication and clarified by centrifugation. Supernatants containing His-tagged Tsa1 or Tsa2 were loaded onto Ni-NTA agarose columns. The columns were eluted with 100 mM NaxHyPO4, pH 8.0, 200 mM imidazole and 300 mM NaCl at 4 °C. Supernatants containing Strep-tagged Tsa1 were loaded onto StrepTactin Superflow columns. The columns were eluted with 100 mM Tris–HCl, pH 8.0, 150 mM NaCl, 1 mM EDTA and 1 mM desthiobiotin at 4 °C. For the tandem purification using pCOLADuet constructs, supernatants were first purified by Ni-NTA and fractions were separately purified by StrepTactin as described above. For the reverse tandem purification, supernatants were purified by StrepTactin and fractions were then purified by Ni-NTA.
Thioredoxin and thioredoxin reductase for kinetics
His-ScTrx1 was produced in E. coli strain XL1-Blue at 37 °C in LB38. Cells were induced with 0.5 mM IPTG at an optical density of 0.5, grown for 4 h at 37 °C, collected by centrifugation and purified by Ni-NTA. His-ScTrr1 was produced in E. coli strain SHuffle T7 Express in LB containing 40 µM FAD. Cells were first grown at 30 °C until an optical density of 0.8, cooled in an ice-water bath, induced with 0.1 mM IPTG, grown overnight at 16 °C, collected by centrifugation and purified by Ni-NTA.
Nanobody (Nbsyn2.20)
His6-Nbsyn2.20 was expressed and purified as described32, with a final Superdex75 step (PBS pH 7.4). Purified protein was stored at −20 °C.
Stopped-flow kinetics
Proteins were reduced with 5 mM DTT for 30 min on ice and desalted into assay buffer (100 mM NaxHyPO₄, pH 7.4, 0.1 mM DTPA at 25 °C). Oxidized enzymes were generated with equimolar H2O2 for 30 min on ice. Reactions were recorded at 25 °C on a thermostated SX-20 spectrofluorometer (Ex = 295 nm, slit width = 2 mm, total emission). For oxidation, 1 or 10 µM reduced Tsa1 and/or Tsa2 was mixed with H2O2. For reduction, 2 µM oxidized peroxiredoxins were mixed with reduced ScTrx1. Traces (≥3 technical replicates) were fit (double/triple exponential) in SigmaPlot 13.0 to obtain kobs; three biological replicates were fit (linear/hyperbolic) to derive rate constants.
Steady-state kinetics
Coupled steady-state kinetic assays with recombinant His-tagged ScTrr1, ScTrx1, Tsa1, Tsa2 or copurified Strep-Tsa1–His-Tsa2 were carried out in assay buffer (100 mM NaxHyPO4, 0.1 mM DTPA, pH 7.4 at 25 °C) at 25 °C using a thermostated Jasco V-650 UV–visual spectrophotometer. The consumption of NADPH was monitored at 340 nm (ε = 6.22 mM−1 cm−1). Stock solutions of 4 mM NADPH, 0.98 mM H2O2 and all enzymes were freshly prepared in assay buffer before each experiment. The activity of ScTrr1 in U ml−1 was determined with 100 μM NADPH and 20 µM ScTrx1. Briefly, NADPH and ScTrr1 were mixed in assay buffer, a baseline was recorded for 30 s and the ScTrxR assay was started by the addition of ScTrx1. Peroxidase assays were optimized for ΔAbs/min values of 0.02‒0.2 and contained 150 μM NADPH, 1 µM ScTrr1 (corresponding to 0.4 mU ml⁻1), 5, 10 or 15 µM ScTrx1, 0.5‒100 µM H2O2, and 50 nM Tsa1, 10 nM Tsa2 or 10 nM Strep-Tsa1–His-Tsa2. After a baseline was recorded for 30 s, peroxidase assays were started by the simultaneous addition of peroxide and peroxidase. For the determination of apparent kcat and Km values, initial activities were corrected by subtracting the final slope of the baseline using the Spectra Analysis program (Spectra Manager (v2), Jasco). Negative controls included the omission of each of the assay components. Controls with variable ScTrr1 concentrations confirmed that the detection system was not rate-limiting at all substrate concentrations tested. Kinetic data from triplicate measurements from independent protein purifications were analyzed according to Michaelis–Menten theory and by linear regression according to Lineweaver–Burk, Eadie–Hofstee and Hanes theory in SigmaPlot (v11.0, Systat) to identify outliers and deviations from Michaelis–Menten kinetics.
