Abstract
Sex disparities have been epidemiologically demonstrated in non-reproductive cancers, yet how the sex-specific intrinsic microbiome orchestrates the immune system to affect these disparities is unclear. Here we identify a subpopulation of RETNLG+LCN2+ senescence-like neutrophils (RLSNs) that preferentially accumulate in the male tumor microenvironment and exert a strong immunosuppressive effect to limit antitumor immunity, resulting in poor prognosis for patients with bladder cancer. This difference in enrichment of RLSNs between sexes can be attributed to intestinal bacterium Alistipes shahii, which preferentially populates in females rather than males. A. shahii-associated metabolite lurasidone directly targets iron sequestrator LCN2 in RLSNs. By freeing Fe2+, lurasidone induces ferroptosis, thereby eliminating RLSNs and promoting antitumor immunity in females. In males lacking A. shahii and lurasidone, RLSNs have a survival advantage. Together, these findings demonstrate that a microbiota–lurasidone–LCN2 circuit regulates sexual disparity in bladder cancer and indicates the therapeutic potential of lurasidone for male cancer patients.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The scRNA-seq data have been deposited in the Gene Expression Omnibus under accession codes GSE240189, GSE287661 and GSE287682, and 16S rRNA-seq data have been deposited in the National Center for Biotechnology Information BioProject under accession code PRJNA1001867. The scRNA-seq data for human bladder cancer were obtained from the CNGB database (CNP 0000460). Source data are provided with this paper.
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Kwon, H. et al. Androgen conspires with the CD8+ T cell exhaustion program and contributes to sex bias in cancer. Sci. Immunol. 7, eabq2630 (2022).
Li, J. et al. Histone demethylase KDM5D upregulation drives sex differences in colon cancer. Nature 619, 632–639 (2023).
Yang, C. et al. Androgen receptor-mediated CD8+ T cell stemness programs drive sex differences in antitumor immunity. Immunity 55, 1268–1283.e1269 (2022).
Abdel-Hafiz, H. A. et al. Y chromosome loss in cancer drives growth by evasion of adaptive immunity. Nature 619, 624–631 (2023).
Bramble, M. S., Lipson, A., Vashist, N. & Vilain, E. Effects of chromosomal sex and hormonal influences on shaping sex differences in brain and behavior: lessons from cases of disorders of sex development. J. Neurosci. Res. 95, 65–74 (2017).
Zhao, R. et al. A GPR174-CCL21 module imparts sexual dimorphism to humoral immunity. Nature 577, 416–420 (2020).
Ng, L. G., Ostuni, R. & Hidalgo, A. Heterogeneity of neutrophils. Nat. Rev. Immunol. 19, 255–265 (2019).
Hidalgo, A., Chilvers, E. R., Summers, C. & Koenderman, L. The neutrophil life cycle. Trends Immunol. 40, 584–597 (2019).
Soehnlein, O., Steffens, S., Hidalgo, A. & Weber, C. Neutrophils as protagonists and targets in chronic inflammation. Nat. Rev. Immunol. 17, 248–261 (2017).
Wang, T. W. et al. Blocking PD-L1–PD-1 improves senescence surveillance and ageing phenotypes. Nature 611, 358–364 (2022).
Baker, D. J. et al. Naturally occurring p16Ink4a-positive cells shorten healthy lifespan. Nature 530, 184–189 (2016).
Debacq-Chainiaux, F., Erusalimsky, J. D., Campisi, J. & Toussaint, O. Protocols to detect senescence-associated beta-galactosidase (SA-βgal) activity, a biomarker of senescent cells in culture and in vivo. Nat. Protoc. 4, 1798–1806 (2009).
Bancaro, N. et al. Apolipoprotein E induces pathogenic senescent-like myeloid cells in prostate cancer. Cancer Cell 41, 602–619.e611 (2023).
Adrover, J. M. et al. A neutrophil timer coordinates immune defense and vascular protection. Immunity 50, 390–402.e310 (2019).
