Abstract
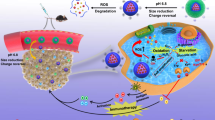
Reactive oxygen species (ROS) dynamics exhibits rhythmic oscillations in cancer cells but how this rhythm influences tumorigenesis and therapeutic responses remains unclear. Here we found coexistence of ROS rhythmicity and rhythm loss in tumor samples. Under low-ROS conditions, indoleamine 2,3-dioxygenase 1 (IDO1), an immune-checkpoint molecule, binds to KEAP1 for proteasomal degradation in the nucleus. In contrast, elevated ROS levels drive IDO1 translocation into the cytosol, where it binds mitochondria-released heme to form an active holoenzyme. This holoenzyme catalyzes tryptophan to kynurenine that allosterically activates glucose-6-phosphate dehydrogenase, enhancing NADPH production and promoting ROS clearance. However, in hypoxic tumor microenvironments, ROS rhythmicity is lost. Compensating for this, hypoxic tumor cells mobilize the sulfenylated aryl hydrocarbon receptor (AhR)-mediated glycogenolysis pathway to manage disordered ROS accumulation, maintaining elevated ROS levels that favor tumor growth. Dual inhibition of IDO1 and AhR significantly prolongs survival of NSG mice, highlighting enforced disruption of ROS rhythm as a common therapeutic strategy.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper or Supplementary Information. All data supporting the findings of this study are available from the corresponding author on request. Source data are provided with this paper.
References
Vercellino, I. & Sazanov, L. A. The assembly, regulation and function of the mitochondrial respiratory chain. Nat. Rev. Mol. Cell Biol. 23, 141–161 (2022).
Bennett, C. F., Latorre-Muro, P. & Puigserver, P. Mechanisms of mitochondrial respiratory adaptation. Nat. Rev. Mol. Cell Biol. 23, 817–835 (2022).
Shadel, G. S. & Horvath, T. L. Mitochondrial ROS signaling in organismal homeostasis. Cell 163, 560–569 (2015).
Sies, H. & Jones, D. P. Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat. Rev. Mol. Cell Biol. 21, 363–383 (2020).
Bock, F. J. & Tait, S. W. G. Mitochondria as multifaceted regulators of cell death. Nat. Rev. Mol. Cell Biol. 21, 85–100 (2020).
Halliwell, B. Understanding mechanisms of antioxidant action in health and disease. Nat. Rev. Mol. Cell Biol. 25, 13–33 (2024).
Sies, H. et al. Defining roles of specific reactive oxygen species (ROS) in cell biology and physiology. Nat. Rev. Mol. Cell Biol. 23, 499–515 (2022).
O’Neill, J. S. & Reddy, A. B. Circadian clocks in human red blood cells. Nature 469, 498–503 (2011).
Nordberg, J. & Arnér, E. S. Reactive oxygen species, antioxidants, and the mammalian thioredoxin system. Free Radic. Biol. Med. 31, 1287–1312 (2001).
Chandel, N. S. NADPH—the forgotten reducing equivalent. Cold Spring Harb. Perspect. Biol. 13, a040550 (2021).
TeSlaa, T., Ralser, M., Fan, J. & Rabinowitz, J. D. The pentose phosphate pathway in health and disease. Nat. Metab. 5, 1275–1289 (2023).
Meireson, A., Devos, M. & Brochez, L. I. D. O. Expression in cancer: different compartment, different functionality? Front. Immunol. 11, 531491 (2020).
van den Eynde, B. J., van Baren, N. & Baurain, J.-F. Is there a clinical future for IDO1 inhibitors after the failure of epacadostat in melanoma? Annu. Rev. Cancer Biol. 4, 241–256 (2020).
Tang, K., Wu, Y.-H., Song, Y. & Yu, B. Indoleamine 2,3-dioxygenase 1 (IDO1) inhibitors in clinical trials for cancer immunotherapy. J. Hematol. Oncol. 14, 68 (2021).
Shadboorestan, A., Koual, M., Dairou, J. & Coumoul, X. The role of the kynurenine/AhR pathway in diseases related to metabolism and cancer. Int. J. Tryptophan Res. 16, 11786469231185102 (2023).
