Abstract
Aging accounts for increased risk and dismal outcome of ischemic stroke. Here, we investigated the impact of age-related changes in the immune system on stroke. Upon experimental stroke, compared with young mice, aged mice had increased neutrophil clogging of the ischemic brain microcirculation, leading to worse no-reflow and outcomes. Aged mice showed an enhanced granulopoietic response to stroke that led to the accumulation of CD101+CD62Llo mature and CD177hiCD101loCD62Llo and CD177loCD101loCD62Lhi immature atypical neutrophils in the blood, endowed with increased oxidative stress, phagocytosis and procoagulant features. Production of CXCL3 by CD62Llo neutrophils of the aged had a key role in the development and pathogenicity of aging-associated neutrophils. Hematopoietic stem cell rejuvenation reverted aging-associated neutropoiesis and improved stroke outcome. In elderly patients with ischemic stroke, single-cell proteome profile of blood leukocytes identified CD62Llo neutrophil subsets associated with worse reperfusion and outcome. Our results unveil how stroke in aging leads to a dysregulated emergency granulopoiesis impacting neurological outcome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. All sequencing data generated in this study have been deposited at NCBI’s Gene Expression Omnibus (GEO) repository and are accessible through GEO Series accession number GSE174440. Source data are provided with this paper. Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, M. Bacigaluppi (bacigaluppi.marco@hsr.it). This study did not generate new or unique reagents.
Code availability
Code that supports the findings of this study are available from the corresponding author upon request. All sequencing code generated in this study have been deposited at NCBI’s GEO repository and are accessible through GEO Series accession number GSE174440.
References
Benjamin, E. J. et al. Heart disease and stroke statistics—2019 update: a report from the American Heart Association. Circulation 139, e56–e528 (2019).
Feigin, V. L., Lawes, C. M., Bennett, D. A. & Anderson, C. S. Stroke epidemiology: a review of population-based studies of incidence, prevalence, and case-fatality in the late 20th century. Lancet Neurol. 2, 43–53 (2003).
Iadecola, C., Buckwalter, M. S. & Anrather, J. Immune responses to stroke: mechanisms, modulation, and therapeutic potential. J. Clin. Investig. 130, 2777–2788 (2020).
Gelderblom, M. et al. Temporal and spatial dynamics of cerebral immune cell accumulation in stroke. Stroke 40, 1849–1857 (2009).
Courties, G. et al. Ischemic stroke activates hematopoietic bone marrow stem cells. Circ. Res. 116, 407–417 (2015).
Semerano, A. et al. Leukocyte counts and ratios are predictive of stroke outcome and hemorrhagic complications independently of infections. Front. Neurol. 11, 201 (2020).
Adrover, J. M. et al. A neutrophil timer coordinates immune defense and vascular protection. Immunity 50, 390–402 e310 (2019).
Jordan, J. E., Zhao, Z. Q. & Vinten-Johansen, J. The role of neutrophils in myocardial ischemia-reperfusion injury. Cardiovasc. Res. 43, 860–878 (1999).
El Amki, M. et al. Neutrophils Obstructing brain capillaries are a major cause of no-reflow in ischemic stroke. Cell Rep. 33, 108260 (2020).
Rossi, D. J. et al. Cell intrinsic alterations underlie hematopoietic stem cell aging. Proc. Natl Acad. Sci. USA 102, 9194–9199 (2005).
Maryanovich, M. et al. Adrenergic nerve degeneration in bone marrow drives aging of the hematopoietic stem cell niche. Nat. Med. 24, 782–791 (2018).
Akunuru, S. & Geiger, H. Aging, clonality, and rejuvenation of hematopoietic stem cells. Trends Mol. Med. 22, 701–712 (2016).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Jaiswal, S. et al. Clonal hematopoiesis and risk of atherosclerotic cardiovascular disease. N. Engl. J. Med. 377, 111–121 (2017).
Yemisci, M. et al. Pericyte contraction induced by oxidative-nitrative stress impairs capillary reflow despite successful opening of an occluded cerebral artery. Nat. Med. 15, 1031–1037 (2009).
Nortley, R. et al. Amyloid beta oligomers constrict human capillaries in Alzheimer’s disease via signaling to pericytes. Science 365, eaav9518 (2019).
Xie, X. et al. Single-cell transcriptome profiling reveals neutrophil heterogeneity in homeostasis and infection. Nat. Immunol. 21, 1119–1133 (2020).
Vafadarnejad, E. et al. Dynamics of cardiac neutrophil diversity in murine myocardial infarction. Circ. Res. 127, e232–e249 (2020).
Zilionis, R. et al. Single-cell transcriptomics of human and mouse lung cancers reveals conserved myeloid populations across individuals and species. Immunity 50, 1317–1334 e1310 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
Casanova-Acebes, M. et al. Rhythmic modulation of the hematopoietic niche through neutrophil clearance. Cell 153, 1025–1035 (2013).
Evrard, M. et al. Developmental analysis of bone marrow neutrophils reveals populations specialized in expansion, trafficking, and effector functions. Immunity 48, 364–379 e368 (2018).
Grassi, L. et al. Dynamics of transcription regulation in human bone marrow myeloid differentiation to mature blood neutrophils. Cell Rep. 24, 2784–2794 (2018).