Zhang, D. et al. Neutrophil ageing is regulated by the microbiome. Nature 525, 528–532 (2015).
Summerhayes, I. C. & Franks, L. M. Effects of donor age on neoplastic transformation of adult mouse bladder epithelium in vitro. J. Natl Cancer Inst. 62, 1017–1023 (1979).
Ma, Z. et al. Interferon-dependent SLC14A1+ cancer-associated fibroblasts promote cancer stemness via WNT5A in bladder cancer. Cancer Cell 40, 1550–1565.e1557 (2022).
Crow, M., Paul, A., Ballouz, S., Huang, Z. J. & Gillis, J. Characterizing the replicability of cell types defined by single cell RNA-sequencing data using MetaNeighbor. Nat. Commun. 9, 884 (2018).
Ng, M. S. F. et al. Deterministic reprogramming of neutrophils within tumors. Science 383, eadf6493 (2024).
Gullotta, G. S., De Feo, D., Friebel, E., Semerano, A. & Scotti, G. M. Age-induced alterations of granulopoiesis generate atypical neutrophils that aggravate stroke pathology. Nat. Immunol. 24, 925–940 (2023).
Mollaoglu, G. et al. The lineage-defining transcription factors SOX2 and NKX2-1 determine lung cancer cell fate and shape the tumor immune microenvironment. Immunity 49, 764–779.e769 (2018).
Xie, X. et al. Single-cell transcriptome profiling reveals neutrophil heterogeneity in homeostasis and infection. Nat. Immunol. 21, 1119–1133 (2020).
Bottazzi, B., Riboli, E. & Mantovani, A. Aging, inflammation and cancer. Semin. Immunol. 40, 74–82 (2018).
Olson, B. et al. Lipocalin 2 mediates appetite suppression during pancreatic cancer cachexia. Nat. Commun. 12, 2057 (2021).
Zhang, X. et al. Sex- and age-related trajectories of the adult human gut microbiota shared across populations of different ethnicities. Nat. Aging 1, 87–100 (2021).
Loebel, A. et al. Lurasidone as adjunctive therapy with lithium or valproate for the treatment of bipolar I depression: a randomized, double-blind, placebo-controlled study. Am. J. Psychiatry 171, 169–177 (2014).
Kato, T. et al. Double-blind, placebo-controlled study of lurasidone monotherapy for the treatment of bipolar I depression. Psychiatry Clin. Neurosci. 74, 635–644 (2020).
Keks, N. A., Hope, J. & Castle, D. Lurasidone: an antipsychotic with antidepressant effects in bipolar depression? Australas. Psychiatry 24, 289–291 (2016).
Corsello, S. M. et al. Discovering the anti-cancer potential of non-oncology drugs by systematic viability profiling. Nat. Cancer 1, 235–248 (2020).
Dempster, J. M. et al. Chronos: a cell population dynamics model of CRISPR experiments that improves inference of gene fitness effects. Genome Biol. 22, 343 (2021).
Ghandi, M. et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569, 503–508 (2019).
Holmes, M. A., Paulsene, W., Jide, X., Ratledge, C. & Strong, R. K. Siderocalin (Lcn 2) also binds carboxymycobactins, potentially defending against mycobacterial infections through iron sequestration. Structure 13, 29–41 (2005).
Yao, F. et al. A targetable LIFR–NF–κB–LCN2 axis controls liver tumorigenesis and vulnerability to ferroptosis. Nat. Commun. 12, 7333 (2021).
Jung, B. K. et al. Reduced secretion of LCN2 (lipocalin 2) from reactive astrocytes through autophagic and proteasomal regulation alleviates inflammatory stress and neuronal damage. Autophagy 19, 2296–2317 (2023).
Flo, T. H. et al. Lipocalin 2 mediates an innate immune response to bacterial infection by sequestrating iron. Nature 432, 917–921 (2004).