Western blotting
Samples (±reducing/nonreducing) were denatured in Laemmli buffer (95 °C, 5 min), separated by SDS–PAGE or clear native PAGE, transferred to PVDF, stained and probed with anti-His, anti-EPEA or anti-Strep, as well as secondary antibody. For tandem-purified His6-Tsa2/Strep-Tsa1ΔCR, 10 mM NEM (1 h, ice) prevented artificial disulfides. Band intensities were analyzed or quantified in ImageJ against calibrated standards.
BLI
For the BLI assay on Octet R8 system (Sartorius), biotinylated ligands (Strep-Tsa1 and His6-Tsa2-EPEA, homo-oligomer and hetero-oligomer, positive-control anti-EPEA nanobody, negative-control BSA) were loaded on Streptavidin (SA) Biosensors (10 µg ml−1, 100 s, 25 °C). The concentration of Nbsyn2.20 (analyte) was fixed (50 nM) in 10 mM HEPES/NaOH pH 8, 137 mM NaCl and 2 mM KCl with 1% BSA/0.05% Tween 20. Association and dissociation phases were recorded (600 s each at 25 °C). Data were reference-subtracted, filtered and fit (1:1 local) to extract kinetics (n = 3).
Nanodifferential scanning fluorimetry
Dialyzed proteins (5 µM) in 10 mM HEPES/NaOH pH 8, 137 mM NaCl and 2 mM KCl were heated to 100 °C (2 °C min−1). Fluorescence ratio was measured at 350/330 nm in a Prometheus spectrophotometer (NanoTemper) and the inflection point values gave Tm (n = 3).
CD
Proteins were buffer-exchanged into 10 mM sodium phosphate pH 8, 140 mM NaF, diluted to 0.35–0.40 mg ml−1 and measured from 190 to 260 nm (1-mm path, 50 nm min−1, 1-nm bandwidth) in a BioLogic MOS-500 CD spectropolarimeter (BioLogic). Five spectra were averaged at temperatures selected from nanoDSF inflection points.
Mass photometry
Dialyzed samples (5 µM) were incubated 15 min at 20 °C, 30 °C, 40 °C or 45 °C, diluted 25× into 10 mM HEPES/NaOH pH 8, 137 mM NaCl and 2 mM KCl, and measured on a Refeyn OneMP (6,000 frames, 60 s). Contrast-to-mass was calibrated (MassFerence P1, 88–344 kDa). Counts were binned and plotted versus mass; mean molecular weight ± s.d. were calculated from triplicates. Relative abundance (%) of LMW species and decameric oligomers was calculated.
Negative-stain EM
Formvar/Carbon 400 Mesh, Cu grids were glow discharged at 4–5 mA and 0.3 mbar vacuum for 30 s. Three microliters of freshly diluted sample (0.02 mg ml−1) in 10 mM HEPES/NaOH pH 8, 137 mM NaCl and 2 mM KCl were incubated on the grids for 30 s, followed by staining with 2% uranyl-acetate. A total of 20 micrographs were collected on a JEOL 1400+ microscope, equipped with a LaB6 filament operating at 120 kV. Micrographs were recorded using a TVIPS F416 CCD camera using a nominal magnification of 60,000, corresponding to a magnified pixel size of 1.94 Å px−1 and a defocus range of 0 to −1.5 μm. The micrographs were processed using CryoSparc (v4.6.0). After running patch contrast transfer function estimation, particles were picked by blob-picker and extracted using a box size of 256 px. These particles were subjected to two-dimensional classification. Particle diameters were measured by TVIPS imaging software.
Hetero-oligomer–nanobody complex
Strep-Tsa1/His6-Tsa2-EPEA (5 µM) and Nbsyn2.20 (6 µM) were incubated for 20 min at room temperature in 10 mM HEPES/NaOH pH 8, 137 mM NaCl and 2 mM KCl, and analyzed by mass photometry and negative-stain EM.