Zhou, N. et al. Aryl hydrocarbon receptor sulfenylation promotes glycogenolysis and rescues cancer chemoresistance. J. Clin. Invest. 133, e170753 (2023).
Ermakova, Y. G. et al. Red fluorescent genetically encoded indicator for intracellular hydrogen peroxide. Nat. Commun. 5, 5222 (2014).
Rey, G. & Reddy, A. B. Interplay between cellular redox oscillations and circadian clocks. Diabetes Obes. Metab. 17, 55–64 (2015).
Dunlap, J. C. Molecular bases for circadian clocks. Cell 96, 271–290 (1999).
Edgar, R. S. et al. Peroxiredoxins are conserved markers of circadian rhythms. Nature 485, 459–464 (2012).
Vomund, S., Schäfer, A., Parnham, M. J., Brüne, B. & von Knethen, A. Nrf2, the master regulator of anti-oxidative responses. Int. J. Mol. Sci. 18, 2772 (2017).
Ma, R. et al. A Pck1-directed glycogen metabolic program regulates formation and maintenance of memory CD8+ T cells. Nat. Cell Biol. 20, 21–27 (2018).
Zhang, H. et al. Ketogenesis-generated β-hydroxybutyrate is an epigenetic regulator of CD8+ T-cell memory development. Nat. Cell Biol. 22, 18–25 (2020).
Ma, J. et al. Glycogen metabolism regulates macrophage-mediated acute inflammatory responses. Nat. Commun. 11, 1769 (2020).
Larigot, L. et al. Aryl hydrocarbon receptor and its diverse ligands and functions: an exposome receptor. Annu. Rev. Pharmacol. Toxicol. 62, 383–404 (2022).
Liu, Y. et al. Tumor-repopulating cells induce PD-1 expression in CD8+ T cells by transferring kynurenine and AhR activation. Cancer Cell 33, 480–494 (2018).
Rashid, F. et al. Mechanomemory in protein diffusivity of chromatin and nucleoplasm after force cessation. Proc. Natl Acad. Sci. USA 120, e2221432120 (2023).
Manford, A. G. et al. A cellular mechanism to detect and alleviate reductive stress. Cell 183, 46–61 (2020).
Ma, Q. Role of Nrf2 in oxidative stress and toxicity. Annu. Rev. Pharmacol. Toxicol. 53, 401–426 (2013).
Guo, X. Localized proteasomal degradation: from the nucleus to cell periphery. Biomolecules 12, 229 (2022).
Baird, L. & Yamamoto, M. The molecular mechanisms regulating the KEAP1–Nrf2 pathway. Mol. Cell. Biol. 40, e00099-20 (2020).
Azmi, A. S., Uddin, M. H. & Mohammad, R. M. The nuclear export protein XPO1—from biology to targeted therapy. Nat. Rev. Clin. Oncol. 18, 152–169 (2021).
Nelp, M. T. et al. Immune-modulating enzyme indoleamine 2,3-dioxygenase is effectively inhibited by targeting its apo-form. Proc. Natl Acad. Sci. USA 115, 3249–3254 (2018).
Fleming, M. D. & Hamza, I. Mitochondrial heme: an exit strategy at last. J. Clin. Invest. 122, 4328–4330 (2012).
Chiabrando, D. et al. The mitochondrial heme exporter FLVCR1b mediates erythroid differentiation. J. Clin. Invest. 122, 4569–4579 (2012).
Zhou, L. et al. Tumor cell-released kynurenine biases MEP differentiation into megakaryocytes in individuals with cancer by activating AhR–RUNX1. Nat. Immunol. 24, 2042–2052 (2023).
Newmeyer, D. D. & Ferguson-Miller, S. Mitochondria: releasing power for life and unleashing the machineries of death. Cell 112, 481–490 (2003).
Balaban, R. S., Nemoto, S. & Finkel, T. Mitochondria, oxidants, and aging. Cell 120, 483–495 (2005).
D’Autréaux, B. & Toledano, M. B. ROS as signalling molecules: mechanisms that generate specificity in ROS homeostasis. Nat. Rev. Mol. Cell Biol. 8, 813–824 (2007).
Pei, J. F. et al. Diurnal oscillations of endogenous H2O2 sustained by p66Shc regulate circadian clocks. Nat. Cell Biol. 21, 1553–1564 (2019).