Moschen, A. R. et al. Lipocalin 2 protects from inflammation and tumorigenesis associated with gut microbiota alterations. Cell Host Microbe 19, 455–469 (2016).
Antoni, S. et al. Bladder cancer incidence and mortality: a global overview and recent trends. Eur. Urol. 71, 96–108 (2017).
Zilionis, R. et al. Single-cell transcriptomics of human and mouse lung cancers reveals conserved myeloid populations across individuals and species. Immunity 50, 1317–1334.e1310 (2019).
López-Otín, C., Pietrocola, F., Roiz-Valle, D., Galluzzi, L. & Kroemer, G. Meta-hallmarks of aging and cancer. Cell Metab. 35, 12–35 (2023).
Gómez-Moreno, D. & Adrover, J. M. Neutrophils as effectors of vascular inflammation. Eur. J. Clin. Invest. 48, e12940 (2018).
Kjeldsen, L., Johnsen, A. H., Sengeløv, H. & Borregaard, N. Isolation and primary structure of NGAL, a novel protein associated with human neutrophil gelatinase. J. Biol. Chem. 268, 10425–10432 (1993).
Wang, D. et al. LCN2 secreted by tissue-infiltrating neutrophils induces the ferroptosis and wasting of adipose and muscle tissues in lung cancer cachexia. J. Hematol. Oncol. 16, 30 (2023).
Cramer, E. P. et al. Lipocalin-2 from both myeloid cells and the epithelium combats Klebsiella pneumoniae lung infection in mice. Blood 129, 2813–2817 (2017).
Feng, Q. et al. Lactate increases stemness of CD8 + T cells to augment anti-tumor immunity. Nat. Commun. 13, 4981 (2022).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147 (2022).
Alshetaiwi, H. et al. Defining the emergence of myeloid-derived suppressor cells in breast cancer using single-cell transcriptomics. Sci. Immunol. 5, eaay6017 (2020).
Chen, P. B. et al. Directed remodeling of the mouse gut microbiome inhibits the development of atherosclerosis. Nat. Biotechnol. 38, 1288–1297 (2020).
Acknowledgements
This research was supported by grants from the CAS Project for Young Scientists in Basic Research (YSBR-076), the National Natural Science Foundation of China (82425026, 82030041, 82441041, 82201924, 82425103, 82202932, 32025022, 32230062), the National Key R&D Program of China (2024YFA1803103, 2018YFA0902703), Strategic Priority Research Program of the Chinese Academy of Sciences (XDB39030300, XDB0480000), Yishan Research Project of Jiangsu Cancer Hospital (YSPY202402), Jiangsu Natural Science Foundation (BK20220735), and CAS Key Laboratory of Tissue Microenvironment and Tumor. We also acknowledge assistance from the personnel of the animal, flow cytometry and other platform core facilities at the Shanghai Institute of Nutrition and Health, Chinese Academy of Sciences. Finally, we thank NovelBio Co., Ltd. for support with bioinformatics analysis using their NovelBrain Cloud Analysis Platform (www.novelbrain.com), F. Wang (Third Military Medical University, Chongqing, China) for providing Lcn2-KO mice and J. Zhou (Institute of Neuroscience, Chinese Academy of Sciences, Shanghai, China) for providing Drd2-KO mice.
Author information
Authors and Affiliations
Contributions
Q.Z. and G.Z. designed and performed the experiments, prepared the figures, and wrote part of the paper; M.C. and M. Zhan contributed to human sample collection and analysis; H.H. and S.P. contributed to analysis and interpretation of scRNA-seq data; J. Xing and M. Zheng contributed to the analysis of lurasidone target prediction and lurasidone–LCN2 docking; X.D., J.L. and G.M. contributed to the anaerobic culture of A. shahii. J. Xu contributed to the experiments and data analysis; D.H. and Z.Z. contributed to the microbiota and lurasidone analyses; D.D. and G.H. contributed to data analysis; C.L. and J.Q. supervised a specific subset of the experiments and analyses; and Y.X. initiated, designed and supervised this study, prepared the figures, and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
Y.X., Q.Z, G.Z. and J. Xu have filed a patent application regarding the application of lurasidone in the treatment of male tumor patients (application number: 202311069387.1). The other authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Nick Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Neutrophils regulates gender difference of bladder tumor.