MALDI-TOF MS
Samples were exchanged into 0.1% TFA; monomers were obtained by reduction with DTT (1:20 molar ratio) and rebuffering (TFA + 1 mM DTT). Furthermore, 1:1:1 mixture of protein sample, 2,5-dihydroxyacetophenone (DHAP) matrix (Bruker) and 2% vol/vol TFA were spotted in duplicate on an MTP ground steel plate. Spectra were acquired on an Ultraflextreme enhanced MALDI-TOF/TOF MS system (Bruker) in linear positive mode (range = 5,000–50,000 m/z) and processed using FlexAnalysis, with two replicates per sample. All acquisition methods were provided by the manufacturer and optimized and calibrated with an in-house calibration standard (15–122 kDa, five calibrants).
LC–MS/MS
Samples were digested using S-Trap mini spin columns according to the manufacturer’s instructions (Protifi). Intact peptides were detected in the Orbitrap Fusion Lumos at a resolution of 120,000. Peptides were selected for MS/MS using HCD setting at 30, and ion fragments were detected in the IonTrap. A data-dependent procedure that alternated between 1 MS scan followed by MS/MS scans was applied for 3 s for ions above a threshold ion count of 1.0 × 104 in the MS survey scan with 30.0 s dynamic exclusion. MS1 spectra were obtained with an automatic gain control target of 4 × 105 ions and a maximum injection time set to auto, and MS2 spectra were acquired with an automatic gain control target of 1 × 104 ions and a maximum injection set to auto. Oxidation of methionine was set as a variable modification, and the thiomethylation of cysteine was fixed. Trypsin specificity with semispecific cleavage was applied, allowing up to two missed cleavages.
Yeast strains and induction
The detailed information on strains is presented in Supplementary Table 7. The TSA1::ROGFP2-TSA1, TSA1::ROGFP2-TSA1 Δtsa2, TSA2::ROGFP2-TSA2 and TSA2::ROGFP2-TSA2 Δtsa1 strains were generated by standard homologous recombination approaches63. TSA1 and TSA2 genes were first replaced with a URA3 cassette, selecting for uracil auxotrophy on Hartwell’s Complete (HC) agar plates lacking uracil. The URA3 cassette was replaced by ROGFP2-TSA1 or ROGFP2-TSA2 with selection on HC plates containing 0.1% wt/vol 5-fluoroorotic acid (5-FOA). Plates were incubated at 30 °C for 48 h. Colonies were picked and screened by PCR.
Yeast peroxiredoxin hetero-oligomer induction
Yeast strains were grown at 30 °C in HC medium for 24 h, diluted to an D600 = 1 in fresh medium and grown for 1 h. Cultures were treated for the indicated time points with 1 mM H2O2. At these time points, 25 D600 units were collected by centrifugation at 800g for 3 min at room temperature. Cells were resuspended in 50 mM Tris–HCl, pH 7.7, 50 mM NaCl, 10% (vol/vol) glycerol, 20 mM NEM, 100 µM DTPA, 1× protease inhibitor cocktail and lysed by glass-bead homogenization. Lysates were cleared by centrifugation at 10,000g for 1 min at 5 °C. Protein concentration was determined by Bradford assays and 20 µg protein was loaded onto a 3–12% Clear-Native gel. Gels were imaged for GFP fluorescence. Cell lysates were also analyzed for total GFP fluorescence using a BMG Labtech CLARIOstar plate-reader.
roGFP2 activity assays (yeast)
BY4742 Δtsa1Δtsa2 cells were transformed with p415TEF and p416TEF plasmids for the expression of roGFP2-peroxiredoxin fusion constructs and unfused peroxiredoxin variants, respectively (Supplementary Table 7). Cells were grown to late-logarithmic phase (D600 = 3–4) in HC medium lacking the appropriate amino acids for plasmid selection. Cells were collected at room temperature and resuspended to a final concentration of 7.5 D600 U ml−1. Cells were transferred to a flat-bottomed 96-well imaging plate (BD Falcon, 353219), with 200 µl cell suspension per well. Fully oxidized and fully reduced controls were established by the addition of 20 mM diamide and 100 mM DTT, respectively64. Fluorescence was monitored in a BMG Labtech CLARIOstar plate-reader at 400 nm and 488 nm, and emission at 510 nm. The experiment was initiated by H2O2 at the indicated concentration. Degree of oxidation (OxD) roGFP2 was calculated according to equation (1).