Kwok, I. et al. Combinatorial single-cell analyses of granulocyte-monocyte progenitor heterogeneity reveals an early uni-potent neutrophil progenitor. Immunity 53, 303–318 e305 (2020).
Jonsson, H., Allen, P. & Peng, S. L. Inflammatory arthritis requires Foxo3a to prevent Fas ligand-induced neutrophil apoptosis. Nat. Med. 11, 666–671 (2005).
Yan, K. et al. Haploinsufficiency of PSMD12 causes proteasome dysfunction and subclinical autoinflammation. Arthritis Rheumatol. 74, 1083–1090 (2022).
Fischer, J. et al. Safeguard function of PU.1 shapes the inflammatory epigenome of neutrophils. Nat. Immunol. 20, 546–558 (2019).
Bjerregaard, M. D., Jurlander, J., Klausen, P., Borregaard, N. & Cowland, J. B. The in vivo profile of transcription factors during neutrophil differentiation in human bone marrow. Blood 101, 4322–4332 (2003).
Khoyratty, T. E. et al. Distinct transcription factor networks control neutrophil-driven inflammation. Nat. Immunol. 22, 1093–1106 (2021).
Zhang, D. E. et al. Absence of granulocyte colony-stimulating factor signaling and neutrophil development in CCAAT enhancer binding protein alpha-deficient mice. Proc. Natl Acad. Sci. USA 94, 569–574 (1997).
Tabula Muris, C. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590–595 (2020).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Giladi, A. et al. Single-cell characterization of haematopoietic progenitors and their trajectories in homeostasis and perturbed haematopoiesis. Nat. Cell Biol. 20, 836–846 (2018).
Kallquist, L. et al. The tetraspanin CD63 is involved in granule targeting of neutrophil elastase. Blood 112, 3444–3454 (2008).
Flohr Svendsen, A. et al. A comprehensive transcriptome signature of murine hematopoietic stem cell aging. Blood 138, 439–451 (2021).
Pillay, J., Tak, T., Kamp, V. M. & Koenderman, L. Immune suppression by neutrophils and granulocytic myeloid-derived suppressor cells: similarities and differences. Cell Mol. Life Sci. 70, 3813–3827 (2013).
Frenette, P. S., Mayadas, T. N., Rayburn, H., Hynes, R. O. & Wagner, D. D. Susceptibility to infection and altered hematopoiesis in mice deficient in both P- and E-selectins. Cell 84, 563–574 (1996).
Tamassia, N. et al. Induction of OCT2 contributes to regulate the gene expression program in human neutrophils activated via TLR8. Cell Rep. 35, 109143 (2021).
Khanna-Gupta, A. et al. Growth factor independence-1 (Gfi-1) plays a role in mediating specific granule deficiency (SGD) in a patient lacking a gene-inactivating mutation in the C/EBPepsilon gene. Blood 109, 4181–4190 (2007).
Shojaei, F. et al. Bv8 regulates myeloid-cell-dependent tumour angiogenesis. Nature 450, 825–831 (2007).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Van Gassen, S. et al. FlowSOM: using self-organizing maps for visualization and interpretation of cytometry data. Cytometry A 87, 636–645 (2015).
Hartmann, F. J. et al. High-dimensional single-cell analysis reveals the immune signature of narcolepsy. J. Exp. Med. 213, 2621–2633 (2016).
Niccoli, G., Burzotta, F., Galiuto, L. & Crea, F. Myocardial no-reflow in humans. J. Am. Coll. Cardiol. 54, 281–292 (2009).
Cruz Hernandez, J. C. et al. Neutrophil adhesion in brain capillaries reduces cortical blood flow and impairs memory function in Alzheimer’s disease mouse models. Nat. Neurosci. 22, 413–420 (2019).
Ng, L. G., Ostuni, R. & Hidalgo, A. Heterogeneity of neutrophils. Nat. Rev. Immunol. 19, 255–265 (2019).
Montaldo, E. et al. Cellular and transcriptional dynamics of human neutrophils at steady state and upon stress. Nat. Immunol. 23, 1470–1483 (2022).
Zhang, D. et al. Neutrophil ageing is regulated by the microbiome. Nature 525, 528–532 (2015).
Dong, Y. et al. Neutrophil hyperactivation correlates with Alzheimer’s disease progression. Ann. Neurol. 83, 387–405 (2018).
Salib, C. & Teruya-Feldstein, J. Hypersegmented granulocytes and COVID-19 infection. Blood 135, 2196 (2020).
Schulte-Schrepping, J. et al. Severe COVID-19 Is marked by a dysregulated myeloid cell compartment. Cell 182, 1419–1440 e1423 (2020).
Silvin, A. et al. Elevated calprotectin and abnormal myeloid cell subsets discriminate severe from mild COVID-19. Cell 182, 1401–1418 e1418 (2020).
Middleton, E. A. et al. Neutrophil extracellular traps contribute to immunothrombosis in COVID-19 acute respiratory distress syndrome. Blood 136, 1169–1179 (2020).
Sokulsky, L. A. et al. A critical role for the CXCL3/CXCL5/CXCR2 neutrophilic chemotactic axis in the regulation of type 2 responses in a model of rhinoviral-induced asthma exacerbation. J. Immunol. 205, 2468–2478 (2020).
Eash, K. J., Greenbaum, A. M., Gopalan, P. K. & Link, D. C. CXCR2 and CXCR4 antagonistically regulate neutrophil trafficking from murine bone marrow. J. Clin. Investig. 120, 2423–2431 (2010).