(a) UMAP visualization of cross-species analysis of CD45+ immune cells from mouse MB49 tumors and human bladder tumor tissues of different sexes, colored by inferred sample. (b, c) Dot plot showing the expression of selected marker genes in different cell types of mouse (b) and human (c) immune cells. (d) MB49 tumor growth in male and female C57BL/6 mice (n = 10). (e) Absolute cell numbers of the tumor-infiltrating immune cells in the MB49 tumor-bearing male and female mice (Ma and Neu, n = 9; others, n = 10). (f, g) Representative flow plots (left) and quantification of the proportions (right) of intratumor IFNγ+ CD8+ T cells (mouse M, n = 7; mouse F, n = 8; human M, n = 65; human F, n = 22) and CD4+ T cells (mouse, n = 6; human M, n = 43; human F, n = 17) between males and females MB49 tumor-bearing mice (f) and bladder cancer patients (g). (h–j) MB49 growth in male and female mice intravenous injection with IgG control or anti-CSF1R antibody (n = 8) (h), or in male and female Rag2−/− mice (n = 6) (i), or in male and female Rag2−/− mice that were intravenously injected with total T cells from male and female mice (M-T to M, n = 7; F-T to M, n = 6; others, n = 8) (j). (k, l) Quantification of the proportions of blood CD11b+ Gr1high neutrophils (k) and CD11b+ CD177+ cells (l) in male mice that were intravenously injected with IgG control or αLy6G one time every three days, the blood was collected every two days to perform flow analysis (n = 5). (m) Flow cytometric analysis about the proportion of intratumor Ly6G+ neutrophils, IFNγ+ CD8+ T cells and IFNγ+ CD4+ T cells in male and female mice that were intravenously injected with IgG control or αLy6G (IgG-M, n = 8; IgG-F, n = 7; others, n = 5). Bar graphs are presented as mean ± s.e.m. A two-tailed Student’s t-test was performed for comparisons. The data are representative of at least three independent experiments.
Extended Data Fig. 2 Identification of senescence-like neutrophil subset in the TME.
(a, b) Dot plots displaying the expression of selected marker genes of 4 neutrophils subpopulations from human (a) and mouse (b) bladder tumors. (c) Heatmap displaying the differentially expressed genes that were highly expressed both in human and mouse N3 neutrophil subpopulations. (d) Ridge distribution of mouse intratumor neutrophil with the genes expressed along pseudotime. (e) Relative genes expression of branches of N2 and N3 subpopulations from mouse neutrophils along pseudotime.
Extended Data Fig. 3 Validation the senescence-associated characteristics of N3 neutrophil.
(a) The tSNE visualization of 4 neutrophils subpopulations and genes expression of Tnfrsf23 (encoding dcTRAIL-R1), Vegfa, and Hk2 in our identified four neutrophils subpopulations. (b) Dot plots displaying the expression of T3-specific signature genes among our identified four neutrophil subsets. (c) Violin plots of N3 signature score between 3 neutrophil clusters in pancreatic cancer (Ng et al.), violin plots of T3 signature score between 4 neutrophil clusters we identified. (d) Violin plots of neutrophil aging score between 3 neutrophil clusters in pancreatic cancer (Ng et al.). (e) GO pathway analysis revealing the pathways enriched in N3 and T3 neutrophils. (f, g) UMAP visualization (f) and distribution comparison (g) of 4 neutrophils populations in peripheral blood from 2-month-old (2 m) and 24-month-old (24 m) mice based on our unpresented data, colored by inferred cluster’s identity. (h) Similarity analysis of our newly identified mouse neutrophil subpopulations (N0-N3) with our unpresented blood neutrophil clusters (Neu1-Neu4). (i–n) Similarity analysis of our newly identified mouse neutrophil subpopulations with the published neutrophil clusters, including neutrophils clusters from young and aged stroke mice (i), peripheral mouse neutrophils clusters under homeostasis and infection (k), and neutrophils clusters from mouse lung tumor (m), which were obtained from the published scRNA-seq data. The aging scores of the published neutrophil clusters were also re-analyzed and presented (j, l, n). (o, p) Violin plot showing the Trem2 expression in 4 neutrophils subpopulations from human (o) and mouse (p) bladder tumors. The statistical significance of differences in the gene set score in (j, l, n) were assessed using Wilcoxon test.