Mammalian PRDX1/PRDX2 hetero-oligomers
HEK293 FLP-In/T-REx WT, PRDX1 KO and PRDX2 KO cells were generated previously in Jan Riemer’s lab43. Cell lysates (20 mM Tris–HCl, pH 9.2, 20 mM NaCl) were fractionated on a HiTrap Q column with a 0–250 mM NaCl gradient. Fractions (0.5 ml) were trichloroacetic acid-precipitated, washed with acetone, resolubilized in reducing Laemmli buffer and analyzed by SDS–PAGE/WB.
Bioinformatics
Peroxiredoxin candidates were identified using iterative BLASTp (NCBI; v2.16.0) with BLOSUM45 on the nonredundant protein database (GenPept, Swiss-Prot, PIR, PDF, PDB, RefSeq; 9 Jan 2025), restricted to genomes at assembly level ‘chromosome’ or ‘complete’. New queries were generated from top-scoring hits in phylogenetic adjacent eukaryotic taxa to broaden and ensure coverage. Sequences containing characteristic peroxiredoxin PFAM domains (PF00578, PF08534, PF10417), identified through HMMER, were retained, and highly similar entries (>99% identity) within the same species were collapsed. Subcellular localization was predicted using DeepLoc 2.1 with default parameters.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All experimental data generated in this study are presented within the main text and Supplementary Information. Raw data relating to LC–MS experiments have been deposited in the PRIDE database, accession PXD060819, and can be accessed with the following login: reviewer_pxd060819@ebi.ac.uk and password: CK9xDcVBX9pU. Source data for all main and supplementary figures are provided. Source data are provided with this paper.
References
Wood, Z. A., Poole, L. B. & Karplus, P. A. Peroxiredoxin evolution and the regulation of hydrogen peroxide signaling. Science 300, 650–653 (2003).
Dietz, K. J. Plant peroxiredoxins. Annu. Rev. Plant Biol. 54, 93–107 (2003).
Wood, Z. A., Schroder, E., Robin Harris, J. & Poole, L. B. Structure, mechanism and regulation of peroxiredoxins. Trends Biochem. Sci. 28, 32–40 (2003).
Stocker, S., van Laer, K., Mijuskovic, A. & Dick, T. P. The conundrum of hydrogen peroxide signaling and the emerging role of peroxiredoxins as redox relay hubs. Antioxid. Redox Signal. 28, 558–573 (2018).
Rhee, S. G. & Kil, I. S. Multiple functions and regulation of mammalian peroxiredoxins. Annu. Rev. Biochem. 86, 749–775 (2017).
Wood, Z. A., Poole, L. B., Hantgan, R. R. & Karplus, P. A. Dimers to doughnuts: redox-sensitive oligomerization of 2-cysteine peroxiredoxins. Biochemistry 41, 5493–5504 (2002).
Nelson, K. J. et al. Analysis of the peroxiredoxin family: using active-site structure and sequence information for global classification and residue analysis. Proteins 79, 947–964 (2011).
Hall, A., Nelson, K., Poole, L. B. & Karplus, P. A. Structure-based insights into the catalytic power and conformational dexterity of peroxiredoxins. Antioxid. Redox Signal. 15, 795–815 (2011).
Phalen, T. J. et al. Oxidation state governs structural transitions in peroxiredoxin II that correlate with cell cycle arrest and recovery. J. Cell Biol. 175, 779–789 (2006).
Liebthal, M., Kushwah, M. S., Kukura, P. & Dietz, K. J. Single molecule mass photometry reveals the dynamic oligomerization of human and plant peroxiredoxins. iScience 24, 103258 (2021).
Barranco-Medina, S., Lazaro, J. J. & Dietz, K. J. The oligomeric conformation of peroxiredoxins links redox state to function. FEBS Lett. 583, 1809–1816 (2009).
Pastor-Flores, D., Talwar, D., Pedre, B. & Dick, T. P. Real-time monitoring of peroxiredoxin oligomerization dynamics in living cells. Proc. Natl Acad. Sci. USA 117, 16313–16323 (2020).
Angelucci, F. et al. Switching between the alternative structures and functions of a 2-Cys peroxiredoxin, by site-directed mutagenesis. J. Mol. Biol. 425, 4556–4568 (2013).
Park, J. W., Piszczek, G., Rhee, S. G. & Chock, P. B. Glutathionylation of peroxiredoxin I induces decamer to dimers dissociation with concomitant loss of chaperone activity. Biochemistry 50, 3204–3210 (2011).