Bacigaluppi, M. et al. Delayed post-ischaemic neuroprotection following systemic neural stem cell transplantation involves multiple mechanisms. Brain 132, 2239–2251 (2009).
De Feo, D. et al. Neural precursor cell-secreted TGF-beta2 redirects inflammatory monocyte-derived cells in CNS autoimmunity. J. Clin. Investig. 127, 3937–3953 (2017).
Lelios, I. & Greter, M. Isolation of leukocytes from mouse central nervous system. Methods Mol. Biol. 1193, 15–19 (2014).
Capotondo, A. et al. Brain conditioning is instrumental for successful microglia reconstitution following hematopoietic stem cell transplantation. Proc. Natl Acad. Sci. USA 109, 15018–15023 (2012).
Gavillet, M. et al. Flow cytometric assay for direct quantification of neutrophil extracellular traps in blood samples. Am. J. Hematol. 90, 1155–1158 (2015).
Perdomo, J. et al. Neutrophil activation and NETosis are the major drivers of thrombosis in heparin-induced thrombocytopenia. Nat. Commun. 10, 1322 (2019).
Poli, V. et al. Inhibition of transcription factor NFAT activity in activated platelets enhances their aggregation and exacerbates gram-negative bacterial septicemia. Immunity 55, 224–236 e225 (2022).
Chua, R. Y. & Wong, S. H. SNX3 recruits to phagosomes and negatively regulates phagocytosis in dendritic cells. Immunology 139, 30–47 (2013).
Friebel, E. et al. Single-Cell mapping of human brain cancer reveals tumor-specific instruction of tissue-invading leukocytes. Cell 18, 1626–1642.e20 (2020).
Zunder, E. R. et al. Palladium-based mass tag cell barcoding with a doublet-filtering scheme and single-cell deconvolution algorithm. Nat. Protoc. 10, 316–333 (2015).
Mei, H. E., Leipold, M. D. & Maecker, H. T. Platinum-conjugated antibodies for application in mass cytometry. Cytometry A 89, 292–300 (2016).
Rahman, A. H., Tordesillas, L. & Berin, M. C. Heparin reduces nonspecific eosinophil staining artifacts in mass cytometry experiments. Cytometry A 89, 601–607 (2016).
Finck, R. et al. Normalization of mass cytom- etry data with bead standards. Cytometry A 83, 483–494 (2013).
Ellis, B. et al. flowCore: Basic structures for flow cytometry data. R package version 2.3.0. https://bioconductor.org/packages/flowCore/ (2019).
Finak, G. flowWorkspaceData: A data package containing two flowJo, one diva xml workspace and the associated fcs files as well as three GatingSets for testing the flowWorkspace, openCyto and CytoML packages. https://bioconductor.org/packages/release/data/experiment/html/flowWorkspaceData.html (2018).
Bendall, S. C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
D’Angelo, A. et al. Effect of clot-detection methods and reagents on activated partial thromboplastin time (APTT). Implications in heparin monitoring by APTT. Am. J. Clin. Pathol. 94, 297–306 (1990).
D’Angelo, S. V., Gilardoni, F. & D’Angelo, A. Evaluation of coagulometric assays in the assessment of protein C anticoagulant activity; variable sensitivity of commercial APTT reagents to the cofactor effect of protein S. Thromb. Haemost. 62, 861–867 (1989).
Hemker, H. C. & Kremers, R. Data management in thrombin generation. Thromb. Res. 131, 3–11 (2013).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Cowland, J. B. & Borregaard, N. Granulopoiesis and granules of human neutrophils. Immunol. Rev. 273, 11–28 (2016).
Wang, J. et al. Excessive neutrophils and neutrophil extracellular traps in COVID-19. Front. Immunol. 11, 2063 (2020).
The Gene Ontology, C. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Escobar, G. et al. Interferon gene therapy reprograms the leukemia microenvironment inducing protective immunity to multiple tumor antigens. Nat. Commun. 9, 2896 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Butti, E. et al. Subventricular zone neural progenitors protect striatal neurons from glutamatergic excitotoxicity. Brain 135, 3320–3335 (2012).
Bacigaluppi, M. et al. Neural stem cell transplantation induces stroke recovery by upregulating glutamate transporter GLT-1 in astrocytes. J. Neurosci. 36, 10529–10544 (2016).
Houwen, B. Blood film preparation and staining procedures. Clin. Lab. Med. 22, 1–14 (2002).
Adewoyin, A. S. & Nwogoh, B. Peripheral blood film—a review. Ann. Ib. Postgrad. Med. 12, 71–79 (2014).
Bain, B. J. Diagnosis from the blood smear. N. Engl. J. Med. 353, 498–507 (2005).
Kolde, R. pheatmap: pretty heatmaps. https://rdrr.io/cran/pheatmap/ (2019).
McInnes, L., Healy, J. & Melville, J. Umap: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Moon, K. R. et al. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 37, 1482–1492 (2019).