Hourihan, J. M., Moronetti Mazzeo, L. E., Fernandez-Cardenas, L. P. & Blackwell, T. K. Cysteine sulfenylation directs IRE-1 to activate the SKN-1/Nrf2 antioxidant response. Mol. Cell 63, 553–566 (2016).
Wu, F. et al. Acetylation-dependent coupling between G6PD activity and apoptotic signaling. Nat. Commun. 14, 6208 (2023).
Ye, Z., Yue, L., Shi, J., Shao, M. & Wu, T. Role of IDO and TDO in cancers and related diseases and the therapeutic implications. J. Cancer 10, 2771–2782 (2019).
Zangar, R. C., Davydov, D. R. & Verma, S. Mechanisms that regulate production of reactive oxygen species by cytochrome P450. Toxicol. Appl. Pharmacol. 199, 316–331 (2004).
Panfili, E. et al. The catalytic inhibitor epacadostat can affect the non-enzymatic function of IDO1. Front. Immunol. 14, 1134551 (2023).
Yu, L. et al. IDO1 inhibition promotes activation of tumor-intrinsic STAT3 pathway and induces adverse tumor-protective effects. J. Immunol. 212, 1232–1243 (2024).
Sun, F. et al. HRG-9 homologues regulate haem trafficking from haem-enriched compartments. Nature 610, 768–774 (2022).
Klomsiri, C. et al. Use of dimedone-based chemical probes for sulfenic acid detection evaluation of conditions affecting probe incorporation into redox-sensitive proteins. Methods Enzymol. 473, 77–94 (2010).
Acknowledgements
This work was supported by National Natural Science Foundation of China (82388201 and 82350111), National Key Research and Development Program of China (2022YFA1206000), Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (2021-I2M-1-021 and 2023-I2M-2-005), Noncommunicable Chronic Diseases National Science and Technology Major Project (2025ZD0552500), China Postdoctoral Science Foundation (2024M750249 and 2025T180405) and Haihe Laboratory of Cell Ecosystem Innovation Fund (22HHXBSS00009).
Author information
Authors and Affiliations
Contributions
B.H. conceptualized the project. N.Z., Z.L., X.C. and C.Z. performed the majority of the experiments and contributed to data interpretation. N.Z. and L. Zhang conducted analyses including molecular docking and molecular dynamics stimulation. D.Y. performed the MS analysis. N.Z. performed ROS rhythm testing experiments. Z.L. and X.C. carried out the BLI, protein purification and animal treatment experiments. N.Z., Z.L., X.C. and C.Z. contributed to data collection and discussion. J.C., C.Z., Z.W., D.W., Y.Z. and L. Zhou reviewed and edited the paper. J.M., K.T., H.Z. and J.L. provided technical and material support and provided scientific advice. B.H. and N.N.Z. wrote the paper. All authors approved the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Pranjal Biswas, Daqian Xu, Yi Xu and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 ROS rhythm is regulated by IDO1 in human tumor cells.
(a) The intensity of HyPerRed fluorescence of synchronized SKOV3 and HCT116 cells were detected by flow cytometry for 96 h. Time 0 was the first time point after synchronization, and cells were collected according to different time points for detection. (b, c) The intensity of HyPerRed fluorescence and GFP fluorescence of an individual synchronized MCF-7 or U2OS cells were detected using time-lapse microscopy for 48 h. Scale bar, 10 μm (b). The intensity of HyPerRed fluorescence and GFP fluorescence were recorded(c). (d) The intensity of HyPerRed fluorescence and SypHer fluorescence in synchronized MCF-7 cells at different time points were recorded. HyPerRed, Ex/Em=575/605 nm; SypHer, Ex/Em=445/510 nm. (e, g, i) The intensity of HyPerRed fluorescence of synchronized MCF-7 and U2OS cells pre-treated with 20 μM CLK8 for 12 h were detected by flow cytometry for 48 h.(e), synchronized BMAL1 knockout MCF-7 and U2OS cells (g), synchronized MCF-7 and U2OS cells transfected with si-NC or si-PRDX1 (i). (f) The expression of BMAL1 in MCF-7 and U2OS cells targeted BMAL1 by CRISPR/Cas9 was analyzed by Western blot. (h) The expression of PRDX1 in MCF-7 and U2OS cells transfected with si-NC, si-PRDX1 were analyzed by Western blot. For a-i, n = 3 biological independent experiments. JTK_CYCLE analysis was used to determine rhythmicity (a), and P < 0.05 was considered to be rhythmic.