Barranco-Medina, S., Kakorin, S., Lazaro, J. J. & Dietz, K. J. Thermodynamics of the dimer-decamer transition of reduced human and plant 2-Cys peroxiredoxin. Biochemistry 47, 7196–7204 (2008).
Morais, M. A. et al. How pH modulates the dimer-decamer interconversion of 2-Cys peroxiredoxins from the Prx1 subfamily. J. Biol. Chem. 290, 8582–8590 (2015).
Hofmann, B., Hecht, H. J. & Flohe, L. Peroxiredoxins. Biol. Chem. 383, 347–364 (2002).
Jeong, J. S., Kwon, S. J., Kang, S. W., Rhee, S. G. & Kim, K. Purification and characterization of a second type thioredoxin peroxidase (type II TPx) from Saccharomyces cerevisiae. Biochemistry 38, 776–783 (1999).
Chae, H. Z., Chung, S. J. & Rhee, S. G. Thioredoxin-dependent peroxide reductase from yeast. J. Biol. Chem. 269, 27670–27678 (1994).
Kim, K., Kim, I. H., Lee, K. Y., Rhee, S. G. & Stadtman, E. R. The isolation and purification of a specific ‘protector’ protein which inhibits enzyme inactivation by a thiol/Fe(III)/O2 mixed-function oxidation system. J. Biol. Chem. 263, 4704–4711 (1988).
Prosperi, M. T., Ferbus, D., Karczinski, I. & Goubin, G. A human cDNA corresponding to a gene overexpressed during cell proliferation encodes a product sharing homology with amoebic and bacterial proteins. J. Biol. Chem. 268, 11050–11056 (1993).
Chae, H. Z. et al. Cloning and sequencing of thiol-specific antioxidant from mammalian brain: alkyl hydroperoxide reductase and thiol-specific antioxidant define a large family of antioxidant enzymes. Proc. Natl Acad. Sci. USA 91, 7017–7021 (1994).
Castro, H. et al. The cytosolic hyperoxidation-sensitive and -robust Leishmania peroxiredoxins cPRX1 and cPRX2 are both dispensable for parasite infectivity. Redox Biol. 71, 103122 (2024).
Lee, W. et al. Human peroxiredoxin 1 and 2 are not duplicate proteins: the unique presence of CYS83 in Prx1 underscores the structural and functional differences between Prx1 and Prx2. J. Biol. Chem. 282, 22011–22022 (2007).
Van Dam, L. et al. The human 2-Cys peroxiredoxins form widespread, cysteine-dependent- and isoform-specific protein–protein interactions. Antioxidants 10, 627 (2021).
Stocker, S., Maurer, M., Ruppert, T. & Dick, T. P. A role for 2-Cys peroxiredoxins in facilitating cytosolic protein thiol oxidation. Nat. Chem. Biol. 14, 148–155 (2018).
Tairum, C. A. et al. Catalytic Thr or Ser residue modulates structural switches in 2-Cys peroxiredoxin by distinct mechanisms. Sci. Rep. 6, 33133 (2016).
Dalla Rizza, J., Randall, L. M., Santos, J., Ferrer-Sueta, G. & Denicola, A. Differential parameters between cytosolic 2-Cys peroxiredoxins, PRDX1 and PRDX2. Protein Sci. 28, 191–201 (2019).
Morgan, B. et al. Real-time monitoring of basal H2O2 levels with peroxiredoxin-based probes. Nat. Chem. Biol. 12, 437–443 (2016).
Roma, L. P., Deponte, M., Riemer, J. & Morgan, B. Mechanisms and applications of redox-sensitive green fluorescent protein-based hydrogen peroxide probes. Antioxid. Redox Signal. 29, 552–568 (2018).
Villar, S. F., Ferrer-Sueta, G. & Denicola, A. Different oligomerization dynamics of reduced and oxidized human peroxiredoxin 1 and 2. Biochem. Biophys. Res. Commun. 750, 151392 (2025).
De Genst, E. J. et al. Structure and properties of a complex of α-synuclein and a single-domain camelid antibody. J. Mol. Biol. 402, 326–343 (2010).
Pardon, E., Steyaert, J. & Wyns, L. Epitope tag for affinity-based applications. US patent US20130115635A1 (2013); https://patents.google.com/patent/US20130115635A1/en
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Hong, S. K., Cha, M. K., Choi, Y. S., Kim, W. C. & Kim, I. H. Msn2p/Msn4p act as a key transcriptional activator of yeast cytoplasmic thiol peroxidase II. J. Biol. Chem. 277, 12109–12117 (2002).