Acknowledgements
We thank A. Hidalgo for scientific advice, counseling and critical reading of the manuscript; P. Frenette for PEdKO mice; R. Ostuni and M. Genua for advice and help on blood scRNA-seq; C. Doglioni and F. Sanvito for advice; Advanced Light and Electron Microscopy BioImaging Center (ALEMBIC); F. Simeoni, D. Lazarevic and M. Morelli, Center for Translational Genomics and BioInformatics, IRCCS San Raffaele Hospital, for support in bioinformatics analysis; P.Panina-Bordignon and C. Bordignon for critical reading of the manuscript; M. Raso, hematological and chemistry laboratory service, Mouse Clinic, IRCCS, San Raffaele Hospital. This work has been supported in part by: TargetBrain (EU Framework 7 project HEALTH-F2-2012-279017) to G.M. and to B.B.; Cariplo, (‘Ricerca Medica Giovani ricercatori’ 2015–1057) to M.B. and A.U.; Ministero della Salute Italiana (Progetto Giovani Ricercatori 58/GR-2011−02348160) to M.B.; European Union (EU) funding within the NextGenerationEU-MUR PNRR Extended Partnership initiative on Consequences and challenges of ageing, Ageing Well in an Ageing Society (Project no. PE00000015, Age-It) to G.M.; European Research Council (ERC) under the EU’s Horizon 2020 research and innovation program grant agreement no. 882424 to B.B.; Swiss National Science Foundation (733 310030_170320, 310030_188450 and CRSII5_183478 to B.B.; Swiss Multiple Sclerosis Society to B.B. and to D.D.F.); University of Zurich postdoctoral fellowship to D.D.F.
Author information
Authors and Affiliations
Contributions
M.B. and D.D.F. conceptualized the study. G.S.G., D.D.F., E.F., A.S., N.M., and M.B. developed the methodology. D.D.F., E.F. and G.M.S. wrote the software. G.S.G., D.D.F., E.F., A.S. and M.B. performed formal analyses. G.S.G., D.D.F., E.F., A.S., N.M., E.Butti., E.Brambilla, A.B., T.V., M.G., A.dʼA., P.D.V., P.R., S.C., M.P., A.C., M.G.S. and M.B. performed the investigation. G.S.G., A.S., A.G., G.M., B.B., A.U., L.R., G.C. and M.B. provided resources. G.S.G., D.D.F., E.F., G.M.S. and M.B. curated the data. D.D.F., G.S.G. and M.B. wrote the original draft of the manuscript. D.D.F., G.S.G., M.B., B.B. and G.M. contributed to writing, review and editing of the manuscript. G.S.G., D.D.F. and M.B. worked on visualization. B.B., G.M. and M.B. were responsible for funding acquisition and supervision of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Steffen Massberg, Andres Hidalgo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Worse stroke outcome in aged male and female mice.
a, mNSS scores of young and aged male mice up to 7 days poststroke (n = 10 young, n = 6 aged, Mixed- effect Model, with Bonferroni post-hoc, one experiment, *p ≤ 0.05, **p ≤ 0.01). b, Kaplan-Meier survival curve showing mortality up to 8 days poststroke (n = 10 young and n = 8 aged male mice, Mantel-Cox test, *p = 0.0341). c- e, Stroke in young (2–4 months) and aged (18–24 –months) female mice. c, mNSS of young (n = 7) and aged (n = 4) female mice measured with the mNSS at 24h poststroke (2-tailed, Mann-Whitney test, **p = 0.003) and Kaplan-Meier survival curve up to 3days poststroke (n = 7 young, n = 5 aged female mice; Mantel-Cox test, **p = 0.0053). d, Staining for neurons (NeuN+ in brown) at 3 days poststroke and quantification of neurons (NeuN+) in the peri-lesional area (n = 7 young, n = 5 aged female mice, ***p = 0.001). e, Quantification of ischemic volume by Cresyl Violet staining at 3 days poststroke (n = 7 young n = 8 aged female mice). f-i, Representative images, and density of DHE+ (in red) astrocytes (GFAP+, green in f), neurons (NeuN+, green in g), blood vessels (CD31+, green in h) and pericytes (PDGFRb+, green in i) in the peri-lesional area at 24h poststroke in male young and aged mice (n = 3 mice per group *p = 0.0358). Cell nuclei are stained in blue (DAPI). j, Representative confocal microscopy images of 3NT+ cells (in green) in the ischemic cortex, and density of 3NT+ cells in the ischemic area of young and aged mice at 3 days poststroke (n = 8 young, n = 4 aged mice, **p = 0.0094). In the scatterplots each dot corresponds to a single mouse; mean ± SEM is shown. Scale bar in d, f, g, h, i, j, 50µm. 2-tailed t-test if not otherwise indicated.
Extended Data Fig. 2 Aged mice with stroke have increased brain and blood neutrophils.
UMAP displaying 17 resident and infiltrating immune cell subsets in the ischemic brain from young and aged mice at 24h poststroke analyzed by 19-color flow cytometry. b, Frequency of immune cell subsets identified in a (as percentage among total CD45+ cells, n = 5 mice per group, *p ≤ 0.05, **p ≤ 0.01, ****p ≤ 0.0001). c, Graph showing the number of eosinophils (SiglecF+) in the lesional and perilesional areas at 24h poststroke (n = 5 mice per group). d, UMAP of 18-color flow cytometry analysis of blood leukocytes in young and aged mice at steady state conditions (0h), at 6h and 24h poststroke (n = 5 mice per group). e, Heatmap showing the MAE values for the clustering markers for each cell cluster. f, Frequencies of blood leukocyte sub-populations among total CD45+ cells in young and aged mice (groups as in d, Mixed- effect Model, Benjamini-Hochberg post-hoc test; *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, one of two independent experiments is shown). In the scatterplots in b each dot corresponds to a single mouse; in f the boxes extend from the 25th to 75th percentiles, the midline denotes the median while whiskers plot min and max. Two tailed t-test if not otherwise indicated.