Extended Data Fig. 2 ROS rhythm is regulated by IDO1 in human tumor cells.
(a) The expression of AHR or NRF2 in MCF-7 and U2OS cells targeted AHR and NRF2 by CRISPR/Cas9 was analyzed by Western blot. (b, e, f, i, j) The intensity of HyPerRed fluorescence of synchronized AHR or NRF2 knockout MCF-7 and U2OS cells (b), synchronized U2OS cells treated with Linrodostat (20 μM) or Epacadostat (20 μM) (e), synchronized IDO1 knockout or IDO1 overexpression-U2OS cells (f), and synchronized MCF-7 and U2OS cells transfected with si-IDO2 or si-TDO2 (i, j) were detected by flow cytometry for 48 h. (c) The IDO1 expression in IDO1 knockout (KO) or overexpression (OE) in MCF-7 or U2OS tumor cells were analyzed by Western Blot. (d) The inhibitory efficacy of the small-molecule IDO1 inhibitors (Linrodostat and Epacadostat) in MCF-7 tumor cells were measured by Indoleamine 2,3-Dioxygenase 1 Activity Assay Kit. (g) The relative mRNA level of IDO1, IDO2 and TDO2 in MCF-7, U2OS, HCT116 and SKOV3 cells was analyzed by real time qPCR. (h) The expression of IDO2 or TDO2 in MCF-7 and U2OS cells transfected with si-NC, si-IDO2 or si-TDO2, respectively, were analyzed by Western blot. For a-j, n = 3 biological independent experiments. All error bars are mean ± SD. P values were calculated by two-tailed Student’s t test (d).
Extended Data Fig. 3 Nuclear IDO1 is degraded upon low ROS.
(a) The intensity of HyPerRed fluorescence of synchronized MCF-7 and U2OS cells. (b)The relative mRNA level of IDO1 in MCF-7 and U2OS cells was analyzed by real time qPCR. (c, d) The expression of cytosolic or nuclear IDO1 in synchronized U2OS cells at different time points was analyzed by Western blot (c) and the normalized IDO1 intensity was calculated (d). (e-g) The IDO1 location of synchronized U2OS cells at different time points were observed by confocal microscope (e). Scale bar, 10 μm. The ratio of nuclear/cytosolic IDO1 fluorescence intensity (f) and the relative HyPerRed fluorescence intensity were calculated (g). (h) The expression of cytosolic or nuclear IDO1 in synchronized MCF-7 cells at different time points was analyzed by Western blot. (i) The location of IDO1 mRNA in MCF-7 and U2OS cells was analyzed by RNA-Scope and the relative IDO1 mRNA were calculated. (j) The expression of cytosolic or nuclear IDO1 in MCF-7 and U2OS cells treated with 100 ng/mL WGA or 20 μM Pitstop2 for 12 h was analyzed by Western blot. (k, l) The IDO1 location of MCF-7 and U2OS cells treated with 100 ng/mL WGA or 20 μM Pitstop2 for 12 h were observed by confocal microscope (k). Scale bar, 10 μm. The ratio of intensity of cytosolic and nuclear IDO1 was calculated (l). n = 6 technical replicates. (m) The intensity of HyPerRed fluorescence of Synchronized MCF-7 treated with 20 μM Pitstop2 for 12 h were detected by flow cytometry for 48 h. For a-j, m, n = 3 biological independent experiments. All error bars are mean ± SD. P values were calculated by two-tailed Student’s t test (i) and one-way ANOVA followed by Bonferroni’s test (l).
Extended Data Fig. 4 Nuclear IDO1 is degraded upon low ROS.