Wong, C. M., Zhou, Y., Ng, R. W., Kung Hf, H. F. & Jin, D. Y. Cooperation of yeast peroxiredoxins Tsa1p and Tsa2p in the cellular defense against oxidative and nitrosative stress. J. Biol. Chem. 277, 5385–5394 (2002).
Rhee, S. G. & Woo, H. A. Multiple functions of 2-Cys peroxiredoxins, I and II, and their regulations via post-translational modifications. Free Radic. Biol. Med. 152, 107–115 (2020).
Zimmermann, J. et al. Tsa1 is the dominant peroxide scavenger and a source of H2O2-dependent GSSG production in yeast. Free Radic. Biol. Med. 226, 408–420 (2025).
Kriznik, A. et al. Dynamics of a key conformational transition in the mechanism of peroxiredoxin sulfinylation. ACS Catal. 10, 3326–3339 (2020).
Mathieu, J. et al. The dual role of active site hydroxylated residue in peroxiredoxin sulfinylation catalysis. Antioxid. Redox Signal. 43, 1–13 (2025).
Gutscher, M. et al. Proximity-based protein thiol oxidation by H2O2-scavenging peroxidases. J. Biol. Chem. 284, 31532–31540 (2009).
Schwarzlander, M., Dick, T. P., Meyer, A. J. & Morgan, B. Dissecting redox biology using fluorescent protein sensors. Antioxid. Redox Signal. 24, 680–712 (2016).
Hoehne, M. N. et al. Spatial and temporal control of mitochondrial H2O2 release in intact human cells. EMBO J. 41, e109169 (2022).
Baier, M. & Dietz, K. J. The plant 2-Cys peroxiredoxin BAS1 is a nuclear-encoded chloroplast protein: its expressional regulation, phylogenetic origin, and implications for its specific physiological function in plants. Plant J. 12, 179–190 (1997).
Horling, F. Divergent light-, ascorbate-, and oxidative stress-dependent regulation of expression of the peroxiredoxin gene family in Arabidopsis. Plant Physiol. 131, 317–325 (2003).
Perkins, A., Nelson, K. J., Parsonage, D., Poole, L. B. & Karplus, P. A. Peroxiredoxins: guardians against oxidative stress and modulators of peroxide signaling. Trends Biochem. Sci. 40, 435–445 (2015).
Bolduc, J. et al. Peroxiredoxins wear many hats: factors that fashion their peroxide sensing personalities. Redox Biol. 42, 101959 (2021).
Teixeira, F. et al. Mitochondrial peroxiredoxin functions as crucial chaperone reservoir in Leishmania infantum. Proc. Natl Acad. Sci. USA 112, E616–E624 (2015).
Teixeira, F. et al. Chaperone activation and client binding of a 2-cysteine peroxiredoxin. Nat. Commun. 10, 659 (2019).
Troussicot, L., Burmann, B. M. & Molin, M. Structural determinants of multimerization and dissociation in 2-Cys peroxiredoxin chaperone function. Structure 29, 640–654 (2021).
Hanzen, S. et al. Lifespan control by redox-dependent recruitment of chaperones to misfolded proteins. Cell 166, 140–151 (2016).
MacDiarmid, C. W. et al. Peroxiredoxin chaperone activity is critical for protein homeostasis in zinc-deficient yeast. J. Biol. Chem. 288, 31313–31327 (2013).
Jang, H. H. et al. Two enzymes in one; two yeast peroxiredoxins display oxidative stress-dependent switching from a peroxidase to a molecular chaperone function. Cell 117, 625–635 (2004).
Meissner, U., Schroder, E., Scheffler, D., Martin, A. G. & Harris, J. R. Formation, TEM study and 3D reconstruction of the human erythrocyte peroxiredoxin-2 dodecahedral higher-order assembly. Micron 38, 29–39 (2007).
Koo, K. H. et al. Regulation of thioredoxin peroxidase activity by C-terminal truncation. Arch. Biochem. Biophys. 397, 312–318 (2002).
Randall, L. et al. Structural changes upon peroxynitrite-mediated nitration of peroxiredoxin 2; nitrated Prx2 resembles its disulfide-oxidized form. Arch. Biochem. Biophys. 590, 101–108 (2016).