Extended Data Fig. 3 Neutrophil depletion and scRNAseq on blood leukocytes in stroke of the aged.
a, Neutrophil depletion by intravenous injection of anti-Ly6G antibody(1A8) or isotype-control (iso) in aged mice (a-f); MCFC plots for blood CD11b+CXCR2+neutrophils (gated on CD45+7/4+cells) and quantification of neutrophils 1.5h after treatment (n = 4 mice/group, **p = 0.0048). b, LDF-recordings (n = 15 Aged+1A8, n = 13 Aged+iso, 2-ANOVA, Benjamini-Hochberg post-test, **p ≤ 0.01). c, Immunofluorescence images and quantification of capillaries (t.lectin+vessels, white) with entrapped erythrocytes (NaBH4+, red) in the peri-ischemic cortex at 6h poststroke (n = 4, Aged+1A8; n = 5 Aged+iso, **p = 0.029). d, Representative images and density of 7/4+neutrophils (red). Stained are capillaries (t.lectin+, white) and nuclei (DAPI, blue) in the ischemic hemisphere at 3days poststroke (groups described in b, 2-way ANOVA, Sidak’s post-hoc test; n = 8 mice/group; pool of 2-independent experiments; ***p = 0.0001, *p = 0.048). e, mNSS score (n = 12 Aged+iso, n = 11 Aged+1A8; data are pooled from 3-independent experiments, Mixed-Effect-Model, Bonferroni post-hoc test, *p = 0.0319 and *p = 0.038). f, Quantification of peri-ischemic NeuN+neurons at 3days poststroke (n = 7. Aged+iso; n = 9, Aged+1A8, *p = 0.0117, data pooled from 2-independent experiments), ischemic volume and edema (n = 11, Aged+iso; n = 12 Aged+1A8). Dots in scatterplots correspond to a single mouse; mean±SEM is shown. 2-tailed t-test if not otherwise indicated. Scale bar c, d, 25µm. g-n, Analysis of blood neutrophils at 24h poststroke by scRNAseq. g, Scatterplots of differentially expressed genes between neutrophil clusters (as in Fig. 3). h-j, Violin plots showing, h, neutrophil activation score, i, neutrophil aging-score (from genes reported in Xie et al.17). j, chemotaxis score for neutrophil cluster as in Fig. 3b. k, Heatmap showing row-scaled expression of granule-related genes and related l, violin plots of azurophil, specific, gelatinase and secretory scores for neutrophil clusters (as in Fig. 3b). In h, i, j, l Bonferroni-corrected Wilcoxon rank sum test, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001; ****p ≤ 0.0001. All comparisons in Supplementary Table 3; boxes in violin plots extend from the 25th-75th percentiles, midline at median. m, n, UMAPs showing the weighted average expression for the signature of NADPH Oxidase, Neutrophil activation, ROS Production and NETs associated genes (as in Fig. 3b) corresponding to violin plots shown in Extended Data Fig. 3h and Fig. 3l, m. In n the correlations between the functional scores within the cluster scNeu1 (as in Fig. 3b) are shown.
Extended Data Fig. 4 Neutrophil subsets in the blood, brain and BM of young and aged stroke mice.
a, Quantification of circulating neutrophil subsets by 18-color MCFC at 0h, 6h and 24h poststroke, (n = 4 to 6 mice/group/ timepoint, 2-ANOVA, Bonferroni-post-hoc test, **p ≤ 0.01; ****p≤0.0001). b, Heatmap displaying the median antigen intensity in blood neutrophil clusters analyzed by MCFC at 24h poststroke and dotplot showing the scaled expression of selected genes, corresponding to markers of MCFC, of the scRNAseq stroke blood neutrophil subsets at 24h poststroke (neutrophil subsets from Fig. 3). The table shows the nomenclature and marker antigens and genes for major neutrophil subsets and GSVA based comparison (shown in Fig. 3g) with the nomenclature used by Xie et al.17. c, Histograms, and barplots showing the median fluorescence intensity (MFI) of CD177, CD11b and CD101 on neutrophils infiltrating the ischemic lesion analyzed by MCFC at 24h poststroke in young and aged mice (n = 5 mice/group, 2-tailed, Wilcoxon Mann-Whitney, *p = 0.031, **p = 0.0079, each dot corresponds to a single mouse). d, Representative IF images of CD177, CD101 or CD14 (in red) on neutrophils (Ly6G+in green, or white and 7/4+ in white) infiltrating the ischemic lesion of aged mice at 24h poststroke (blood vessels, stained by t-lectin, white) Scale bar, 25µm. e, Quantification of neutrophils and progenitors in the BM by MCFC at 0h and 24h poststroke of young and aged mice (n = 4- 6 mice per group per timepoint, 2-ANOVA Bonferroni post-hoc test; *p ≤ 0.05; **p ≤ 0.01; ****p≤0.0001). f, Barplots showing frequencies among mature BM neutrophils or BM neutrophil at 6h poststroke (n = 3 young and 4 aged mice, 2-tailed t-test, *p ≤ 0.05). g, ELISA for selected blood cytokines and chemokines at 0h and 48h poststroke; (n = 5 to 7 mice per group per timepoint, 2-ANOVA Bonferroni post-hoc test *p ≤ 0.05; **p≤0.01, ***p ≤ 0.001; §p≤0.055). In a, c, e and g, the box extends from the 25th- 75th percentiles, midline at median, whiskers plot min and max.