(a) The expression of Nup153 or Nup214 in MCF-7 cells transfected with si-NC, si-Nup153, si-Nup214 was analyzed by Western blot. (b) The expression of cytosolic or nuclear IDO1 in MCF-7 cells transfected with si-NC, si-Nup153, si-Nup214 was analyzed by Western blot. (c) The IDO1 location of synchronized MCF-7 cells transfected with si-NC, si-Nup153, si-Nup214 was observed by confocal microscope. Scale bar, 10 μm. The ratio of intensity of cytosolic and nuclear IDO1 was calculated. n = 6 technical replicates. (d) The intensity of HyPerRed fluorescence in synchronized MCF-7 cells transfected with si-NC, si-Nup153, si-Nup214 were analyzed. (e) The concentration of endogenous H2O2 in synchronized MCF-7 cells at different time points were analyzed. (f) The concentration of intracellular H2O2 in MCF-7 cells treated with 50 or 100 μM H2O2 were analyzed. (g, h) The fluorescence intensity of NucPE1 indicating nuclear ROS in MCF-7 cells (g) or HyPerRed fluorescence in MCF-7 cells stably expressing HyPerRed- nuclear localization sequence (NLS) (h) exposed with 100 μM H2O2 was detected. n = 6 technical replicates. For a, b, d-f, n = 3 biological independent experiments. All error bars represent mean ± SD. P values were calculated by one-way ANOVA followed by Bonferroni’s test (c, f) or two-tailed Student’s t test (g, h).
Extended Data Fig. 5 Nuclear IDO1 is degraded upon low ROS.
(a) The expression of cytosolic or nuclear IDO1 in MCF-7 cells treated with 50 or 100 μM H2O2, or combined with 5 mM NAC was analyzed by Western blot. (b) The expression of cytosolic or nuclear IDO1 in U2OS cells treated with 100 μM H2O2 or combined with pre-treated with 5 mM NAC for 12 h was analyzed by Western blot. (c) KEAP1, TRIM21 and TRIM25 in immuno-precipitations of IDO1 were analyzed by Western blot. (d, e) The expression of nuclear IDO1 in TRIM21 KO MCF-7 (d) or TRIM25 KO MCF-7 cells (e) were analyzed by Western blot. (f) The ubiquitination of nuclear IDO1 in CUL3 KO MCF-7 cells was analyzed by Western blot. (g) The expression of nuclear IDO1 in MCF-7 cells transfected with IDO151-54 mutant plasmid was analyzed by Western blot. (h) The ubiquitination of nuclear IDO1, the binding of IDO1 and KEAP1 in synchronized MCF-7 cells at different time points was analyzed by Western blot. (i) The ubiquitination of cytosolic IDO1 in synchronized MCF-7 cells at different time points was analyzed by Western blot. For a-i, n = 3 biological independent experiments.
Extended Data Fig. 6 Nuclear IDO1 is translocated to the cytosol by increased ROS.
(a) Ubiquitination of nuclear IDO1 and the binding with KEAP1 in MCF-7 cells treated with 100 μM H2O2 was analyzed by Western blot. (b) XPO1 and KEAP1 in immuno-precipitates of IDO1 in MCF-7 and U2OS cells at 0 h and 12 h was analyzed by Western blot. (c) The expression of XPO1 in MCF-7 and U2OS cells targeted XPO1 gene knockout by CRISPR/Cas9 was analyzed by Western blot. (d) The IDO1 location of XPO1 knockout MCF-7 and U2OS cells or MCF-7 and U2OS cells treated with selinexor were observed by confocal microscope. Scale bar, 10 μm. The ratio of Cyto-IDO1 /Nuc-IDO1 MFI was calculated. n = 6 technical replicates. (e) The expression of KEAP1 or XPO1 in MCF-7 cells targeted KEAP1 and XPO1 gene knockout by CRISPR/Cas9, or targeted KEAP1 or XPO1 gene expression by lentiviral infection were analyzed by Western blot. (f) The binding between purified IDO1 and XPO1 treated with H2O2 was measured by BLI. (g, h) The recombinant KEAP1 protein (g) and XPO1(h) were stained with Coomassie Brilliant Blue. For a-c, e, f, n = 3 biological independent experiments. All error bars represent mean ± SD. P values were calculated by two-tailed Student’s t test (d).
Extended Data Fig. 7 IDO1-catalyzed Kyn activates G6PD to clear ROS via NADPH.