Engelman, R. et al. Multilevel regulation of 2-Cys peroxiredoxin reaction cycle by S-nitrosylation. J. Biol. Chem. 288, 11312–11324 (2013).
Jang, H. H. et al. Phosphorylation and concomitant structural changes in human 2-Cys peroxiredoxin isotype I differentially regulate its peroxidase and molecular chaperone functions. FEBS Lett. 580, 351–355 (2006).
Sobotta, M. C. et al. Peroxiredoxin-2 and STAT3 form a redox relay for H2O2 signaling. Nat. Chem. Biol. 11, 64–70 (2015).
Talwar, D., Messens, J. & Dick, T. P. A role for annexin A2 in scaffolding the peroxiredoxin 2-STAT3 redox relay complex. Nat. Commun. 11, 4512 (2020).
Jarvis, R. M., Hughes, S. M. & Ledgerwood, E. C. Peroxiredoxin 1 functions as a signal peroxidase to receive, transduce, and transmit peroxide signals in mammalian cells. Free Radic. Biol. Med. 53, 1522–1530 (2012).
Lang, L. et al. Substrate promiscuity and hyperoxidation susceptibility as potential driving forces for the co-evolution of Prx5-type and Prx6-type 1-Cys peroxiredoxin mechanisms. ACS Catal. 13, 3627–3643 (2023).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Morgan, B., Sobotta, M. C. & Dick, T. P. Measuring E(GSH) and H2O2 with roGFP2-based redox probes. Free Radic. Biol. Med. 51, 1943–1951 (2011).
Acknowledgements
B.M. and M.D. gratefully acknowledge funding from the Deutsche Forschungsgemeinschaft (German Research Foundation; grants MO 2774/6-1 project 505680640, MO 2774/7-1 and DE 1431/19-1 project 508372800). J.M. was supported by a VIB grant. J.M.P. was supported by an FWO fellowship (1193524N). The Deutsche Forschungsgemeinschaft funds research in the Laboratory of J.R. through the grants RI2150/5-1 project 435235019, RTG2550/2 project 411422114, SPP2453 project 541742459, CRC1218 project 269925409 and CRC1678 project 520471345. A.S. acknowledges funding from COMPETE 2020—Operational Programme for Competitiveness and Internationalisation—and from Portuguese national funds through Fundação para a Ciência e a Tecnologia (FCT; projects UIDB/04539/2020, UIDP/04539/2020, LA/P/0058/2020, UIDB/00313/2020 and UIDP/00313/2020). We thank E. Pardon (Steyaert Lab, VIB-VUB Center for Structural Biology) for providing the Nbsyn2.20 nanobody. We thank J. Dechenne (Louvain Drug Research Institute, The Medicinal Chemistry Group, UCL) for his help with nanoDSF experiments. We also thank the BECM VIB–VUB cryo-EM imaging facility in Brussels and M. Fislage for the support during negative staining EM imaging and processing. The BLI work was supported by a research infrastructure grant from the VUB (OZR4292). A.M.T. acknowledges support from national funds through FCT, I.P. (project UIDB/04293/2020).
Funding
Open access funding provided by Universität des Saarlandes.