Extended Data Fig. 5 MCFC analysis of BM hematopoietic stem cells and progenitors after stroke.
a, 18-color MCFC analysis of live CD45+Lin-Ly6G-CD11b-ckit+ BM progenitor cells from young and aged mice at steady-state (0h) and 24h poststroke (n = 6young, n = 4 aged at 0h; n = 6 mice per group at 24h). Shown is the PHATE-algorithm based map showing BM progenitor clustering. b, Individual PHATE plots are overlaid with the clustering markers from the panel and c, corresponding quantification (groups as in a). The box plots extend from the 25th to 75th percentiles, the midline denotes the median while whiskers plot min and max, each dot corresponds to a single mouse (n = 4 to 6 mice per group per timepoint). 2-ANOVA, Benjamini-Hochberg post-hoc test *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001.
Extended Data Fig. 6 scRNAseq of BM and blood granulopoiesis of young and aged mice at steady-state and after stroke.
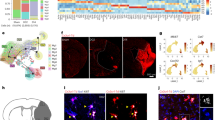
Integrated analysis of scRNAseq datasets of blood and BM cells from young (2-4months) and aged (18–24 months) male mice at steady state (0h) and 24h poststroke. a, Combined UMAP of the extracted neutrophil and neutrophil precursor subsets (total of 18’154 cells, see Supplementary Fig. 2, Supplementary Table 4) from blood and BM (see inset) of young and aged male mice at steady state and 24h poststroke colored by Seurat guided cluster identity. In brackets the inferred neutrophil population name following the nomenclature of Xie et al.17. b. The Blood and BM neutrophil precursors and neutrophil subsets shown in panel a, are here displayed divided for condition (steady state or 24h poststroke) and for age (young and aged mice). c, Dot plot for scaled expression of the gene signature for each neutrophil and neutrophil progenitor cluster (as in a). d, GSVA based comparison of the indicated neutrophil seurat clusters (as in a) with scRNA-seq-defined neutrophil populations reported by Xie et al.17 (HSPC, G0 to G5); GSVA score calculated on the list of top genes expressed per cluster75. The score of indicated defined clusters (HSPC, G0-G5) for each group of neutrophil subsets is shown (cluster 4, cMOP, is not computed). e, Graphs showing the proportion of cells for each cluster in young or aged mice, for the blood or BM at steady state and 24h poststroke (cluster 4, cMOP, not computed). f, Projection of pseudo-time generated with Monocle version 3 to infer the potential lineage differentiation trajectory onto the combined UMAP; the selected starting point is cluster 14 (HSPC). g, Expression of selected genes along pseudotime for granulocyte differentiation (Cluster 14, 6, 3, 9, 12, 10, 7, 8, 2, 0, 13, 1, 5) either at steady state or at 24h poststroke in young and aged mice.
Extended Data Fig. 7 Differentially expressed genes of granulopoietic clusters in the BM and blood among young and aged mice.
a, b, Heatmap showing the log2 [fold change] of selected differential expressed genes of aged over young mice in steady state (a) and 24h poststroke (b) for each main cluster of the BM (similar clusters have been computed together, 9 and 12 (G1) and 7 and 10 (G2)). Genes are marked with an asterisk if their expression changed significantly, as identified by 2-tailed t-test, Bonferroni-corrected p value, *p ≤ 0.05). c, Dotplot showing the scaled expression of signature genes for each cluster in the BM 24h poststroke in young (Y) and aged (A) mice, colored by the average expression of each gene in each cluster scaled across the indicated clusters (as in a). Dot size represents the percentage of cells in each cluster with more than one read of the corresponding gene. d, Projection of the stem-score, proliferation-score and G-CSF score signatures33 onto the combined UMAP (as in Extended Data Fig. 6a) comprehending blood and BM neutrophils and neutrophil progenitors of young and aged mice at steady state and 24h poststroke. The dotplots displayed below each score report for HSPC (cluster 14), G0_CMP (cluster 6), G0_GMP (cluster 3), G1 (cluster 9 and 12) and G2 (cluster 7 and 10) and G3 (cluster 8) the mean signature score. e, Scatterplots showing the differentially expressed genes in the blood of aged stroke mice compared to young stroke mice within the four mostly represented neutrophil clusters of the blood (cluster 2, 0, 1, 5 as in Extended Data Fig. 6a). Colored dots display an adjusted p- value p≤0.05, as identified by 2-tailed t-test, Bonferroni-corrected p value *p ≤ 0.05.