(a, b) The relative FLVCR1b mRNA level (a) and heme concentration of IDO1 (b) in MCF-7 and U2OS cells transfected with si-FLVCR1b was analyzed. (c, d) The relative FLVCR1b mRNA level (c) and the heme concentration (d) in MCF-7 cells treated with 100 μM H2O2 or 5 mM NAC were measured. (e) The relative FLVCR1b mRNA level and the heme concentration in MCF-7 cells during ROS oscillation were measured. (f) The intensity of HyPerRed fluorescence and the concentration of NO in synchronized MCF-7 cells were measured. (g) Mapping of IDO1 mutant site (H346A). (h) The IDO1 expression in WT, IDO1 knockout (KO), or with IDO1 and IDO1H346A overexpression (OE) MCF-7 cells by lentiviral infection were by Western blot. (i) HyPerRed fluorescence of synchronized WT, IDO1 KO, IDO1 KO with IDO1-OE or with IDO1H346A -OE MCF-7 cells were detected. (j, k) Kyn concentration (j) and NADPH concentration (k) in MCF-7 and U2OS cells treated with 10 μM Linrodostat were measured. (l) ROS level in MCF-7 and U2OS cells treated with 10 μM Linrodostat were measured. (m) The relative mRNA level of G6PD in synchronized MCF-7 cells was analyzed. (n) The expression of G6PD in synchronized MCF-7 cells at different time points was analyzed by Western blot. The normalized IDO1 intensity was calculated. (o) NADPH concentration in MCF-7 and U2OS cells treated with 100 μM Kyn or 50 μM 6-AN was measured. (p, q) The expression of G6PD, ME1, ME2, ME3 and MTHFD2 in MCF-7 and U2OS cells transfected with si-NC, si-G6PD, si-MEs or si-MTHFD2 was analyzed by Western blot. (r) NADPH concentration in transfected MCF-7 cells were measured. For a-f, h-r, n = 3 biological independent experiments. All error bars represent mean ± SD. P values were calculated by one-way ANOVA followed by Bonferroni’s test (a, b, c, d, e, o, r) or Student’s t test (j, k, l). NS, no significant, på 0.05.
Extended Data Fig. 8 IDO1-catalyzed Kyn activates G6PD to clear ROS via NADPH.
(a) Molecular docking and Molecular dynamics simulation of G6PD and G6P with Kyn. (b) Recombinant G6PD proteins were stained with Coomassie Brilliant Blue. (c) MCF-7 cells cultured in 13C-glucose for 5 days and switched to 12C-glucose combined with 100 μM kyn treatment for 6 h. 13C-labeled R5P was detected by LC-MS/MS. n = 6 technical replicates. (d) The NADPH concentration in MCF-7 cells treated with 100 μM Kyn for 6 h was measured. n = 6 technical replicates. (e, f) The intensity of HyPerRed fluorescence of synchronized U2OS cells at different time points was detected by flow cytometry for 48 h, and Kyn concentration (e) and the NADPH/NADP+ level (f) were measured by chromatography and NADPH assay kit. (g, h) The concentration of NADPH/NADP+ level (g) and Kyn concentration (h) in synchronized IDO1 knockout MCF-7 cells were analyzed. (i) The intensity of HyPerRed fluorescence and the NADPH/NADP+ level of synchronized G6PD knockdown MCF-7 cells was analyzed. For e-i, n = 3 biological independent experiments. All error bars are mean ± SD. P values were calculated by two-tailed Student’s t test (c, d).
Extended Data Fig. 9 Hypoxia disrupts ROS rhythm in tumor cells of patients.