Extended Data Fig. 4 The N3 neutrophil subset predicts poor survival.
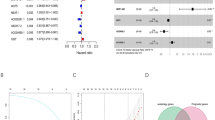
(a, c) Prognostic value of specific human N3 (a) and mouse N3 (c) enriched genes for overall survival in bladder cancer patients from TCGA datasets. Forest plots showing HR and 95% CI derived from univariate Cox analysis in male and female bladder cancer patients. Within the forest plot, HR for each variable is depicted as a box, and 95% CIs are shown as horizontal. The vertical line crossing the value of 1.0 represents a non-statistically significant effect, and odds greater than one indicate worse effects. (b, d) Distribution of risk score, survival time, specific human N3 (b) and mouse N3 (d) enriched genes involved in the prognostic model among bladder cancer patients in TCGA datasets. (e) QPCR analysis of mRNA expression of the genes including Anxa2, Rdh11, S100a8, St3gd16, Taldo1 and Timp2 in tumor-infiltrated senescence-like fluorescein di-β-D-galactopyranoside (FDG)+ and non-senescence FDG− neutrophils (n = 3). Bar graphs are presented as mean ± s.e.m. A two-tailed Student’s t-test was performed for comparisons. The data are representative of at least three independent experiments.
Extended Data Fig. 5 RLSNs exhibited more immunosuppressive activity.
(a–c) Violin plot showing Lcn2 and Retnlg expression in different neutrophils clusters from three published scRNA-seq data, including neutrophils clusters from young and aged stroke mice (a), peripheral mouse neutrophils clusters under homeostasis and infection (b), and neutrophils clusters from mouse lung tumor (c). (d) The mRNA (upper) (n = 4) and proteins expression (lower) of Lcn2 among flow-sorted tumor cells, dendritic cells (DCs), macrophage (Ma), T cells, Lcn2+ neutrophils (Lcn2+ Neu) and Lcn2− neutrophils (Lcn2− Neu). (e) ELISA analysis of supernatant Lcn2 concentration of Ex-vivo cultured intratumoral Lcn2− and Lcn2+ neutrophils (n = 3). (f, g) Absolute cell numbers calculation by FACS analysis (n = 6) (f) and fraction of total neutrophils with ScRNA-Seq and FACS analysis (g) of intratumoral Lcn2+ and Lcn2− neutrophils (n = 6). (h) QPCR analysis of mRNA express of peripheral blood and spleen FACS-sorted Lcn2− or Lcn2+ neutrophils from 2-month-old MB49-bearing mice (left) or 24-month-old mice (right) (n = 4). (i, j) Representative flow plots of Lcn2+ neutrophils and ELISA analysis of serum (n = 6) and spleen Lcn2 (n = 5) expression between young (2-month-old) and old (24-month-old) mice. (k, l) Representative flow cytometry histogram of fluorescein di(β-D-galactopyranoside) (FDG) fluorescence and quantification of FDG mean fluorescence intensity (MFI) of neutrophils in spleen, blood and tumor from MB49-bearing young male mice (n = 4). (m, n) Representative flow plots and quantification of Lcn2+ neutrophils in spleen, blood and tumor from MB49-bearing young male mice (n = 4). (o) Representative flow plot of the percentages of IFNγ+ CD8+ T cells, GzmB+ CD8+ T cells, proliferating CD8+ T cells, PD1+ CD8+ T cells and TIM3+ CD8+ T cells by co-culturing intratumor Lcn2− or Lcn2+ neutrophils with splenic CD8+ T cells. Bar graphs are presented as mean ± s.e.m. A two-tailed Student’s t-test was performed for comparisons. The data are representative of at least three independent experiments.