Extended Data Fig. 8 Neutrophils’ functional assays and heterochronic neutrophil transfer in stroke.
a, ROS production by blood neutrophil subsets (as defined in Fig. 4) in young and aged mice 24h poststroke assessed by MCFC. b, Phagocytosis of GFP+ E. coli by mature BM neutrophil subsets in young and aged mice. c, Neutrophil-platelet aggregates (NPAs) (Ly6G+ CD41+ cells) in the blood at 24h poststroke. d, Representative MCFC-plots for NPA in Neu3a and quantification of NPAs among neutrophils. e, NETting neutrophils (citH3+MPO+ cells) in the blood of young and aged mice at 24h poststroke. f, Representative MCFC plots of the NETting Neu2 subset and frequencies of citH3+MPO+NETting neutrophils subsets. In a-f n = 4 to 5 mice/group). g, MCFC plots of CD62LhiCXCR4lo and CD62LloCXCR4hi gated either on Ly6G+CD101+ or on Ly6G+CD101lo blood neutrophils of young and aged mice 24h poststroke. Microscopy images of sorted neutrophils stained by May-Grunwald/Giemsa. h, Quantification of neutrophils, band cells and hypersegmented neutrophils, on blood smears of young and aged mice at 0h, 1h, 24h poststroke (insets are May-Grünwald/Giemsa stained cells, n = 3 to 15 mice per group per timepoint). In h, the box extends from the 25th - 75th percentiles, midline at median, whiskers plot min and max. In a, b, d, f, h 2-ANOVA, Sidak’s post-hoc test, in c, e two-tailed t-test. *p ≤ 0.05; **p ≤ 0.01. i-j, MCFC at 48h post-stroke on the inflammatory brain infiltrate of CD45.1+ young recipients intravenously transferred with BM neutrophils from young or aged CD45.2+ donors. i, MCFC plots of inflammatory cells within the ipsilateral (ipsi) and contralateral (contra) hemispheres and quantification (n = 4 mice per group, contra-hemispheres were pooled in groups of two). j, Representative immunofluorescence image of labeled (CellTracker+ cells, green) aged-donor neutrophils (Ly6G+, red) in the ischemic lesion of young stroke recipients. Blood vessels (t.lectin+, white). k-m, Adoptive intravenous transfer of blood leukocytes from young(YtoY) or aged (AtoY) or aged donors with neutrophil depleted leukocytes(AnoNEUtoY) at 24h poststroke to young stroke recipients. k, LDF-recordings, l, mNSS-score, m, quantification of peri-ischemic NeuN+neurons, ischemic volume and edema at 72h poststroke. In k, l, 2-ANOVA, Tukey post-hoc test; m, 1-ANOVA, Tukey post-hoc test; n = 6 YtoY, n = 6 AtoA, n = 5 AnoNEUtoY, except in k n = 4 mice; *p ≤ 0.05; **p ≤ 0.01. n-o, Adoptive intravenous-transfer of neutrophils from wild-type donors (NtoY), or PEdKO donors (NKOtoY), or neutrophil-depleted PEdKO donors (NKO+𝜶Gr-1toY) into young stroke mice. n, MCFC plots of transferred cells; LDF-recordings and mNSS-score and o, immunofluorescence-images and quantification of peri-ischemic NeuN+neurons (green), and ischemic volume evaluation at 3 days poststroke. (n = 5 to 6 NtoY, n = 5 to 6 NKOtoY and n = 4 NKO+𝜶Gr-1toY). In n 2-ANOVA, Tukey post-hoc test; in o, 1-ANOVA, Tukey post hoc test; *p ≤ 0.05; ***p ≤ 0.001. In the scatterplots each dot corresponds to a single mouse; mean±SEM is shown. Scale bar, h, 5µm, j, o, 50µm. Nuclei (DAPI, blue).
Extended Data Fig. 9 CXCL3 worsens stroke outcome.
a, GSEA-enrichment curve for IL8 and CXCR2 signaling in aged over young mice at 24h poststroke calculated on bulk RNA-seq data (Fig. 7) and violin plot showing CXCR2 signature score in scRNAseq neutrophil subsets (as in Fig. 3b). b, Bar graphs of CD62L, CXCR4, CXCR2 and CD11b MCFC-based MFI on blood neutrophils after in vitro CXCL3 (or vehicle) stimulation (2-tailed t-test; n = 4 mice/group, *p ≤ 0.05) and c, histograms. d-g, In vivo intravenous injection of CXCL3 (or vehicle) in young mice. d, Analysis of total blood neutrophils 2h post-injection, e, representative MCFC plots, f, quantification of neutrophil subsets (as in Fig. 4d). In d-g n = 5 to 6 mice per group; one of three independent experiments is shown; two-tailed t-test; ***p ≤ 0.001, **p ≤ 0.01,. h-k, In vivo intravenous injection of CXCL3 (or vehicle) to young mice and stroke induction. h, LDF-recordings (n = 9 vehicle vs n = 8 CXCL3, 2-ANOVA, Benjamin-Hochberg post hoc test), i, mNSS-score, and j, quantification of peri-ischemic NeuN+neurons and ischemic volume 3 days poststroke. k, Immunofluorescence images and quantification of perilesional Ly6G+neutrophils (green) at 3 days poststroke. Blood vessels (t.lectin+, red), nuclei (DAPI, blue). In i-k, n = 5 to 8 mice per group, two-tailed t-test except for mNSS, 2-ANOVA, Sidak’s post hoc test. l-p, In vivo intraperitoneal injection of anti-CXCL3 antibody (A+αCXCL3) or isotype (A+isotype) in aged mice undergoing stroke. l, LDF (n = 9 A+isotype; n = 8, A+αCXCL3, 2-ANOVA, Benjamin-Hochberg post hoc test), m, mNSS-score and n, quantification of peri-ischemic NeuN+neurons, o, ischemic volume and edema at 3 days poststroke. p, Quantification of perilesional Ly6G+neutrophils 3days poststroke. In m-p, n = 5 mice per group, 2-tailed t-test, except for mNSS, Mixed-effects model, Holm-Sidak’s post hoc test. q-u, Adoptive transfer of BM neutrophils from aged donors into young mice with stroke, pre-treated with anti-CXCL3 (AtoY+αCXCL3). The control groups YtoY and AtoY are reported from Fig. 6c. q, LDF, r, mNSS-score and s, quantification of peri-ischemic NeuN+neurons, t, ischemic volume, and edema. u, Quantification of perilesional Ly6G+ neutrophils at 3days poststroke. n = 5 to 8 mice per group; 1-ANOVA, Tukey post-hoc test; for mNSS, Mixed-effects model, Sidak’s post hoc test; for LDF, 2-ANOVA Tukey post-hoc test. Dots in scatterplots corresponds to a single mouse; mean±SEM is shown. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001.