(a) HyPerRed fluorescence distribution in sorted MCF-7 and U2OS tumor cells. (b) The expression of IDO1 in sorted HyPerRedhigh IDO1-OE and HyPerRedlow Sh-IDO1-transfected MCF-7 cells were analyzed by Western blot. (c, d) HyPerRed fluorescence in synchronized IDO1-OE HyPerRedhigh and Sh-IDO1 transfected HyPerRedlow MCF-7 cells were detected. (e) The gate strategy for sorting tumor cells from transplanted MCF-7 tumor tissue. (f) The expression of IDO1 in sorted MCF-7 cells were measured by digital western blot. (g) HyPerRed fluorescence in synchronized sorted MCF-7 cells were analyzed. (h) The ratio of nuclear/cytosolic IDO1 in isolated ROShigh and ROSlow MCF-7 cells from tumor were analyzed (n = 6 technical replicates). scale bar, 10 μm. (i, j) HyPerRed fluorescence, Kyn concentration (i) and NADPH/NADP⁺ ratio (j) in synchronized HyPerRedlow IDO1high MCF-7/U2OS cells were measured. (k, l) Kyn concentration (k) and NADPH/NADP⁺ ratio (l) in sorted ROShigh and ROSlow MCF-7 cells were measured (n = 6 technical replicates). (m) The ratio of nuclear/cytosolic IDO1 in isolated ROShigh and ROSlow cells from human breast tumor were analyzed (n = 6 patients per group). Scale bar, 10 μm. (n-p) The expression of IDO1 in IDO1-OE ROShigh and Sh-IDO1 ROSlow human tumor cells were analyzed by Western blot. (q) IDO1 immunofluorescence in chemo-responsive/non-responsive breast cancer tissues (n = 6 patients per group) with relative IDO1 level and nuclear/cytosolic ratio calculated. Scale bar, 10 μm. (r, s) IDO1 immunofluorescence (r) in IgG/anti-VEGFR2-treated mouse tumors with relative level and nuclear/cytosolic ratio calculated (s). n = 5 mice per group; scale bar, 10 μm. (t, u) The level of STAT1, p-STAT1 and IDO1 in hypoxic MCF-7 cells and sorted HyPerRedhigh cells and HyPerRedlow MCF-7 cells were analyzed Western blot. For a-d, g, i, j, n-p, t, u, n = 3 biological independent experiments. All error bars are mean ± SD. P values were calculated by two-tailed Student’s t test (h, k, l, m, q, s).
Extended Data Fig. 10 Blockade of IDO1 and AhR exerts a superior anti-tumor effect.
(a) The relative cell growth of MCF-7 cells treated with IDO1 inhibitors (Lin, 1 μM or 5 μM; 1-MT, 50 μM or 200 μM) was recorded; the relative cell growth of WT or IDO1 knockout MCF-7 cells was recorded. (b) Immunoblot with dimedone of immunoprecipitations of AHR in sorted of HyPerRedhighIDO1low or HyPerRedlowIDO1high U2OS tumor cells were analyzed by Western blot. (c) Immunoblot with AHR and PP1c of immuno-precipitates of PTG in sorted of HyPerRedhighIDO1low or HyPerRedlowIDO1high U2OS tumor cells were analyzed by Western blot. (d) Immunoblot with PYGL and pPYGL of sorted HyPerRedhigh IDO1low or HyPerRedlow IDO1high U2OS tumor cells were analyzed by Western blot. (e) Immunoblot with dimedone of immunoprecipitations of AHR in MCF-7 cells treated with IDO1 inhibitor (Linrodostat (Lin), 5 μM) was analyzed by Western blot. (f) Immunoblot with AHR, p-PYGL and PP1c of immuno-precipitates of PTG in MCF-7 cells treated with IDO1 inhibitor (Lin, 5 μM) was analyzed by Western blot. (g) Cell death and HyPerRed fluorescence of HyPerRedhigh IDO1low U2OS tumor cells transfected with Sh-NC or Sh-AHR were detected by flow cytometry. (h, i) ROS level (h) and cell death (i) of sorted HyPerRedhighIDO1low MCF-7 and U2OS tumor cells treated with 10 μM SR1 were detected. (j) HyPerRed fluorescence of sorted HyPerRedlowIDO1high MCF-7 and U2OS cells treated with 10 μM linrodostat or epacadostat were analyzed by flow cytometry. (k) Cell death of sorted HyPerRedlow IDO1high MCF-7 and U2OS cells treated with 10 μM linrodostat or Linrodostat with 10 μM SR1 were analyzed by flow cytometry. For a-k, n = 3 biological independent experiments. All error bars are mean ± SD. P values were calculated by one-way ANOVA followed by Bonferroni’s test (a, g, j, k) or two-tailed Student’s t test (h, i).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1 and 2, and Tables 1–8.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, N., Ling, Z., Cao, X. et al. IDO1 regulating ROS rhythm reveals glycogenolysis/PPP as a cancer treatment target. Nat Chem Biol (2026). https://doi.org/10.1038/s41589-026-02161-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41589-026-02161-w