Extended Data Fig. 6 Lcn2 was crucial for N3 neutrophil and its immunosuppressive function.
(a) The experimental protocol of chimeric mice, 9.5 Gy irradiated female mice receiving male WT and Lcn2 KO bone marrow, then inoculated with MB49 tumor cells 8 weeks after BM reconstitution. (b, c) UMAP visualization and distribution comparison of cell populations among CD45+ immune cells (b), UMAP visualization of 4 clusters of neutrophils (c) from mouse MB49 tumors from (a), colored by inferred cluster’s identity. (d) Violin plots of cytotoxic score of CD8+ T cells from WT and Lcn2 KO chimeric MB49-bearing mice. (e–h) Representative flow plot and quantification of the percentages of IFNγ+ CD8+ T cells, GzmB+ CD8+ T cells and proliferating CD8+ T cells by co-culturing intratumor neutrophils from WT and Lcn2 KO chimeric MB49-bearing mice with splenic CD8+ T cells (n = 3). Bar graphs are presented as mean ± s.e.m. A two-tailed Student’s t-test was performed for comparisons. The statistical significance of differences in the gene set score in (d) were assessed using Wilcoxon test. The data are representative of at least three independent experiments.
Extended Data Fig. 7 Lcn2 and Retnlg were specifically and highly co-expressed in N3.
(a) Construction strategy of Retnlg-DTR-GFP mice. (b) Flow analysis of Retnlg-GFP expression among flow-sorted tumor cells, dendritic cells (DCs), macrophage (Ma), T cells, Lcn2+ neutrophils and Lcn2− neutrophils from Retnlg-DTR-GFP MB49 tumor-bearing mice (n = 5). (c) QPCR validation of Retnlg and Lcn2 mRNA expression in tumor-infiltrated GFP+ and GFP− neutrophils, and ELISA analysis of supernatant Lcn2 concentration of ex-vivo cultured tumor-infiltrated GFP+ and GFP− neutrophils from Retnlg-DTR-GFP MB49 tumor-bearing mice (n = 3). (d) Representative histogram of Retnlg-GFP fluorescence and quantification of Retnlg-GFP mean fluorescence intensity (MFI) in intratumor Lcn2+ and Lcn2− neutrophils from Retnlg-DTR-GFP MB49 tumor-bearing mice (n = 11). (e) Histogram showing the Retnlg-GFP expression in tumor-infiltrated CD11b+ Ly6G+ neutrophils from the diphtheria toxin (DT)-treated wild-type (WT) and Retnlg-DTR-GFP mice and PBS-treated Retnlg-DTR-GFP mice. (f) Quantification of Retnlg-GFP expression (n = 3), percentages of Lcn2+ neutrophils, absolute cell numbers of the tumor-infiltrating Lcn2+ and Lcn2− neutrophils (n = 6), and ELISA assay of serum Lcn2 (PBS, n = 8; DT, n = 6) from PBS or DT treated Retnlg-DTR-GFP male mice. (g) Quantification of the proportions of intratumor IFNγ+ CD4+ T cells in WT and Retnlg-DTR-GFP tumor-bearing mice that were intraperitoneally injected with DT between male and female (M-WT, n = 8; M-DTR, n = 7; F-WT, n = 6; F-DTR, n = 5). (h–j) Tumor growth (n = 7) (h), flow analysis of Lcn2+ neutrophil proportion in tumor (n = 6) (i), spleen and blood, and BM neutrophil Lcn2 MFI (n = 6) (j) in male Retnlg-DTR tumor-bearing mice intratumorally injected with diphtheria toxin (DT) or PBS. Bar graphs are presented as mean ± s.e.m. A two-tailed Student’s t-test was performed for comparisons. The data are representative of at least three independent experiments.