Extended Data Fig. 10 Leukocytes subsets in young and old stroke patients.
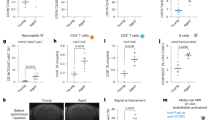
a-d. Mass cytometry analysis of blood leukocytes (groups as in Fig. 8a). a, Representative UMAP map showing the FlowSOM-guided meta-clustering of blood leukocytes. b, Heatmap displaying the MAE of markers used to generate the UMAP. c, Bar graphs showing frequencies of leukocyte subsets, and donut graphs of non-neutrophil leukocytes in MC and in SP at the indicated timepoints. d, Violin plots displaying quantifications of circulating leukocytes in MC and in SP at D1, D3 and D7 after stroke. e, Flow cytometry analyis of total white blood cells (WBCs), neutrophils, lymphocytes, and neutrophil-to-lymphocytes ratio (NLR) (groups as in Fig. 8f). f, Average thrombogram of control conditions for tissue factor-triggered thrombin generation in FV and FVIII-deficient plasma (depleted), in normal pool plasma (NPP), in mixtures of factor V and VIII-deficient plasma and NPP with final factor V and VIII concentrations of 1% and 0.5% (NPP0.5% and NPP 1%), and ellagic acid-triggered thrombin generation in factor V and VIII-deficient plasma (depleted_FSL). No ellagic acid-triggered thrombin generation was observed in factor V and VIII-deficient plasma-neutrophils mixtures (any neutrophil preparation). Thrombin generation did not occur with a reagent containing phospholipid vesicles in the absence of tissue factor. g, Representative flow cytometry plots of purified human neutrophils. In d, e 2-tailed multiple Mann-Whitney Wilcoxon test; *refers to comparisons between young-SP and old-SP. For comparison of young-SP or old-SP with respectively young-MC or old-MC Kruskall-Wallis with Dunn’s post-hoc test: ^refers to comparisons with MC-young; # refers to comparisons with MC-old; *,^,#p ≤ 0.05, **,^^,## p ≤ 0.01, ***,^^^,### p ≤ 0.001, ****,#### p ≤ 0.0001.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1,7 and 8 and Figs 1–5.
Supplementary Table 2 (download XLSX )
Flow cytometry leukocytes counts of blood from young and aged mice.
Supplementary Table 3 (download XLSX )
Stroke blood scRNA-seq quality control (QC) parameters, cell cluster signature genes and gene lists used for enrichment scores with related P values.
Supplementary Table 4 (download XLSX )
Blood steady state (0 h), blood stroke (24 h), BM steady state (0 h) and BM stroke (24 h) scRNA-seq.
Supplementary Table 5 (download XLSX )
Tabula Muris Senis.
Supplementary Table 6 (download XLSX )
Bulk RNA-seq.
Source data
Source Data Fig. 1 (download XLSX )
Statistical Source Data.
Source Data Fig. 2 (download XLSX )
Statistical Source Data.
Source Data Fig. 4 (download XLSX )
Statistical Source Data.
Source Data Fig. 5 (download XLSX )
Statistical Source Data.
Source Data Fig. 6 (download XLSX )
Statistical Source Data.
Source Data Fig. 7 (download XLSX )
Statistical Source Data.
Source Data Fig. 8 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical Source Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gullotta, G.S., De Feo, D., Friebel, E. et al. Age-induced alterations of granulopoiesis generate atypical neutrophils that aggravate stroke pathology. Nat Immunol 24, 925–940 (2023). https://doi.org/10.1038/s41590-023-01505-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-023-01505-1
This article is cited by
-
The ageing immune system as a driver of systemic ageing
Nature Reviews Immunology (2026)
-
The role of vascular aging, bone marrow and immune system in hypertension
Cell Death Discovery (2026)
-
High Mobility Group Protein B1 Mediates the Role of the Neutrophil Extracellular Traps in the Progression of Acute Myocardial Infarction
Cardiovascular Drugs and Therapy (2026)
-
Neutrophil extracellular traps released by CD177+ neutrophils aggravated inflammation and neuronal impairment post-SCI
Cell Communication and Signaling (2025)
-
Microbiota-shaped neutrophil senescence regulates sexual dimorphism in bladder cancer
Nature Immunology (2025)