Author information
Authors and Affiliations
Contributions
J.Z. performed initial yeast construct construction, yeast hetero-oligomer induction assays and native-PAGE analyses. L. Lang performed, evaluated and supervised the stopped-flow kinetic measurements. J.M.P. expressed and purified Tsa1 and Tsa2 recombinant proteins, performed nanodifferential scanning fluorimetry and CD measurements, mass photometry analyses, negative-stain EM imaging, and WBs and native-PAGE analyses, and wrote the paper. M.R. cloned and (co)purified recombinant Tsa1, Tsa2 and Tsa1–Tsa2, and performed and evaluated the stopped-flow kinetic measurements. K.W. expressed and purified Tsa1, Tsa2 and Nbsyn2.20 recombinant proteins, and performed BLI, nanodifferential scanning fluorimetry and CD experiments, and WBs and native-PAGE analyses. D.S. performed the ion-exchange and WB analyses on HEK293T cell lysates. L. Leiskau (co)purified recombinant Tsa1, Tsa2 and Tsa1–Tsa2, and performed and evaluated the steady-state kinetic measurements. E.A. performed WBs to assess hyperoxidation in yeast induction assays. C.L. performed the bioinformatics analyses. S.J. performed MALDI-TOF MS and LC–MS/MS measurements and data analysis. D.V. performed LC–MS/MS measurements and data analysis. S.L. optimized roGFP2 assays to test for enzymatic activity of peroxiredoxin pairs heterologously expressed in yeast. F.H. designed and cloned E. coli and yeast expression constructs, and optimized conditions for recombinant protein purification. S.T. performed HADDOCK-based analyses of peroxiredoxin heterodimers. H.C. established the roGFP assay to monitor Leishmania peroxiredoxin activity and performed initial experiments. V.H. supervised in silico HADDOCK-based analyses. A.M.T. and J.M.H. supervised experiments and analyzed data. A.S. performed the mathematical determination for number of possible distinct hetero-oligomers. T.M. designed and supervised the bioinformatics analyses. J.R. conceived and supervised the ion-exchange experiments in HEK293T cell lysates and assisted with data analysis. M.D. and B.M. conceived and developed the concept of peroxiredoxin hetero-oligomerization. J.M., M.D. and B.M. designed and supervised experiments, assisted with data analysis, and wrote and revised the paper. All authors read and approved the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Gerardo Ferrer-Sueta and the other, anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The oligomeric state of peroxiredoxins is closely linked to their functions and is affected by numerous factors.
Prx1/AhpC-type peroxiredoxins (Prx) exist in a dynamic equilibrium between dimeric and decameric forms of the (α2)₅-type depicted as a ribbon diagram (top left), space-filling model (top middle), and space-filling side-view (top right). The oligomeric assembly is formed through A-type (dimer–dimer) and B-type (intradimer) interfaces. Prx dimers (bottom left) transition between reduced (Prx(SH)2) and oxidized (Prx(SS)) forms via reaction with H2O2, followed by reduction facilitated by thioredoxins (Trx) or glutaredoxins (Grx). Hyperoxidation leads to Prx(SO2) formation, reversible via sulfiredoxin (Srx) in an ATP-dependent reaction. The function of Prx1–AhpC-type peroxiredoxins is intricately coupled to their oligomeric state and includes peroxide detoxification, redox signaling, and chaperone activity.
Extended Data Fig. 2 Enumeration of the distinct decamers that can be formed from two types of monomers.
a, Distinct decamers as a function of the stoichiometric ratio between monomer types. b, Number of unique hetero-decamer configurations as a function of the number of possible states each monomer type can adopt. The horizontal dashed line indicates the estimated number of peroxiredoxin decamers in a typical mammalian cell. Please note the logarithmic scale on the vertical axis. Calculations are explained in the Supplementary Note.
Extended Data Fig. 3 Coupled steady-state peroxidase assays for His6-Tsa1, His6-Tsa2, or copurified Strep-Tsa1 and His6-Tsa2.
a, Controls to estimate the component-dependent background-consumption of NADPH. b, Control to exclude that the TrxR detection system was rate-limiting. c‒e, Direct Michaelis–Menten (top) and linearized Lineweaver–Burk (bottom) plots at variable H2O2 concentrations for 50 nM Tsa1, 10 nM Tsa2, or 10 nM copurified Tsa1 and Tsa2, respectively. Squares were omitted for regression analysis. All data were obtained from three (Tsa1) or four (Tsa2 and copurified Tsa1 and Tsa2) independent biological protein purifications and measurements. f, Secondary plot for the Trx1-dependent rate constants from c‒e. The kcatapp values for the oxidation by H2O2 and the rate constants for the Trx1-dependent reduction of oxidized Tsa1, Tsa2, or copurified Tsa1 and Tsa2 are listed in Supplementary Tables 4 and 5 and were estimated from the slopes in c‒e and f, respectively.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–15, supporting data for Figs. 3, 10, 11, 13 and 14, Tables 1–7, and Note.
Source data
Source Data Fig. 1 (download PDF )
Unprocessed gels.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed gels.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed gels.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download PDF )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Unprocessed gels.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zimmermann, J., Lang, L., Malo Pueyo, J. et al. Hetero-oligomerization drives structural plasticity of eukaryotic peroxiredoxins. Nat Chem Biol 22, 580–592 (2026). https://doi.org/10.1038/s41589-026-02157-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-026-02157-6