Extended Data Fig. 8 Microbiota and metabolites difference between male and female mice.
(a) Cladogram generated from LEfSe of cecal feces in mice from M-Ctrl, F-Ctrl, M-FMT and F-FMT. (b) The relative abundance of bacterium species selected with an average species abundance greater than 0.5% in at least one of the groups according to the 16 s sequencing data (n = 5). (c) QPCR analysis of A. Shahii abundance of cecal feces in male mice post sham (Ctrl) or surgical castration (Cas), or intraperitoneal injection of enzalutamide (Enza) (Enza, n = 8; others, n = 7). (d) Quantification of the proportions of intratumor neutrophils in male and female MB49 tumor-bearing mice orally administered with PBS (Ctrl) or A. Shahii (A. sha) (M- A. sha, n = 8; F- A. sha, n = 6; others, n = 7). (e) Bar graphs showing the relative concentration of selected metabolites (n = 6). (f, g) The serum concentration of lurasidone (f), correlation analysis between serum lurasidone and relative abundance of A. Shahii (g) in male and female MB49 tumor-bearing control or ABX-treated mice administered with PBS (Ctrl) or A. Shahii (A. sha) via oral gavage (F- A. sha, n = 9; others, n = 10). (h, i) Representative flow plots and quantification of the proportions of intratumor IFNγ+/TNFα+ CD4+ T cells (M-Ctrl, n = 7; others, n = 6) and CD8+ T cells (F-Ctrl, n = 5; others, n = 6) in the dissected orthotopic MB49 tumors from male mice intraperitoneally injected daily with vehicle or lurasidone and in female mice intraperitoneally injected with vehicle. (j) Subcutaneous MB49 tumor growth in male mice intraperitoneally injected daily with vehicle, enzalutamide, lurasidone (1 mg/kg) or in combination, and in female mice intraperitoneal injected daily with vehicle or lurasidone (M-Veh, M/F-Lura, n = 9; others, n = 10). (k) B16-F10 tumor growth in male and female mice intraperitoneal injection daily with vehicle or lurasidone (1 mg/kg) (M-Veh, n = 7; F-Veh, n = 8; M-Lura, n = 6; F-Lura, n = 9). Bar graphs are presented as mean ± s.e.m., The statistical significance of differences between the M-Ctrl and F-Ctrl, M-FMT and F-FMT groups in (b) were assessed using the Wilcoxon test. A two-tailed Student’s t-test was performed for comparisons of other figures. The data are representative of at least three independent experiments.
Extended Data Fig. 9 Lcn2 is the target of lurasidone.
(a, b) Confocal microscopy images showing TUNEL+ Ly6G+ dead neutrophils (Neu) in male and female MB49 tumors (a) and quantification of the proportion of TUNEL+ Ly6G+ dead neutrophils (b) (n = 8). Scale bars, 10 µm. (c) Violin plots comparing the signatures scores of different death pathway including ferroptosis, apoptosis, necroptosis, pyroptosis and autophagy in male and female intratumor neutrophils. (d) Violin plots comparing the signatures scores of ferroptosis in 4 neutrophil clusters. (e) Relative lipid ROS showing the ferroptosis levels of intratumor neutrophils from male and female mouse MB49 tumor (n = 7). (f) Relative lipid ROS showing ferroptosis level of the in vitro cultured MB49 tumor cells, tumor-infiltrated CD8+ T cells and CD4+ T cells treated with DMSO (DM) or lurasidone (Lura) (n = 3). (g) Quantification of the proportion of intratumor total Ly6G+ neutrophils in orthotopic (M-Ctrl, n = 8; others, n = 5) and subcutaneous (F-Ctrl, n = 8; others, n = 6) MB49 tumors from the male and female mice that treated daily with vehicle (17% Polyethylene glycol 12-hydroxystearate in PBS, Ctrl) or lurasidone (1 mg/kg). (h) MB49 tumor growth in 9.5 Gy-irradiated male mice that reconstituted with the bone marrows from male wild-type (WT) and Drd2-KO mice, inoculated with MB49 tumor cells 8 weeks after bone marrow reconstitution, then intraperitoneally injected daily with vehicle or lurasidone (1 mg/kg) (WT-Veh, n = 9; Drd2 KO-Lura, n = 7; others, n = 8). (i) LCN2 basal mRNA expression in different lurasidone-sensitive cell line including not sensitive, weakly sensitive and sensitive. (j, k) Flow analysis of proportion of live neutrophils, Intracellular FerroOrange MFI, relative lipid ROS of mouse neutrophils treated with DMSO, lurasidone and Lcn2 inhibitor ZINC00640089, ZINC00784494 (n = 3). Bar graphs are presented as mean ± s.e.m. A two-tailed Student’s t-test was performed for comparisons. The statistical significance of differences in the gene set score in (c) were assessed using Wilcoxon test. The data are representative of at least three independent experiments.
Extended Data Fig. 10 Lcn2 eliminates gut bacterium A. Shahii through iron sequestration.
(a–c) ELISA analysis of serum Lcn2 protein concentration in male and female MB49 tumor-bearing mice (n = 7) (a), and in male and female MB49 tumor-bearing mice undergo castration (Cas), treated with enzalutamide (Enza), Lurasidone (Lura) and anti-Ly6G antibody (αLy6G) (F-Ctrl, n = 6; F-Lura, n = 5; M-Cas/F-αLy6G, n = 4; others, n = 7) (b), and in wild-type (WT) and Retnlg-DTR-GFP (DTR) MB49 tumor-bearing mice that were intraperitoneally injected with diphtheria toxin (DT) between male and female (M-WT, n = 8; F-WT, n = 9; M-DTR, n = 6; F-DTR, n = 5) (c). (d) Relative abundance of A. Shahii treated with PBS or LCN2 proteins (n = 4). (e–g) The free Fe2+ concentration (e) and relative abundance of A. Shahii (f) in stool, relationship between stool Fe2+ and relative abundance of A. Shahii (g) in 9.5 Gy irradiated female mice receiving male WT, male Lcn2 KO and female WT bone marrow, then inoculated with MB49 tumor cells 8 weeks after BM reconstitution (n = 7). (h–k) Lcn2 concentration in serum (h), relative abundance of A. Shahii in stool (i), lurasidone concentration in serum (j) and relationship between serum lurasidone concentration and relative abundance of A. Shahii (k) in male and female mice injected with PBS and female mice injected with Lcn2 one time every three days (n = 8). (l, m) Flow analysis of intratumoral Lcn2+ neutrophil proportion (n = 8) (l) and tumor growth (F-Lcn2, n = 9; others, n = 8) (m) in male and female mice injected with PBS and female mice injected with Lcn2 one time every three days. Bar graphs are presented as mean ± s.e.m. A two-tailed Student’s t-test was performed for comparisons. The data are representative of at least three independent experiments.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1 and Tables 1–7.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data and unprocessed western blots.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhu, Q., Zhang, G., Cao, M. et al. Microbiota-shaped neutrophil senescence regulates sexual dimorphism in bladder cancer. Nat Immunol 26, 722–736 (2025). https://doi.org/10.1038/s41590-025-02126-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02126-6
This article is cited by
-
Lipocalin 2: a double-edged sword in cellular ferroptosis
Cell Biology and Toxicology (2026)
-
Cellular senescence in cancer: from mechanism paradoxes to precision therapeutics
Molecular Cancer (2025)
-
The association between 4-HPR-mediated LCN2 suppression and reduced intestinal cell senescence in ulcerative colitis
Clinical and Experimental Medicine (2025)